The transcriptional regulator BZR1 mediates trade-off between plant innate immunity and growth
Figures
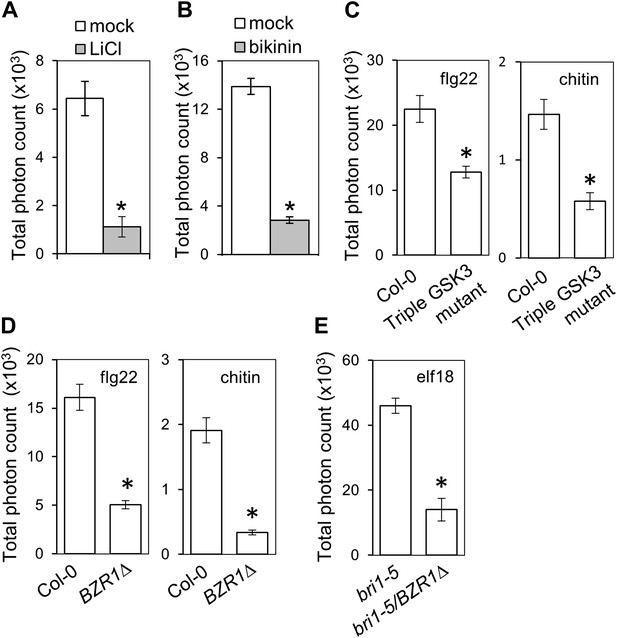
Activation of BZR1 is sufficient to inhibit the PAMP-triggered ROS burst.
(A) and (B) Flg22-triggered ROS burst after LiCl (A) or bikinin (B) treatment. Leaf discs were pre-treated with a 10 mM LiCl solution for 5 hr or with a 50 μM bikinin solution for 16 hr. (C) Flg22- or chitin-triggered ROS burst in Col-0 and the triple GSK3 mutant plants. (D) Flg22- or chitin-induced ROS burst in Col-0 and BZR1Δ plants. (E) Elf18-triggered ROS burst in bri1-5 and bri1-5/BZR1Δ plants. In all cases, bars represent SE of n = 28 rosette leaf discs. Asterisks indicate a statistically significant difference compared to the corresponding control (mock treatment [A and B], Col-0 [C and D] or bri1-5 [E]), according to a Student’s t-test (p<0.05). Leaf discs of four- to five-week-old Arabidopsis plants were used in these assays. Flg22 and elf18 were used at a concentration of 50 nM; chitin was used at a concentration of 1 mg/ml. Total photon counts were integrated between minutes two and 40 after PAMP treatment. All experiments were repeated at least three times with similar results.
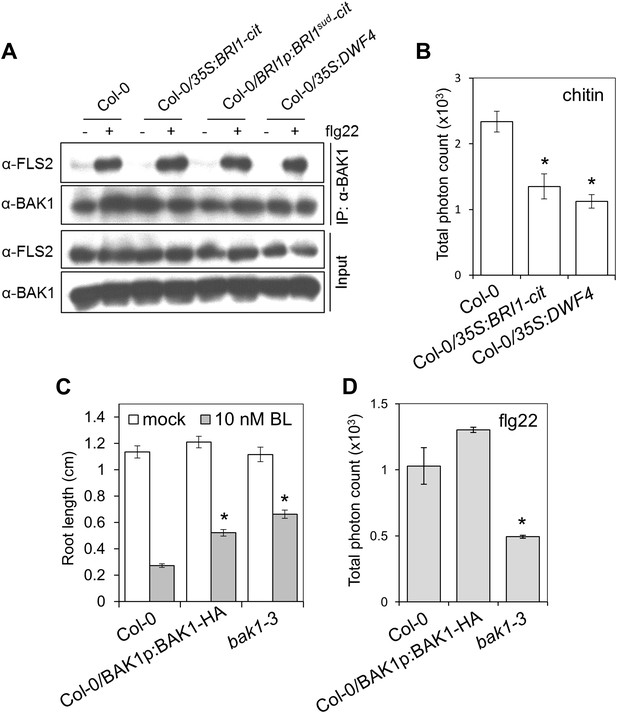
The BR-mediated suppression of PTI can be triggered independently of a competition for BAK1.
(A) Co-IP of BAK1 and FLS2 in Col-0, 35S:BRI1-cit, 35S:BRI1sud-cit and 35S:DWF4 seedlings treated with 1 μM flg22 for 10 min. Coimmunoprecipitated proteins were analyzed by using anti-FLS2 or anti-BAK1 antibodies. (B) Chitin-triggered ROS burst in Col-0, 35S:BRI1-cit and 35S:DWF4 plants. Chitin was used at a concentration of 1 mg/ml. Total photon counts were integrated between minutes two and 40 after PAMP treatment. Bars represent SE of n = 28 rosette leaf discs. (C) Root length of seven-day-old Col-0, BAK1p:BAK1-HA (in Col-0 WT background) or bak1-3 seedlings grown on medium supplemented or not with 10 nM BL. Bars represent SE of 12 ≤ n ≤ 17. Asterisks indicate a statistically significant difference between treatments according to a Student's t-test (p<0.05). (D) Flg22-triggered ROS burst in Col-0, BAK1p:BAK1-HA (in Col-0 WT background) or bak1-3 plants. Leaf discs of four- to five-week-old Arabidopsis plants were used in these assays. Flg22 was used at a concentration of 50 nM. Total photon counts were integrated between minutes two and 40 after PAMP treatment. Bars represent SE of n = 28 rosette leaf discs. All experiments were repeated at least twice with similar results.
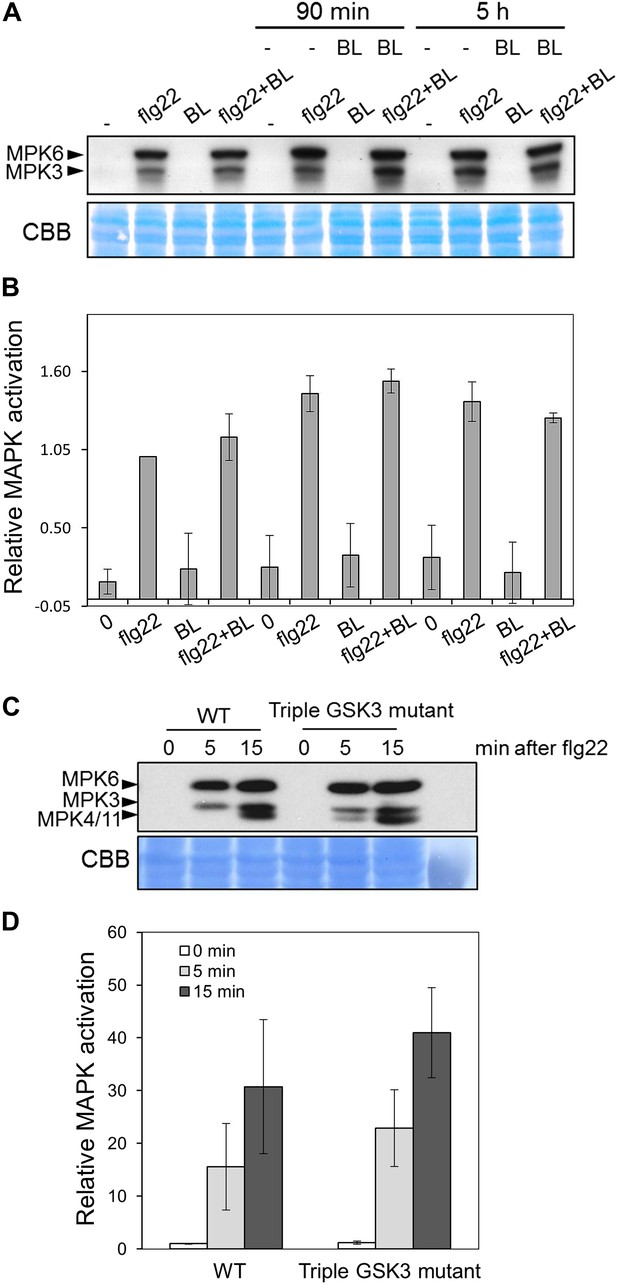
PAMP-triggered MAPK activation is not impaired upon activation of BR signaling.
(A) MAPK activation in Col-0 seedlings upon treatment with 1 μM flg22 (F) and/or epiBL (B) for 10 min (with or without a 90-min or 5-hr BL pre-treatment). (B) Quantification of total MAPK activation in the experiment shown in (A), measured as pixel intensity using ImageJ. Results are the average of two independent blots, corresponding to two independent biological replicated. (C) MAPK activation in Col-0 and Triple GSK3 mutant seedlings upon treatment with 1 μM flg22. (D) Quantification of total MAPK activation in the experiment shown in (C), measured as pixel intensity using ImageJ. Results are the average of two independent blots, corresponding to two independent biological replicated. Proteins were separated in a 10% acrylamide gel and transferred to PVDF membranes. Membranes were blotted with phospho-p44/42 MAPK (Erk1/2; Thr202/Tyr204) rabbit monoclonal antibodies. Bars represent SD.
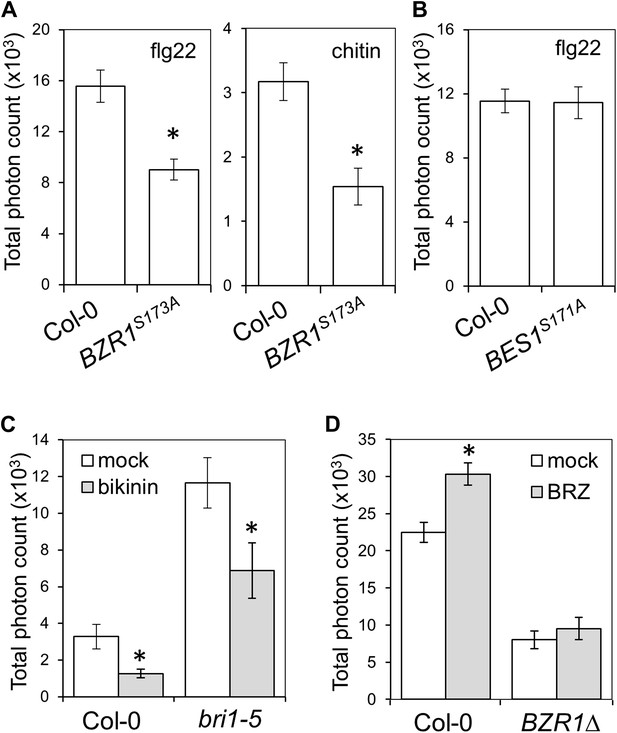
Activation of BZR1, but not BES1, is sufficient to inhibit the PAMP-triggered ROS burst.
(A) Flg22- or chitin-triggered ROS burst in BZR1S173A plants. (B) Flg22-triggered ROS burst in BES1S171A plants. (C) Flg22-triggered ROS burst in mock- or bikinin-treated Col-0 or bri1-5 plants. Leaf discs were pre-treated with a 50 μM bikinin solution for 16 hr. (D) Flg22-triggered ROS burst in mock- or BRZ-treated Col-0 or BZR1Δ plants. Leaf discs were pre-treated with a 2.5 μM BRZ solution for 16 hr. In all cases, bars represent SE of 21 ≤ n ≤ 28. Asterisks indicate a statistically significant difference compared to Col-0 (A and B) or mock-treatment (C and D) according to a Student's t-test (p<0.05). Flg22 was used at a concentration of 50 nM; chitin was used at a concentration of 1 mg/ml. Total photon counts were integrated between minutes two and 40 after PAMP treatment.
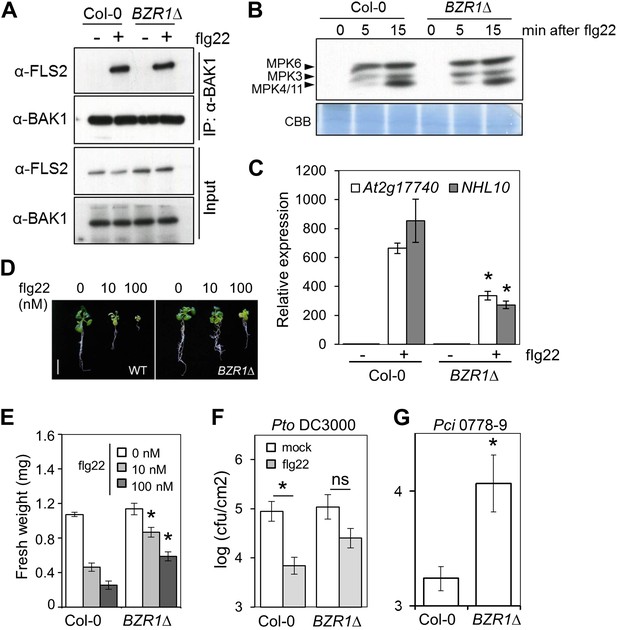
Activation of BZR1 results in the suppression of specific PTI outputs.
(A) Co-immunoprecipitation (Co-IP) of BAK1 and FLS2 in Col-0 and BZR1Δ seedlings after 10 min mock (−) or 1 μM flg22 (+) treatment. Proteins were separated in a 10% acrylamide gel and transferred to PVDF membranes. Membranes were blotted with anti-FLS2 or anti-BAK1 antibodies. (B) MAPK activation in Col-0 and BZR1Δ seedlings upon 1 μM flg22 treatment. Proteins were separated in a 10% acrylamide gel and transferred to PVDF membranes. Membranes were blotted with phospho-p44/42 MAPK (Erk1/2; Thr202/Tyr204) rabbit monoclonal antibodies. CBB: Coomassie brilliant blue. (C) Marker gene (At2g17700 and NHL10) expression in Col-0 and BZR1Δ seedlings after 1 hr mock (−) or 1 μM flg22 (+) treatment, as determined by qPCR. Bars represent SE of n = 3. (D) and (E) Seedling growth inhibition of 10-day-old Col-0 or BZR1Δ seedlings induced by increasing concentrations of flg22, as indicated. Scale bar (D), 1 cm. Bars (E) represent SE of 8 ≤ n ≤ 16. (F) Flg22-induced resistance to P. syringae pv. tomato DC3000 in Col-0 and BZR1Δ plants. Plants were pre-treated with 1 μM flg22 or water 24 hr prior to bacterial infiltration. Bars represent SE of n = 4. This experiment was repeated seven times with similar results. (G) Susceptibility of Col-0 and BZR1Δ plants to P. syringae pv. cilantro 0788-9. Bars represent SE of n = 4. Asterisks indicate a statistically significant difference compared to Col-0 according to a Student’s t-test (p<0.05); ns = not significant. All experiments were repeated at least twice with similar results unless otherwise stated.
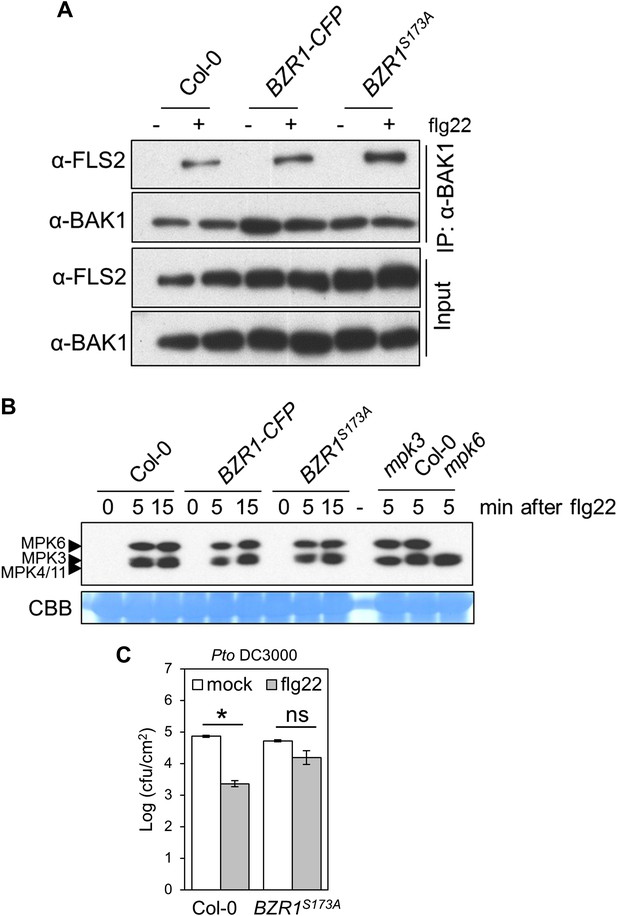
Expression of the constitutively active BZR1S173A results in the suppression of specific PTI outputs.
(A) Co-IP of BAK1 and FLS2 in Col-0 and BZR1S173A seedlings treated with 1 μM flg22 for 10 min. Co-immunoprecipitated proteins were analyzed by using anti-FLS2 or anti-BAK1 antibodies. (B) MAPK activation in Col-0 and BZR1S173A seedlings upon treatment with 1 μM flg22. Membranes were blotted with phospho-p44/42 MAPK (Erk1/2; Thr202/Tyr204) rabbit monoclonal antibodies. (C) Flg22-induced resistance to Pto DC3000 in BZR1S173A plants. Plants were pre-treated with 1 μM flg22 or water 24 hr prior to bacterial inoculation. Bars represent SE of n = 4. Asterisks indicate a statistically significant difference compared to mock-treated plants according to a Student's t-test (p<0.05); ns = not significant. All experiments were repeated at least twice with similar results.
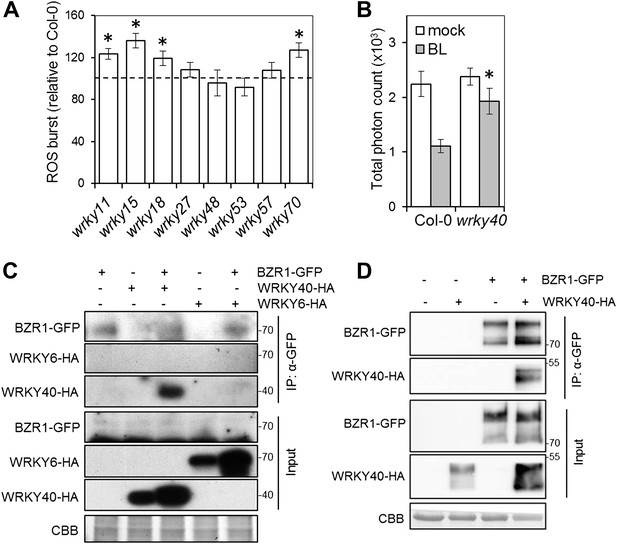
WRKY transcription factors play a dual role on the BR-mediated regulation of PTI signaling.
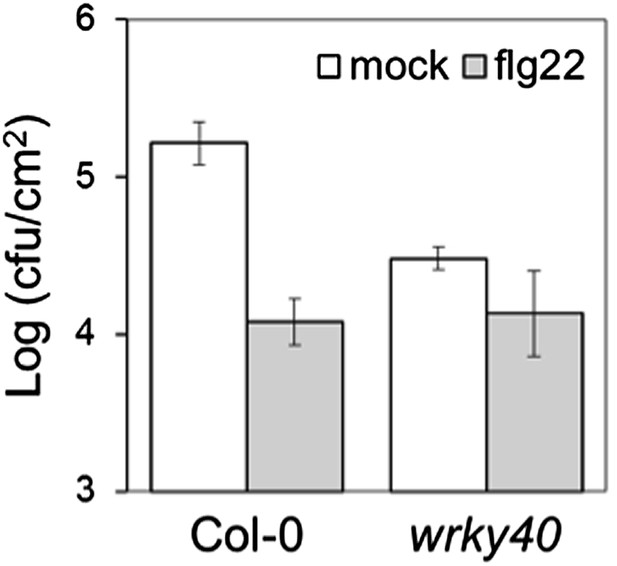
(A) Flg22-triggered ROS burst in mutants in each BR-induced BZR1-targeted WRKY. Leaf discs of four- to five-week-old Arabidopsis plants were used in these assays. Flg22 was used at a concentration of 50 nM. Total photon counts were integrated between minutes two and 40 after PAMP treatment. Bars represent SE of n = 28. Asterisks indicate a statistically significant difference compared to Col-0 according to a Student’s t-test (p<0.05). (B) Flg22-triggered ROS burst in epiBL (BL)- or mock- pre-treated wrky40 mutant or wild-type plants. Leaf discs of four- to five-week-old plants were pre-treated with a 1 μM BL solution or mock solution for 8 hr. Flg22 was used at a concentration of 50 nM. Total photon counts were integrated between minutes two and 40 after PAMP treatment. Bars represent SE of n = 21. Asterisks indicate a statistically significant difference compared to Col-0 according to a Student’s t-test (p<0.05). (C) Co-IP of BZR1-GFP transiently expressed in N. benthamiana, alone or together with WRKY40-HA or WRKY6-HA. BZR1-GFP was immunoprecipitated with an anti-GFP antibody. Immuniprecipitated or total proteins were separated in a 10% acrylamide gel and transferred to PVDF membranes. Membranes were blotted with anti-HA or anti-GFP antibodies. CBB: Coomassie brilliant blue. (D) Co-IP of BZR1-GFP transiently expressed in Arabidopsis protoplasts, alone or together with WRKY40-HA. BZR1-GFP was immunoprecipitated with an anti-GFP antibody. Immuniprecipitated or total proteins were separated in a 10% acrylamide gel and transferred to PVDF membranes. Membranes were blotted with anti-HA or anti-GFP antibodies. CBB: Coomassie brilliant blue. All experiments were repeated at least twice with similar results.
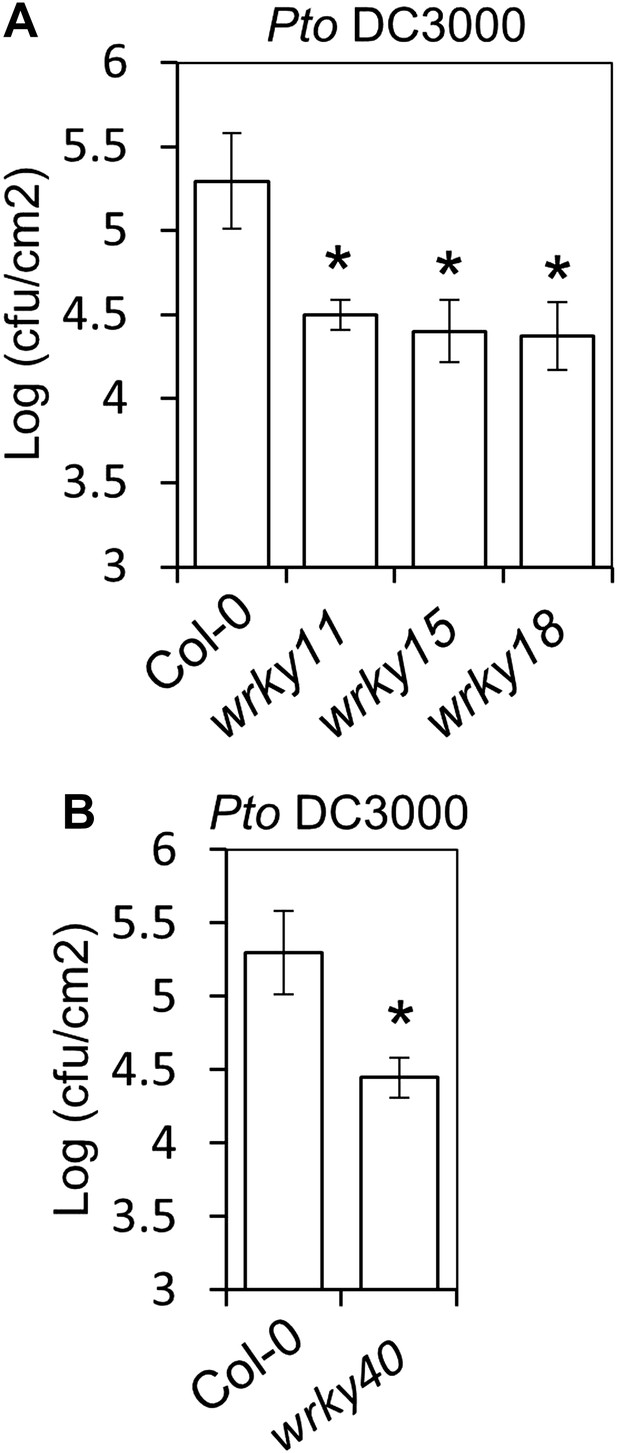
Mutants in WRKY11, WRKY15, WRKY18 and WRKY40 are more resistant to Pto DC3000.
(A) and (B) Pto DC3000 infections in Col-0, wrky11, wrky15, and wrky18 (A) and in Col-0 and wrky40 (B) plants. Bars represent SE of n = 4. Asterisks indicate a statistically significant difference compared to Col-0 plants according to a Student's t-test (p<0.05); ns = not significant. All experiments were repeated at least three times with similar results.
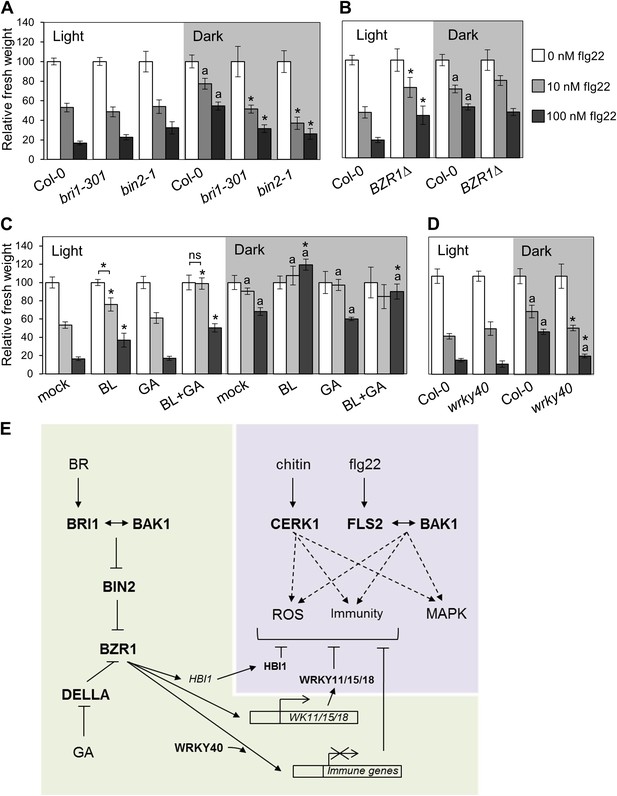
Activation of BR signaling and BZR1 prioritizes growth over immunity in the dark.
(A) and (B) Relative seedling growth inhibition of 10-day-old (A) Col-0, bri1-301 and bin2-1 or (B) Col-0 and BZR1Δ seedlings induced by increasing concentrations of flg22 in either light or dark. (C) Relative seedling growth inhibition of 10-day-old Col-0 seedlings grown on medium supplemented or not with BL (1 μM), GA (1 μM), BL+GA (1 μM + 1 μM) or mock solution in light or dark. (D) Relative seedling growth inhibition of Col-0 or wrky40 seedlings induced by increasing concentrations of flg22 in either light or dark. Bars represent SE of n = 16 (A, B and D) or n = 8 (C) Asterisks indicate a statistically significant difference compared to Col-0 in the same condition (light or dark and same concentration of flg22), according to a Student’s t-test (p<0.05); ‘a’ indicates a statistically significant difference compared to the same genotype/treatment and concentration of flg22 in light, according to a Student’s t-test (p<0.05). All experiments were repeated at least three times with similar results. Values are relative to Col-0 (A, B and D) or mock-treated seedlings (C) (set to 100). Absolute values of these experiments are shown in Figure 4—figure supplement 3. (E) Schematic model depicting the BZR1-mediated inhibition of PTI. Upon BR- and DELLA-dependent activation, BZR1 induces the expression of negative regulators of PTI, such as WRKY11, WRKY15, WRKY18, or HBI1. In addition, BZR1 also inhibits the expression of immune genes, acting cooperatively with WRKY40 and possibly other WRKYs. Ultimately, the BZR1-mediated changes in transcription would lead to the suppression of PTI signaling. The PTI signaling pathway is shadowed in violet; the BR signaling pathway is shadowed in green.
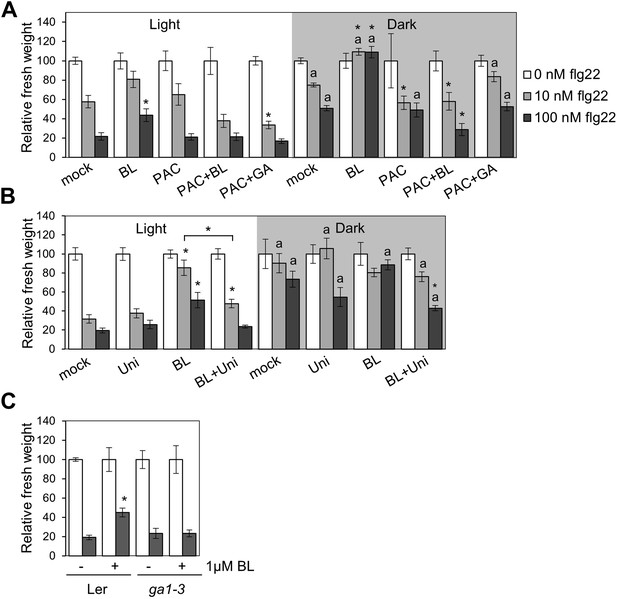
The BR-mediated suppression of seedling growth inhibition in the dark requires GA synthesis.
Seedling growth inhibition of 10-day-old Col-0 seedlings grown on medium supplemented or not with (A) BL (1 μM), paclobutrazol (PAC; 1 μM), BL+PAC (1 μM + 1 μM) and PAC+GA (1 μM + 1 μM), or (B) uniconazole (Uni; 100 μM), BL (1 μM) and Uni+BL (100 μM + 1 μM) induced by increasing concentrations of flg22 in light or dark. (C) Seedling growth inhibition of 10-day-old Ler and ga1-3 seedlings grown on medium supplemented or not with BL (1 μM). Bars represent SE of n = 8. Asterisks indicate a statistically significant difference compared to Col-0 in the same condition (light or dark and same concentration of flg22), according to a Student's t-test (p<0.05); ‘a’ indicates a statistically significant difference compared to the same genotype/treatment and concentration of flg22 in light, according to a Student's t-test (p<0.05). Values are relative to mock-treated seedlings (set to 100). All experiments were repeated at least twice with similar results.
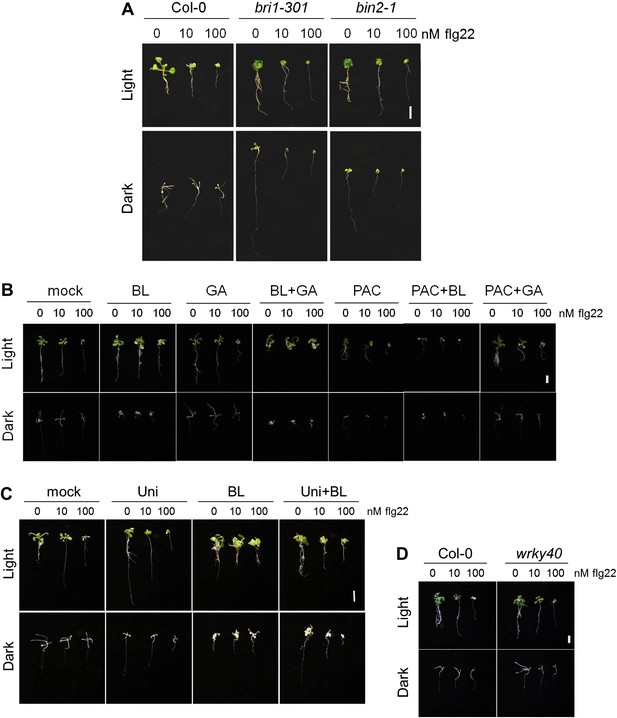
Phenotype of the light- or dark-grown seedlings used in the seedling growth inhibition assays (Figure 4 and Figure 4—figure supplement 1).
Representative seedlings of the seedling growth inhibition experiments depicted in: (A) Figure 4A; (B) Figure 4—figure supplement 1A; (C) Figure 4–figure supplement 1B; (D) Figure 4D. Scale bar, 1 cm.
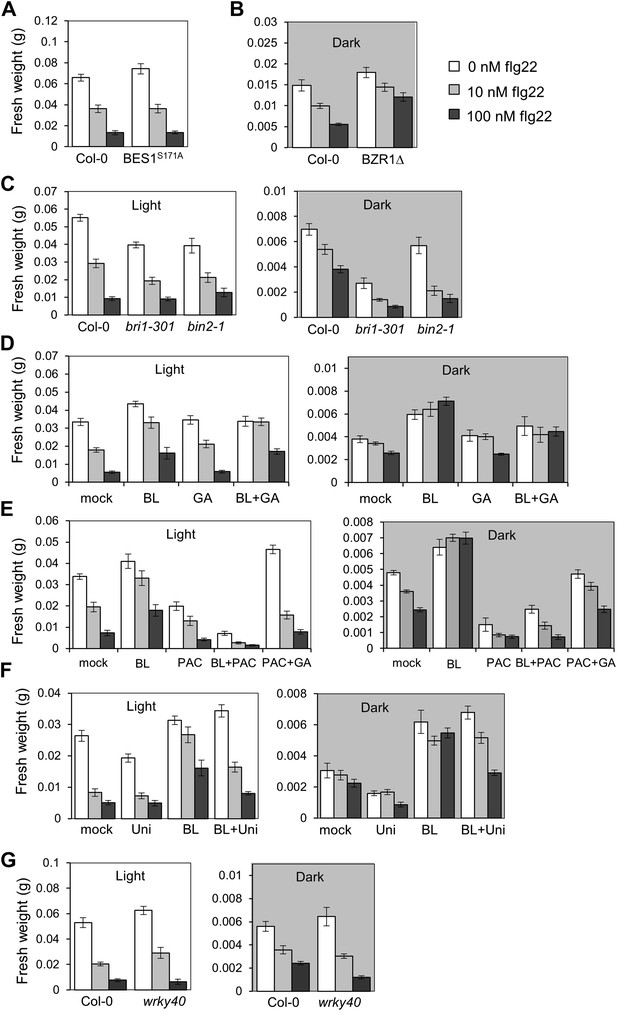
Absolute fresh weight values of seedling growth inhibition assays.
(A) Seedling growth inhibition of 10-day-old Col-0 or BES1S171A seedlings induced by increasing concentrations of flg22. (B) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4B, dark. (C) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4A. (D) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4C. (E) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4—figure supplement 1A. (F) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4—figure supplement 1B. (G) Absolute fresh weight values of the seedling growth inhibition assay depicted in Figure 4D. Error bars represent SE as indicated in Figure 4, Figure 4—figure supplement 1.
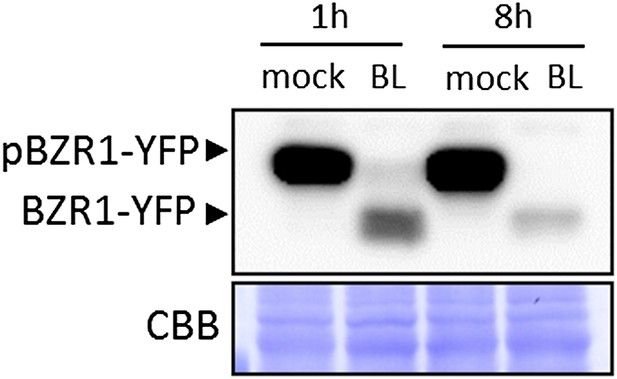
BL-induced dephosphorylation of BZR1 is maintained after an 8-hour BL treatment. Ten-day-old transgenic Arabidopsis seedlings expressing BZR1-YFP were treated with 1μM BL or mock solution for the indicated time. Proteins were detected using an anti-GFP antibody conjugated to HRP.
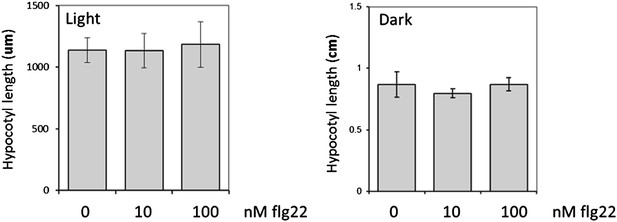
Hypocotyl length of light- or dark-grown Arabidopsis seedlings in increasing concentrations of flg22. Seedlings were grown in MS plates for four days, then transferred to liquid MS supplemented with flg22 at the indicated concentrations. Hypocotyl length was measured four days later. Bars represent SE with n=8.
Tables
Defense-related Gene Ontology terms (Biological Process ontology) over-represented among all BR-regulated genes, BR-regulated BZR1 targets and BR-regulated BES1 targets
| Defense-related GO term | Observed frequency (%) | Expected frequency (%) | p-value |
|---|---|---|---|
| BR-Regulated genes | |||
| response to bacterium | 2.2 | 1 | 3.31 × 10−08 |
| defense response to bacterium | 1.9 | 0.8 | 3.31 × 10−08 |
| response to chitin | 1.4 | 0.5 | 1.78 × 10−07 |
| defense response | 4.7 | 3 | 3.32 × 10−07 |
| response to fungus | 1.5 | 0.7 | 3.4 × 10−06 |
| response to nematode | 0.7 | 0.2 | 0.000532 |
| defense response to fungus | 1 | 0.5 | 0.0035 |
| BR-regulated BZR1 targets | |||
| response to chitin | 2.6 | 0.5 | 9.13 × 10−13 |
| response to bacterium | 2.3 | 1 | 0.00112 |
| defense response to bacterium | 1.9 | 0.8 | 0.00154 |
| response to fungus | 1.6 | 0.7 | 0.00495 |
| BR-regulated BES1 targets | |||
| response to chitin | 2.4 | 0.5 | 0.00439 |
Gene Ontology terms (Molecular Function ontology) over-represented among all BR-regulated BZR1 targets
| Over-represented GO term | Observed frequency (%) | Expected frequency (%) | p value |
|---|---|---|---|
| BR-regulated BZR1 targets | |||
| nucleic acid binding transcription factor activity | 14.8 | 10.2 | 0.000223 |
| transferase activity | 21.6 | 16.8 | 0.00333 |
| kinase activity | 11.6 | 8.1 | 0.00702 |
| transcription repressor activity | 1.1 | 0.3 | 0.01 |
Defense-related Gene Ontology terms (Biological Process ontology) over-represented among the BZR1-target BR-regulated transcription factors
| Defense-related GO Term | Observed frequency (%) | Expected frequency (%) | p value |
|---|---|---|---|
| BZR1-target BR-regulated TFs | |||
| response to chitin | 16.6 | 0.5 | 1.36 × 10−26 |
| defense response to bacterium | 7.6 | 0.8 | 4.71 × 10−07 |
| response to bacterium | 7.6 | 1 | 4.51 × 10−06 |
| regulation of defense response to virus by host | 1.4 | 0 | 0.000964 |
| regulation of immune effector process | 1.4 | 0 | 0.00151 |
| regulation of defense response to virus | 1.4 | 0 | 0.00151 |
| regulation of defense response | 2.8 | 0.3 | 0.00484 |
| defense response | 8.3 | 3 | 0.00603 |
| response to fungus | 3.4 | 0.7 | 0.01 |
| defense response to fungus | 2.8 | 0.5 | 0.02 |
Over-represented cis-acting promoter elements among the defense-related BR-regulated genes according to Athena (http://www.bioinformatics2.wsu.edu/cgi-bin/Athena/cgi/home.pl)
| Enriched TF site | % promoters | p value |
|---|---|---|
| Defense-related BR-regulated genes | ||
| W-box | 72.4 | <10−6 |
BR-regulated BZR1-target WRKY genes
| AGI number | WRKY TF |
|---|---|
| BR-Induced BZR1 targets | |
| AT4G31800 | WRKY18 |
| AT4G31550 | WRKY11 |
| AT4G23810 | WRKY53 |
| AT3G56400 | WRKY70 |
| AT5G49520 | WRKY48 |
| AT5G52830 | WRKY27 |
| AT1G69310 | WRKY57 |
| AT2G23320 | WRKY15 (Yu et al., 2011) |
| BR-repressed BZR1 targets | |
| AT4G01250 | WRKY22 |
| AT1G80840 | WRKY40 |
| AT2G24570 | WRKY17 |
| AT2G23320 | WRKY15 (Sun et al., 2010) |
| AT2G30590 | WRKY21 |
Overlap between the targets of WRKY40 and BZR1
| Known targets of WRKY40 (Pandey et al., 2010) | Targets of BZR1 (Sun et al., 2010) |
|---|---|
| Confirmed by ChIP | |
| EDS1 | Yes |
| RRTF1 | Yes |
| JAZ8 | Yes |
| Putative (according to expression analyses) | |
| LOX2 | Yes |
| AOS | Yes |
| JAZ7 | Yes |
| JAZ10 | Yes |
Representation of the W-box motif among the BR-regulated BZR1 targets according to Athena (http://www.bioinformatics2.wsu.edu/cgi-bin/Athena/cgi/home.pl)
| BZR1 targets | % of promoters with W-box motif(s) | p value |
|---|---|---|
| BR-induced | 66 | <10−10 |
| BR-repressed | 72 | <10−4 |