Extracellular matrix signatures of human mammary carcinoma identify novel metastasis promoters
Figures
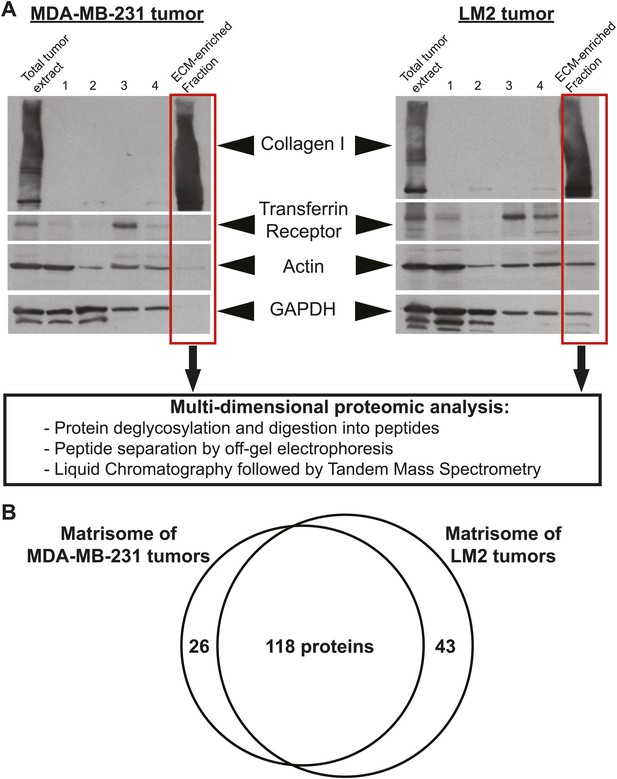
Enrichment of extracellular matrix proteins from human mammary tumor xenografts.
(A) The sequential extraction of intracellular components was monitored by immunoblotting for GAPDH (cytosol), the transferrin receptor (plasma membrane), and actin (cytoskeleton). The remaining insoluble fraction was highly enriched for ECM proteins (collagen I panel) and largely depleted for intracellular components. The ECM-enriched fraction obtained is subsequently submitted to multidimensional proteomic analysis and the matrisome (ECM composition) of each tumor type is defined as the ensemble of proteins present in two replicate samples and with at least two peptides in one of the two replicates. (B) Venn diagram represents the comparison of the matrisomes of MDA-MB-231 and LM2 tumors. In addition to 118 ECM proteins detected in both tumor types, we identified 26 proteins specific to poorly metastatic (MDA-MB-231) tumors and 43 proteins characteristic of highly metastatic tumors (LM2).
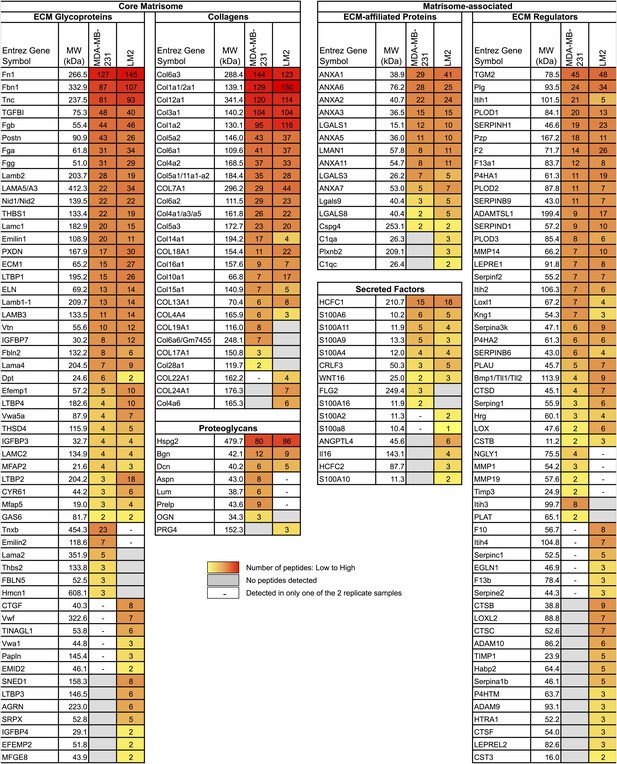
Comparison of the matrisomes of MDA-MB-231 tumors and LM2 tumors identifies ECM proteins characteristic of poorly and highly metastatic tumors.
Color code represents the number of unique peptides for each protein from poorly metastatic (MDA-MB-231) or highly metastatic (LM2) human mammary tumors. Values used to generate the figure were extracted from Figure 2—source data 1, columns P and AA (number of peptides). Grayed cells indicate that no peptides were detected in either of the two replicate samples. A dash (–) indicates that the protein was detected in only one of the two replicate samples of a given tumor type or with only one peptide in both replicate samples.
-
Figure 2—source data 1
Complete MS data set of ECM-enriched fraction from MDA-MB-231 and LM2 human mammary tumor xenografts.
- https://doi.org/10.7554/eLife.01308.005
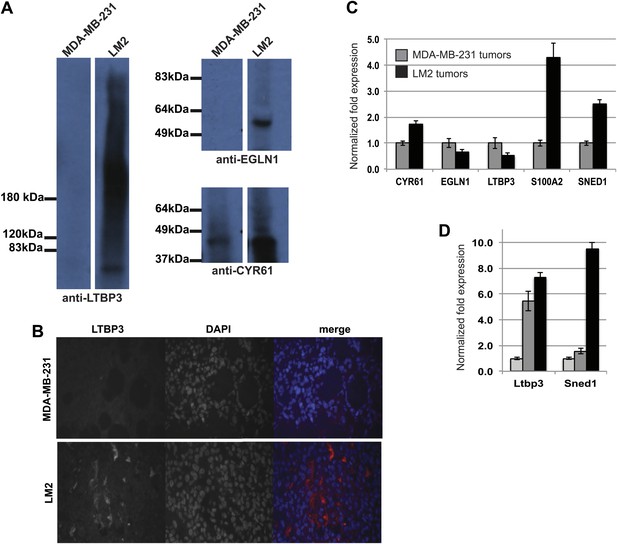
Validation of the differential expression of proteins identified by proteomics.
(A) Immunoblots were performed on ECM-enriched fractions prepared from MDA-MB-231 or LM2 tumors to confirm the differential expression of some of the proteins identified by proteomics. (B) Immunohistochemistry was performed on MDA-MB-231 or LM2 tumor sections to evaluate the expression and localization of LTBP3. LTBP3 is detected in LM2 but not MDA-MB-231 tumors. Note the extracellular distribution of LTBP3. (C) RT-qPCR was used to monitor mRNA expression levels in LM2 tumors as compared to MDA-MB-231 tumors and was conducted in triplicate in two independent tumors per tumor type. Bar charts represent normalized expression of the genes in LM2 tumors (black) as compared to MDA-MB-231 tumors (gray). Actin expression was used to normalize RT-qPCR data. Data are presented as means ±SEM. (D) RNA was extracted from normal murine mammary gland (light grey) or from autochtonous murine mammary tumors (MMTV-PyMT transgenic mice: adenoma, dark grey or adenocarcinoma, black). RT-qPCR was conducted in triplicate. Bar charts represent normalized expression of the genes. Actin expression was used to normalize RT-qPCR data. Data are presented as means ± SEM.
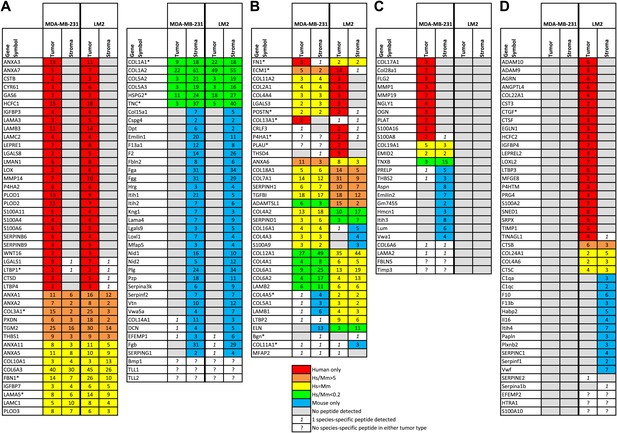
The tumor extracellular matrix is secreted by both tumor cells and stromal cells and differs with the tumor’s metastatic potential.
(A) Proteins expressed by both tumor types and by the same compartment in the two tumor types. (B) Proteins expressed by both tumor types but by different compartments. (C) Proteins secreted by MDA-MB-231 tumors and not by LM2 tumors. (D) Proteins secreted by LM2 tumors and not by MDA-MB-231 tumors. The number of peptides detected for each protein is indicated. For the proteins secreted by both the tumor cells and the stromal cells, the number of peptides listed corresponds to the number of human (tumor-derived)- or murine (stroma-derived)-specific peptides. For the proteins secreted by only one compartment, the number of peptides includes both species-specific and indistinguishable peptides. Proteins are sorted by tumor type and by their origins: tumor (red), stroma (blue), or both (yellow: similar abundance of the human and mouse proteins, orange: human form is at least five times more abundant than the mouse form, green: the mouse form is at least five times more abundant). To determine the relative contributions of the tumor and stromal cells to the secretion of ECM proteins, human-to-mouse peptide abundance ratios were calculated using the values indicated in column P and AB for the MDA-MB-231 and LM2 tumors respectively (Figure 3—source data 1). Proteins for which different isoforms have been detected are indicated with an asterisk (*) and, for simplicity, isoforms are combined here, their UniProt accession numbers can be found in Figure 3—source data 1, column AP. In a few instances, the origin of the protein could not be determined due to the lack of species-specific peptides; these proteins are indicated with a question mark (?).
-
Figure 3—source data 1
Complete MS data set of ECM-enriched fraction from MDA-MB-231 and LM2 human mammary tumor xenografts taking into account the origin of matrisome proteins.
- https://doi.org/10.7554/eLife.01308.008
-
Figure 3—source data 2
Detailed list of all of the confidently identified peptide spectrum matches (PSMs) from the LC-MS/MS runs of the 11 fractions resulting from off-gel electrophoresis of each of the two MDA-MB-231 tumor replicates and each of the two LM2 tumor replicates.
The four tables containing all of the confidently identified peptide spectrum matches (PSMs) from the LC-MS/MS runs of the 11 fractions resulting from off-gel electrophoresis of each of the four tumor samples can be downloaded from MassIVE (http://massive.ucsd.edu) using the identifier: MSV000078535. The file should be accessible at ftp://MSV000078535:a@massive.ucsd.edu/results.
- https://doi.org/10.7554/eLife.01308.009
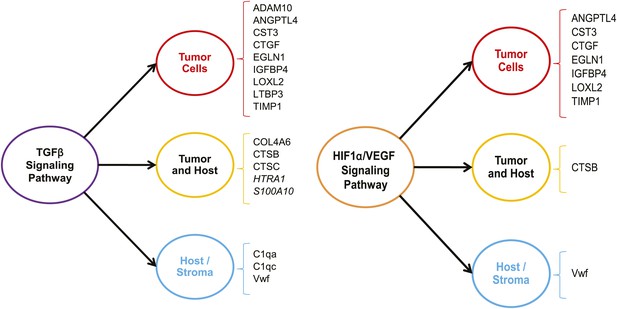
The TGFβ and the HIF1α/VEGF pathways are up-regulated in highly metastatic mammary tumors.
The 43 ECM and ECM-associated proteins (tumor- and/or stroma-derived) unique to highly metastatic mammary tumors were uploaded to the Ingenuity Pathway Analysis software and queried for common upstream regulators. The analysis revealed an enrichment of TGFβ and HIF1α/VEGF targets within our ECM signature of highly metastatic mammary tumors.
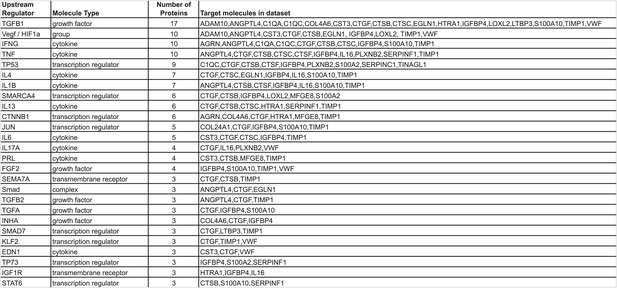
Ingenuity Pathway Analysis.
Identification of regulators (cytokines, growth factors, G-protein-coupled receptors, transcription regulators, or transmembrane receptors) controlling the expression of subsets of the 43 ECM genes characteristic of highly metastatic tumors was searched. Regulators upstream of at least three ECM genes are listed.
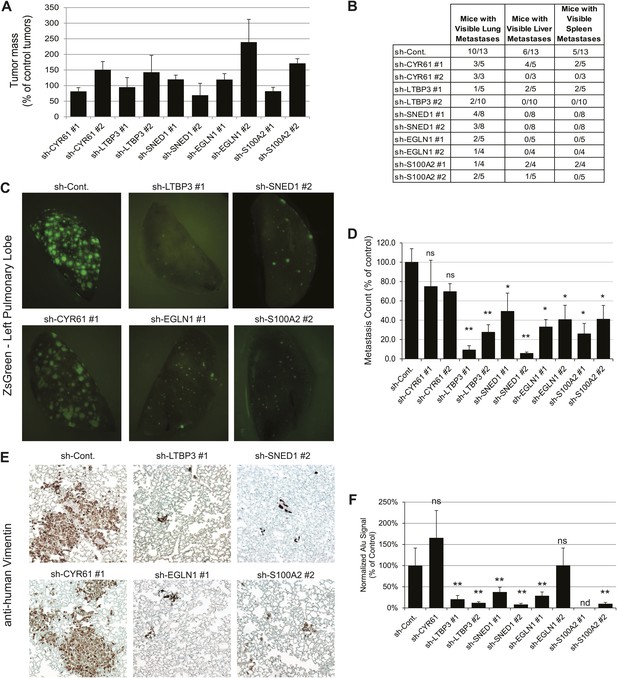
Tumor-cell-derived ECM proteins influence the metastatic dissemination of tumor cells to distant organs.
Mice were injected orthotopically with control or knockdown LM2 cells. Tumors were allowed to grow for 7 ± 0.5 weeks. Number of mice per condition is indicated in Figure 5B. (A) ECM protein knockdown does not inhibit primary tumor growth. At sacrifice, control and knockdown primary tumors were weighed. Bar chart represents the mass of knockdown tumors as a percentage of that of control tumors ± SEM. Student’s t test was performed and none of the genes affected significantly and consistently primary tumor growth. (B) LM2 control tumors metastasize to the lungs, liver and spleen. The number of mice that presented with visible metastases in the indicated organs is indicated. (C) Representative pictures of whole left pulmonary lobe from LM2 (control or knockdown)-tumor-bearing mice with ZsGreen-positive metastatic foci. (D) Numbers of ZsGreen-positive metastatic foci in the left pulmonary lobe were counted. Data are presented as percentage of control ± SEM (Student’s t test, *p<0.05, and **p<0.01). Number of animals per group is indicated in Figure 5B. (E) Lung sections were stained with a human-specific anti-vimentin antibody to detect human tumor cells in the murine lung. (F) Alu PCR was performed on genomic DNA extracted from the lungs of control or knockdown tumor-bearing mice. Data are presented as normalized Alu signal as compared to murine actin signal and as percentage of control ± SEM (Student’s t test, *p<0.05, **p<0.01, ns: not significant, and nd: not determined). Number of animals per group is indicated in Figure 5B.
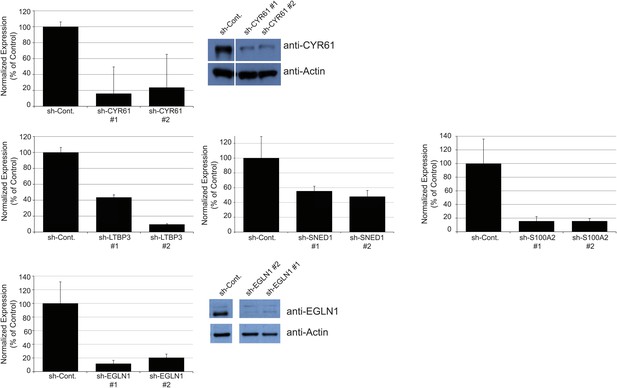
Establishment of stable LM2 knockdown cells.
RT-qPCR was used to monitor knockdown efficiency in cells in culture and was conducted for each cell line in triplicate on three independent sets of RNA collected at three consecutive cell passages. Bar charts represent normalized expression of the genes in knockdown cell lines relative to the control cell line (sh-Cont.). Actin expression was used to normalize RT-qPCR data. Data are presented as means ± SEM. The two hairpins giving the most efficient knockdown are shown. When antibodies were available, knockdown efficiency was also monitored by immunoblotting (see CYR61 and EGLN1 panels).
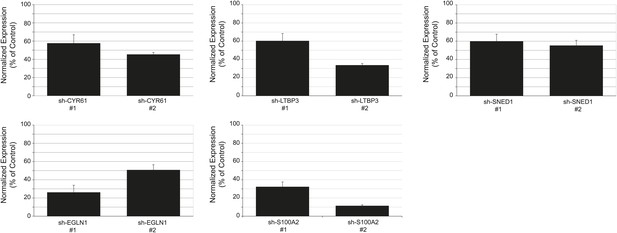
Persistence of gene expression knockdown in tumors.
RT-qPCR was used to monitor knockdown persistence in tumors and was conducted on RNA extracted from individual control or knockdown tumors and in triplicate for each sample. Bar chart shows normalized expression of the genes in knockdown tumors relative to control tumors. Actin expression was used to normalize RT-qPCR data. Data are presented as means ± SEM. The number of mice per group ranged from 3 to 13 and is indicated in Figure 5B.
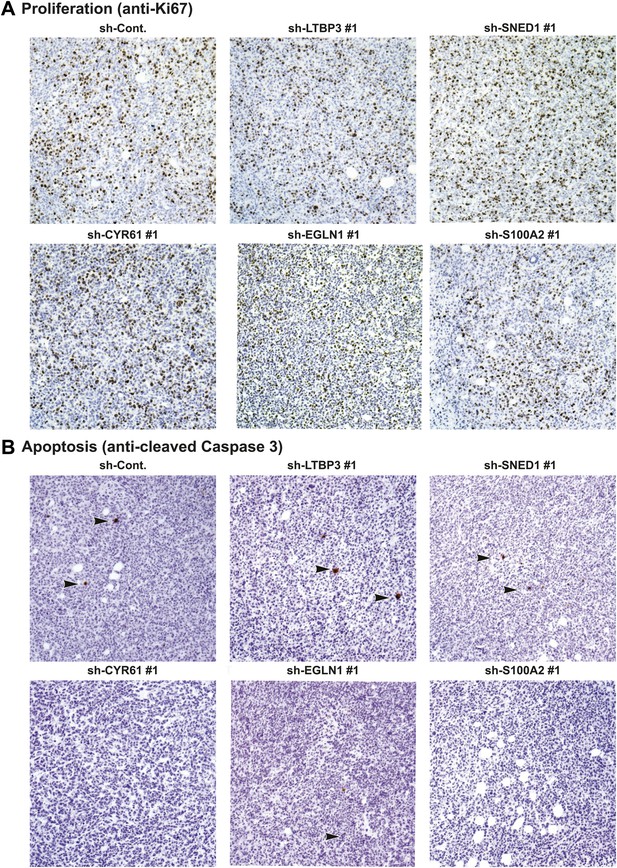
Tumor-cell-derived ECM proteins do not influence proliferation or apoptosis of primary mammary tumors.
Primary tumor sections were stained with the anti-Ki67 antibody (A) to evaluate cell proliferation or anti-cleaved caspase 3 (B) to evaluate apoptosis. Knockdown of tumor cell-derived ECM proteins (CYR61, LTBP3, SNED1, EGLN1, or S100A2) did not affect tumor cell proliferation (Ki67 staining, A) or apoptosis (cleaved caspase 3, B).
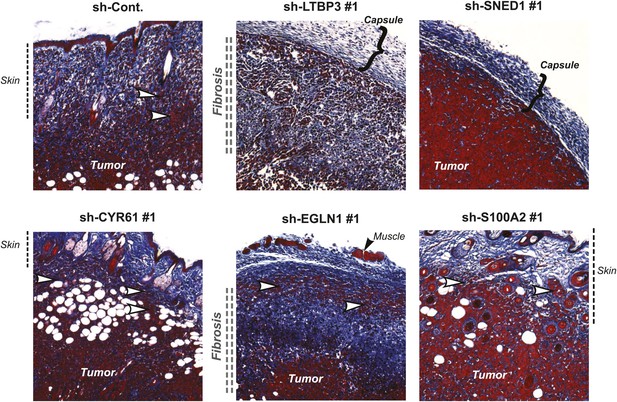
Tumor-cell-derived ECM proteins influence the invasiveness of primary mammary tumors.
Primary tumor sections were stained with Masson’s trichrome (blue: collagen fibers, red: cells) to evaluate fibrosis (indicated by vertical double-dashed line) and encapsulation (bracket) or invasiveness (white arrowhead) into the skin of the primary tumors. Note that tumors in which LTBP3 or SNED1 are knocked down are less invasive and more encapsulated than in the control tumors. CYR61, EGLN1, or S100A2 knockdown did not affect tumors’ invasiveness.
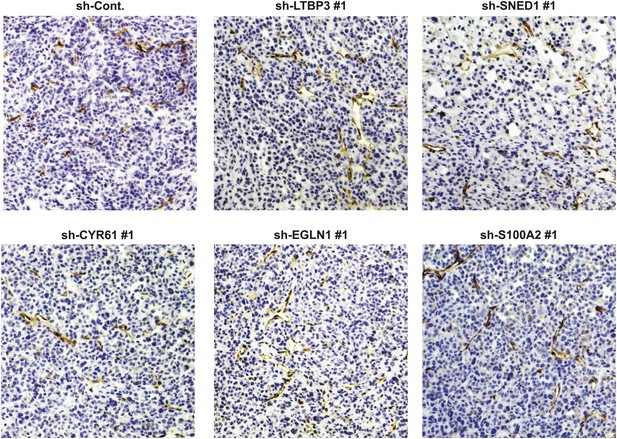
Tumor-cell-derived ECM proteins do not influence vascularization of primary mammary tumors.
Primary tumor sections were stained with an anti-CD31 (endothelial cell marker) to visualize tumor vasculature.
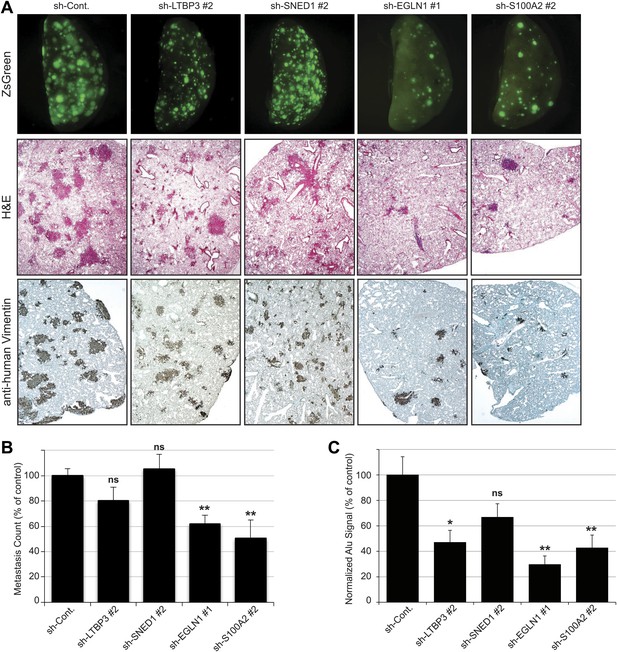
Tail vein metastasis assay.
Control or knockdown cells were injected via the tail vein and the formation of lung metastases was evaluated. (A) Upper panel: representative pictures of the whole left pulmonary lobe with ZsGreen-positive metastatic foci from mice injected with LM2 (control or knockdown) cells (upper panel). Lung sections were stained with hematoxylin and eosin (H&E, middle panel) or with human-specific anti-vimentin antibody (lower panel) to visualize the metastatic foci. (B) Numbers of ZsGreen-positive metastatic foci in the left pulmonary lobe. Data are presented as percentage of control ± SEM (Student’s t test, *p<0.05, **p<0.01, ns: not significant). Number of animals per group: sh-Cont.: 17 mice, sh-LTBP3: 10 mice, sh-SNED1: 10 mice, sh-EGLN1: 8 mice, sh-S100A2: 8 mice. (C) Alu PCR was performed to monitor the presence of human tumor cells in the murine lung. Data are presented as percentage of control ± SEM (Student’s t test, *p<0.05, **p<0.01, ns: not significant). Number of animals per group: sh-Cont.: 17 mice, sh-LTBP3: 10 mice, sh-SNED1: 10 mice, sh-EGLN1: 8 mice, sh-S100A2: 8 mice.
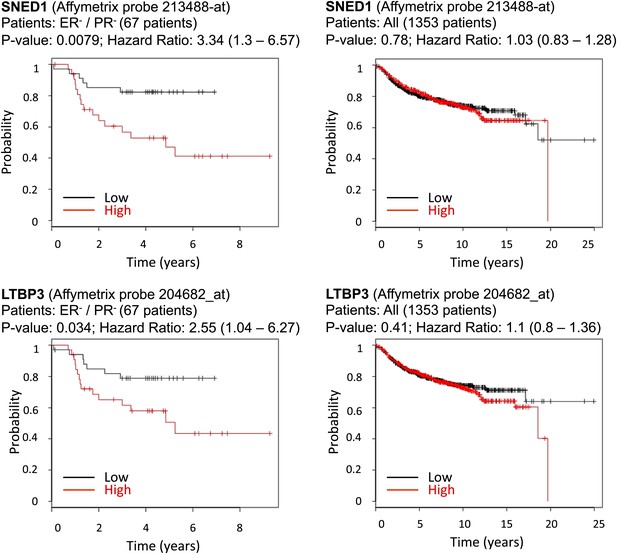
Correlation between SNED1 and LTBP3 expression and distant-metastasis-free survival in breast cancer patients.
We tested the predictive value of the four ECM genes studied using the online assessment tool Kaplan–Meier Plotter. Whereas the expression of none of the genes studied correlated with distant metastasis-free survival for the entire population of breast cancer patients, SNED1 and LTBP3 expression inversely correlated with the survival of estrogen-receptor-negative and progesterone-receptor-negative (ER−/PR−) breast cancer patients.
Additional files
-
Supplementary file 1
Cell Profiler pipeline to quantify ZsGreen-positive metastatic foci. The Cell Profiler pipeline designed to quantify ZsGreen-positive metastatic foci in whole organs is provided as a .cp file. The file can be opened in any text editor software or using Cell Profiler.
- https://doi.org/10.7554/eLife.01308.020
-
Supplementary file 2
shRNA sequences.
- https://doi.org/10.7554/eLife.01308.021
-
Supplementary file 3
qPCR primer sequences.
- https://doi.org/10.7554/eLife.01308.022





