Scaffold nucleoporins Nup188 and Nup192 share structural and functional properties with nuclear transport receptors
Figures
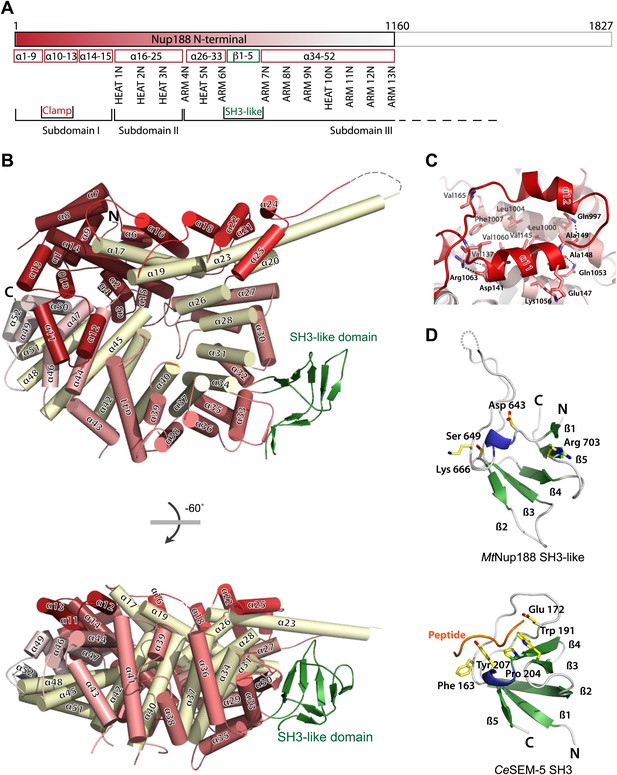
Crystal structures of Nup188N.
(A) Schematic diagram of full-length Myceliophthora thermophila Nup188N with details about the subdomain arrangement. The crystallized fragment is boxed and gradient-colored in red with helical HEAT- and ARM-repeats indicated. (B) The crystal structure of Nup188N is shown in cartoon representation. The helices are gradient-colored as in (A). The inner helical ring is in pale-yellow to highlight the superhelical organization of the protein. The SH3-like domain insert is shown in green. (C) Close-up of the ring-closing latch. The clamping helices α11 and α12 (red) contact a substantial surface area formed by helices 44, 46, and 47. (D) The SH3-like domain of Nup188 compared to the canonical, peptide-bound SH3 domain of Sem-5 from C. elegans (PDB 1SEM). Conserved residues important for peptide interaction are labeled. Note that these residues are not conserved in the SH3-like domain of Nup188. In addition, the SH3-like domain is a circular permutation of the canonical SH3-domain.
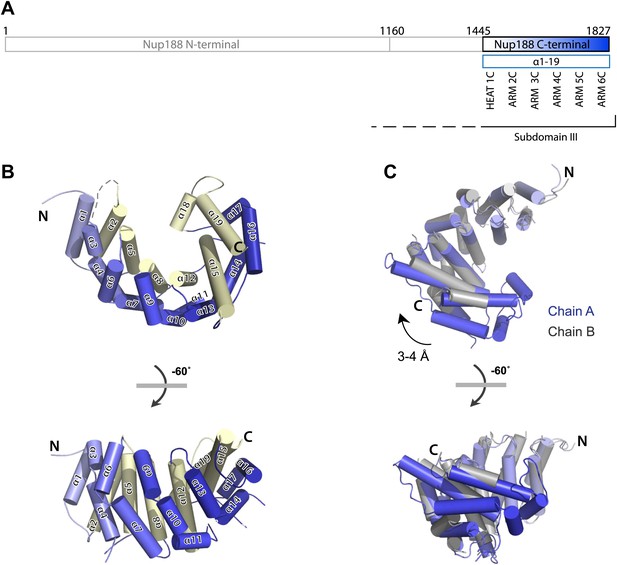
Crystal structures of Nup188C.
(A) Schematic diagram of full-length Myceliophthora thermophila Nup188C with details about the subdomain arrangement. The crystallized fragments is boxed and gradient-colored in blue with helical HEAT- and ARM-repeats indicated. (B) Crystal structure of the C-terminal part of Nup188, gradient-colored as in (A), with the inner helices in pale-yellow. (C) Superposition of chains A (blue) and B (gray) of Nup188C shows the flexibility seen in the crystal, in which the outer helices of chain B move by 3–4 Å relative to the helices in chain A.
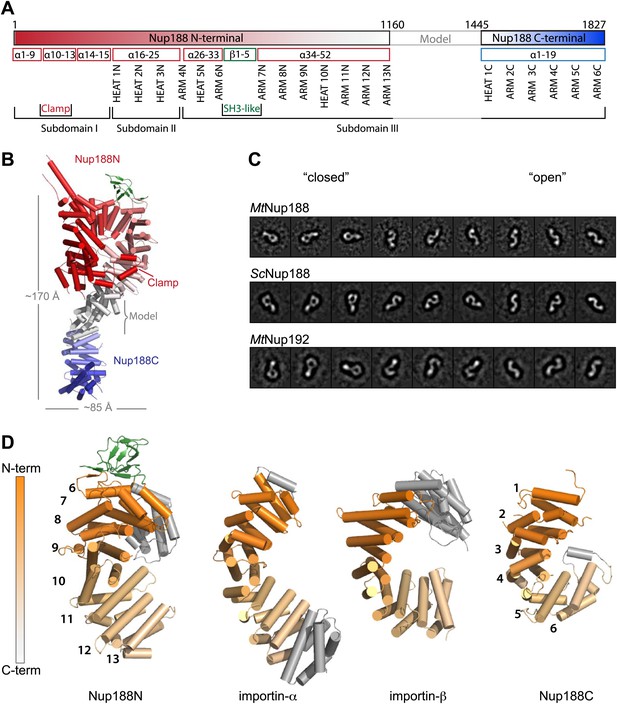
Structural details of Nup188 and comparison to other proteins.
(A) Schematic diagram of full-length Myceliophthora thermophila Nup188 with details about the subdomain arrangement for the entire protein. (B) Full-length Nup188 gradient-colored in red-gray-blue. The modeled region and the overall dimensions are indicated. (C). Single-particle negative stain EM class averages of full-length MtNup188, ScNup188 and MtNup192. Classes are roughly grouped and are ordered from more ‘closed’ (left) to ‘open’ conformations (right). The three proteins are evidently structurally related. (D) Structural similarity between Nup188 and importin-α and -β. The aligned areas of Nup188N, Nup188C, importin-α (3L3Q) and importin-β (1QGR) are gradient-colored in orange from N to C terminus. Non-aligned helices are in gray and the repeat numbering is indicated.
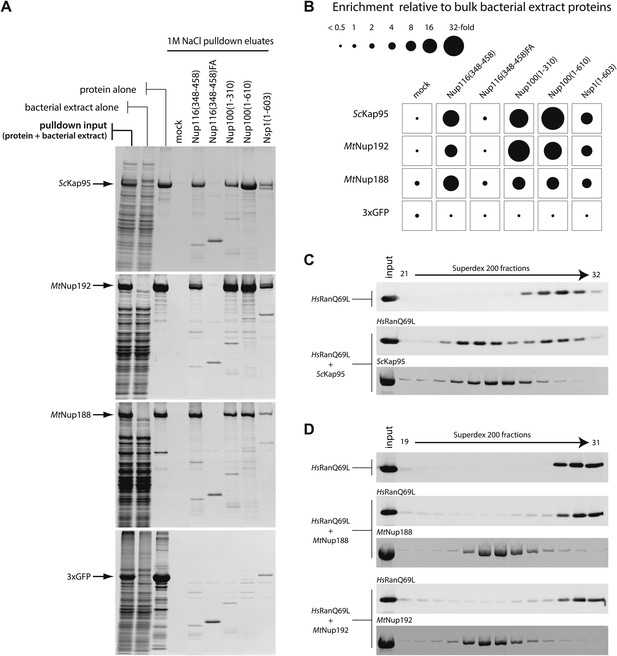
Biochemical properties of Nup188 and Nup192.
(A and B) Binding of MtNup188 and MtNup192 to various FG-coated beads monitored by in vitro pulldown assays. (A) Standard volumes of the soluble protein mixtures (pulldown input), separate components of the mixtures (protein alone, bacterial extract alone) and the 1M NaCl pulldown eluates form various FG-coated beads (1M NaCl pulldown elates) were precipitated by methanol-chloroform and re-solubilized in a standard volume of SDS-sample buffer. The samples were separated by SDS-PAGE and the gels were stained with SYPRO-Ruby. (B) The diagram representing relative enrichments of ScKap95, MtNup188, MtNup192 and 3xGFP in various FG-pulldown eluates, as compared to bulk bacterial extract proteins. The enrichment values (ratios between the eluted and input protein amounts) were computed using protein band intensities from the SYPRO Ruby stained gels shown in (A). (C) HsRanQ69L alone or an equimolar mixture of HsRanQ69L with ScKap95 were subjected to gel filtration using a Superdex 200 column, and the eluted fractions were separated by SDS-PAGE. Ran was visualized by Western blotting against the ZZ-tag, and ScKap95 was visualized by SYPRO Ruby staining. (D) Superdex 200 elution profiles of ZZ-HsRanQ69L (ZZ-tag Western blot) with or without the addition of MtNup188 or MtNup192 (SYPRO Ruby) analyzed as described in (C).
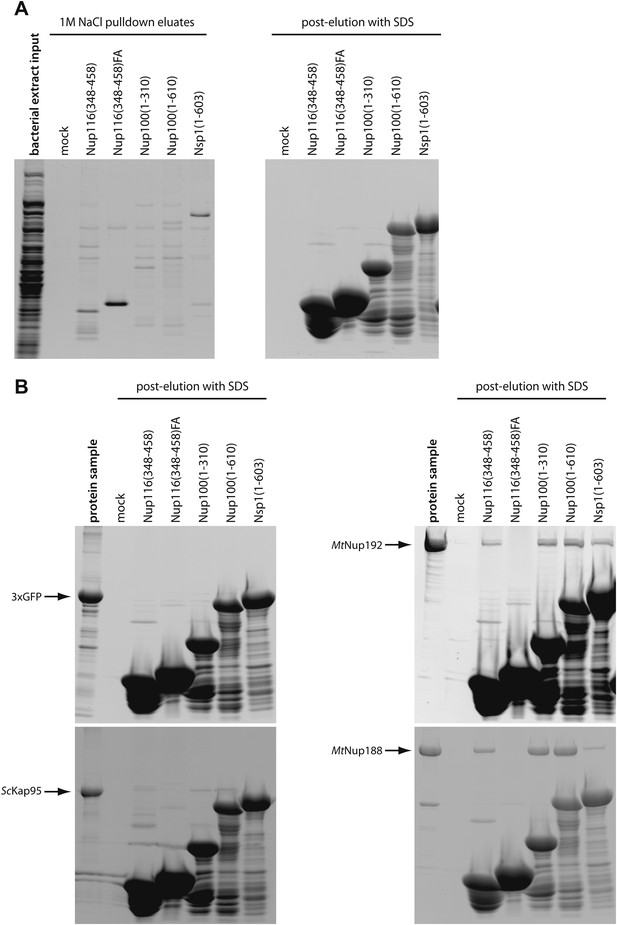
Characterization of the FG-repeat interactions in pulldown assays.
(A) The specificity of the FG-pulldown assay conditions were examined by subjecting bacterial extract proteins alone to pulldown assays with various FG- coated beads. Bound proteins were eluted with 1M NaCl sufficient to disrupt the expected transient interactions (left panel), followed by SDS sample buffer (right panel). The input bacterial extract sample and 1M NaCl eluates were precipitated by methanol-chloroform and re-solubilized in SDS-sample buffer prior to separation by SDS-PAGE. The gels were stained with SYPRO Ruby. (B) Proteins resistant to 1M NaCl elution in the experiment described in Figure 4A were stripped-off by SDS-sample buffer and separated by SDS-PAGE followed by SYPRO Ruby gel staining. The ‘protein sample’ lanes are the same as ‘protein alone’ in Figure 4A, and are included as references for the protein amounts remaining after 1 M NaCl elution.
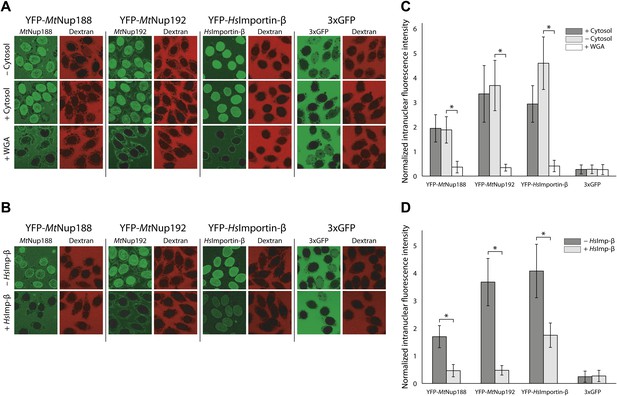
Nuclear translocation of Nup188 and Nup192 in digitonin-permeabilized HeLa cells.
(A and B) The nuclear translocation of YFP-MtNup188 and YFP-MtNup192 (green) was monitored in digitonin-permeabilized HeLa cells in transport buffer alone (−cytosol), in transport buffer containing Xenopus cytosol and an energy regenerating mix (+cytosol), or in transport buffer after pre-treatment of nuclei with WGA (+WGA) (A). Similar assays were performed in transport buffer with or without the addition of unlabeled HsImportin-β (B). 155kD TRITC-dextran (red) was used to check nuclear integrity. YFP-HsImportin-β and 3xGFP were used as positive and negative controls for facilitated NPC translocation, respectively. (C and D) Quantitative analysis of the protein translocation experiments described in (A and B), respectively. The intranuclear fluorescence intensities of the proteins were quantified and normalized against the background (extranuclear) fluorescence. Bar graphs represent mean values ± standard deviations. Asterisks (*) indicate a significant difference in median value using the Mann-Whitney U test (p<0.01; >50 nuclei for each condition).
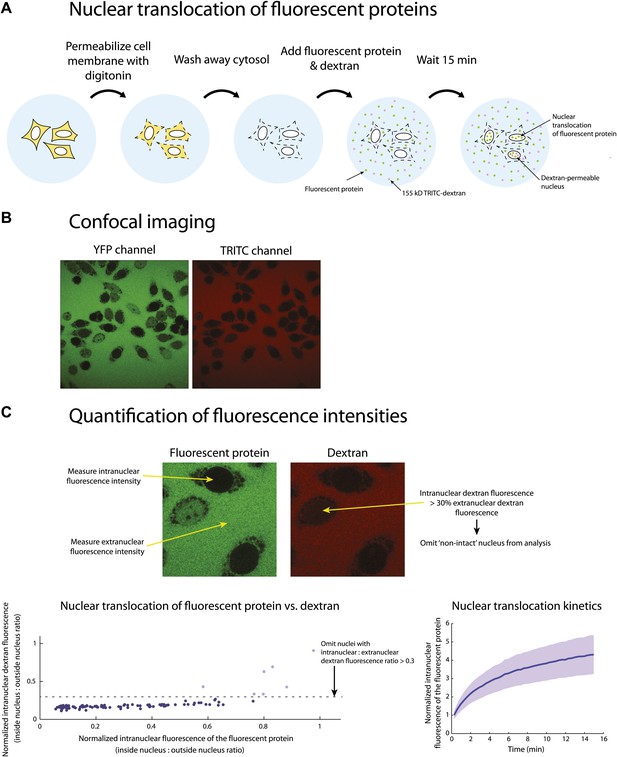
Outline of the nuclear translocation assay.
(A) Illustration of the protocol used for the nuclear translocation assays. (B) An example of two-channel confocal images used to simultaneously monitor the nuclear translocation of fluorescent protein fusions (YFP channel) and 155 kDa TRITC-dextran (TRITC channel). (C) Schematic representation of the procedure used to quantify the nuclear translocation of fluorescent protein fusions into intact nuclei. Fluorescence intensities inside and outside the nuclei were quantified using custom written code in MATLAB. The normalized intranuclear fluorescence intensity for each nucleus was calculated by taking the ratio of the fluorescence inside to outside the nucleus. All nuclei whose normalized TRITC fluorescence exceeded 0.3 were considered ‘non-intact’ and omitted from the analysis.
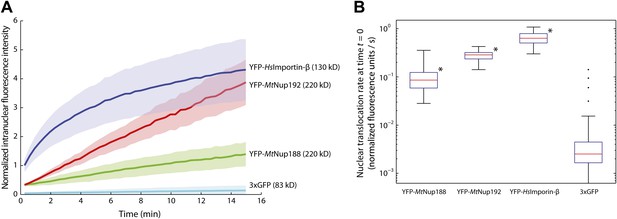
Kinetics of Nup188 and Nup192 nuclear translocation.
(A) The nuclear accumulation kinetics of the scaffold nucleoporins in transport buffer as compared to YFP-HsImportin-β and 3xGFP. Nuclear translocation reactions were imaged immediately after the addition of the fluorescent proteins and TRITC-dextran. Non-intact nuclei permeable to TRITC-dextran were excluded from the analysis. The graph represents mean values ± standard deviations. (B) Quantitative analysis of the initial nuclear translocation rates (equal to the total protein translocation rate through NPCs) for the experiments described in (A). The nuclear accumulation dataset for each nucleus (>30 for each condition) was separately fitted to a single-exponential curve, and the derivative at time t = 0 representing the initial nuclear translocation rate were computed. Asterisks (*) denote a significant difference in median value from the 3xGFP rate using the Mann-Whitney U test (p<0.01). Box plots show the median, first and third quartiles, and non-outlier extrema. Values greater than six standard deviations from the mean are marked as outliers.
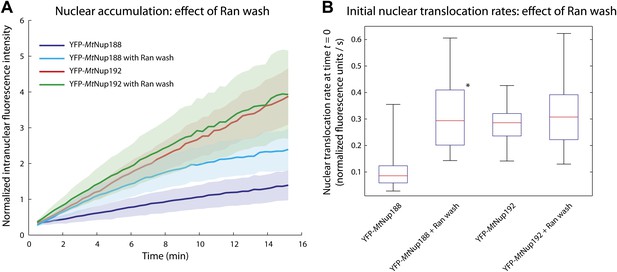
Effect of RanGTP on the nuclear translocation kinetics of Nup188 and Nup192.
(A) Nuclear translocation kinetics of the scaffold nucleoporins in permeabilized HeLa cells pre-incubated with Ran and an energy-regenerating system were obtained as described in Figure 6A. The kinetic plots for MtNup188 and MtNup192 from Figure 6A obtained without a Ran pre-wash are shown as a reference. (B) Quantitative analysis of the initial nuclear translocation rates for the experiments described in (A). The analysis was performed as described in Figure 6B. The asterisk (*) denotes a significant difference in median values in YFP-MtNup188 translocation rates between the no Ran pre-wash and Ran pre-wash conditions using the Mann-Whitney U test (p<0.01). Box plots show the median, first and third quartiles, and non-outlier extrema.
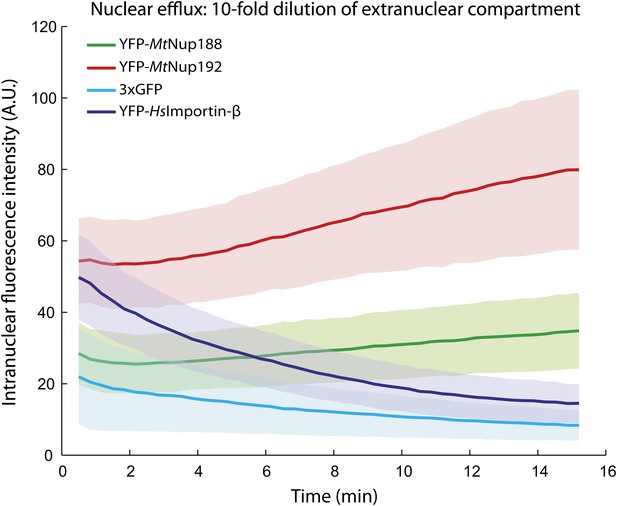
Nuclear accumulation of Nup188 and Nup192 upon 10-fold dilution.
YFP-MtNup188, YFP-MtNup192, YFP-HsImportin-β and 3xGFP were allowed to translocate into the nuclei of permeabilized HeLa cells for 15 min, and then their nuclear fluorescence was followed after a 10-fold dilution with transport buffer as described in Figure 6A. Note that unlike YFP-HsImportin-β and 3xGFP, YFP-MtNup188 and YFP-MtNup192 continue to accumulate even after the 10-fold dilution.
Tables
X-Ray data collection and refinement statistics
| MtNup188N (1–1160) | MtNup188N (1–1160) | MtNup188C (1445–1827) | |
|---|---|---|---|
| PDB code | 4KF7 | 4KF8 | |
| Data collection | |||
| Data set | Native | Selenomethionine | Selenomethionine |
| Space group | C2 | C2 | P212121 |
| a, b, c (Å) | 169.0, 94.8, 91.6 | 168.3, 94.6, 91.4 | 64.4, 66.7, 162.5 |
| α, β, γ (°) | 90, 98.9, 90 | 90, 98.4, 90 | 90, 90, 90 |
| Wavelength (Å) | 0.9792 | 0.9792 | 0.9792 |
| Resolution range (Å) | 66.8–2.65 (2.72–2.65) | 90.4–2.90 (3.02–2.90) | 81.1–3.00 (3.11–3.00) |
| Total reflections | 170,859 | 209,695 | 50,886 |
| Unique reflections | 41,673 | 58,249 | 14,539 |
| Completeness (%) | 100 (99.9) | 99.9 (98.8) | 99.9 (99.7) |
| Redundancy | 4.1 (3.8) | 3.6 (3.1) | 3.5 (3.1) |
| Rmerge(%) | 3.1 (77.2) | 3.0 (32.8) | 2.7 (56.1) |
| Rp.i.m.(%) | 2.0 (51.0) | 4.1 (26.3) | 1.6 (27.0) |
| I /σ(I) | 31.3 (1.8) | 9.6 (1.2) | 21.4 (4.8) |
| Wilson B factor (Å2) | 51.6 | 50.0 | 75.0 |
| Refinement | |||
| Resolution range (Å) | 66.8–2.65 | 81.1–3.00 | |
| Rwork (%) | 18.1 | 27.5 | |
| Rfree (%) | 22.5 | 29.4 | |
| Number of reflections | |||
| Total | 41,661 | 14,536 | |
| Rfree reflections | 1919 | 1139 | |
| Number of atoms | |||
| Protein | 8516 | 4366 | |
| Water | 128 | 0 | |
| B factors (Å2) | |||
| Protein | 64.4 | 92.3 | |
| Water | 44.9 | ||
| RMSD | |||
| Bond length (Å) | 0.004 | 0.003 | |
| Bond angles (°) | 0.904 | 0.769 | |
| Ramachandran plot | |||
| Favored (%) | 95.0 | 92.2 | |
| Allowed (%) | 4.1 | 6.8 | |
| Outliers (%) | 0.9 | 1.0 | |
-
The highest resolution shell is in parenthesis. Rmerge is the merging R factor. Rp.i.m. is the precision-indicating merging R factor. For definitions, see Weiss (2001).
Additional files
-
Supplementary file 1
List of constructs used in this study.
- https://doi.org/10.7554/eLife.00745.014





