Patient-specific iPSC-derived photoreceptor precursor cells as a means to investigate retinitis pigmentosa
Figures
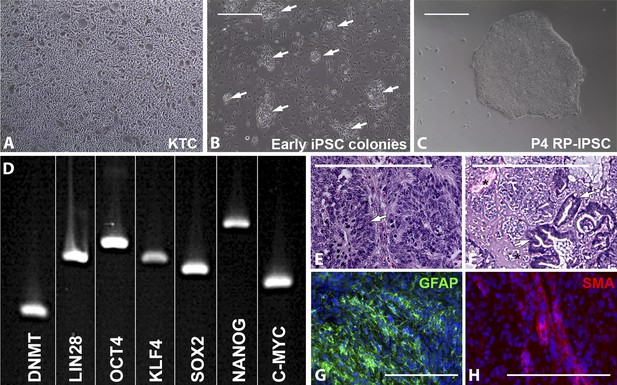
Derivation of iPSCs from keratinocytes of a patient affected with USH2A-associated RP.
(A–H) Microscopic analysis of human keratinocytes (A), early keratinocyte-derived iPSC colonies (B, arrows), and purified keratinocyte-derived iPSC cultures (C). At 2–3 weeks post-viral transduction, ES-cell-like iPSC colonies begin to emerge (B, arrows). iPSC colonies isolated, subcultured, and expanded on Synthemax cell culture surfaces maintain a pluripotent morphology (C) express the pluripotency markers DNMT, LIN28, OCT4, KLF4, SOX2, NANOG, and C-MYC (D), and form teratomas consisting of tissues of ectoderm (E, arrow and G, GFAP in green), mesoderm (F, asterisk and H, SMA in red), and endoderm (F, arrows) each of the three embryonic germ layers (E–H). Scale bar = 400 μm.
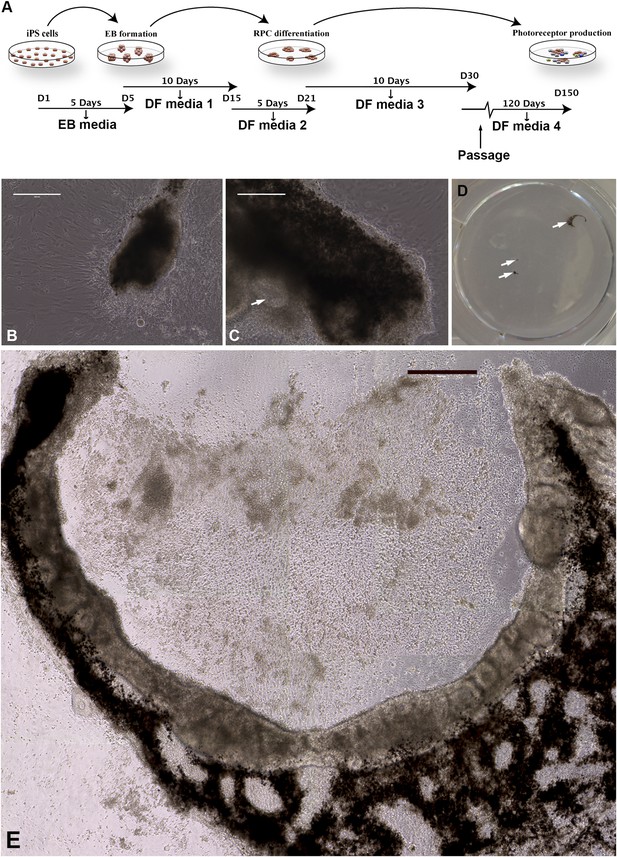
Differentiation of human USH2A-associated iPSCs into eyecup-like structures.
(A) Schematic diagram illustrating the differentiation paradigm utilized to generate human eyecup-like structures. (B–E) Morphological analysis of USH2A-associated iPSC-derived eyecups. iPSC-derived eyecups form pigmented cell clumps (B) that extend and wrap in a C shape around newly formed neural rossettes (C). Following this protocol, a typical six-well cell culture dish will have two to four eyecups/well, each at slightly different stages of development (D, arrows, a low magnification image of a typical well of a six-well plate with developing eyecups). At 150 days post-differentiation, complete eyecups with clearly defined neural retina and RPE layers can be identified (D, top right arrow, and E). Scale bar, B and C = 200 μm, D = 400 μm.
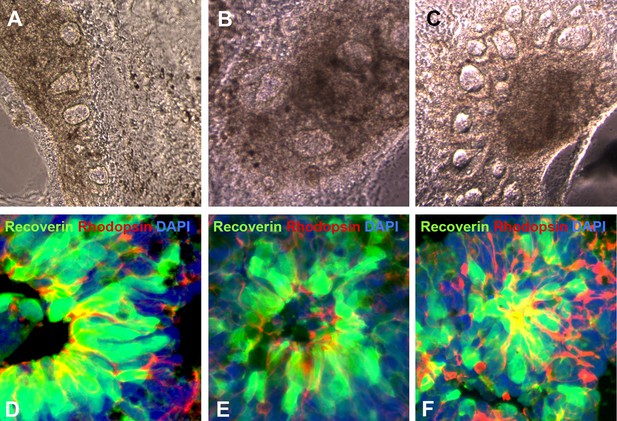
Additional examples of human USH2A-associated iPSC-derived eyecup-like structures.
(A–C) At 150 days post-differentiation, extensive neural rosette formation was present. (D–F) Immunocytochemical analysis targeted against recoverin and rhodopsin confirms that neural rosettes consisted predominantly of rod photoreceptor precursor cells.
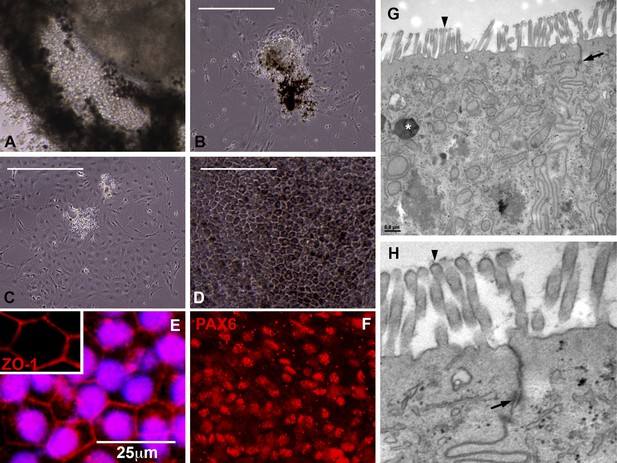
Cells contained within the pigmented layer of USH2A-associated eyecups are of RPE origin.
(A) A high magnification phase image of the RPE layer of an USH2A eyecup prior to biopsy and subculture. (B–D) Area of the RPE presented in panel A was picked and subcultured in fresh RPE culture media on collagen, laminin, and fibronectin coated six-well culture dishes. 24 hr after plating, RPE cells spread and take on a fibroblastic morphology (B). By 72 hr post-plating, RPE cells lose their pigmentation and begin to form cell–cell contacts (C). 2 weeks post-plating, a confluent monolayer of RPE cells are present that have taken on the typical cuboidal RPE morphology and regained pigmentation (D). (E–F) Immunocytochemical analysis of USH2A-associated RPE cells with antibodies targeted against the tight junction marker ZO1 (E) and the transcription factor PAX6 (F). (G–H) TEM analysis of RPE cells within the intact RPE layer of USH2A eyecups (H is a high magnification view of the upper right corner of panel G). RPE cells are polarized, have apical microvilli, make tight junctions with neighboring RPE cells (G and H, arrows) and contain pigment granules within their cytoplasm (G, asterisk). Scale bar, B–D = 200 μm, G = 0.5 μm.
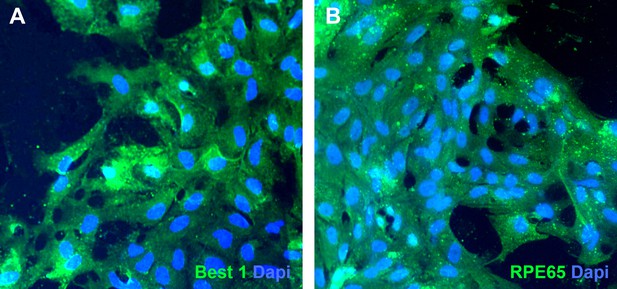
Pigmented cells isolated from USH2A-associated eyecups express bestrophin 1 and RPE65.
(A and B) Immunocytochemical analysis of subcultured USH2A-associated RPE cells with antibodies targeted against bestrophin 1 (A) and RPE65 (B).
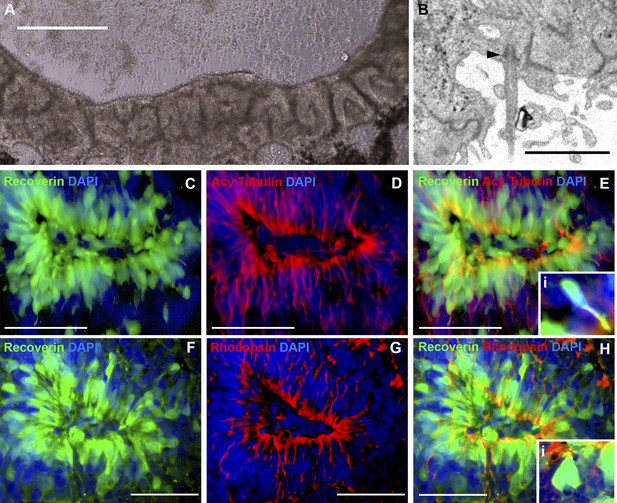
iPSC-derived USH2A-associated neural retinal rosettes consist predominantly of rod photoreceptor cells.
(A) Morphological depiction of the neural retina at 120 days post-differentiation. (B) TEM analysis of neural rosettes demonstrates the existence of cilia with clearly identifiable basal bodies. (C–H) Immunocytochemical analysis targeted against the rod photoreceptor markers recoverin and rhodopsin (C–E and Ei-high magnification inlay), and the rod photoreceptor marker recoverin and the connecting cilia marker acetylated tubulin (F–H and Hi-high magnification inlay).
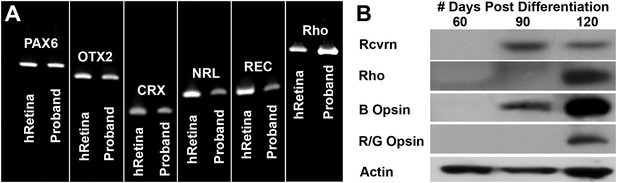
Developmental timeline of neural retina marker expression.
(A) RT-PCR analysis of USH2A and human control neural retina for expression of the retinal transcription factors/photoreceptor markers PAX6, OTX2, CRX, NRL, recoverin, and rhodopsin at 60 days post-differentiation. (B) Western blot analysis of USH2A neural retina for expression of the retinal photoreceptor markers recoverin, rhodopsin, blue cone opsin and red/green cone opsin at 60, 90, and 120 days post-differentiation. Although retinal transcripts can be detected as early as 60 days post-differentiation, mature photoreceptor proteins such as recoverin, rhodopsin, and the cone opsins could not be detected until 90 to 120 days post-differentiation.
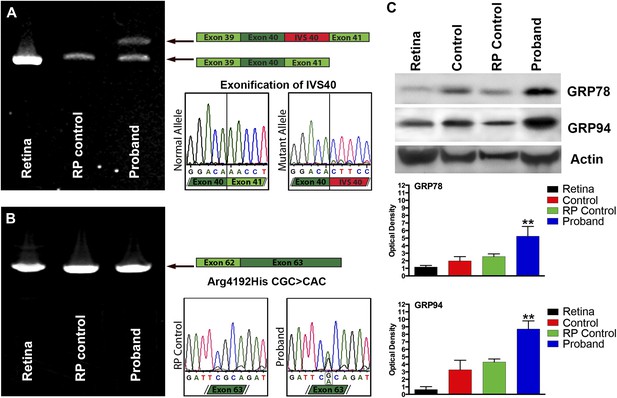
Confirmation of genomic USH2A variants in iPSC-derived neural retina.
(A) RT-PCR analysis of USH2A exons 39 to 41 in human control retina (DePristo et al., 2011), human control iPSC-derived neural retina (Baux et al., 2007), and human RP iPSC-derived neural retina. An intronic splice site mutation in intervening sequence 40 of the USH2A gene results in the introduction of a pseudoexon (IVS40 Red) causing a translation frameshift and a premature stop codon. (B) RT-PCR analysis of USH2A exon 62 to 63 in human control human retina (DePristo et al., 2011), human control iPSC-derived neural retina (Baux et al., 2007), and human RP iPSC-derived neural retina. A single heterozygous point mutation identified by whole exome sequencing (Arg4192His) was confirmed in the proband’s retinal transcript. (C) Western blot analysis of protein isolated from human control retina and iPSC-derived photoreceptor precursor cells obtained from the proband, an unaffected control, and a separate RP patient with known disease pathophysiology for expression of the ER-stress markers GRP78 and GRP94. Elevated expression of both GRP78 and GRP94 suggests that the mutations identified within the proband result in protein misfolding and ER-stress. **p<0.001
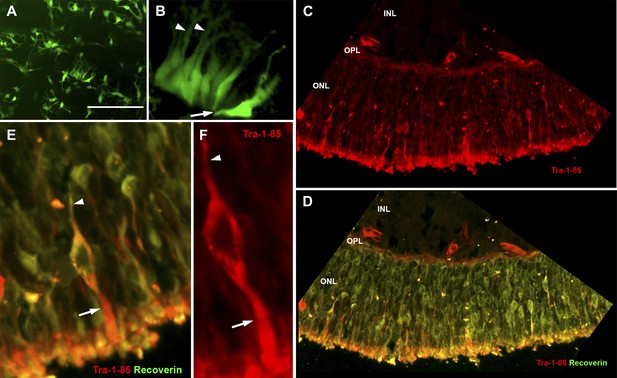
USH2A-associated photoreceptor precursor cells integrate into the dystrophic mouse retina and develop into mature photoreceptor cells.
(A and B) Microscopic analysis of rhodopsin kinase GFP expression in 150-day photoreceptor precursor cells at 14 days post-plating. (C–E) Immunocytochemical analysis performed on the retinas of Rag−/− x Crb1−/− degenerative eyes 14 days after receiving subretinal injections of patient-specific photoreceptor precursor cells targeted against expression of the human cell antigen Tra-1-85 (C–F) and the photoreceptor marker recoverin (D and E). Scale bar = 50 μm.

USH2A-associated photoreceptor precursor cells integrate into the dystrophic mouse retina.
Immunocytochemical analysis performed on the Rag−/− x Crb1−/− degenerative eyes 14 days after receiving subretinal sham injections targeted against expression of the human cell antigen Tra-1-85. ONL, outer nuclear layer; OPL, outer plexiform layer; and INL, inner nuclear layer.
Additional files
-
Supplementary file 1
(A) Prioritization of exome variants. (B) Gene-specific primer sequences used for rt-PCR. F = forward primer and R= reverse primer.
- https://doi.org/10.7554/eLife.00824.013





