The splicing regulator PTBP2 controls a program of embryonic splicing required for neuronal maturation
Figures
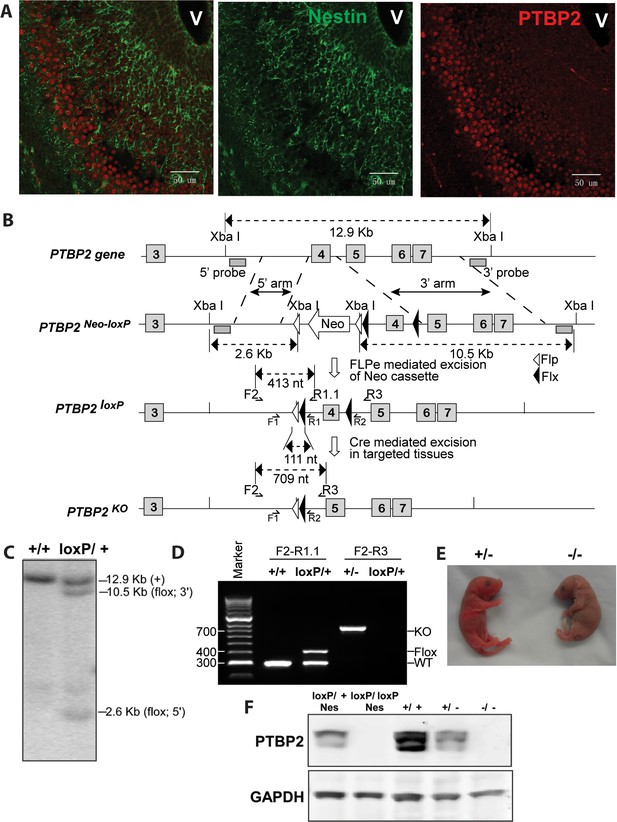
Generation and validation of the Ptbp2 conditional mutation.
(A) Expression of PTBP2 in maturing neurons of E15.5 brain. Nestin-positive cells (green, middle panel) of the subventricular zone express only limited PTBP2 (red, right panel), but as these cells mature and migrate to outer layers, PTBP2 is induced (Overlay panel to the left). V indicates the ventricle. (B) The targeting construct carrying loxP sites flanking exon 4 of Ptbp2 was integrated into the endogenous Ptbp2 locus by homologous recombination. The neomycin (Neo) selection cassette flanked by Frt sites was removed by crossing founder mice (Ptbp2Neo-loxP/+) with 129S4/SvJaeSor-Gt(ROSA)26Sortm1(FLP1)Dym/J mice (Jackson) carrying Flp recombinase. The resulting Ptbp2loxP/+ mice were bred to C57Bl/6 Cre transgenic strains to generate Ptbp2+/− animals in targeted tissues (C and D). Genotypes were verified by Southern blot of genomic DNA digested with XbaI (C), and by PCR (D). (E) Ptbp2 null mice display cyanosis and die immediately after birth due to respiratory failure. (F) Immunoblot for PTBP2 confirms that the disrupted allele eliminates expression of PTBP2.
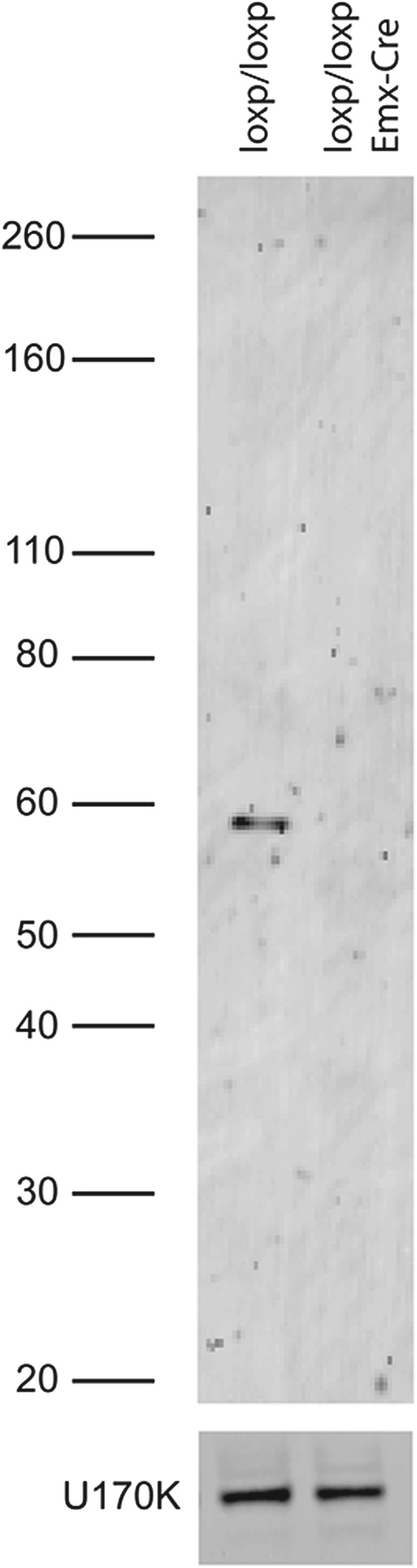
Immunoblot of PTBP2 in Emx-knockout and wild-type brain. Truncated proteins are not observed.
https://doi.org/10.7554/eLife.01201.004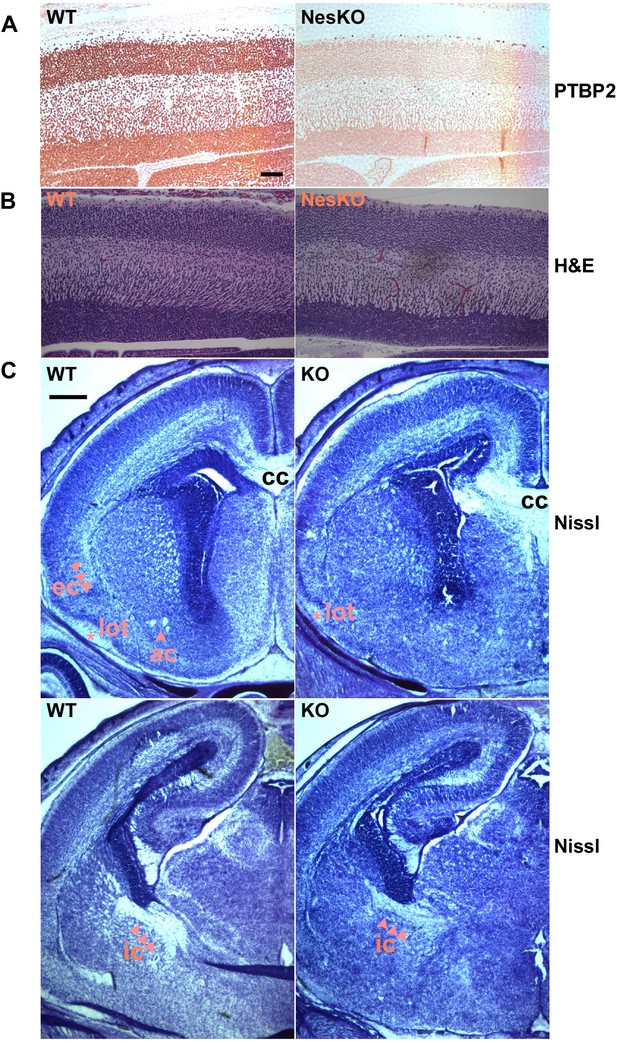
Embryonic brain development appears largely normal in the absence of PTBP2.
(A and B) Sagittal sections of E18.5 cortex from control and NesKO mice. (A) Staining with anti-PTBP2 antibody, control cortex shows PTBP2 expression in all neuronal nuclei, whereas NesKO cortex has lost PTBP2 expression in the vast majority of neurons. (B) H&E staining showing that NesKO cortex has similar thickness and largely normal morphology. Scale bar = 100 μM. (C) Nissl stained coronal sections of E18.5 cortex of wild-type and NesKO brain at two rostral–caudal planes. In the knockout brain, several major axonal tracks including the lateral olfactory tract (lot), internal capsule (ic) and external capsule (ec) were reduced, and the anterior commisure (ac) was missing. Corpus collosum, the major axon bundle connecting the two hemispheres, was present in the knockout brain. Scale bar = 250 μM. Staining was done on at least four knockout mice and four wild-type littermate controls from two litters.
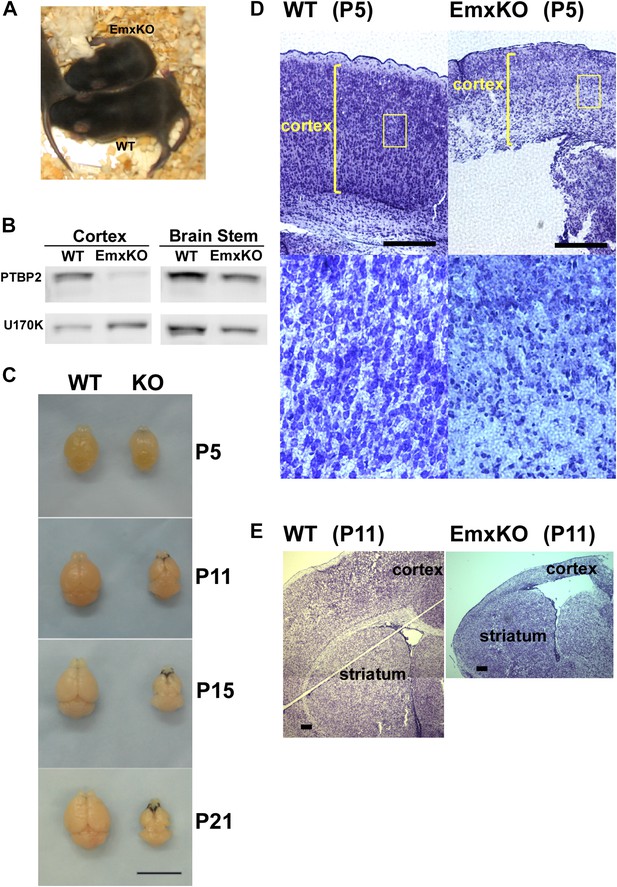
PTBP2 is required for postnatal cortical development.
(A) Ptbp2 EmxKO mutants display slow growth (shown at P11) and die around weaning. (B) Immunoblot for PTBP2 at P21 confirms that its expression is largely eliminated in cortex, but remains unchanged in other structures such as brain stem. (C) Postnatal development of the cortex was disrupted in EmxKO mice. Mutant cortical tissue failed to thicken as in wild type and degenerated. Scale bar = 1 cm. (D) Nissl stained coronal sections of control and EmxKO brain at P5 showing loss of cell density and degeneration. Enlarged panels show a loss of nuclei in the EmxKO tissue. (E) Coronal sections at P11, similar to (D). Scale bars in D and E indicate 0.5 mm. Staining was done on at least four knockout mice and four wild-type littermate controls from two litters.
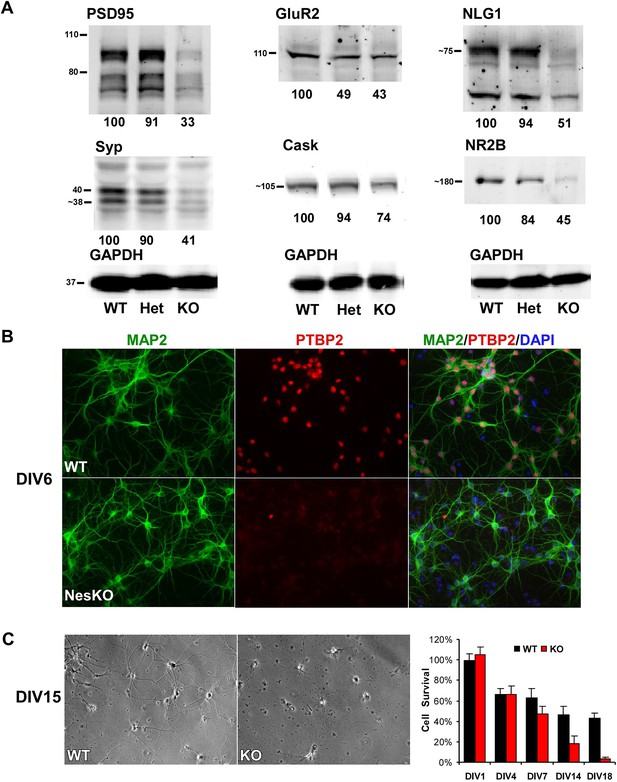
Loss of PTBP2 leads to reduced synaptic protein expression in vivo and to early cell death in primary cell culture.
(A) Immunoblots of whole brain lysates from wild type, heterozygous (+/−), and knockout (−/−) mouse at E18.5 show substantial reduction of synaptic proteins in knockout brain. (B) Dissociated cortical cultures (Days in vitro, DIV6) of wild-type and Ptbp2-knockout neurons stained for PTBP2 and the neurite marker MAP2. Neurons lacking PTBP2 survive plating and short-term culture with similar efficiency to wild type, and extend multiple MAP2-positive neurites. (C) PTBP2-deficient hippocampal cultures show increasing neuronal death starting from second week of culture and do not survive past 3 weeks. A phase contrast image of cells at DIV15 is shown, with the quantification of cell numbers plotted to the right. Starting viable cell numbers averaged 417 cells from wt and 442 cells from knockout based on cultures from four embryos of each genotype at DIV1. Percent Survival is the fraction of live cells at each time point relative to the wild type average at DIV1. Error bars are standard deviations derived from four cultures of each genotype at each time point.
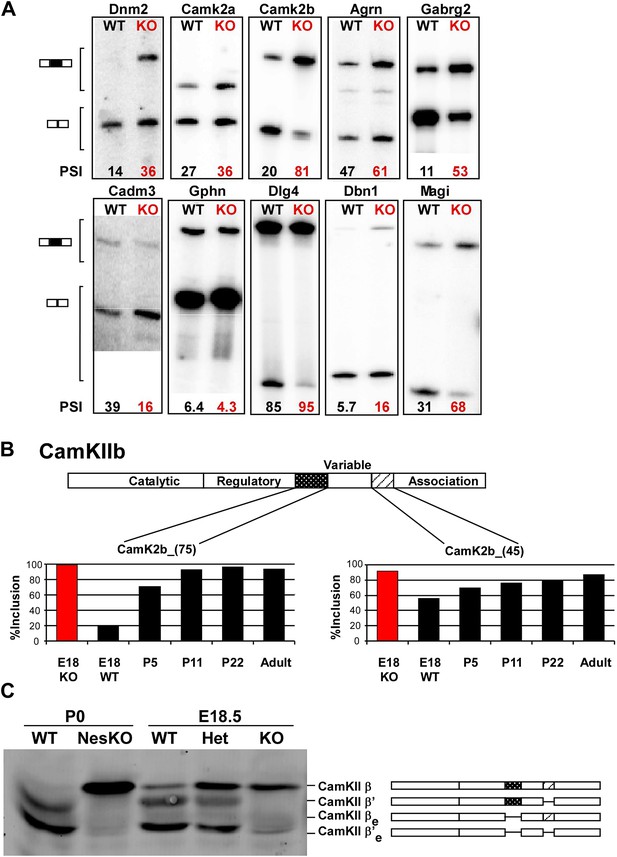
PTBP2 regulates a large set of splicing targets important to neuronal development and function.
(A) Sample RT-PCR gels showing aberrant splicing ratios for selected targets important for neuronal development and function. (B) Two alternative exons in Camk2b are developmentally regulated. In plots of RT/PCR measurements, normal mouse brain shows low inclusion of these exons in the embryo (E18) and a postnatal increase (P5, P11 and P22) in splicing to near full inclusion in adult brain. In the PTBP2 null brain these exons exhibit adult levels of splicing at E18. (C) The premature switch in CAMK2B protein isoforms is observed by immunoblot in both Ptbp2 null (KO) and NesKO mice (E18.5-P0).
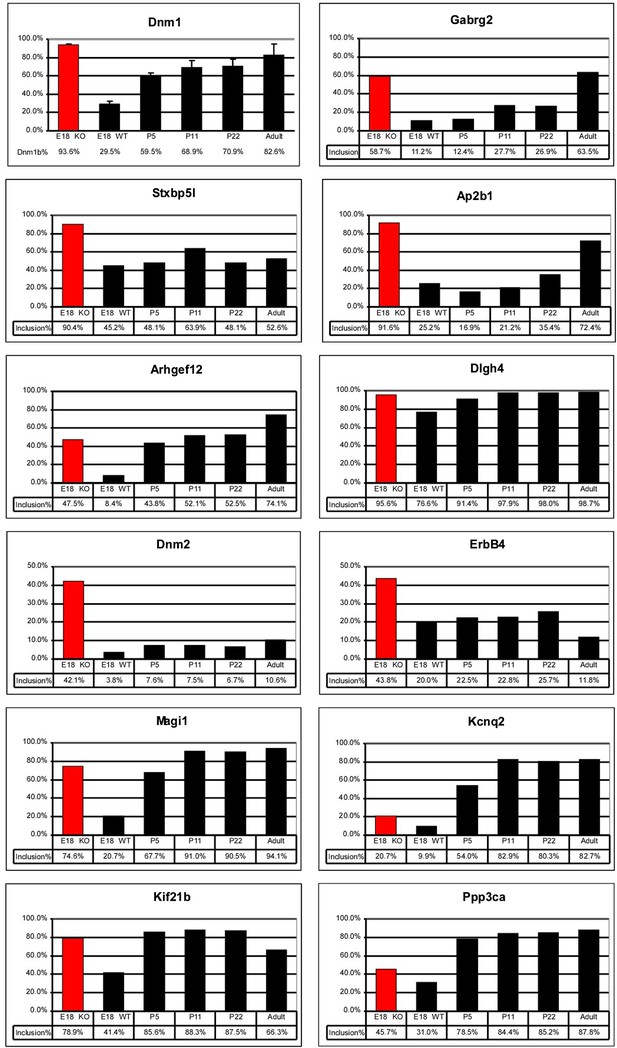
Quantification of RT/PCR analysis for multiple exons mis-regulated in Ptbp2 Nes-KO brain.
In many cases, exon inclusion is abnormally high in the embryo for an exon that normally increases in postnatal development.
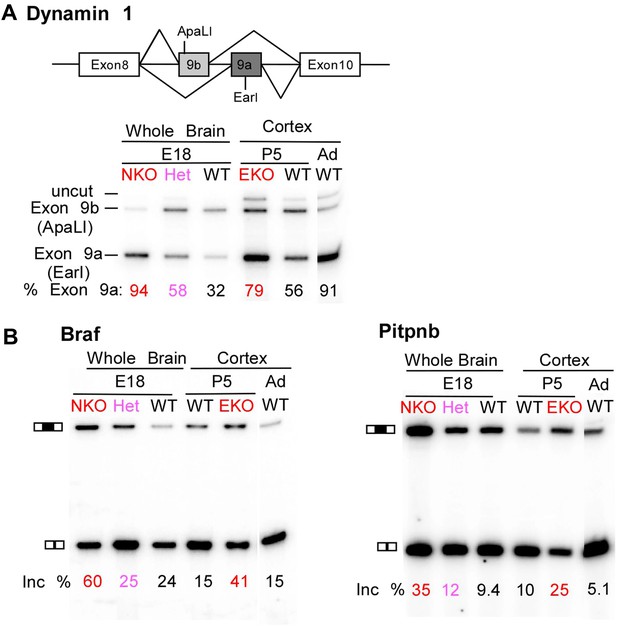
Premature and aberrantly high splicing of PTBP2 target exons in E18 Nes-KO whole brain and P5 Emx-KO cortex.
(A) RT/PCR of Dynamin1. Dynamin1 contains a pair of mutually exclusive exons 9a and 9b that normally switch from 9b to 9a during development. In both the Nes-KO and Emx-KO there is a premature switch to the 9a isoform. Note that the heterozygous Nes-KO exhibits an intermediate level of exon 9a splicing. The high level of 9a splicing can be seen in the wild-type adult cortex (Ad WT). (B) RT/PCR of Braf (left) and Pitpnb (right). Braf and Pitpnb each contain a PTBP2 repressed exon that is spliced at aberrantly high levels in the Ptbp2 KO mice. Braf exon 9 and Pitpnb exon 11 are spliced into the mRNAs two to three times more frequently in the embryonic (E18) or postnatal (P5) mutant mice (NKO and EKO), and at higher levels than normally seen in the wild-type adult (Ad WT). The percent exon inclusion is shown at the bottom.
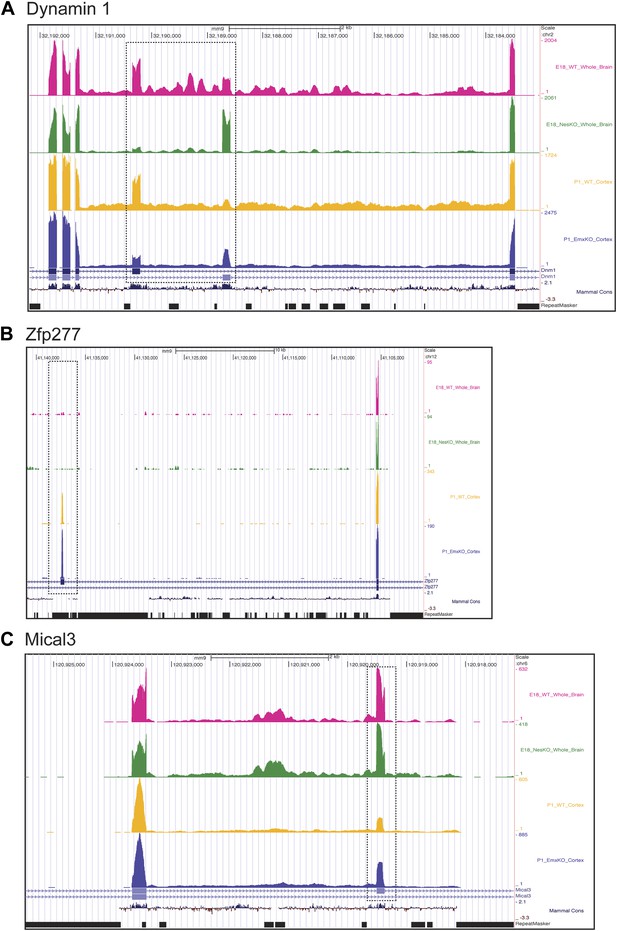
Exons can respond to the loss of PTBP2 differently in E18 whole brain or P1 cortex.
Genome browser tracks of aligned RNAseq reads from E18 whole brain, NesKO (green) and wild type (pink), and from P1 cortex, EmxKO (blue) and wild type (yellow). (A) For Dynamin 1, exons 9a and 9b are boxed. The two knockout samples show a different degree of shift towards the use of downstream exon 9a. (B) An exon in the Zinc finger protein Zfp277 (boxed) is entirely excluded in E18 brain. In postnatal cortex this exon has been strongly induced, but is still regulated by PTBP2 as indicated by the stronger induction of the exon in the Emx-KO. (C) An exon in the microtubule associated monooxygenase Mical3 is fully spliced at E18, regardless of the presence of PTBP2. In postnatal cortex this exon becomes repressed and this repression is in part dependent on PTBP2.
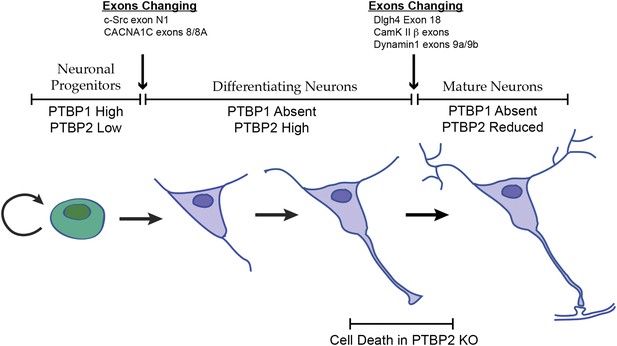
Changes in the expression of the two PTB proteins define three splicing regulatory states during neuronal differentiation.
Neuronal progenitor cells primarily express PTBP1. When these cells are induced to differentiate, PTBP1 is repressed and PTBP2 is induced. This switch in RNA binding proteins causes changes in the splicing of exons that are more sensitive to PTBP1 such as exon 8 of the Cacna1c calcium channel transcript. Other exons that are sensitive to both proteins maintain their repression during the early stages of neuron differentiation, when PTBP2 is high. When these cells finally mature and form synapses, PTBP2 is downregulated. This leads to changes in a splicing regulatory program that includes exons in Cam Kinase 2 and Dynamin 1. These exons are found in mRNA isoforms associated with adult brain and are precociously expressed in PTBP2 knockout brains, leading to cell death prior to final maturation.
Additional files
-
Supplementary file 9
The genomic DNA sequence of PTBP2 loxP.
- https://doi.org/10.7554/eLife.01201.015





