In situ structural analysis of the Yersinia enterocolitica injectisome
Figures
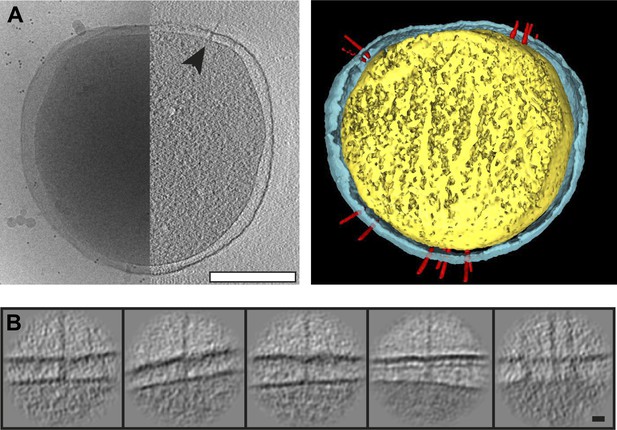
Visualization of Y. enterocolitica injectisomes in situ.
(A) Left: cryo-EM image of a Yersinia enterocolitica bacteria (left half of the panel) and a 20-nm thick slice through a tomogram of the bacteria, showing an injectisome (right half of the panel, black arrowhead); Right: volume rendering of the same Y. enterocolitica bacteria showing the inner (yellow) and outer (blue) membranes, and injectisomes (red). (B) Example images of individual injectisomes, illustrating different types of observed injectisomes. Left to right: regular, tilted, with dim basal body, denser peptidoglycan layer, and clustered injectisomes. Scale bars: A: 300 nm, B: 20 nm.
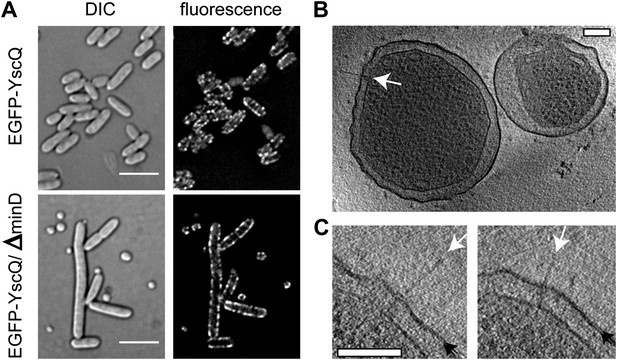
Y. enterocolitica minicells.
(A) Differential interference contrast (DIC) (left) and fluorescence (right) imaging of regular (top) and minD mutant (bottom) Y. enterocolitica cells. Minicells appear together with extra-long bacteria. For the fluorescent microscopy, EGFP was coupled to YscQ. Scale bar: 5 μm. (B) A 22-nm thick slice through a tomogram of two minicells showing an injectisome (white arrow). Scale bar: 100 nm. (C) Injectisomes from minicells showing the outer membrane as resolved bilayer (double black arrows) and the injectisome needle (white arrows). Scale bar: 100 nm.
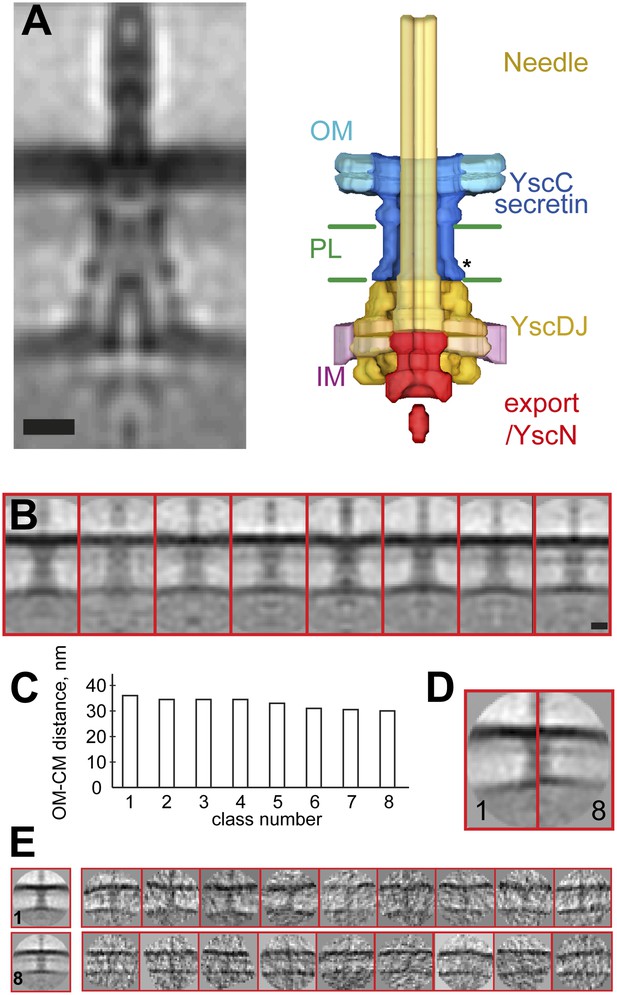
Structure of the Y. enterocolitica injectisome in situ.
(A) Slice through the average 3D structure of the injectisome and a model with indicated components. OM–outer membrane, PL–peptidoglycan layers, IM–inner membrane, * indicates the junction between YscC and YscD. (B) 8 class averages of injectisomes from wt cells obtained by MRA classification; their length varies significantly. (C) Intermembrane distances for the corresponding class averages from (B), the longest class has a distance of 36 nm, the shortest of 30 nm. (D) Overlay of the longest and the shortest class aligned by the OM. (E) Class averages 1 (longest) and 8 (shortest), together with representative individual injectisomes for the two classes, all at the same scale. Scale bars: 10 nm; box heights for B,D,E: 96 nm.
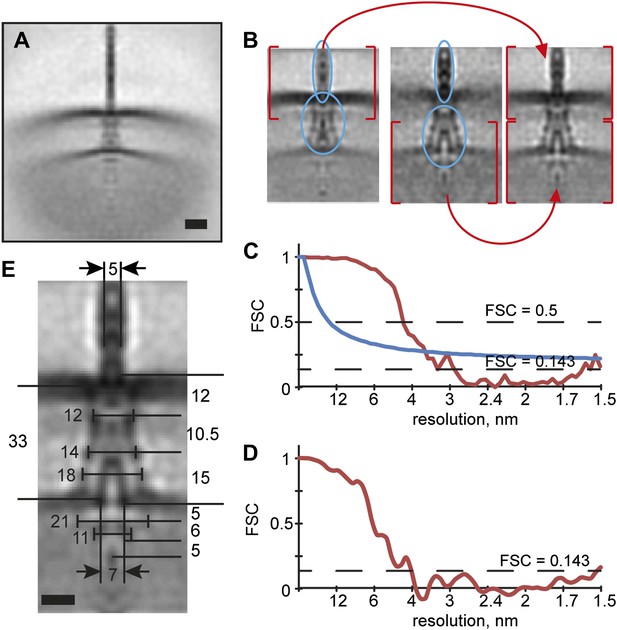
Structural elasticity of the injectisome.
(A) Initial alignment of the injectisomes to the common origin. Scale bar: 20 nm. (B) A composite average is produced by merging two independent alignments focusing on regions at the outer membrane (left) respectively the inner membrane (middle). Selected low-defocus and minicell particles were processed together. Height of the box: 96 nm. (C) Fourier shell correlation (FSC) plot between half-populations of the aligned particles. This curve is an average of the FSC curves of the alignments for the two masks from (B) (red line), indicating a resolution of 3.7 nm, using the 1 bit information threshold (blue line) (van Heel and Schatz, 2005). (D) FSC between two structures derived from independent processing of two sub-sets of the particles (‘gold standard’ method, ‘Materials and methods’), indicating a resolution of 4 nm (E) Average structure of the injectisome; the dimensions of the various regions are indicated. The large ring-like structure on the cytoplasmic side of the IM is 21 nm wide. The smaller, ring-like structure proposed to be the export apparatus, is 11 nm wide and ∼6 nm high. Scale bar: 10 nm.
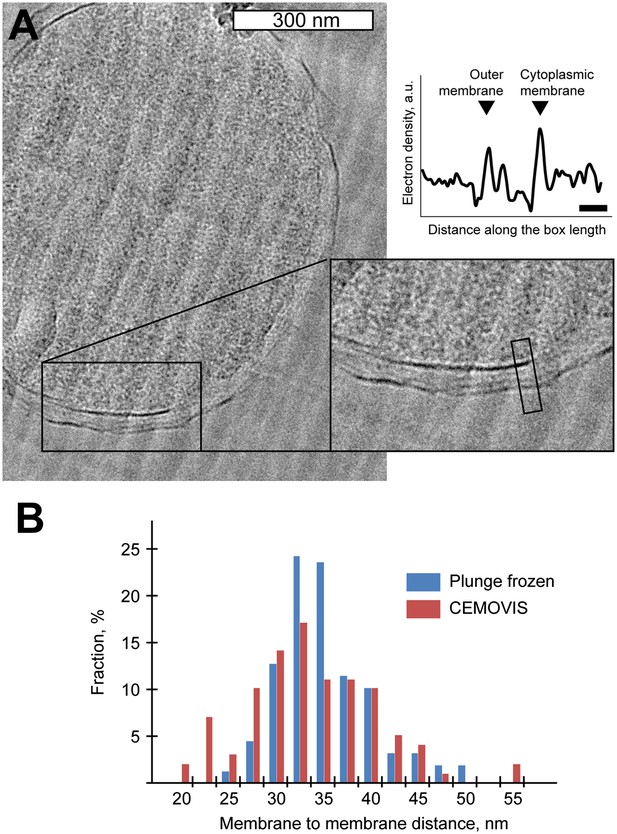
Comparison of membrane-to-membrane distance using CEMOVIS and cryo-ET of plunge frozen Y. enterocolitica.
(A) A micrograph from cryo-electron microscopy of vitrified sections (CEMOVIS) of high-pressure frozen bulk solution of Y. enterocolitica bacteria, showing the distance between the membranes. Inset: 2× zoomed area indicated in the main panel. The graph on the right shows the laterally projected electron density along the black box in the inset. Scale bar in the micrograph: 300 nm, in the graph: 20 nm. (B) Distribution of center-to-center distances between the outer and inner membranes for plunge-frozen Y. enterocolitica in the area close to the injectisomes (blue) and in CEMOVIS images in arbitrarily chosen positions (red). Numbers of measurements: 154 and 97 correspondingly.
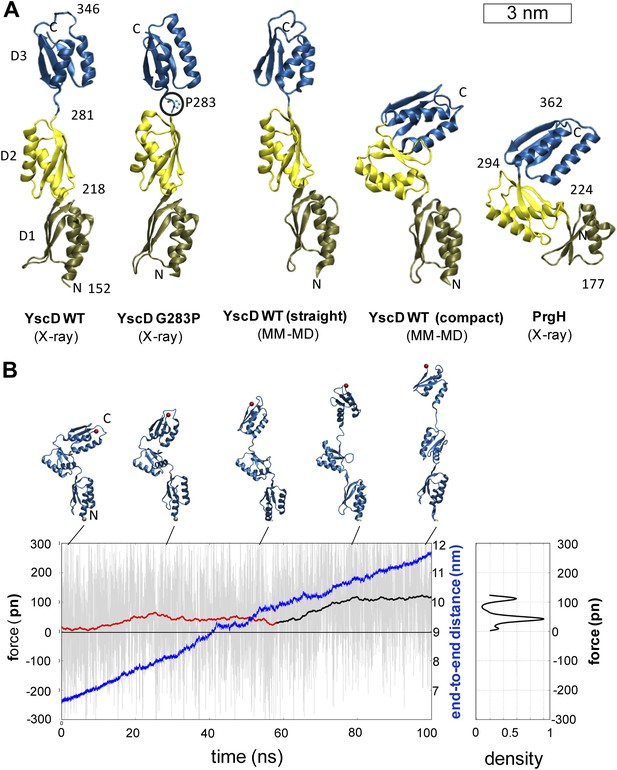
Structural elasticity of YscD.
(A) Comparison of different conformers of YscD with the structure of PrgH (Spreter et al., 2009); from left to right: X-ray structures of wild type and G283P mutant (mutation highlighted by a black circle) of YscD, two representative conformations of the elongated and contracted YscD monomer obtained from MD simulations, and X-ray structure of PrgH. (B) Force-extension profile from steered MD simulations stretching an YscD monomer; raw data obtained using a pulling velocity v = 0.05 nm/ns are reported (light gray) together with the running average (red and black for extension and unfolding, respectively); end-to-end distance is reported in blue. The inset on the right reports the normalized kernel density profile of the forces: the two distinct peaks correspond, respectively, to the barrier-free extension from the compact to the elongated form, and to the beginning of the unfolding of YscD’s third domain.
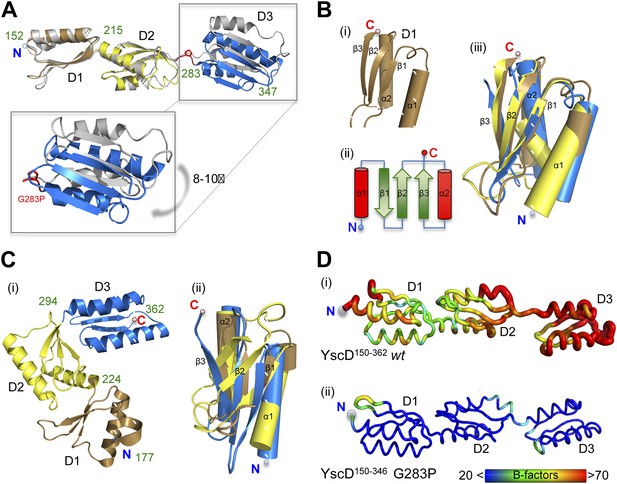
Comparison of YscD150–362 wt, YscD150–347 G283P, and PrgH crystal structures.
(A) Superposition of YscD150–362 (grey) wt and YscD150–347 G283P (periplasmic domain D1, D2 and D3 are colored in brown, yellow and blue, respectively). Domain boundaries are labeled with green numbers. Mutation G283P is labeled in red. Periplasmic domain 3 (D3) of YscD150–347 G283P is shifted by 8–10° in reference to wt D3 (see close up). (B) Secondary structure composition of the periplasmic domain 1–3 of YscD. Each domain comprises the αββαβ-ring building motif (i, ii) (Lilic et al., 2010). All three domains superpose well (iii). Coloring for each domain D1–D3 was preserved as in (A) for YscD150–347 G283P. (C) PrgH (S. typhimurium) domain arrangement (PDB code: 3GR0). (i) Domains 1–3 (D1–D3) are colored and labeled as in (A). (ii) Although the three PrgH domains have the same αββαβ-ring building motif, their superposition is poor. (D) Comparison of dynamic mobility of YscD150–362 (i) wt and YscD150–347 G283P (ii) in the crystal visualized with a B-factor putty representation in rainbow colors. Thin blue tubes represent low B-values, while broad red tubes represent high B-values (see also scale bar).
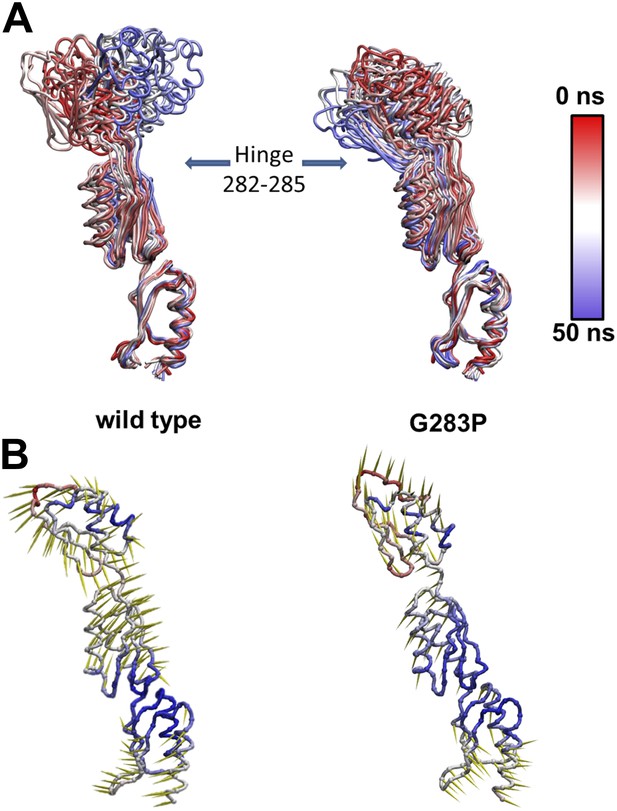
Effect of G238P mutation on YscD elasticity.
(A) Collection of snapshots taken from 50 ns long molecular dynamics simulation of wild type (left) and G238P mutant (right) YscD systems. Color is used to indicate the position of the selected snapshot along the trajectory (red: first, blue: last). Structures are aligned on the first domain. (B) Porcupine plot of the first normal mode, calculated applying a standard normal mode analysis (NMA) on the molecular dynamics trajectories of wild type (left) and G238P mutant (right) YscD systems. The length of arrows (yellow) is proportional to the contribution of each residue to the overall motion for the first normal mode. Color is used to indicate fluctuation of each Cα atom along dynamics (red: high fluctuation, blue: low fluctuation)
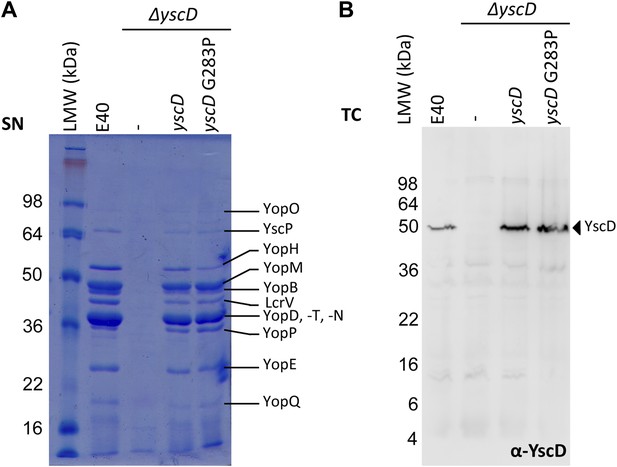
Mutation G283P in YscD has no impact on type III secretion compared to E40 (WT) and in trans complemented YscD.
(A) SDS-PAGE analysis of Yop secretion pattern and (B) immunoblot analysis (α-YscD) of total cells in secretion-permissive conditions. Culture supernatant (SN) and total cells (TC) were separated on a 12% and 15% SDS-PAGE gel, respectively. Strains and plasmids used for the secretion assay (Supplementary file 1B, C): E40 (WT); ΔyscD (AD4051), non-complemented or complemented in trans with yscD (from pMA11), or with yscD G283P (from pUW22). Both constructs fully complement effector secretion in the ΔyscD strain.
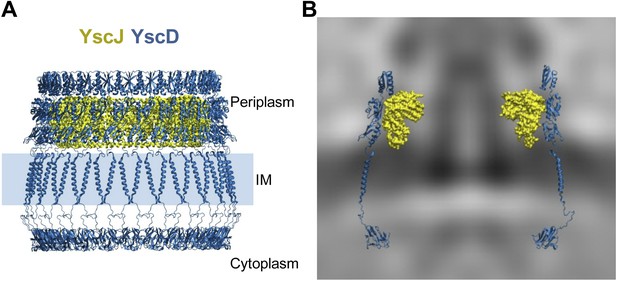
Structural assembly of YscDJ ring at the basal body.
(A) Side view of the generated 24-mer ring model of YscD (blue) and YscJ (yellow). Each YscD subunit has been extended by the transmembrane helix segment and the N-term cytosolic domain. The position of the IM is indicated by a blue area and is positioned according to the IM observed in cryo-ET on the right (B). (B) Overlay of the injectisome cryo-ET map with the atomistic YscDJ ring model from (A). Only two YscDJ units have been superimposed on the cryo-ET map section for clarity.
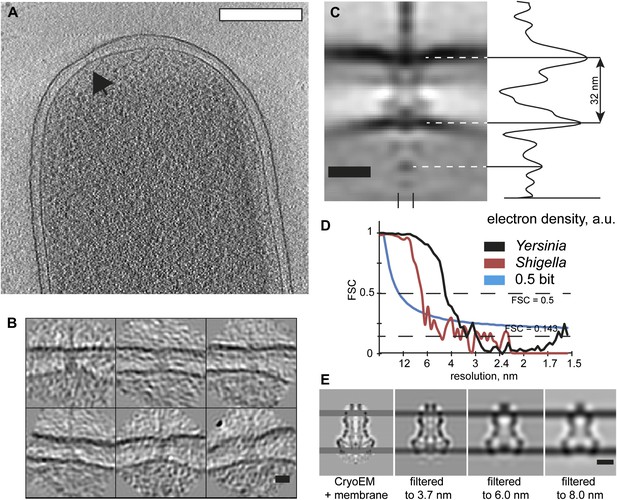
Visualization and structure of the Shigella flexneri injectisomes in situ.
(A) 30-nm thick section through a tomogram of an S. flexneri cell. Arrow points to the basal body of an injectisome. Scale bar: 300 nm. (B) Typical views of S. flexneri injectisomes oriented vertically. Scale bar: 20 nm. (C) Left: average structure of the S. flexneri injectisome in situ. Scale bar: 20 nm; right: electron density along the 8 nm profile indicated with two dashes (bottom). (D) Right: comparison of Fourier shell correlation for S. flexneri (red) and Y. enterocolitica (black); Blue line: 0.5 bit information threshold. Resolution of S. flexneri is 7 nm, Y. enterocolitica is 4 nm (0.5 criterion). (E) Resolution limitation applied to the single particle cryo-EM map (EMD 1871) placed between two added membrane densities. A lower resolution does not affect the visible inter-membrane distance. Scale bar: 10 nm.
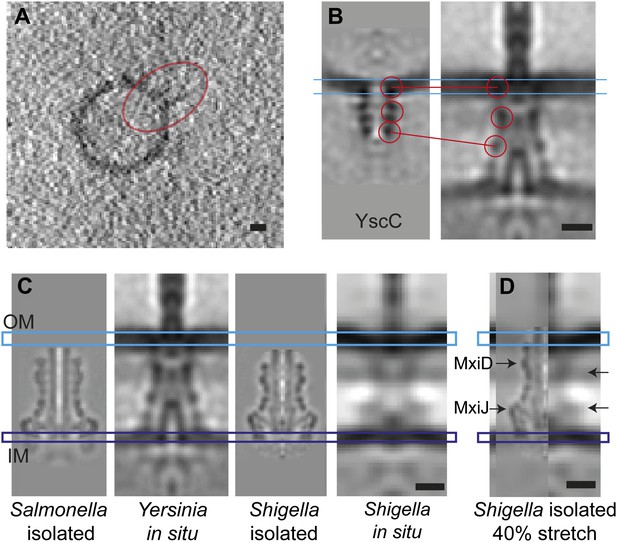
Elongation of the in situ structure over the isolated versions.
(A) 8-nm thick section through a single tomogram of an YscC multimer reconstituted into a lipid vesicle. (B) Average structure of liposome-reconstituted YscC (left), and matching densities in the Y. enterocolitica injectisome (right). (C) Comparison of the Y. enterocolitica and S.flexneri in situ injectisomes with high-resolution single particle structures of Salmonella enterica SPI-1 (EM Data Bank entry EMD 1617), and Shigella flexneri (EMD 1871). Blue and purple bars indicate the outer (OM) and inner (IM) membranes. (D) The 40% stretched structure of the isolated S. flexneri injectisome (right) overlaid onto the S. flexneri injectisome in situ (left). Arrows indicate positions of recognizable densities of MxiDJ in the in situ and the stretched-isolated structures. Scale bars: 10 nm.
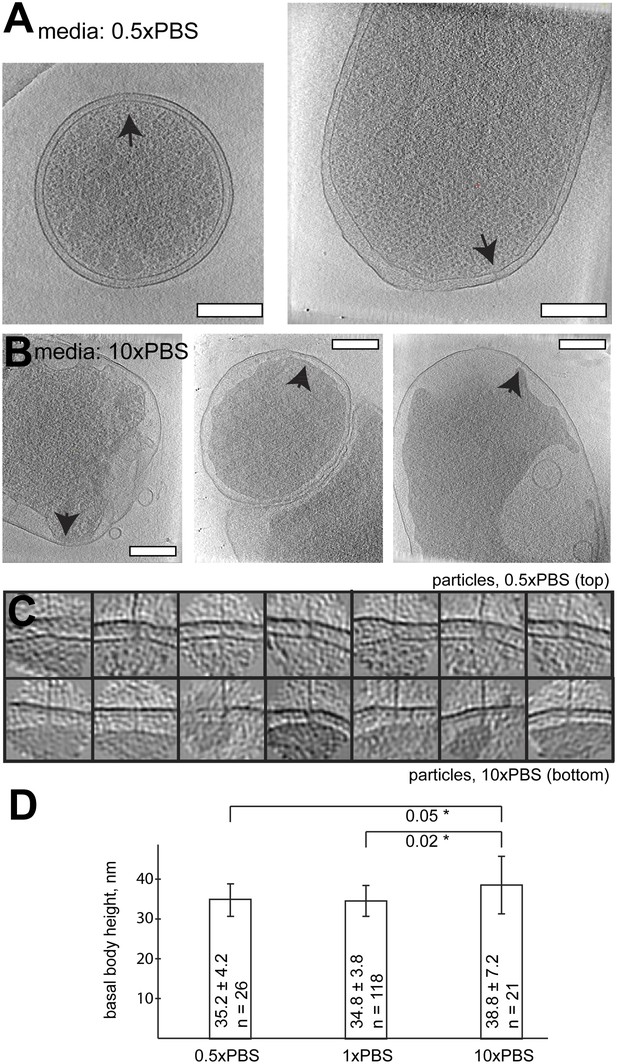
Length of basal bodies of injectisomes from Y. enterocolitica exposed to media with different osmolarities.
(A, B) Slices though tomograms of bacteria placed into 0.5× PBS diluted with H20 1:1 (A) and into concentrated 10× PBS (B). Black arrowheads point to injectisomes. Scale bar in the micrograph: 300 nm. (C) 15-nm thick sections though sub-tomograms with roughly vertically aligned particles from 10× PBS media (top) and 0.5× PBS (bottom). Box size: 192 nm. (D) Average membrane-to-membrane distances around basal bodies in nanometers with significance tests. Differences between 10× PBS and 0.5× PBS and between 10× PBS and PBS are significant (p=0.05 and p=0.02), while the difference between PBS and 0.5× PBS is not significant (p=0.6).
Additional files
-
Supplementary file 1
- https://doi.org/10.7554/eLife.00792.016





