The autoregulation of a eukaryotic DNA transposon
Figures
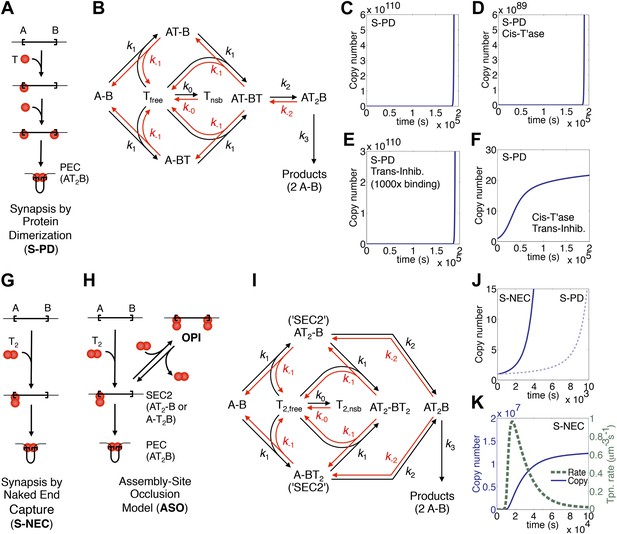
Dynamic models for genome invasions.
(A) The mechanism of transpososome assembly in Tn10/5. In the bacterial elements Tn10 and Tn5 synapsis is mediated by the dimerization of monomers bound to either end of the transposon. T, transposase; T2, transposase dimer; A and B, transposon ends; PEC, paired ends complex. (B) The kinetic model embedded in the computer simulation for the S-PD mechanism of synapsis. Abbreviations are as given in part (A); Tfree and Tnsb, free and non-specifically bound transposase. (C) Simulation of the S-PD model. Kinetic parameters: k0, 9.9 × 106 M−1s−1; k−0, 139 s−1; k1, 3.8 × 108 M−1s−1; k−1, 1.2 × 10−2 s−1; k2, 12.7 s−1; k−2, 1 × 10−10 s−1; k3, 1.4 × 10−3 s−1. Sources of the values are given in Table 2. (D) As in part (C) but the transposase is 99% cis-acting. 1% of the transposase leaks into the bulk phase and acts on all copies of the element. (E) As in part (C) but the transposon expresses a trans-acting inhibitor which reduces the transposase concentration. The inhibitor is 1000-fold more active than the transposase, which approximates the situation in Tn10 where the inhibitor is an antisense RNA. (F) The cis-acting transposase and the trans-acting inhibitor from parts (D) and (E) are combined. (G) The mechanism of transpososome assembly in mariner. A transposase dimer bound to one transposon end recruits a naked transposon end. Abbreviations as in part (A). (H) The assembly-site occlusion model. When the transposase concentration is low, most transposons are occupied by only one transposase dimer, leading to productive synapsis (bottom left). When the transposase concentration is high, a transposon may be occupied by two transposase dimers, causing overproduction inhibition (OPI, top right). SEC2 is a transposon with a transposase dimer bound to one of the ends. (I) Kinetic model embedded in the computer simulation for the ASO mechanism. Abbreviations as given in parts (A) and (B). (J) Simulation of the S-NEC mechanism and comparison with S-PD. Parameters as in part (C). (K) The time scale of the S-NEC simulation in part (J) is extended.
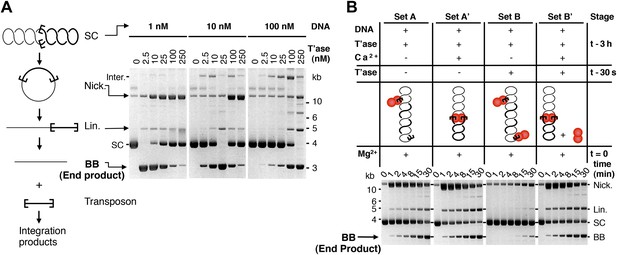
Experimental test of the ASO model for mariner transposition.
(A) The efficiency of transposition depends on the ratio of transposase to DNA. The cartoon to the left of the gels illustrates the mechanism of cut-and-paste transposition using a supercoiled (SC) substrate. First strand nicking generates an open circular product (Nick.). Second strand nicking at one end yields the linear single-end break product (Lin.). A similar set of nicks at the other transposon end yield the plasmid backbone (BB) plus the excised transposon. Transposase was titrated into reactions with three different substrate concentrations. Reactions contained 1.5 µg of supercoiled substrate plasmid in a volume of 500 µl, 50 µl and 5 µl, which provided a final concentration of 1 nM, 10 nM and 100 nM, respectively. Transposase was diluted so that the addition of one tenth of the respective reaction volumes achieved the indicated concentrations. Reactions were incubated for 4 hr at 37 °C and deproteinated. Photographs of ethidium bromide stained agarose gels are shown. Consumption of the supercoiled substrate and production of the plasmid backbone both provide a measure of the efficiency of the reaction. Inter., product of intermolecular integration of the transposon into an unreacted substrate. (B) Preassembly of the paired-end complex protects from OPI. Four sets of staged reactions were assembled as indicated. Set A was a standard transposition reaction with 6.7 nM plasmid substrate and 20 nM of transposase except that the components were pre-incubated for 3 hr before the addition of the catalytic Mg2+ ion. Set A′ was identical except that Ca2+ was present during the 3 hr pre-incubation period, which enhances PEC assembly. Sets B and B′ were identical to A and A′ except that the mixture was supplemented with 50 nM transposase 30 s before the addition of Mg2+. OPI inhibited the reaction in Set B. In contrast, the stable PEC assembly supported by the presence of Ca2+ protected Set B′ from the inhibitory effects of the excess transposase added just before the catalytic Mg2+ ion. Photographs of ethidium bromide stained agarose gels are shown. See also Figure 2—figure supplement 1.
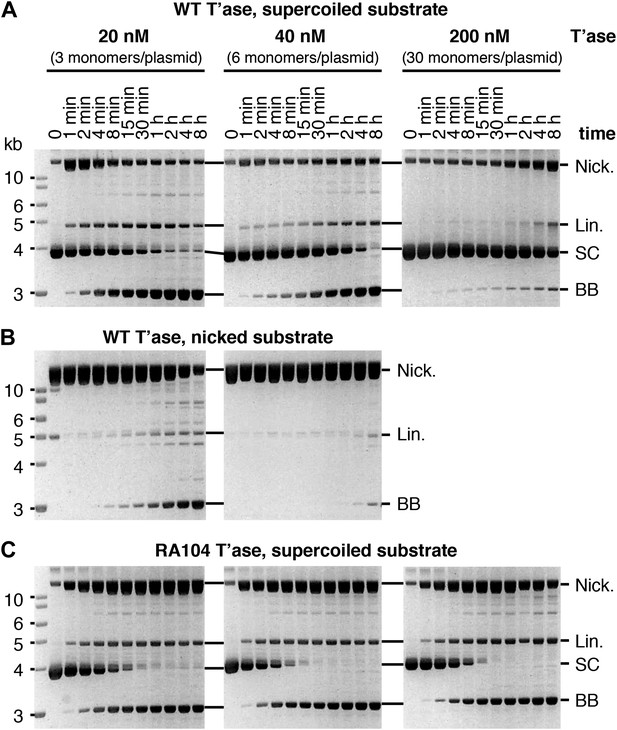
Kinetic analyses of OPI and interpretation of in vitro transposition reactions.
The kinetics of the transposition reaction were analyzed in standard reactions containing 6.7 nM plasmid substrate and the indicated concentrations of the transposases. Photographs of ethidium bromide stained agarose gels are shown. (A) The effects of the ASO mechanism complicates the interpretation of the gels, particularly the kinetic analyses. In the reactions with 20 nM transposase about half of the substrate is converted to the nicked intermediate in the first 2 min. However, it takes an hour or more to consume the remainder of the substrate. This biphasic behavior arises from the ASO mechanism, which operates in any reaction that contains more than one dimer of transposase. Thus, even if the transposase was 100% active and the reaction was performed with the optimum ratio of one dimer to one transposon, only half of the substrate would react initially. At the start of the reaction half of the transposons would be occupied by a dimer and would react, a quarter would be occupied by two dimers and would suffer OPI and the other quarter would be completely unoccupied. The slow phase of consumption corresponds to the redistribution of dimers from transposons which initially suffered OPI to those that were completely unbound. Even under OPI conditions, when both ends of the transposon are occupied by transposase dimers, their occasional dissociation provides a window of opportunity for synapsis. OPI can therefore be overcome by extending the incubation period so that the accumulated windows of opportunity eventually suffice to complete the reaction. Thus, the reaction with 40 nM transposase goes on to reach completion during the 8 hr incubation, despite initially suffering from OPI. With 200 nM transposase, the extended incubation is unable to overcome OPI because the windows of opportunity provided by unoccupied transposon ends are shorter at this concentration. (B) The nicked substrate is more sensitive to OPI and is inhibited at only 40 nM transposase. This is because any free transposon ends, made available by the dissociation of a transposase dimer, are re-bound before they can achieve synapsis. In other words, the slow synapsis of the nicked substrate requires a longer window of opportunity than is available at 40 nM transposase. Time points and transposase concentrations as in part (A). (C) The RA104 transposase mutant is resistant to OPI because the higher OFF-rate provides more windows of opportunity for synapsis (see Figure 4C and text for details). Time points and transposase concentrations as in part (A).
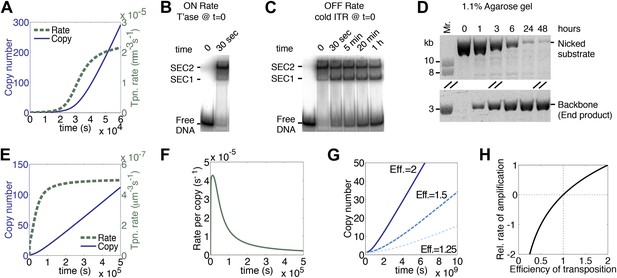
A semi-quantitative description of mariner transposition.
(A) The ASO model for mariner transposition was simulated after taking account of the slow diffusion which prevails in vivo: k0, 5 × 106 M−1s−1; k1, 1.9 × 108 M−1s−1; k2, 4.5 × 10−4 s−1. Other parameters were as in Figure 1K. (B) Transposase binds the transposon end rapidly and tightly. Binding reactions (20 µl) contained 40 fmol of 32P-labeled transposon end and 120 fmol of transposase. The reactions were incubated at 37 °C and separated on a native polyacrylamide gel. The lane indicated as time zero contains no transposase. The 30 s time interval was the time required to mix the sample, load the gel and apply the voltage. SEC1 and SEC2 represent a single transposon end bound by a transposase monomer and dimer, respectively. An autoradiogram is shown. (C) Binding reactions were set up and analyzed as in part (B). After 5 min incubation, to allow the complexes to form, a 20-fold molar excess of unlabeled transposon end ('cold ITR') was added. The lane indicated as time zero contains transposase but no cold competitor. The rate of transposase dissociation can be estimated from the amount of free DNA released from the complexes. (D) The rate of transposon excision from a nicked substrate provides an estimate of the rate of synapsis. The kinetics of a transposition reaction was analyzed in standard reactions containing 6.7 nM of nicked plasmid substrate and 20 nM transposase. A photograph of an ethidium bromide stained agarose gels is shown. (E) The ASO model was simulated as in Figure 1K but taking account of the effects of allostery, as described in the main text and ‘Materials and methods’. Parameters as in Figure 1K except k2 = 9.6 × 10−5 s−1; k−1 = 5.8 × 104 s−1. (F) As in part (E) but the rate of transposition is divided by the transposon copy number. (G) As in part (A), but taking account of allostery and slow diffusion in vivo: k0, 5.0 × 106 M−1s−1; k1, 1.9 × 108 M−1s−1; k2, 3.4 × 10−9 s−1. The effects of changing transposition efficiency are shown (see text for definition of efficiency). (H) The relationship between transposition efficiency and the relative rate of transposition is plotted with the maximum rate scaled to 1: y = 1.4427 ln(x). See also Figure 3—figure supplements 1 and 2.
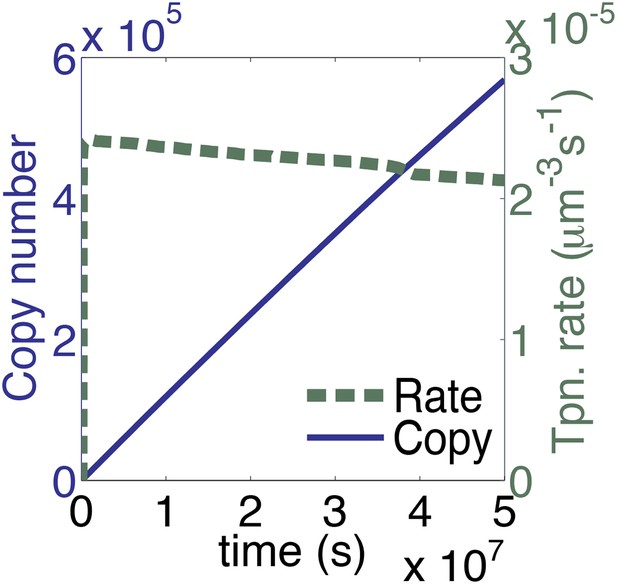
The time scale of the graph in Figure 3A is extended.
The parameters take account of the slow diffusion in vivo, but take no account of allosteric interactions between transposase subunits. This plot therefore represents the in vivo dynamics of a generic homomeric DNA looping reaction in the absence of allostery.
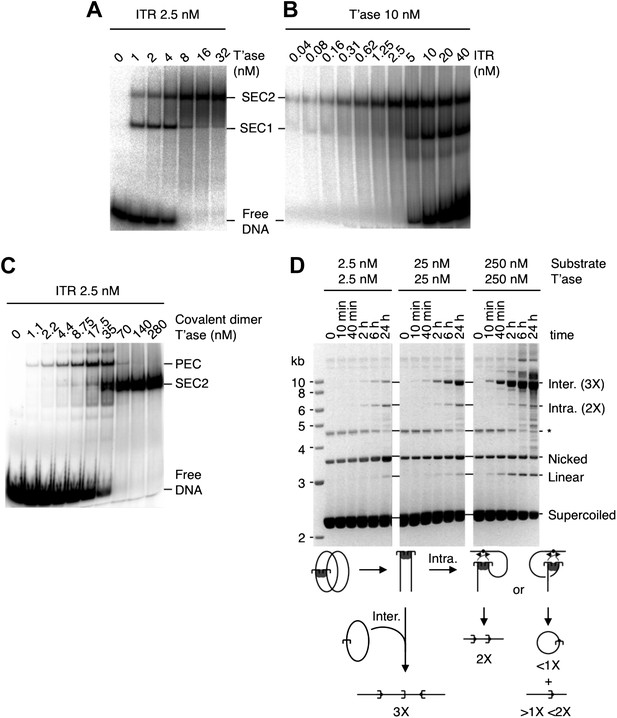
EMSA analysis of transpososome assembly shows that SEC1 arises from dissociation of the PEC.
Transposon ends were encoded on radiolabeled linear DNA fragments. Binding reactions were incubated at 37 °C for 2 hr, separated on a 5% native polyacrylamide gel and recorded by phosphoimaging. (A) SEC1 and SEC2 represent a single transposon end bound by a transposase monomer and dimer, respectively (see main text for details). SEC2 comes to dominate the reaction as the transposase concentration rises. There is a significant transition between 4 and 8 nM transposase when SEC1 largely disappears. This corresponds to the point at which the transposon ends become sub-stoichiometric to the transposase dimers. At this point no free transposon ends remain as they are all bound by transposase. According to the S-NEC mechanism (see Figure 1 for details), SEC2 is converted to the transpososome (=PEC) by recruitment of a naked transposon end. OPI occurs when the transposon ends are sub-stoichiometric to transposase dimers and there is a shortage of free transposon ends available for recruitment. Note that the various species observed in these binding reactions are identical to those observed in reactions with the related Mos1 and Himar1 transposons, which display the same behavior for example (Dawson and Finnegan, 2003; Lipkow et al., 2004). The present data suggests that the PEC in all three of these related systems is unstable in the EMSA and dissociates into two SEC1 complexes soon after the start of electrophoresis. Thus, in agreement with the S-NEC mechanism, SEC1 disappears at the point in the titration when the transposon ends become sub-stoichiometric to transposase dimers. (B) A fixed amount of transposase was titrated with an increasing amount of free transposon ends. The appearance of SEC1 coincides exactly with the appearance of the free transposon ends, which are required for PEC assembly in the S-NEC model. As the amount of transposon ends is increased further, the amount of SEC1 increases. This reflects mass action, which drives PEC assembly by favoring the capture of a free transposon end (see part D below for confirmation). This supports the data in part (A) in suggesting that SEC1 arises from the dissociation of the PEC. (C) Binding reactions were with a single-chain transposase dimer, in which two monomers are concatenated by a linker peptide joining the C-terminus of one to the N-terminus of the other. Concatenation of the subunits stabilizes the PEC, which is now detected in the gel. As the transposase concentration increases, the PEC disappears at the same point as the free DNA and gives way to SEC2. This behavior is identical to SEC1 in parts (A) and (B). The single-chain dimer of transposase is fully proficient for the transposition reaction (not shown), demonstrating that SEC1 is not an obligate intermediate of the reaction. This supports the data in parts (A) and (B) further confirming that SEC1 arises from the dissociation of the PEC. (D) In vitro transposition reactions were performed with a plasmid substrate encoding a single transposon end. Reactions were stopped at the indicated times and deproteinated before analysis on a 1.1% agarose TBE gel stained with ethidium bromide. All three sets of transposition reactions contained the same amounts of transposase and supercoiled plasmid substrate. However, the respective reaction volumes were adjusted to 500 µl, 50 µl and 5 µl to achieve the indicated concentrations. Transpososome assembly requires bimolecular synapsis between ends located on different molecules, as illustrated below the gels. This is inefficient owing to the relatively low concentration of one transposon end with respect to another when on separate molecules. When such a transpososome performs cleavage, followed by integration into an unreacted substrate molecule, the product is a linear molecule three times the size of the substrate (Claeys Bouuaert et al., 2011). There is very little reaction when the substrate concentration is low. This reflects the inefficiency of second end recruitment. At high substrate concentration, mass action drives the reaction by favoring second-end recruitment.
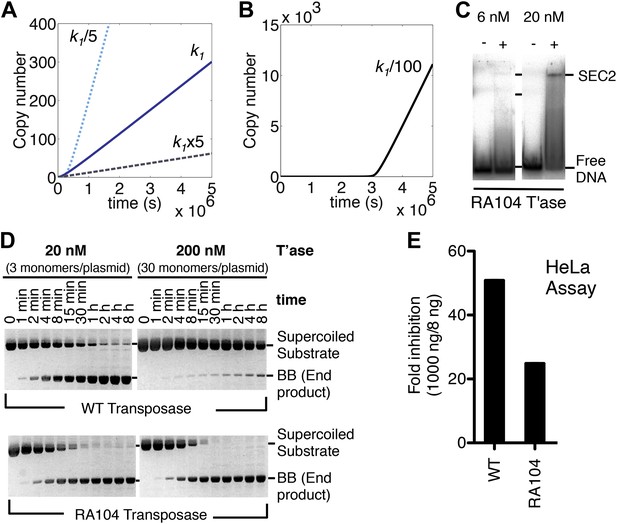
Association and dissociation rate constants.
(A) Simulation as in Figure 3E, with k1 fivefold up or down. We have retained the effects of allostery but here and in subsequent simulations we have ignored the effects of the slow diffusion in vivo. This allows the algorithm to run more smoothly and shortens the scale on the x axis, but does not affect the conclusions, which are based on the differential responses to changing the various parameters. (B) Simulation as in part (A) (solid line), with k1 100-fold down. (C) Binding reactions with the RA104 mutant transposase were set up as in Figure 3B. In the lane with 20 nM transposase the smear between SEC2 and the position of free DNA is probably due to complexes that have dissociated during electrophoresis. Autoradiogram of an EMSA is shown. (D) The kinetics of the transposition reaction were analyzed in standard reactions containing 6.7 nM of supercoiled plasmid substrate and the indicated concentrations of the transposases. Photographs of ethidium bromide stained agarose gels are shown. With 200 nM wild-type transposase, the windows of opportunity for synapsis, provided by periods when one transposon end is unoccupied, are too short to allow for synapsis. The RA104 transposase mutant is resistant to OPI because the higher dissociating rate provides more windows of opportunity for synapsis. (E) Mutant and wild type transposase were assayed in HeLa cell culture with 8 ng or 1000 ng of helper plasmid and 500 ng of neomycin resistant reporter plasmid.
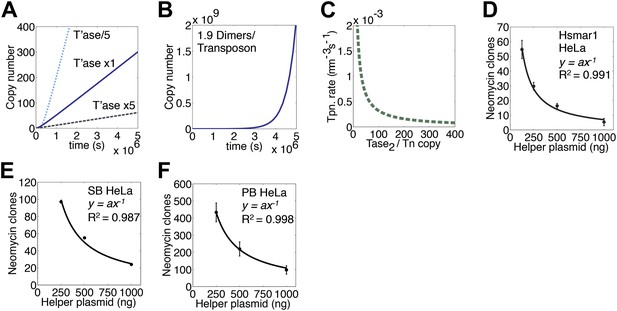
Transposase expression level.
(A) Simulation as in Figure 3E showing the effects of changing the transposase expression fivefold up or down. Unless noted otherwise, each transposon copy produces 500 transposase dimers, which are considered as being contained within a 500 fl nucleus. (B) As in part (A) (1× line) but with a transposase expression level of 1.9 dimers per transposon. (C) Parameters as part (A) (1× line), plotting the relationship between transposase expression and the rate of transposition at the point in the invasion when there are 1000 copies of the transposon present in the genome. (D) HeLa cell assay for Hsmar1 transposition. HeLa cells were transfected with 500 ng of neomycin reporter plasmid and the indicated amount of helper plasmid expressing transposase. The rate of transposition is given by the number of neomycin resistant colonies recovered after drug selection. Bars indicate standard error of the mean, n = 3. R2 is least squares goodness of fit to the line y = ax−1. (E) As in part (D) but with isogenic Sleeping Beauty (SB100X) transposon reporter and helper plasmids. Data points are a mean of three experiments and were extracted and re-plotted from Figure 2A of (Grabundzija et al., 2010). (F) As in part (D) but with isogenic piggyBac transposon reporter and helper plasmids. n = 3. See also Figure 5—figure supplement 1.
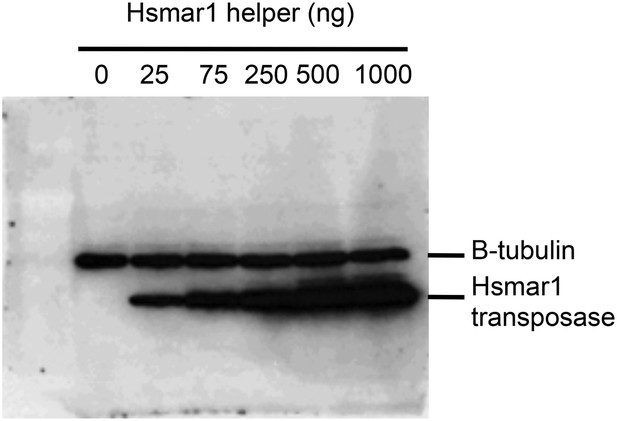
Transposase expression in HeLa cells.
HeLa cells were transfected with an increasing quantity of the helper plasmid, which expresses Hsmar1 transposase from a strong viral promoter. The titration was identical to that shown in Figure 5D. Cells were harvested after two days and analyzed by SDS-PAGE and Western blotting with antibodies against β-tubulin and the Hsmar1 transposase. Transposase expression increased throughout the titration. This indicates that transposase expression is not toxic. If transposase expression had been toxic the amount of transposase recovered from the cells would have plateaued or decreased with higher amounts of helper plasmid transfected.
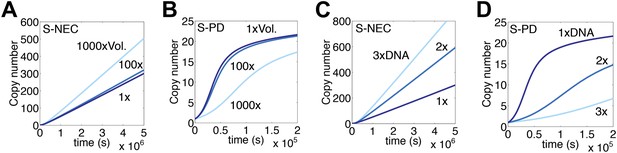
Genome size and nuclear volume.
(A) Simulation as in Figure 3E, changing the nuclear volume by the indicated amounts. (B) Simulation of regulated S-PD mechanism as in Figure 1F, changing the nuclear volume by the indicated amounts. (C) Simulation as in part (A), changing the genome size by the indicated amounts. (D) Simulation as in part (B), changing the genome size by the indicated amounts.
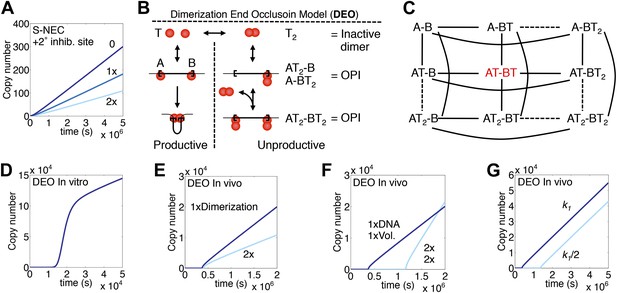
Variations and alternative mechanism of regulation.
(A) The S-NEC mechanism is simulated as in Figure 3E but with a secondary transposase binding site that inhibits transposition when occupied. 0, the secondary site has no affinity for the transposase; 1×, the secondary site has the same affinity for transposase as the primary binding sites at the transposon ends; 2×, secondary site with twice the affinity for transposase. (B) The dimerization end occlusion model for OPI. (C) All possible states of the DEO model are illustrated. A and B are transposon ends, T is an active transposase monomer, T2 is an inhibitory transposase dimer. Solid lines indicate reaction pathways allowed in the model. The active species is shown in red. (D) Simulation of the DEO model using in vitro, non-allosteric parameters as in Figure 1C. Inhibitory dimers bind transposon ends with the same affinity as the active monomers. Active monomers dimerize with the same affinity as they bind transposon ends. (E) DEO model as in part (D) but with in vivo slow diffusion rate as in Figure 3E. Affinity of the monomers in the inhibitory dimer is doubled as indicated. (F) DEO model as in part (E) showing the effect of doubling in the genome size and nuclear volume. (G) DEO model as in part (E) showing the effect of changing the association rate constant of the active monomers and the inhibitory dimers by the indicated amount.
Tables
Kinetic parameters for DNA binding
| Reaction | Kd (M) | k0, k1 (M−1s−1) | k0, k1 (µm3 s−1) | k−0, k−1 (s−1) | t1/2 (s) |
|---|---|---|---|---|---|
| LacI specific binding | 3.1 × 10−11 | 3.8 × 108 * | 0.633 | 1.2 × 10−2 * | 58 † |
| LacI non-specific binding | 1.4 × 10−5 ‡ | 9.9 × 106 § | 0.0166 | 139 § | 5 × 10−3 † |
-
*
From Table 3 of Hsieh et al. (1987).
-
†
This is the experimental estimate for LacI bound to non-specific DNA dissociating to the bulk solution (Elf et al., 2007). During this time it will have visited about 85 non-specific sites by one-dimensional diffusion. However, for the purposes of the simulation it is convenient to subsume the two phases into a single bulk behavior of the system (see text for details).
-
‡
-
§
Estimated from Kd using the relationship Kd = k−1/k1 or Kd = k−0/k0.
Estimation of k2 for transposase-DNA interactions in vitro
| Quantity | Symbol | Value | Source |
|---|---|---|---|
| Reaction 1 rate constant | k1 | 3.8 × 108 M−1s−1 | Table 3 of Hsieh et al. (1987). |
| DNA persistence length | a | 50 nm | Well established (e.g. Petrov et al., 2006). |
| Reaction radius | R | 2.5 nm | The transposase dimeric binding domain is 111 amino acids and weighs 26 kDa. From Note 3 of Andrews (2012), its radius is about 1.9 nm. Add to this about 0.6 nm to account for the 0.34 nm length of 1 DNA basepair (because transposase specificity is accurate to 1 bp), combined with the 1 nm radius of DNA. |
| Segmental diffusion coefficient | Ds | 27 µm2s−1 | Figure 3 of Petrov et al. (2006) for 147 bp, which is 1 DNA persistence length. |
| Transposase diffusion coefficient | Dp | 61 µm2s−1 | Note 3 of Andrews (2012), using a molecular weight of 80.7 kDa for transposase. Assumes DNA center of mass is effectively stationary. |
| Diffusion-limited part of k1 | k1,diff | 1.5 × 109 M−1s−1 | Equation 5. |
| Activation-limited reaction rate | kact | 5.2 × 108 M−1s−1 | Equation 3. |
| Diffusion-limited part of | 4.2 × 108 M−1s−1 | Equation 6. | |
| Reaction 2 rate constant | 2.3 × 108 M−1s−1 | Equation 4. | |
| Effective local DNA site concentration | [B] | 5.5 × 10−8 M | Equation 8, using a 1287 bp transposon length. |
| Transposase-transposon end dissociation rate constant | k−1 | 5.8 × 10−4 s−1 | Estimated from Figure 3C. |
| Reaction 2, pseudo-first order rate constant for synapsis | k2 | 12.7 s−1 | Equation 7. |
| Reaction 2, pseudo-first order rate constant for synapsis with an open circular substrate in vitro. | k2 | 9.6 × 10−5 s−1 | Estimated using the equation t1/2 = ln2/k2. t1/2 is the time taken to consume one quarter of the substrate in Figure 3D. See Figure 2—figure supplement 1 for an explanation of why the time to consume one quarter of the substrate is used, rather than the time to consume half of the substrate. |
Estimation of in vivo transposase-DNA interactions
| Quantity | Symbol | Value | Source |
|---|---|---|---|
| DNA persistence length | a | 50 nm | See Table 2. |
| Reaction radius | R | 2.5 nm | See Table 2. |
| Segmental diffusion coefficient | Ds | 5 × 10−4 µm2s−1 | From Figure 2 caption of Marshall et al. (1997). |
| Transposase diffusion coefficient | Dp | 15 µm2s−1 | ¼ of Table 2 value, based on Note 3 of Andrews, (2012). |
| Diffusion-limited part of k1 | k1,diff | 2.9 × 108 M−1s−1 | Equation 5. |
| Activation-limited reaction rate | kact | 5.2 × 108 M−1s−1 | From Table 2. |
| Diffusion-limited part of | 7.8 × 103 M−1s−1 | Equation 6. | |
| Reaction 1 rate constant | k1 | 1.9 × 108 M−1s−1 | Equation 3. |
| Reaction 2 rate constant | 7.8 × 103 M−1s−1 | Equation 4. | |
| Effective local DNA site concentration | [B] | 5.5 × 10−8 M | See Table 2. |
| Reaction 2 pseudo-first order rate constant | k2 | 4.3 × 10−4 s−1 | Equation 7. |





