Ribosome profiling reveals pervasive and regulated stop codon readthrough in Drosophila melanogaster
Figures
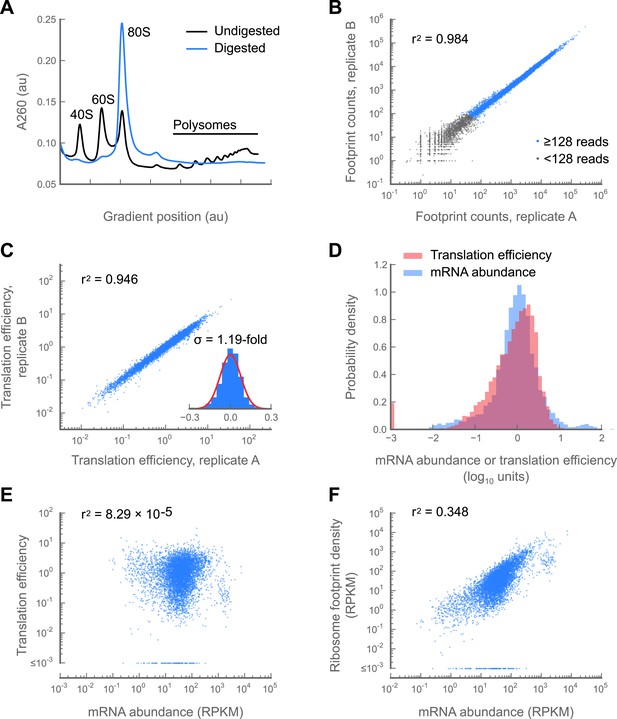
Development and validation of a ribosome profiling assay for Drosophila melanogaster.
(A) Aliquots of polysome lysate from 0–2 hr embryos were fractionated on 10–50% sucrose gradients with or without prior micrococcal nuclease digestion. Digestion of exposed mRNA between ribosomes collapses the polysome peaks into the monosomal (80S) peak. The area under the monosome peak in the digested sample is 1.04-fold the combined area under the monosome and polysome peaks in the undigested sample, indicating quantitative recovery. (B and C) Measurements of translation are reproducible between replicates samples of 0–2 hr embryos. Pearson correlation coefficients (r2) are shown for total ribosome-protected footprint counts in coding regions for all genes sharing at least 128 summed footprint counts between replicates (B), or translation efficiency measurements for all genes sharing 128 summed mRNA fragment counts between replicates (C). Histogram of log10 fold-changes in translational efficiency for each gene between two embryo replicates, along with normal error curve (C, inset). (D–F) Pooled data for genes containing at least 128 summed mRNA counts between both embryo replicates. Median-centered histograms of translation efficiency (pink) and mRNA abundance (blue) (D). Translational efficiency vs mRNA abundance for each gene (E). Ribosome density vs mRNA abundance for each gene (F). Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
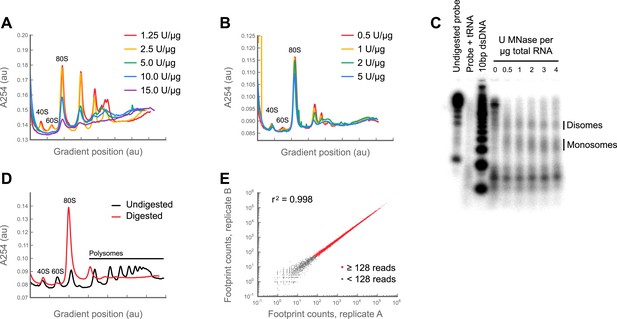
Digestion with micrococcal nuclease yields a robust ribosome profiling assay.
(A) Digestion of polysomes with RNase I degrades ribosomes. A lysate was made from S2 cells using a previous version of our protocol. Aliquots of this lysate were digested with increasing amounts of RNase I, and resolved on 10–50% sucrose gradients. As amounts of RNase I increase, the heights of all peaks—including the monosomal (80S) peak—decrease before polysomes are fully resolved to monosomes. (B) as in (A), but using micrococcal nuclease (MNase) and our current protocol. From 0.5 to 2 U MNase/μg total RNA, monosomes are resolved with no reduction in the size of the monosome peak. This result indicates that Drosophila ribosomes are stable to MNase over a broad range of concentrations, whereas the mRNA between ribosomes is digested. (C) Ribosome protection assay. A 320 nucleotide fragment of enolase (FlyBase accession: FBgn0000579) was amplified using oligos oJGD123 & oJGD124 (Supplementary file 2). A body-labeled probe against this sequence was transcribed from this template using α32P-UTP and the T7 MaxiScript kit (Ambion). S2 cell lysates were prepared as in methods and aliquoted. Aliquots were digested as in methods, except with 0, 0.5, 1, 2, 3 or 4 U MNase/μg total RNA. Monosomes were sedimented through a sucrose cushion, resuspended in 600 μl 10 mM Tris pH 7.0, and their RNAs extracted as in ‘Materials and methods’. Concentrations were determined using a NanoDrop spectrophotometer. 5 μg of each sample was hybridized to 50,000 CPM of probe overnight at 42°C. Single-stranded regions were digested with RNase A/T1 and the remaining footprint: probe duplexes detected using the mirVana micro-RNA detection kit (Ambion), resolved on a 15% TBE-urea gel (Invitrogen), and visualized on a Storm phosphorimager (Molecular Dynamics by GE Healthcare Bio-Sciences, Pittsburgh, PA). For size markers, we end-labeled the Novex 10 bp dsDNA ladder (Invitrogen) with 32P. Over two-fold range of nuclease concentrations, the ∼30 nt peak corresponding to ribosome-protected footprints remains constant in size and intensity, indicating a lack of degradation consistent with the unchanged monosome peak height across this range of digestion conditions in (B). Also visible is a roughly 60 nt band which we infer to be protected by adjacent ribosomes (disomes) that sterically exclude the nuclease. This interpretation is consistent with the presence of a small disome peak in digested samples (c.f. panels B and D, and Figure 1A). (D) A polysome lysate was prepared from S2 cells and resolved in 10–50% sucrose gradients, with or without prior digestion with 3 U MNase/μg total RNA (E) A culture of S2 cells was split into aliquots and processed using our current protocol as if they were independent samples. Total counts aligning to the coding region of each gene were tabulated in each replicate. Genes sharing at least 128 footprint counts between replicates (red) are well-correlated, demonstrating the assay is robust (see full discussion in Figure 1—figure supplement 2). Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
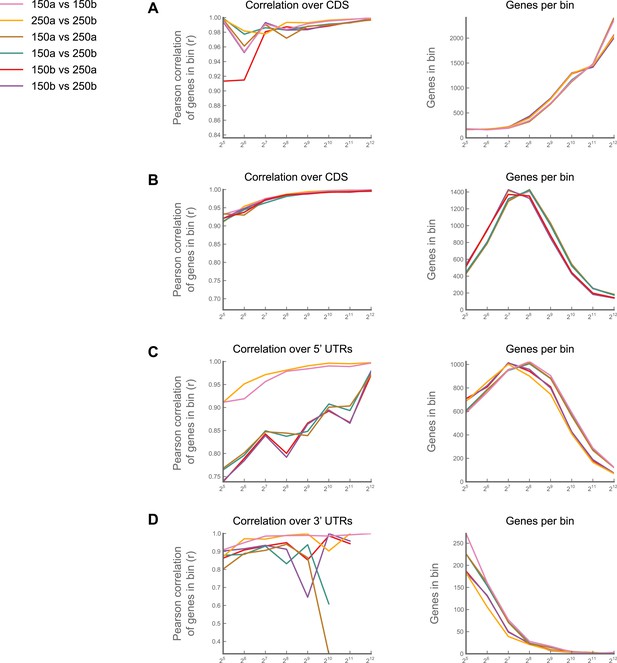
Effects of buffer conditions upon reproducibility.
A culture of S2 cells was divided into four aliquots, and each aliquot carried through the entire ribosome profiling procedure as an independent sample. Two aliquots (‘150a’ and ‘150b’) were processed using our standard lysis buffer with 150 mM Na+ and 5 mM Mg+ and digested with 3 U MNase/μg total RNA as described in ‘Materials and methods’. The other two (‘250a’ and ‘250b’) were processed using an earlier version of our protocol, in which our lysis buffer contained 250 mM Na+ and 15 mM Mg++, and in which we digested lysates with 30 U MNase/μg total RNA. We then calculated ribosome density for each gene over coding regions (A), 5' UTRs (C) and 3' UTRs (D), performed pairwise comparisons between samples. For each comparison, we binned genes based upon the summed number of reads in samples A and B, and calculated the correlation coefficients (Pearson's r) for the RPKM values for each gene in each bin (left column). The number of genes in each bin are also shown (right column). Correlations between samples for coding regions are robust across buffer regions (A), though some salt-dependence is visible in 5′ and 3′ UTRs (C and D). (B) As in (A), but using only 10% of the reads. The high correlation observed at our 128-minimum-count threshold is therefore not a function of the number of genes in each bin Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
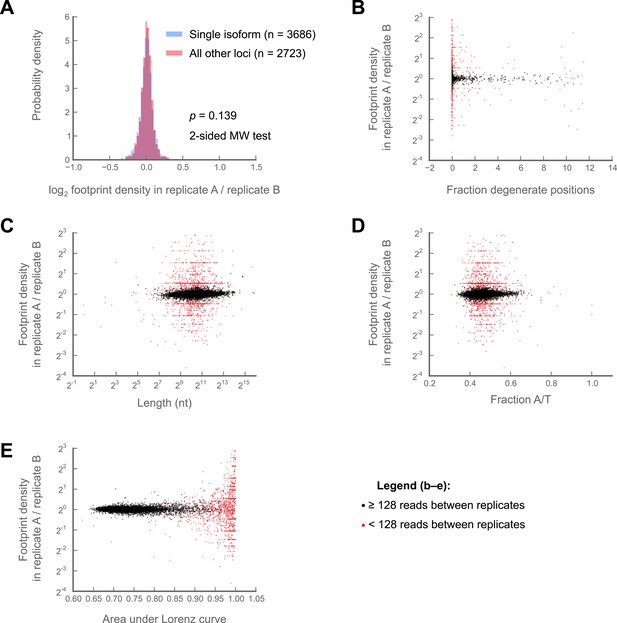
Variability in ribosome footprint density measurements are not correlated with isoform number, sequence degeneracy in the locus of interest, locus length, A/T content, or evenness of coverage.
Comparisons are made between S2 cell technical replicates 150a and 150b (Figure 1—figure supplement 2) (A) Variability of log2 fold-changes in ribosome footprint densities are no greater for multi-isoform loci (pink) than they are for single-isoform loci (blue) (B) Correlation of the fraction degenerate positions in each locus (‘Materials and methods’) with fold-changes in ribosome density between replicates at that locus. Loci with at least 128 counts between replicates are shown in black, those with less in red. (C) as in (B), but correlation of length with inter-replicate fold-changes. (D) as in (B), but correlation of A/T content with inter-replicate fold-changes. (E) as in (B), but correlation of area under Lorenz curve with inter-replicate fold-changes Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
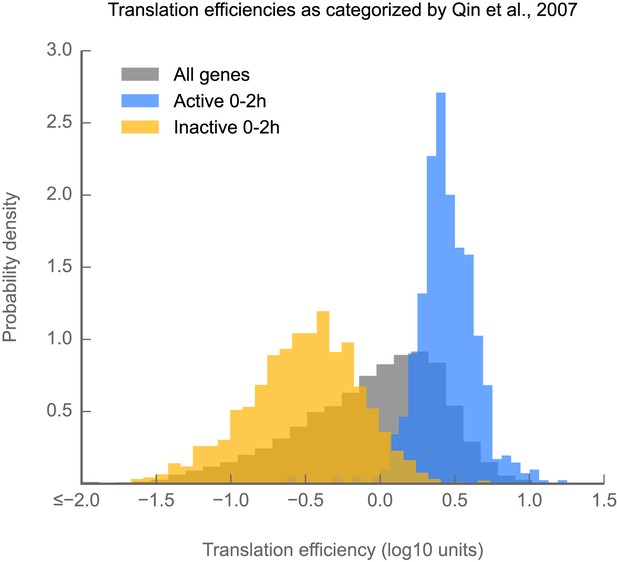
Measurements of translation efficiency obtained via ribosome profiling are consistent with those made using semiquantitative polysome gradients.
Histograms of translation efficiency for genes labeled by Qin et al. (2007) as active (blue) or inactive (yellow) in 0–2 hr embryos. All genes are shown in gray. Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
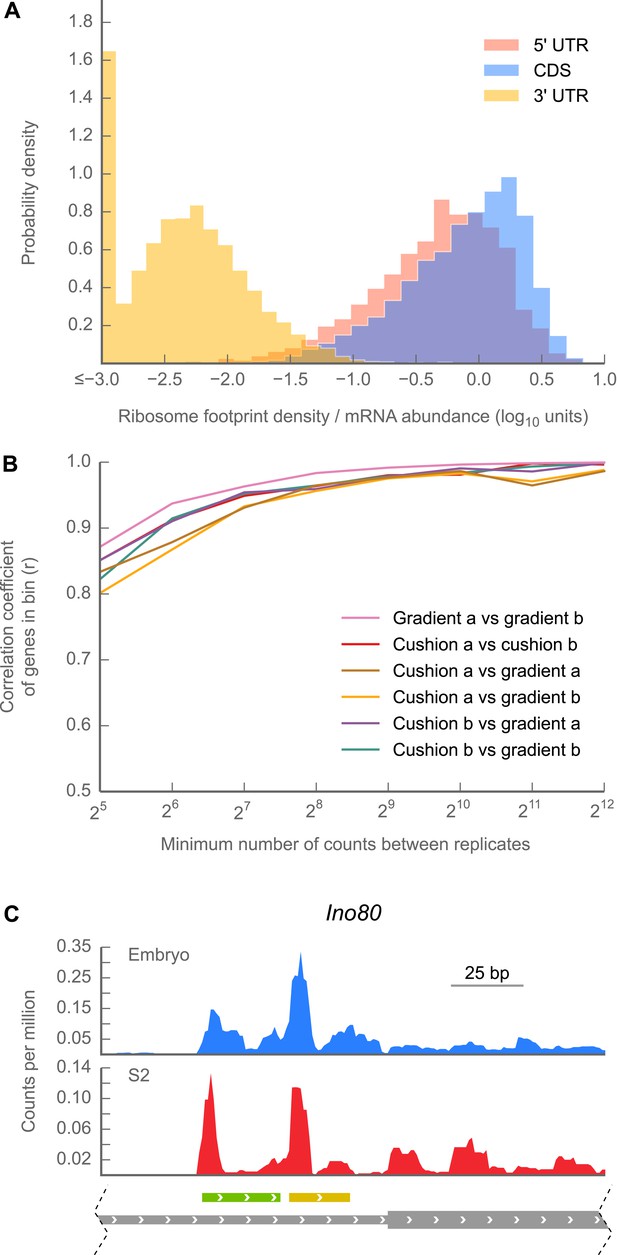
5’ UTRs are translated.
(A) Histograms of ribosome footprint density, corrected by mRNA abundance, for 5’ UTRs, coding regions (CDS), and 3’ UTRs in 0–2 hr embryos. (B) Measurements of ribosome footprint densities of 5’ UTRs agree comparably well across a range of sequencing depths, regardless of whether 80S monosomes are specifically isolated on a sucrose gradient or enriched in a cushion. For each pair of sequencing samples, Pearson correlation coefficients (r) of ribosome footprint density measurements for 5’ UTRs are plotted as a function of sequencing depth. (C) Example of ribosome density in 5’ UTRs corresponding to the locations of uORFs. Roughly ∼200 nt of the genomic locus Ino80 covering portions of the 5’ UTR (thin gray box) and CDS (thick gray box) are shown. In both 0–2 hr embryos and S2 cells, Initiation peaks are visible at the starts of uORFs starting with an ATG codon (green box) and a near-cognate TTG codon (yellow box) as well as at the annotated start codon (beginning of thick gray box). Source data for panels (A) and (B) may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
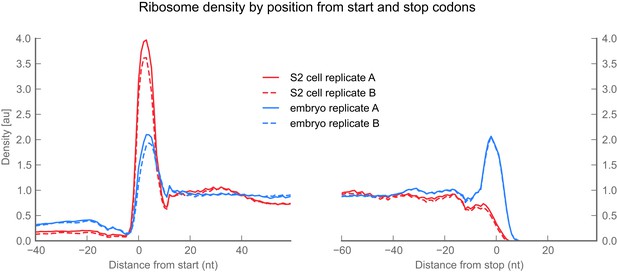
Ribosome density over start and stop codons.
Ribosome density across the average gene or ‘metagene’ reveals peaks of ribosome density at start and stop codons. For this analysis we included all genes that met the following criteria: (a) all transcripts deriving from that gene had one annotated start codon (left panel) or stop codon (right panel), (b) all transcripts deriving from that locus covered identical genomic positions over the region of interest (ROI) shown, (c) all positions within the ROI were non-degenerate (‘Materials and methods’), and (d) at least 10 reads were present in the coding subregion of the ROI. For each ROI meeting these criteria (2800–3200 ROI per sample), we generated a ‘coverage vector’ tallying ribosome density at each nucleotide position. We then normalized each coverage vector to the mean number of footprint reads covering the annotated coding region in the ROI, excluding a 3-codon buffer flanking the start or stop codon to avoid bleedthrough from initiation or termination peaks. We then plotted the median value across all normalized coverage vectors at each position. Peaks are visible in the start and stop codons of embryo samples. Consistent with our previous work, stop codon peaks are missing from S2 cell samples because terminating ribosomes release during our 2-min treatment with translation inhibitors. They are present in our embryo samples, because these are flash-frozen and lysed in the presence of translation inhibitors, which block termination as well as initiation and elongation.
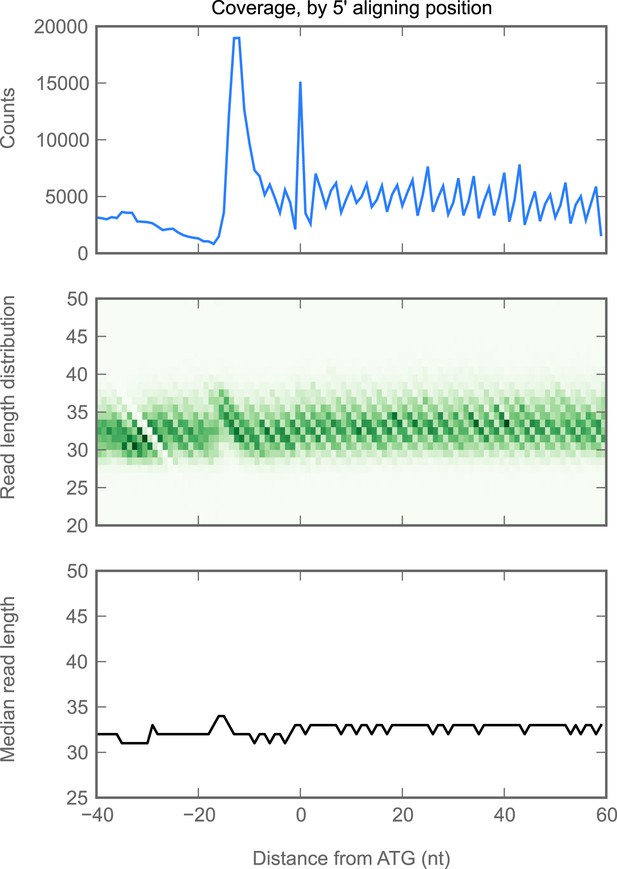
Read lengths are similar in 5’ UTRs and coding regions.
We aggregated all ribosome-protected reads aligning to all genes with a single initiation codon, and in which all annotated isoforms cover the same genomic positions in the ROI shown. We plotted the following statistics as a function of the reads whose 5' end mapped to each position on the x-axis. Top: number of reads (y-axis) aligning at each position. Because the 5' end, rather than the P-site, is plotted, the peak of ribosome density is approximately 13 nucleotides 5' of the start codon (position 0, x-axis). Middle: heatmap of read lengths (y-axis) as a function of position. Bottom: median read length (y-axis) at each position.
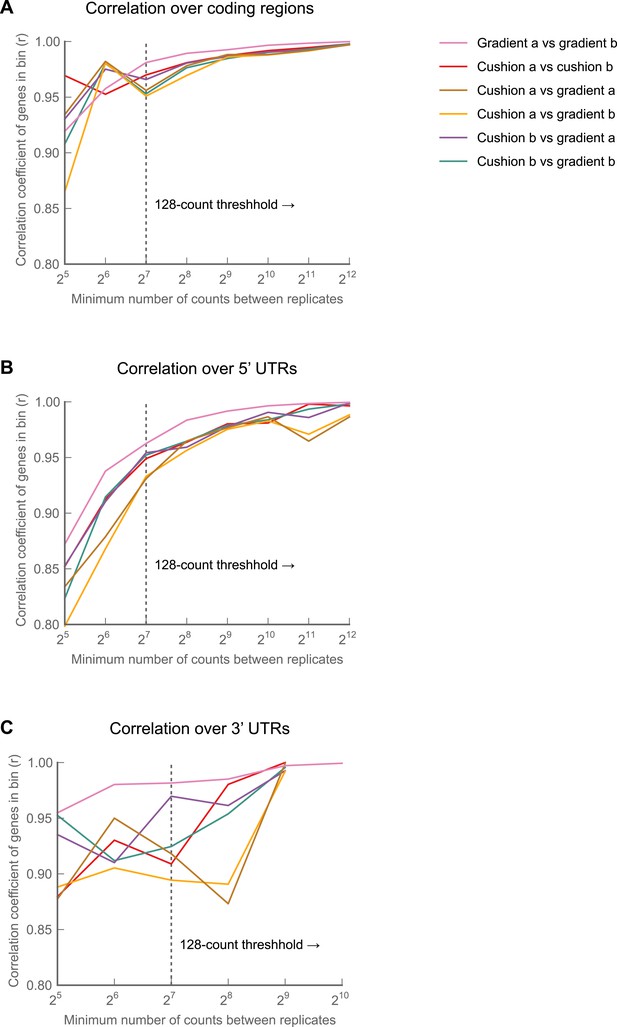
The choice of monosome enrichment technique—sedimentation through sucrose cushions or by fractionation on sucrose gradients—minimally affects of ribosome density across 5’ UTRs and coding regions. 3’ UTR measurements are noisier in samples prepared on cushions rather than gradients.
A polysome lysate was made from collected 0–2 hr embryos, digested with MNase, and split into four aliquots. Monosomes from two aliquots were sedimented through a sucrose cushion and recovered. Monosomes from the remaining two aliquots were fractionated on 10–50% sucrose gradients and collected. All four samples were then independently carried through our protocol, and footprint density was calculated over coding regions, 5' UTRs, and 3' UTRs. Pairwise comparisons were made for each sample as in Figure 1—figure supplement 2 over coding regions (A), 5' UTRs (B), or 3' UTRs (C). Pearson correlations (r) for the regions are plotted as a function of sequencing depth. Source data may be found in supplementary table 1 (at Dryad: Dunn et al., 2013).
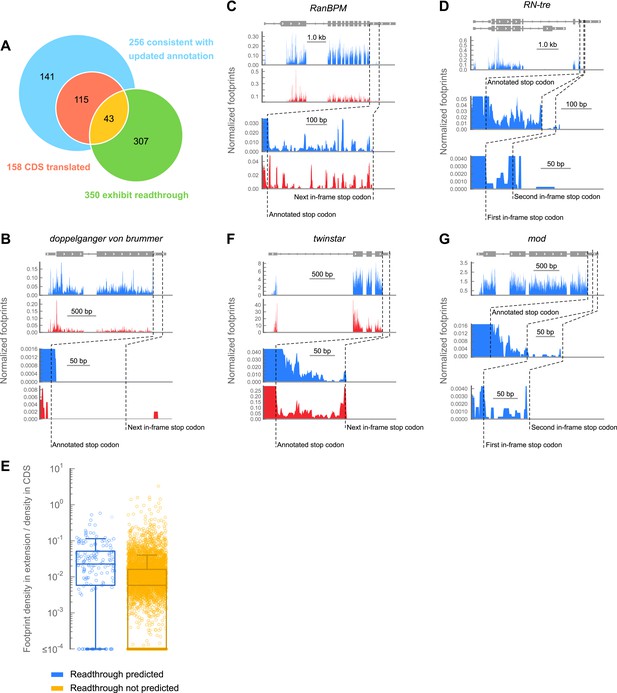
A subset of genes exhibit apparent stop codon readthrough.
(A) Venn diagram summarizing readthrough events. Of 283 predicted extensions, 256 were consistent with FlyBase genome annotation revision 5.43. For 158 of these, the corresponding coding regions were expressed in 0–2 hr embryos. Of this subset, 43 exhibited clear signs of readthrough. Others were ambiguous, untranslated, or could be explained by other mechanisms (Figure 3—figure supplement 1). In addition, we identified 307 examples of readthrough that were not phylogenetically predicted. (B) Example of a gene that does not exhibit readthrough. Top: genomic locus with UTRs (thin boxes), introns (line), and coding regions (thick boxes). Middle: normalized footprint density covering the locus in 0–2 hr embryos (blue) and S2 cells (red) in reads per million. Bottom: magnification of region where a putative C-terminal extension would be found. Dashed lines: annotated and next in-frame stop codons (C) as in (B), except stop codon readthrough creates a C-terminal protein extension in RanBPM, a gene phylogenetically predicted to undergo readthrough (D) as in (B), but an example of phylogenetically predicted double-readthrough. (E) Ratios of the ribosome footprint density in putative extensions to corresponding coding regions. Blue: extensions predicted to undergo readthrough. Yellow: all other possible extensions. Extensions that overlapped any annotated CDS, snoRNA, or snRNA were excluded. Boxes: IQR. Whiskers: 1.5*IQR. (F) as in (C), except this transcript was not predicted to undergo readthrough. (G) as in (D), except this transcript was not predicted to undergo single or double readthrough. Source data may be found in supplementary table 2 (at Dryad: Dunn et al., 2013).
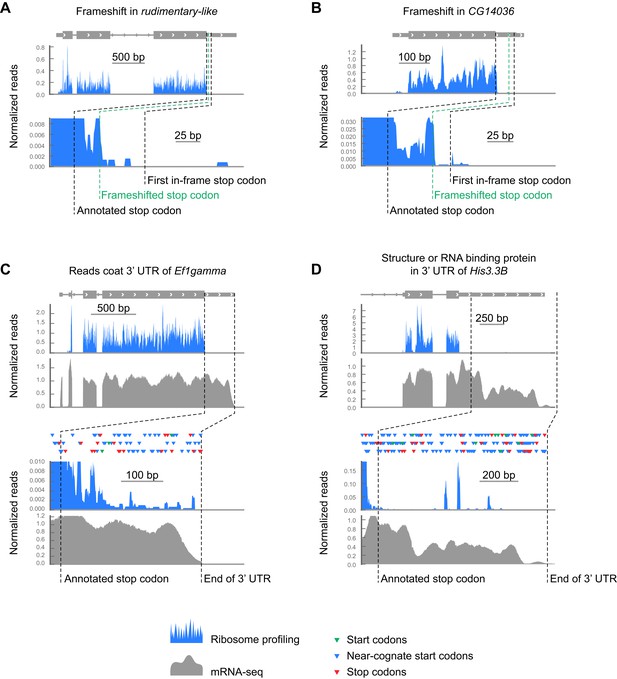
Examples of footprint density in 3’ UTRs attributed to sources other than readthrough.
(A and B) Sample transcripts exhibiting translation in alternate frames. (C) Footprint density, potentially caused by RNA binding proteins or structures, coats the 3' UTR of EF1gamma, passing through stop codons (red triangles) in all three frames reaching the 3' end of the transcript. Colors as in (A and B), but additionally showing RNA-seq data in gray. (D) The 3' UTR of HIS3.3B contains highly localized read density consistent with the presence of an RNA binding protein or mRNA structure, but not with translation of an open reading frame. Colors as in (C).
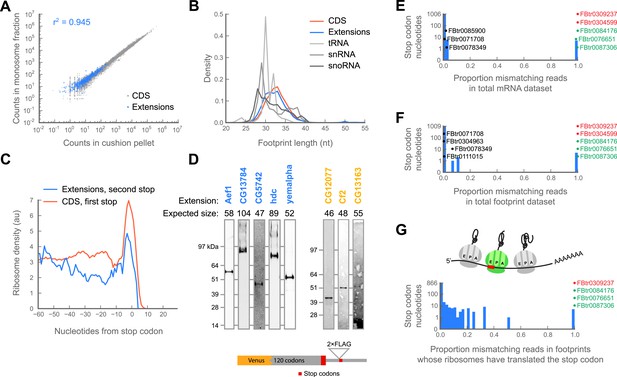
Translation downstream of the stop codon is due to readthrough.
(A) Ribosome footprint counts for each C-terminal extension are well correlated between samples prepared by sedimentation through sucrose cushions or by fractionation on sucrose gradients (blue). For comparison, footprint counts for annotated coding regions in each sample type are plotted (gray). The Pearson correlation coefficient (r2) for C-terminal extensions is shown. (B) Distributions of read lengths for footprints aligning to annotated coding regions (CDS, red) and to C-terminal extensions (blue) are similar, while lengths of footprints aligning to tRNAs, snRNAs, and snoRNAs are quite different. (C) Meta-gene average of ribosome density at the annotated stop codons of coding regions (red), or at the stop codons that terminate extensions (blue). Both averages show characteristic peaks of ribosome density above the stop codon, characteristic of translation termination. (D) Readthrough produces detectable protein products. Bottom: schema of reporters. Reporters containing the GFP variant Venus fused to the 120 C-terminal codons and entire endogenous 3’ UTR of a gene of interested were transfected into S2 cells. To facilitate detection of readthrough products, a double-FLAG epitope was inserted upstream of the stop codon (red) that terminates the putative extension. Top: reporters were immunoprecipitated with anti-GFP antibodies. Immunoprecipitates were then resolved by SDS-PAGE and western blotted with anti-FLAG antibodies to detect protein products of readthrough. Blue: names of genes containing extensions predicted to undergo readthrough. Yellow: names of genes containing novel extensions. (E) For each nucleotide in each stop codon that undergoes readthrough, we counted the fraction of reads containing nucleotide mismatches and present the data as a histogram. Transcripts containing stop codon nucleotides with significantly elevated mismatch rates are explicitly noted. Green: transcripts containing genomic polymorphisms that mutate one stop codon to another. Red: transcripts containing genomic polymorphisms that convert stop codons to sense codons. Black: other transcripts containing significantly elevated proportions of mismatches. (F) as in (E), but for ribosome-protected footprint data. (G) as in (F), but the analysis was restricted to the subset of footprints that both include the sequence of the stop codon and derive from ribosomes that have already translated the stop codon (top, green ribosome in cartoon).
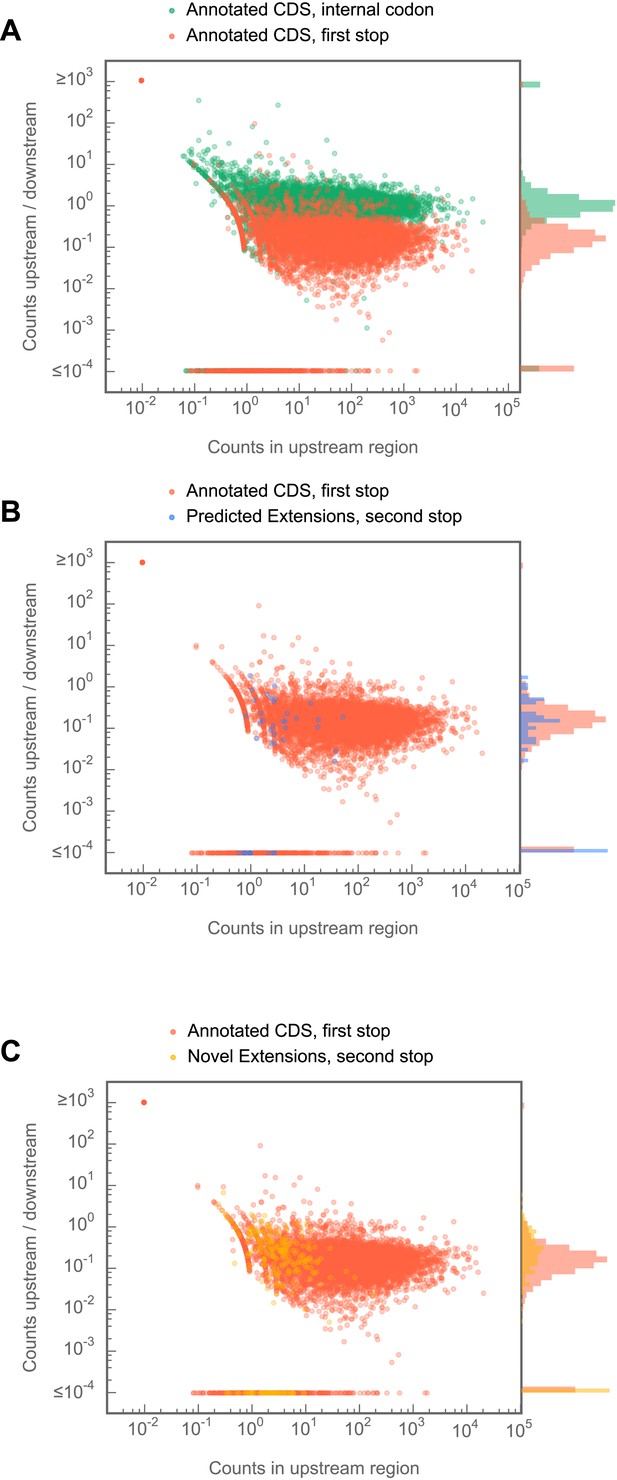
C-terminal extensions in Drosophila melanogaster show ribosome release typical of coding regions, but not of internal codons.
For each region of interest, the total number of reads aligning to 5 codon windows immediately upstream and downstream of that codon were tabulated, and the ratio (downstream counts/upstream counts) plotted against the total number of counts in the upstream window. (A) Comparison of release scores for termination codons of annotated coding regions and form randomly-selected codons internal to (i.e., at least 10 codons from the annotated start or end) annotated coding regions. (B) as in (A), but stop codons that terminate predicted extensions are compared against those that terminate annotated coding regions. (C) as in (A) but stop codons that terminate novel extensions are compared against those that terminate annotated coding regions. Source data may be found in supplementary table 2 (at Dryad: Dunn et al., 2013).
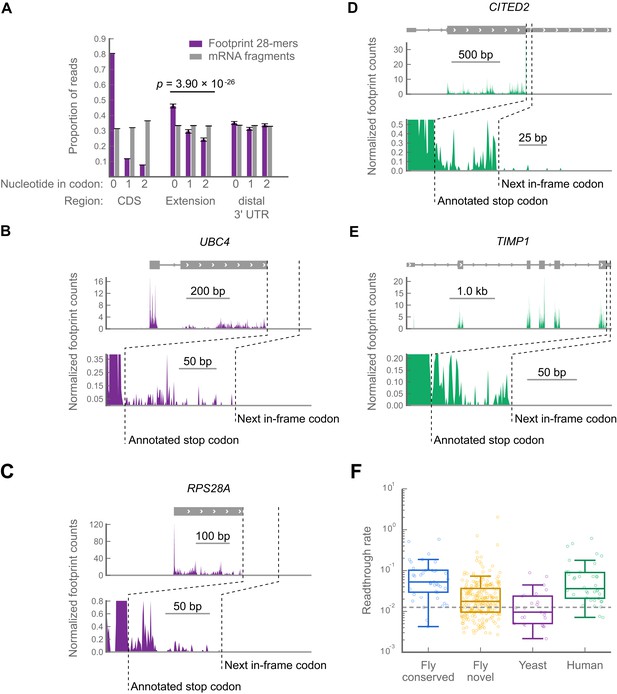
Readthrough occurs at specific stop codons in [psi-] yeast and in human foreskin fibroblasts.
(a) Triplet periodicity of 28-mers from yeast data in all non-overlapping coding regions (CDS), putative C-terminal extensions, and distal 3’ UTRs indicates that a signature of translation readthrough is visible in extensions on a bulk scale. Distal 3’ UTRs were estimated as 40 codon windows following putative extensions. Putative extensions and distal 3’ UTRs that overlap annotated coding regions, snoRNAs, snRNAs, tRNAs or 5’ UTRs were excluded from the analysis. (B and C) Examples of yeast transcripts that undergo readthrough, as in Figure 3B. (D and E) Examples of transcripts that undergo readthrough in human foreskin fibroblasts, as in Figure 3B. (F) Distribution of readthrough rates, by organism, for all extensions of sufficient length not to be covered by bleedthrough from termination peaks (‘Materials and methods’). Dashed line: fifth percentile of readthrough rate in conserved extensions in D. melanogaster, 1.2%. Source data may be found in supplementary tables 2, 3, and 4 (at Dryad: Dunn et al., 2013).
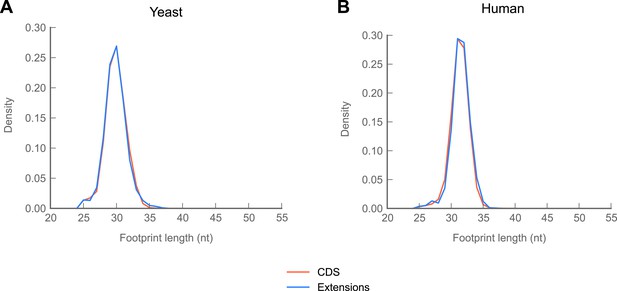
In yeast and humans, reads mapping to C-terminal extensions are drawn from the same length distribution as reads mapping to coding regions.
(A) Length distributions of reads mapping to coding regions and extensions in yeast. (B) Length distributions of reads mapping to coding regions and extensions in human foreskin fibroblasts.
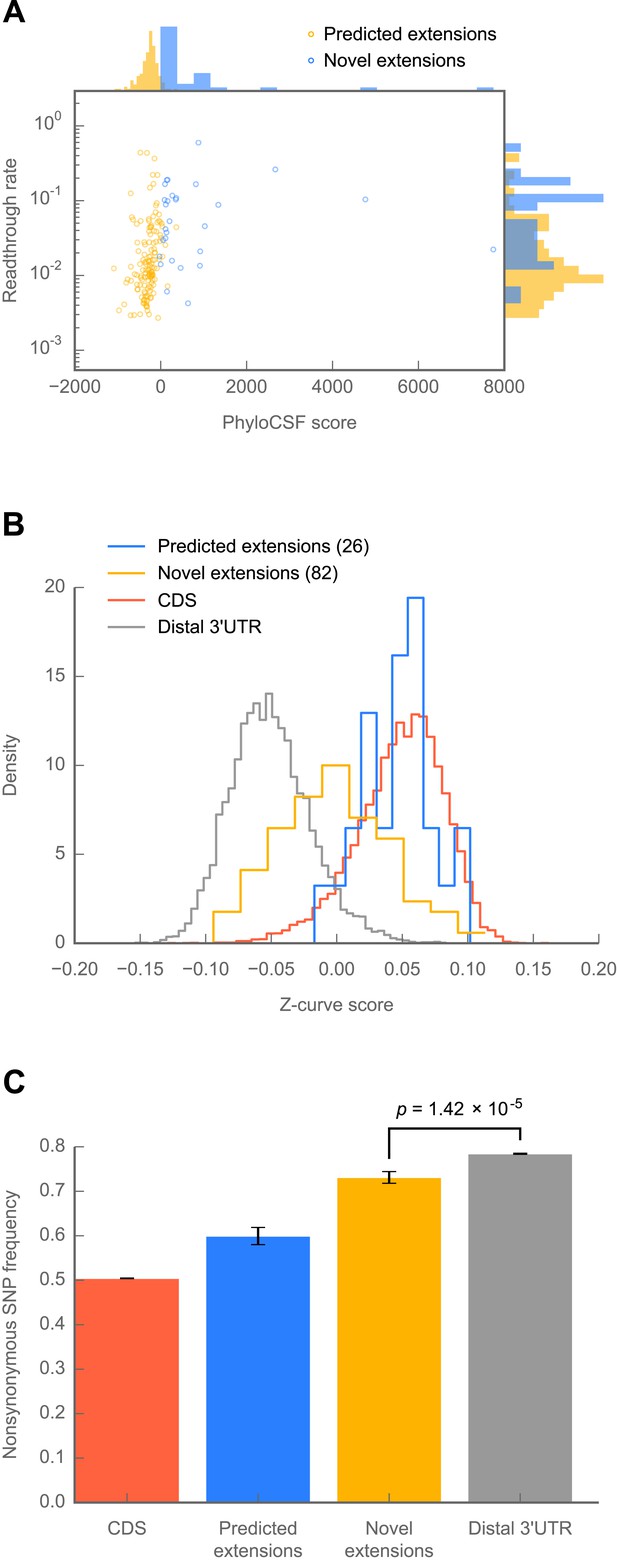
Novel C-terminal extensions in Drosophila melanogaster show signatures of selection within the melanogaster lineage.
(A) Scatter plot comparing readthrough rates for confirmed extensions against PhyloCSF scores. Blue: predicted extensions. Yellow: novel extensions. Datapoints with unreliably measured PhyloCSF scores or readthrough rates are not shown (‘Materials and methods’). (B) Z-curve classifier suggests that novel extensions have a nucleotide character intermediate between distal 3’ UTRs and coding regions. Histograms of Z-curve scores for 81-nucleotide windows drawn from annotated coding regions (CDS), distal 3’ UTRs, predicted extensions, and novel extensions. A single window was selected from each region 81 or more nucleotides long. Shorter regions were excluded from analysis, as they were empirically found to be noisy during classifier training. The Z-curve classifier was trained on windows drawn from CDS and distal 3’ UTRs as described in ‘Materials and methods’. (C) Novel extensions accumulate SNPs with a stronger preference than distal 3’ UTRs. Proportion of SNPs in CDS, predicted extensions, novel extensions, and distal 3’ UTRs which would be nonsynonymous if translated in frame. SNPs were obtained from wild isolates of wild-type flies by the Drosophila Population Genomics Project, and were downloaded from Ensembl (Flicek et al., 2013). Source data may be found in supplementary table 2 (at Dryad: Dunn et al., 2013).
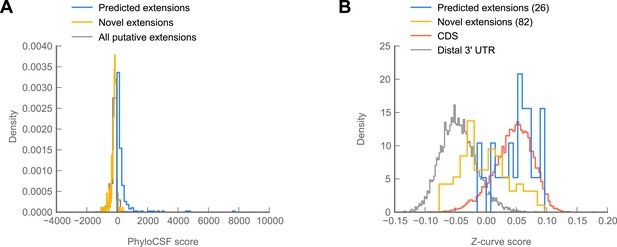
Novel C-terminal extensions in Drosophila melanogaster show signatures of selection within the melanogaster lineage.
(A) Histogram of PhyloCSF scores for C-terminal extensions. Blue: phylogenetically predicted extensions that were confirmed in our datasets. Yellow: unpredicted extensions discovered in our datasets. Gray: global distribution of all potential extensions. The distribution of novel extensions is not substantially different from the global distribution, suggesting that many of these extensions are not phylogenetically conserved beyond melanogaster. Source data may be found in supplementary table 2 (at Dryad: Dunn et al., 2013). (B) A second Z-curve classifier was trained on 81-nucleotide windows of coding regions, and 81-nucleotide windows of distal 3′ UTRs, but excluding the last 50 bases of annotated UTR to remove potential effects of polyadenylation signals upon classifier scoring. As in Figure 6B, predicted extensions overlay coding regions, and novel extensions display a significant shift in median from distal 3′ UTRs (p=3.81 × 10–22, Mann–Whitney U test), indicating the shift identified in Figure 6B is not due to polyadenylation signals.
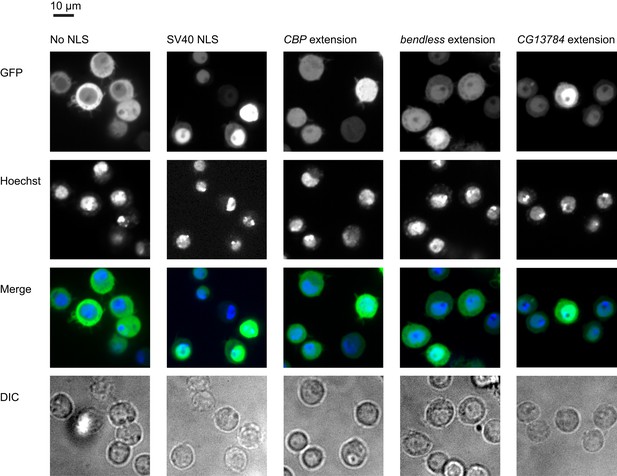
Extensions contain functional localization signals.
Ordinarily, a GFP-mCherry-GST reporter is excluded from the nucleus (first column). When an SV40 NLS is appended to the reporter, it is predominantly nuclear (second column). Three extensions also contain functional NLSes which at least partially relocalize the reporter to the nucleus when constitutively fused to it (remaining columns). First row: GFP reporter. Second row: nuclei stained with Hoechst. Third row: merged GFP and Hoechst. Fourth row: DIC.
Tables
Readthrough is differentially regulated between 0–2 hr embryos and S2 cells
https://doi.org/10.7554/eLife.01179.020| Gene ID | Alias | Embryo readthrough rate | S2 readthrough rate | PhyloCSF score | p value | log10 fold change | Direction of change |
|---|---|---|---|---|---|---|---|
| FBgn0036824 | CG3902 | 7.15E−01 | 2.46E−03 | −241.07 | 6.55E−10 | −2.46 | ↓ |
| FBgn0004362 | HmgD | 8.82E−03 | 1.21E−02 | −747.85 | 7.08E−07 | 0.14 | ↑ |
| FBgn0035432 | ZnT63C | 7.17E−03 | 2.71E−02 | 181.26 | 1.14E−06 | 0.58 | ↑ |
| FBgn0010409 | RpL18A | 1.39E−02 | 2.08E−03 | −197.78 | 5.85E−06 | −0.83 | ↓ |
| FBgn0039218 | Rpb10 | 5.18E−03 | 2.03E−02 | −333.38 | 8.06E−06 | 0.59 | ↑ |
| FBgn0038100 | Paip2 | 2.10E−02 | 4.60E−03 | −497.09 | 3.71E−05 | −0.66 | ↓ |
| FBgn0261790 | SmE | 7.55E−03 | 7.80E−04 | −530.28 | 9.60E−04 | −0.99 | ↓ |
| FBgn0030991 | CG7453 | 2.18E−01 | 5.28E−02 | −164.36 | 2.00E−03 | −0.62 | ↓ |
| FBgn0043796 | CG12219 | 2.85E−01 | 1.90E+00 | −27.83 | 2.11E−03 | 0.82 | ↑ |
-
For each transcript, the number of reads aligning to the CDS and corresponding extension were tabulated in both embryo and S2 cell datasets. p values for significant changes were calculated using Fisher’s Exact Test. The False Discovery Rate was controlled at 5% using the procedures of Benjamini and Hochberg (‘Materials and methods’), yielding nine transcripts with significant p values.
C-terminal extensions contain predicted functional peptide signals
https://doi.org/10.7554/eLife.01179.021| Gene ID | Alias | Extension coordinates | PhyloCSF score | Signal detected |
|---|---|---|---|---|
| FBgn0000173 | ben | X:13892649–13892781(+) | −302.18 | NLS |
| FBgn0005278 | Sam-S | 2L:113542–113647(+) | −195.30 | NLS |
| FBgn0026144 | CBP | X:7235840–7236599(+) | 128.52 | NLS |
| FBgn0031897 | CG13784 | 2L:7206347–7208015(−) | 4775.49 | NLS |
| FBgn0033712 | CG13163 | 2R:8209607–8209934(+) | −675.02 | NLS |
| FBgn0036272 | CG4300 | 3L:12265284–12265557(−) | −193.87 | NLS |
| FBgn0039213 | atl | 3R:20459429-20459720(+) | 28.43 | NLS |
| FBgn0260934 | par-1 | 2R:15370912–15371608(+) | 654.90 | NLS |
| FBgn0261606 | RpL27A | 2L:4457220–4457289^4457374–4457380(−) | −148.56 | NLS |
| FBgn0262114 | RanBPM | 2R:6322727–6323228(+) | 1045.90 | NLS |
| FBgn0031683 | CG4230 | 2L:5098384–5098573(+) | −5.34 | Transmembrane domain |
| FBgn0033712 | CG13163 | 2R:8209607–8209934(+) | −675.02 | Transmembrane domain |
| FBgn0035498 | Fit1 | 3L:4106386–4106518(+) | −323.36 | Transmembrane domain |
| FBgn0036980 | RhoBTB | 3L:20374798–20374821^20374891–20374982(+) | 154.91 | Transmembrane domain |
| FBgn0037321 | CG1172 | 3R:1221902–1222220(+) | −624.55 | Transmembrane domain |
| FBgn0040813 | Nplp2 | 3L:13350197–13350296(+) | −242.85 | Transmembrane domain |
| FBgn0053523 | CG33523 | 3L:5922386-5922854(+) | 383.85 | Transmembrane domain |
| FBgn0263864 | Ark | 2R:12913933-12914062(+) | −123.89 | Transmembrane domain |
| FBgn0039690 | CG1969 | 3R:25567115–25567154(+) | 11.52 | PTS1 |
| FBgn0035540 | Syx17 | 3L:4404848–4404983(+) | 290.83 | Farnesyltransferase signal |
-
Peptide sequences of C-terminal extensions were examined using various prediction servers (see ‘Materials and methods’). Those containing predicted features are shown here. NLS: nuclear localization signal. PTS1: peroxisome localization signal. Coordinates are 0-indexed and half-open. Splice junctions are denoted with carrots (‘^’). Strands are indicated in parentheses.
Additional files
-
Supplementary file 1
Alignment statistics.
Provides statistics on read alignments by sample and genomic region (e.g., CDS, 5’ UTR, 3’ UTR, intergenic, etc; A), as well as by sample and alignment type (e.g., chromosomal, spliced, unaligned; B).
- https://doi.org/10.7554/eLife.01179.023
-
Supplementary file 2
Oligonucleotides used in this study.
For readers who wish to implement the Drosophila ribosome profiling protocol.
- https://doi.org/10.7554/eLife.01179.024





