Discovery of a metabolic alternative to the classical mevalonate pathway
Figures
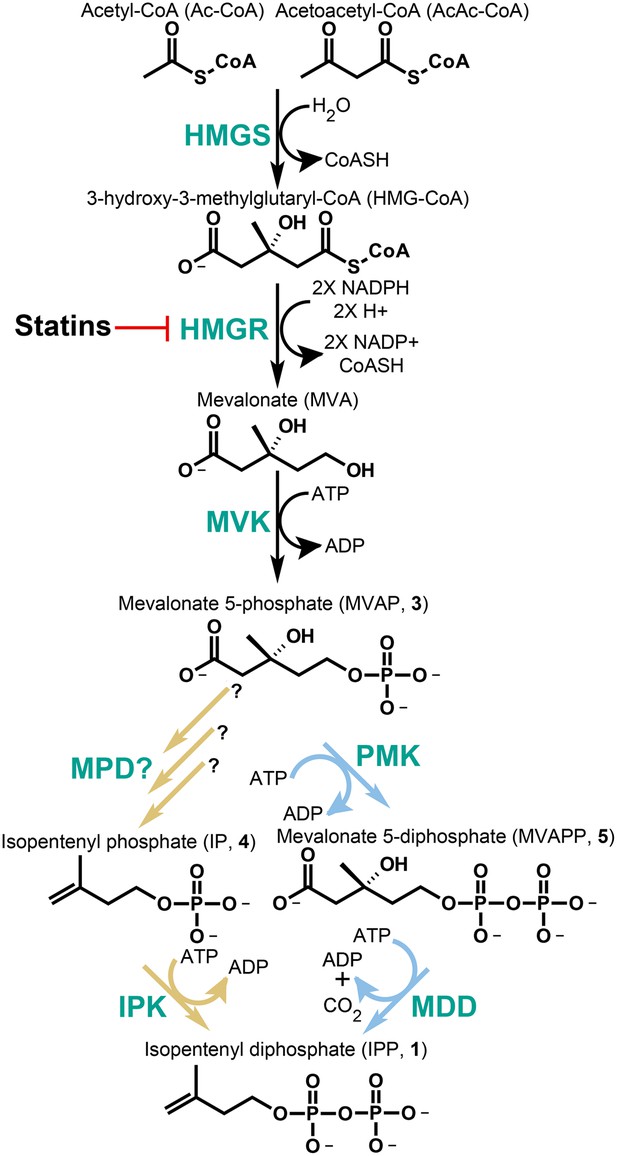
The classical and alternative MVA pathways.
Both branches of the MVA pathway begin with acetyl-CoA (and acetoacetyl-CoA) and proceed through a series of enzymatic reactions involving 3-hydroxy-3-methylglutary-CoA Synthase (HMGS), 3-hydroxy-3-methylglutary-CoA Reductase (HMGR, the presumed early rate-limiting step), and mevalonate kinase (MVK) before branching. At the bifurcation, the canonical MVA pathway, highlighted by light blue arrows, guides MVAP (3) through an additional phosphorylation reaction followed by a phosphorylation-dependent decarboxylation carried out by phosphomevalonate kinase (PMK) and diphosphomevalonate decarboxylase (MDD), respectively. The alternative MVA pathway, highlighted with light brown arrows, hypothetically decarboxylates MVAP (3) prior to the phosphorylation reaction carried out by IPK but the former step has not been discovered until now (Grochowski et al., 2006). The enzymes MVK, PMK, MDD, and IPK all consume ATP during catalysis. All enzymes are shown in green type. Statins serve as inhibitors of HMGR as highlighted.
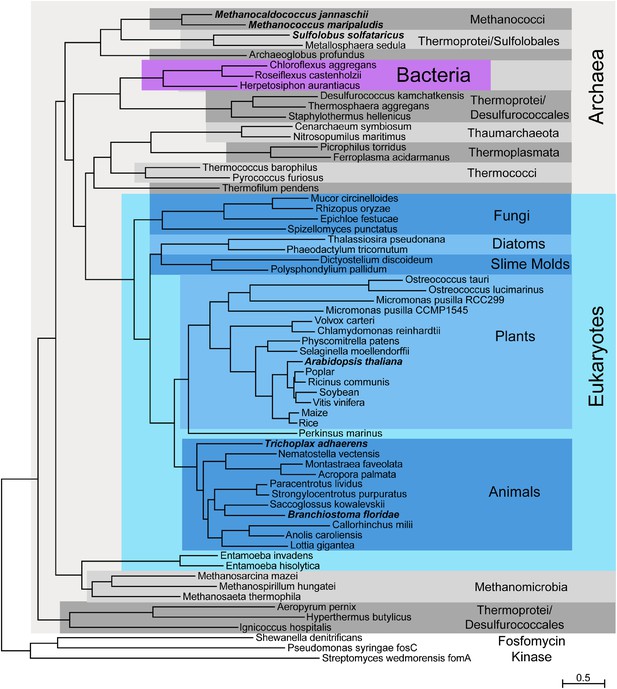
Phylogenetic distribution of IPK across the three domains of life.
Maximum likelihood tree of selected IPK protein sequences. Eukaryotes are highlighted with blues, selected archaeal clades with grays and a small group of bacteria with purple. The tree is anchored by several bacterial fosfomycin kinases. See Figure 2—source data 1 for an alignment of IPK homologs. See Figure 2—source data 2 for a table of IPK homolog sequences.
-
Figure 2—source data 1
Alignment of IPKs from the three domains of life.
- https://doi.org/10.7554/eLife.00672.005
-
Figure 2—source data 2
Additional IPK sequences from an assortment of sequence databases.
- https://doi.org/10.7554/eLife.00672.006
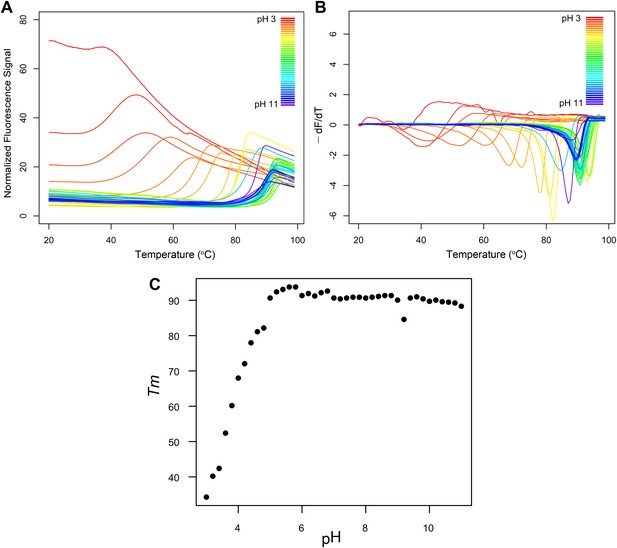
Fluorescence thermal shift assays of Roseiflexus castenholzii MDD-like MPD.
(A) Thermograms for R. castenholzii MPD in 100 mM buffer (pH 3.0–3.8 citric acid; pH 4.0–4.8 sodium acetate; pH 5.0–5.8 sodium citrate; pH 6.0–6.8 sodium cacodylate; pH 7.0–7.8 sodium HEPES; pH 8.0–8.8 Tris-HCl; pH 9.0–11.0 CAPSO) colored from red to violet (acidic to alkaline pH depicted in the inset). (B) Negative derivatives of the thermograms (−dF/dT) color-coded as in (A). (C) Tms for each of the curves show in (A) and (B) plotted as a function of pH. R. castenholzii MPD was unfolded from pH 2.2 to 2.8 at 20°C (data now shown).
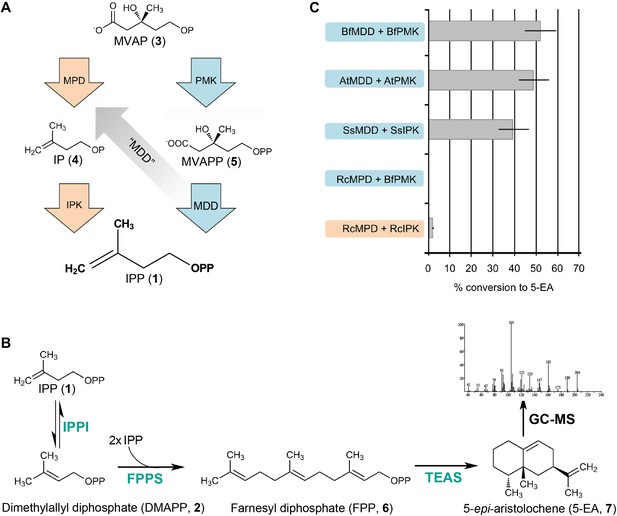
In vitro reconstitution and GC-MS analysis of the alternative and classical MVA pathways.
(A) Terminal steps of the MVA pathways shown with enzymes of the alternative (orange) and classical (blue) pathways depicted as arrows. The putative neofunctionalization of MDD to MPD is highlighted by a grey arrow. (B) In vitro assays include either the alternative or the classical enzymes to produce IPP (1) (as shown in panel A) as well as all downstream enzymes, including isopentenyl phosphate isomerase (IPPI) to produce DMAPP (2), farnesyl diphosphate synthase (FPPS) to produce farnesyl diphosphate (FPP, 6) and tobacco 5-epi-aristolochene synthase (TEAS) to produce the sesquiterpene product, 5-EA (7). Products are separated by GC and detected by MS ionization and fragmentation. Enzymes used are highlighted in turquoise type. (C) Results of the in vitro reconstitutions of various enzyme combinations. The y-axis of the graph represents combinations of enzymes shown in panel A and panel B and the % conversion to the expected sesquiterpene end product, 5-EA (7) shown as grey bars along the x-axis. Abbreviations of organisms are as follows: Bf = Branchiostoma floridae , At = Arabidopsis thaliana, Ss = Sulfolobus solfataricus, and Rc = Roseiflexus castenholzii. Note, RcMPD is colored both orange and blue on the y-axis, depending on whether it is being tested as an MDD (blue) or an MPD (orange).
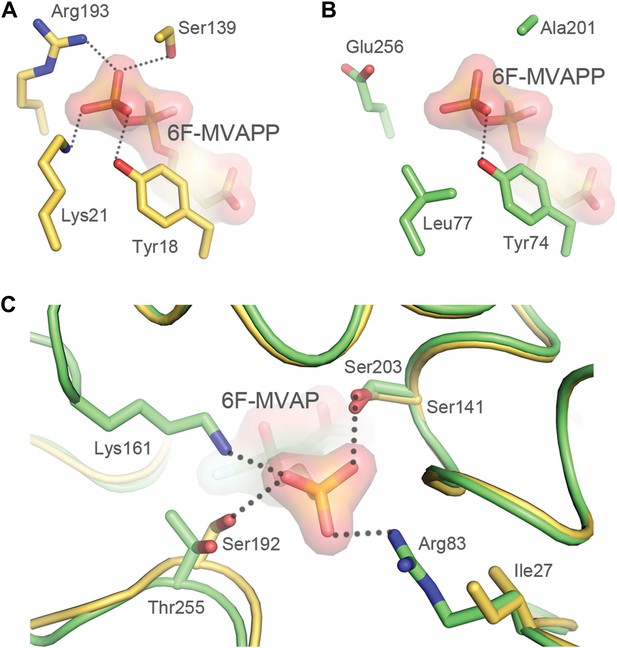
Structural comparisons of a bona fide MDD and MPD.
Interactions between the terminal phosphate of 6F-MVAPP and the surrounding amino acid residues from the crystal structure of S. epidermidis MDD (PDB ID 3QT7(Barta et al., 2011)) and the 3D model of MPD from R. castenholzii. (A) S. epidermidis MDD has multiple interactions with the diphosphate of MVAPP (5). Atoms are colored by type with carbon gold. (B) The active site model of R. castenholzii MPD lacks many of the key interactions shown by S. epidermidis MDD in panel A. Atoms are colored by type with carbon green. (C) Interactions between the monophosphate of modeled 6F-MVAP and the surrounding amino acids in a superposition of the modeled R. castenholzii MPD, backbone atoms and carbon colored green, on the crystal structure of S. epidermidis MDD, backbone atoms and carbon colored gold. In R. castenholzii MPD, two divergent side chains, Arg83 and Lys161, putatively provide additional electrostatic interactions with the single phosphate group of MVAP (3). These amino acid side chains would clash with the second phosphate of MVAPP (5). These models suggest that the predicted active site topology of R. castenholzii MPD facilitates substrate recognition of MVAP (3) through complementary charged and polarized hydrogen bonds and excludes MVAPP (5) through steric incompatibility with its second phosphate.
Tables
Phylogenetic distribution of IPK in Eukarya
| Classification | IPK-bearing species | Species with no IPK |
|---|---|---|
| Mammals | 35 mammalian genomes | |
| Birds | Gallus gallus | |
| Taeniopygia guttata | ||
| Meleagris gallopavo | ||
| Reptiles | Anolis carolensis | |
| Philodryas olfersii (EST)* | ||
| Amphibians | Notophthalmus viridescens (EST) | Xenopus tropicalis |
| Fish | Callorhinchus milii (partial IPK from draft genome) | Danio rerio |
| Oryzias latipes | ||
| Takifugu rubripes | ||
| Tetraodon nigrovidris | ||
| Gasterosteus aculeatus | ||
| Invertebrate chordates | Branchiostoma floridae | Ciona intestinalis |
| Saccoglossus kowalevskii | Ciona savignyi | |
| Echinoderms (Deuterostomes) | Paracentrotus lividus (EST) (almost complete assembly) | |
| Strongylocentrotus purpuratus (almost complete prediction) | ||
| Arthropods | Homarus americanus (EST) | Drosophila sp. (Cassera et al., 2004) |
| Apis mellifera | ||
| Aedes aegypti | ||
| Anopheles gambiae | ||
| Culex quinquefasciatus | ||
| Pediculus humanus | ||
| Ixodes scapularis | ||
| Daphnia pulex | ||
| Other bilaterians | Hirudo medicinalis (EST) | Capitella teleta |
| Eisenia fetida (EST) | Helobdella robusta | |
| Lottia gigantea | Schistosoma mansoni | |
| Aplysia kurodai (EST) | Schistosoma japonicum | |
| Aplysia californica (partial prediction) | Brugia malayi | |
| Meloidogyne incognita | ||
| Pristionchus pacificus | ||
| Cnidarians | Acropora palmate (EST) | Hydra magnapilliata |
| Montastraea faveolata (EST) | ||
| Nematostella vectensis (predicted) | ||
| Other early metazoans | Trichoplax adherens | Amphimedon queenslandica |
| Pre-metazoans (within Holozoa) | Monosiga brevicollis | |
| Salpingoeca rosetta | ||
| Capsaspora owczarzaki | ||
| Fungi | Spizellomyces punctatus | 34 fungal genomes |
| Rhizopus oryzae | ||
| Mucor circinelloides | ||
| Epichloe festucae (EST)‡ | ||
| Green plants | all plants, including: | |
| Selaginella moellendorffii | ||
| Physcomitrella patens | ||
| Ostreococcus tauri | ||
| Ostreococcus lucimarinus | ||
| Micromonas pusilla | ||
| Chlamydomonas reinhardtii | ||
| Volvox carteri | ||
| Amoebozoa | Dictyostelium discoideum | |
| Polysphondylium pallidum | ||
| Entamoeba hislolytica§ | ||
| Entamoeba invadens§ | ||
| Entamoeba dispar§ | ||
| Alveolata | Perkinsus marinus | Plasmodium sp. (Lynen, 1967) |
| Cryptosporidium sp. (Hedden and Kamiya, 1997) | ||
| Tetrahymena thermophila | ||
| Paramecium tetraurelia | ||
| Ichthyophthirius multifilis | ||
| Diatoms | Thalassiosira pseudonana | |
| Phaeodactylum tricornutum | ||
| Kinetoplastida | Trypanosoma sp. (Chew and Bryant, 2007) | |
| Leishmania sp. (Chew and Bryant, 2007) | ||
| Excavates | Giardia lamblia | |
| Trichomonas vaginalis | ||
| Others | Malawimonas jakobiformis (EST) | Naegleria gruberi |
| Alexandrium tamarense |
-
*
EST stands for expressed sequence tag. All other IPKs were found in draft or complete genomes.
-
‡
Since IPK was not found in related fungal genomes, this may be a case of horizontal transfer or even EST library contamination.
-
§
Greater similarity to IPK from a clade of archaea than to eukaryotes.
Conserved residues in IPK
| Highly-conserved residues* | Partially conserved residues |
|---|---|
| K6, G8, G9, K15, G54, H60, P140, G144, D213, T215, G216 | K221†, G253‡, T254‡ |
-
*
Numbering is in accordance with IPK from M. jannaschii.
-
†
Conserved in nearly all species except one possibly due to a gene prediction error.
-
‡
May be invariant, but mis-predicted in several sequences.
Steady-state kinetic constants for enzymes of the MVA pathway*
| Organism | gi number | Enzyme | Substrate | KM (μM) | kcat (s−1) | Ki (μM) | kcat/KM (s−1μM−1) | R2 (†) |
|---|---|---|---|---|---|---|---|---|
| M. jannaschii | 15668214 | IPK | IP | 4.3 (± 0.6)‡ | 1.46 (± 0.03) | –§ | 0.34 (± 0.05) | 0.90 |
| M. maripaludis | 132664414 | IPK | IP | 21.4 (± 4.3) | 15.2 (± 1.4) | 877 (± 550) | 0.71 (± 0.16) | 0.99 |
| S. solfataricus | 15897030 | IPK | IP | 23.6 (± 4.8) | 0.91 (± 0.05) | – | 0.04 (± 0.01) | 0.92 |
| 15899698 | PMK | MVAP | 23.1 (± 3.8) | 1.98 (± 0.10) | 5500 (± 2200) | 0.09 (± 0.01) | 0.98 | |
| R. castenholzii | 156743980 | IPK | IP | 4.3 (± 0.7) | 1.70 (± 0.04) | – | 0.40 (± 0.06) | 0.96 |
| 156740939 | MPD# | MVAP | 152 (± 38) | 1.7 (± 0.1) | – | 0.011 (± 0.003) | 0.98 | |
| 156740939 | MDD | MVAPP | ND# | ND | ND | ND | ND | |
| B. floridae | NA¶ | IPK | IP | 13.3 (± 2.0) | 27.2 (± 1.2) | 2820 (± 1700) | 2.05 (± 0.32) | 0.98 |
| 260829481 | PMK | MVAP | 38.0 (± 7.3) | 12.6 (± 0.4) | – | 0.33 (± 0.06) | 0.93 | |
| 260794527 | MDD | MVAPP | 15.1 (± 3.7) | 1.36 (± 0.09) | – | 0.09 (± 0.02) | 0.89 | |
| A. thaliana | 22329798 | IPK | IP | 0.79 (± 0.35) | 1.9 (± 0.2) | 522 (± 381) | 2.4 (± 1.1) | 0.95 |
| 15222502 | PMK | MVAP | 11.8 (± 2.0) | 20.9 (± 0.7) | – | 1.77 (± 0.31) | 0.97 | |
| 15224931 | MDD | MVAPP | 15.7 (± 5.0) | 2.02 (± 0.13) | – | 0.13 (± 0.04) | 0.89 | |
| T. adhaerens | 195996013 | IPK | IP | 3.1 (± 1.6) | 2.4 (± 0.2) | – | 0.77 (0.40) | 0.79 |
-
*
See Table 3—source data 1, Table 3—source data 2, Table 3—source data 3, and Table 3—source data 4 for Michaelis-Menten fitted kinetic curves for each enzyme.
-
†
R2 represents goodness of fit for the kinetic curve to the experimental data. R2 = 1.0 − (SSreg/SStot), where SSreg = sum of squares, SStot = sum of squares of the distances between each point and a horizontal line passing through the average of all y values.
-
‡
Values in parentheses represent standard error (or propagation of error) for each calculated kinetic constant.
-
§
Ki constant was not calculated or was not applicable.
-
#
Not detected.
-
¶
B. floridae IPK sequence was predicted from scaffold_167 of the genome assembly (Putnam et al., 2008) using genewise and manual annotation.
-
Table 3—source data 1
Steady-state kinetic plots for IPKs listed above each curve.
- https://doi.org/10.7554/eLife.00672.010
-
Table 3—source data 2
Steady-state kinetic plots for PMKs listed above each curve.
- https://doi.org/10.7554/eLife.00672.011
-
Table 3—source data 3
Steady-state kinetic plots for MDDs listed above each curve.
- https://doi.org/10.7554/eLife.00672.012
-
Table 3—source data 4
Stead-state kinetic plots for MPD from R. castenholzii.
- https://doi.org/10.7554/eLife.00672.013
Active site phosphate-binding residues identified in MDDs across Archaea, Bacteria and Eukarya*
| MDD-like active site residues | ||||||
|---|---|---|---|---|---|---|
| S. epidermidis | Tyr18 | Lys21 | Ile27 | Ser139 | Ser141 | Ser192 |
| Bacteria | Tyr | Lys | X† | Ser | Ser | Ser |
| Sulfolobales | Tyr | Lys | Asn | Ser | Ser | Ser |
| Eukarya | Tyr | Lys | Ile/Asn | Ser | Ser | Ser |
| MPD-like active site residues | ||||||
| R.castenholzii | Tyr74 | Leu77 | Arg83 | Ala201 | Ser203 | Thr255 |
| Haloarchaea/Thermoplasmatales | Tyr /Phe | Met/Tyr | Arg | Ser | Ser | Polar‡ |
| Chloroflexi | Tyr | Leu | Arg/Thr | Ala | Ser | Thr |
-
*
See Table 4—source data 1, Table 4—source data 2, Table 4—source data 3, and Table 4—source data 4 for alignments.
-
†
Not conserved among canonical bacterial MDDs.
-
‡
Includes Glu, Asp, Asn, and Ser.
-
Table 4—source data 1
Amino acid sequence alignments of archaeal MDDs.
- https://doi.org/10.7554/eLife.00672.018
-
Table 4—source data 2
Amino acid sequence alignments of bacterial MDDs.
- https://doi.org/10.7554/eLife.00672.019
-
Table 4—source data 3
Amino acid sequence alignments of Chloroflexi MDDs.
- https://doi.org/10.7554/eLife.00672.020
-
Table 4—source data 4
Amino acid sequence alignments of eukaryotic MDDs.
- https://doi.org/10.7554/eLife.00672.021





