Minor introns are embedded molecular switches regulated by highly unstable U6atac snRNA
Figures
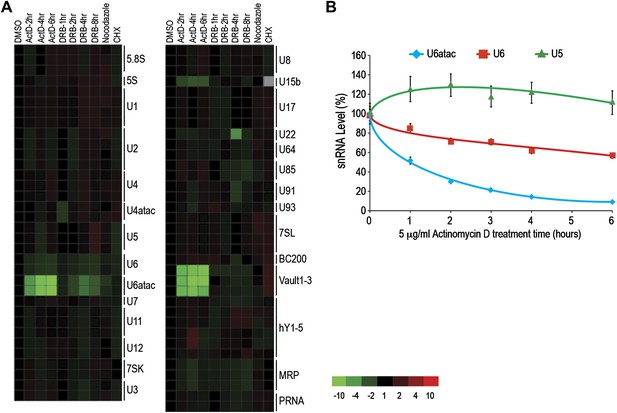
U6atac is an extremely unstable ncRNA.
(A) The heat map summarizes data for HeLa cells treated with inhibitors of several cellular processes, including the transcription inhibitors Actinomycin D (ActD) for 2–6 hr or DRB for 1–8 hr, the cell cycle inhibitor nocodazole for 18 hr, and the translation inhibitor cycloheximide (CHX) for 16 hr. A color scale bar for heat maps (−10 to +10 fold change) is indicated. (B) Real time qPCR verification of U5, U6 and U6atac snRNA level after ActD treatment. Absolute snRNA levels were measured in triplicates and normalized to 5S rRNA. Error bars represent standard deviations of three replicates.
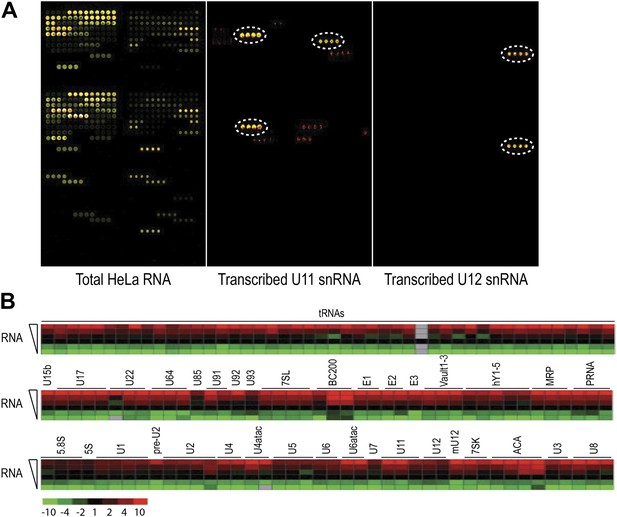
Hybridization of total RNA and in vitro transcribed RNA standards.
Hybridization of total RNA and in vitro transcribed RNA standards. (A) Representative microarray image of total RNA from HeLa cells, in vitro transcribed U11 and U12 snRNA. Total RNA or in vitro transcribed snRNAs of the same RNA sample, labeled with Alexa647 (red) and Alexa546 (green), were hybridized at 48°C. Yellow signifies an overlap of green and red fluorescence. Dotted circles indicate the probe locations for U11 and U12 snRNAs. (B) Total RNA labeled with Alexa647 was hybridized at six different amounts: 50, 100, 250, 500, 1000 and 2500 ng. The heat maps are shown for all probes on the array with signals twofold above background. The 250 ng sample signals were set at 1 (black) while the range covered −10 fold (green) to +10 fold (red).
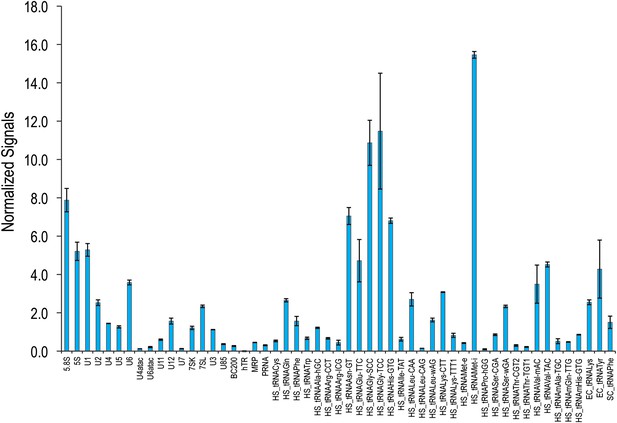
ncRNA microarray is highly reproducible.
Total RNA from three HeLa cell cultures were labeled and hybridized as described in ‘Materials and methods’. The average normalized signals from the three experiments were plotted along with their standard deviations that are presented as error bars.
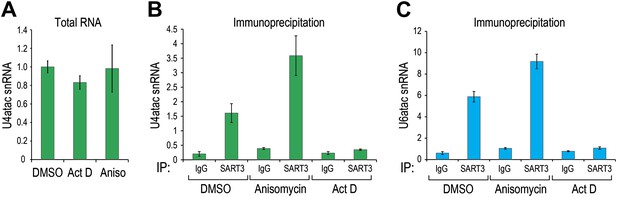
U6atac, not U4atac, is rate limiting for di-snRNP association.
(A) U4atac level was measured by real time PCR (absolute quantification and normalized to 5S) in total RNA extracted from cells treated with actinomycin D or anisomycin for 4 hr. (B) U4atac and (C) U6atac association with di-snRNP specific protein p110/SART3 was measured after immunoprecipitation of SART3 or control IgG from cells treated as in (A).
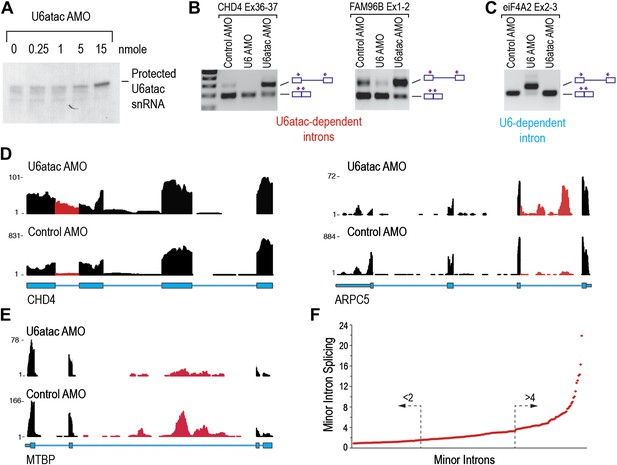
U6atac inactivation shows a wide range of minor intron splicing dependence on U6atac level.
(A) RNase H protection of U6atac snRNA in total cellular RNA after 8 hr treatment with U6atac AMO showing efficient U6atac protection at 15 nmole U6atac AMO. The same RNA sample (15 nmole U6atac AMO) was used for RT-PCR of representative minor (B) and major (C) introns. The diagrams to the right of the gel indicate spliced or unspliced RNA. Arrows show position of the primers used. (D) Representative view on UCSC genome browser for two genes containing highly efficient minor introns (red reads). Numbers on the left represent peak reads count. Gene structures are depicted in blue boxes (exons) and lines (introns). (E) Same as in (D) for low efficiency minor intron. (F) The splicing index is plotted for each expressed minor intron in HeLa cells. The dashed arrows point to minor introns with the lowest (<twofold) and highest (>fourfold) splicing indices.
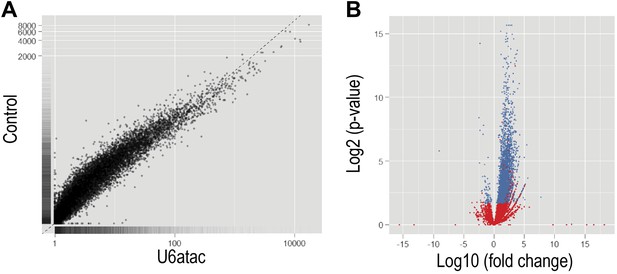
Functional knockdown of U6atac affects a large number of genes.
(A) Scatter plot comparing the FPKM values representing differential expression analysis from Cufflinks between U6atac AMO and Control AMO. (B) Volcano plot of fold change vs significance for genes in U6atac AMO and Control AMO samples. Red dots indicate significant genes (p<0.05), whereas blue dots show genes having no changes.
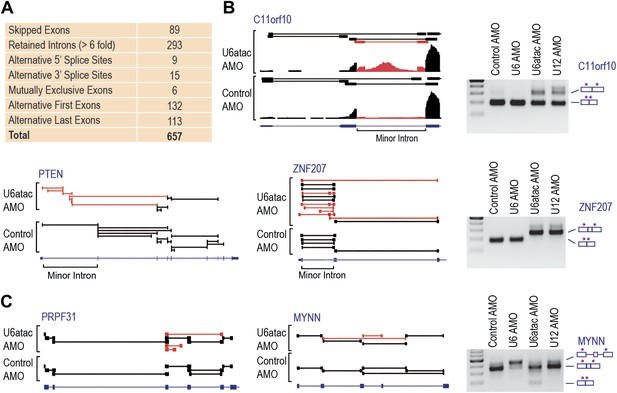
Genome-wide analysis of the minor splicing pathway shows widespread transcriptome changes.
(A) A summary of the splicing effects in the U6atac AMO treated cells identified using MISO. (B) Representative minor intron-containing genes showing alternative splicing pattern changes after U6atac knockdown. Exon junction reads are presented on top. Red junction reads are unique to U6atac AMO sample. RT-PCR confirmations for U6atac AMO as well as U6 and U12 knockdown for comparison are also shown. Arrows show position of the primers used, which are located in the exons. (C) Representative genes that do not contain a minor intron showing alternative splicing pattern changes after U6atac inactivation.
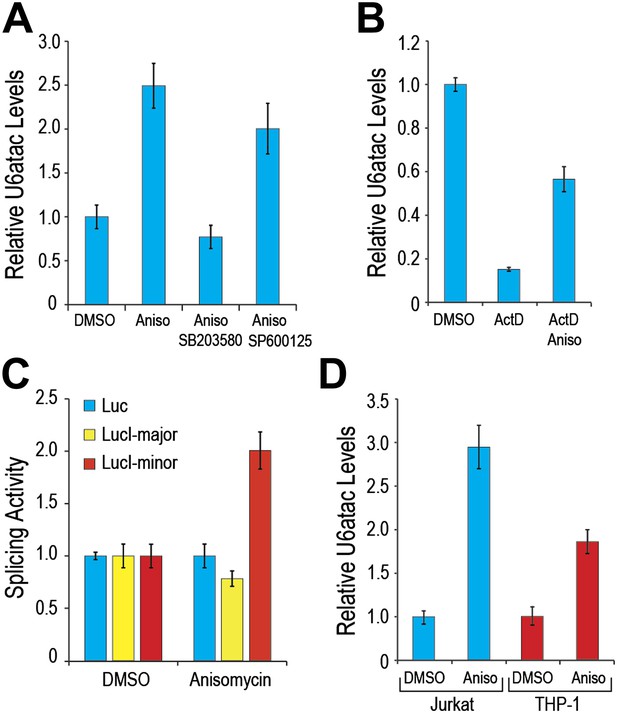
p38MAPK cell signaling up-regulates U6atac level and minor intron splicing.
(A) Real time qPCR of U6atac snRNA after treatment with 1 μg/ml anisomycin (4 hr), a potent p38MAPK activator, in HeLa cells. Cells were treated with 10 μM of p38MAPK inhibitor, SB203580, or JNK inhibitor, SP600125, 30 min prior to anisomycin. (B) HeLa cells were treated with 5 μg/ml Actinomycin D (ActD) or ActD plus 1 μg/ml anisomycin (Aniso) or DMSO as control for 4 hr, followed by measurement of U6atac level. (C) Luciferase activity of the various splicing reporters after cell treatment with DMSO or anisomycin for 4 hr. (D) Real time qPCR of U6atac snRNA after treatment with anisomycin (4 hr) in Jurkat T cells and THP-1 monocytes. Error bars represent the standard deviation of at least three replicates.
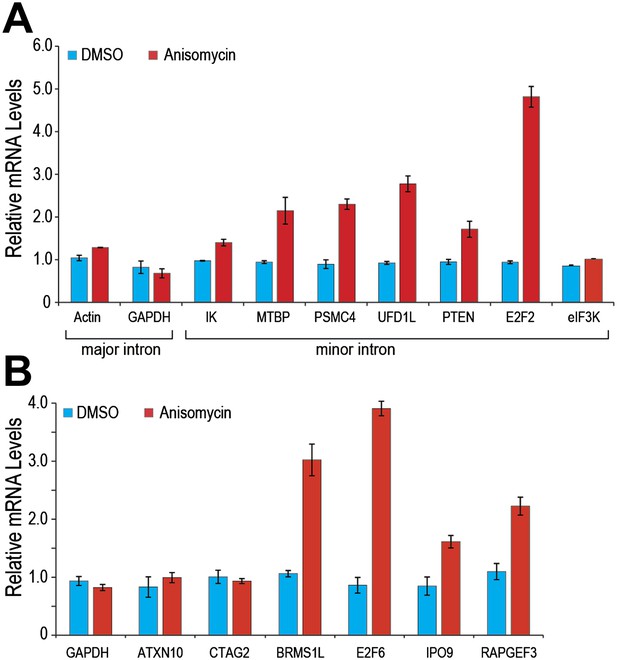
p38MAPK activation increases expression of minor intron-containing genes.
(A) Real time qPCR for a representative set of minor introns with low splicing indices with or without 1 μg/ml anisomycin treatment. All primer sets span the splice junction and thus measure spliced mRNAs. (B) Real time qPCR for representative minor intron-containing genes that are normally suppressed in HeLa cells. Error bars represent the standard deviation of three replicates.
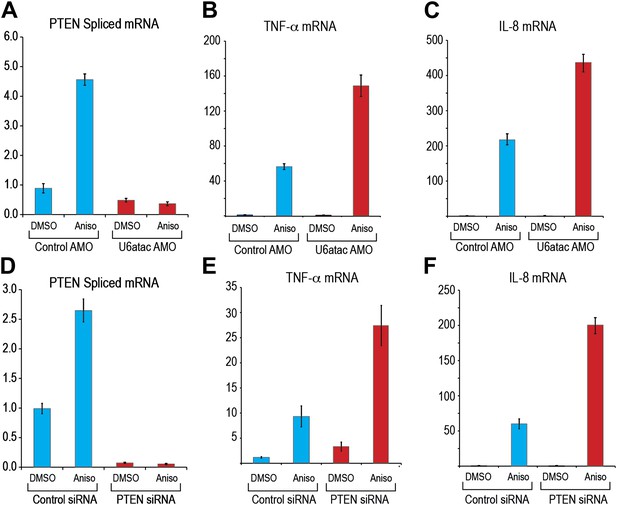
U6atac level regulates PTEN minor intron splicing and buffers cytokine production in response to p38MAPK activation.
(A) Real time qPCR for PTEN in HeLa cells transfected with control or U6atac AMO followed by treatment with DMSO or 1 μg/ml anisomycin for 4 hr. (B) and (C) Real time qPCR for TNF-α and IL-8 after 4 hr of anisomycin treatment in HeLa cells. (D) Real time qPCR for PTEN in HeLa cells transfected with control or PTEN siRNA followed by treatment with DMSO or anisomycin for 4 hr. (E) and (F) Real time qPCR for TNF-α and IL-8 after 4 hr of anisomycin treatment in HeLa cells transfected with control or PTEN siRNA for 48 hr. Error bars represent the standard deviation of three replicates.
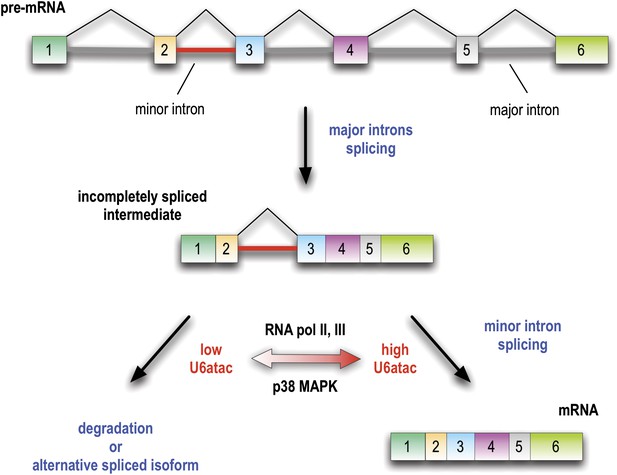
Minor introns are embedded molecular switches regulated by U6atac abundance.
Splicing of minor introns, typically one amidst several major introns, is dependent on the limiting level of U6atac snRNP in cells. Low abundance and fast turnover of U6atac results in incompletely spliced pre-mRNAs that are either degraded or switch into spliced isoforms that do not require the minor spliceosome. While transcription attenuation rapidly lowers U6atac level and limits minor intron-containing gene’s expression, activated p38MAPK rapidly stabilizes U6atac, increasing its level and enhancing minor splicing and production of full-length mRNAs.
Tables
Summary of the RNA-seq statistics
| Control AMO | U6atac AMO | |
|---|---|---|
| Platform | Illumina HiSeq2000 | Illumina HiSeq2000 |
| Read length | 100 base | 100 base |
| Single/paired ends | single end | single end |
| Total number of reads | 120,274,499 | 121,794,728 |
| Mapped reads | 81,198,953 | 85,645,962 |
| Expressed genes (FPKM > 1) | 9799 | 8397 |
| Upregulated genes (> fourfold) | ND | 3 |
| Downregulated genes (< fourfold) | ND | 2088 |
| Expressed U12-dependent genes (FPKM > 1) | 429 | 350 |
Additional files
-
Supplementary file 1
(A) Noncoding RNAs represented on the microarray. (B) Minor intron-containing genes with lowest and highest splicing indices and their function.
- https://doi.org/10.7554/eLife.00780.015





