The life cycle of Drosophila orphan genes
Figures
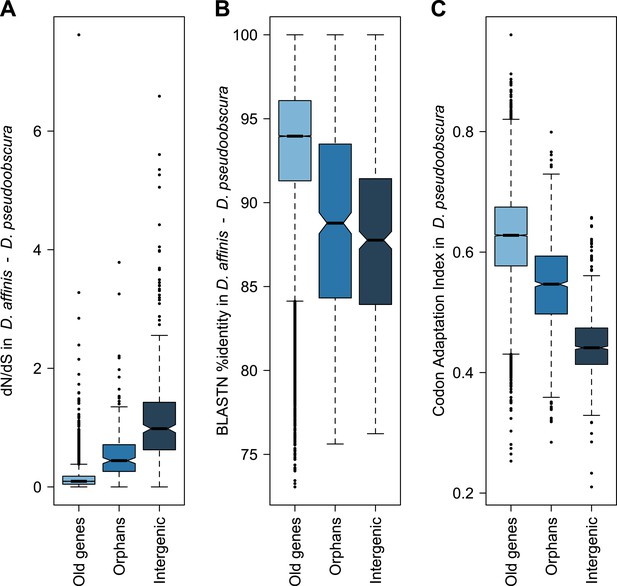
Orphans are subject to purifying selection.
(A) dN/dS of D. pseudoobscura and D. affinis orthologs. dN/dS is lowest for old genes, but also orphan genes have dN/dS smaller than one. A comparison of orphans and intergenic regions shows that dN/dS for orphans is significantly smaller (Mann–Whitney test, p=9.5 × 10−10), indicating purifying selection on orphan genes. Intergenic regions were of similar length and chromosomal position as the orphan genes. (B) Sequence similarity in HSPs obtained from BLASTing D. pseudoobscura genes against the D. affinis genome. Orphans are more conserved than intergenic regions (Mann–Whitney test, p=0.00238) and less conserved than old genes (Mann–Whitney test, p<1.0 × 10−15). (C) Codon usage was measured by the Codon Adaptation Index (Sharp and Li, 1987). The codon usage of orphans is significantly higher than that of intergenic regions (Mann–Whitney test, p<1.0 × 10−15) indicating that orphans are subject to purifying selection. In comparison to old genes, orphans have a significantly lower codon usage bias (Mann–Whitney test–p<1.0 × 10−15). Overall, all three analyses demonstrate that orphans are not annotation artifacts, but evolutionary conserved genes.
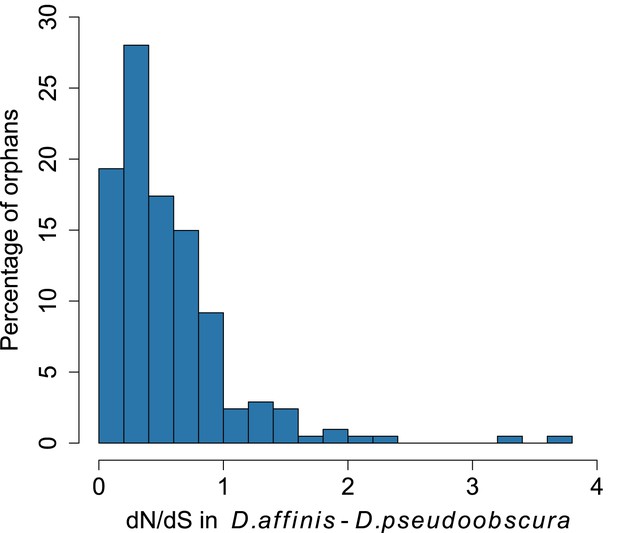
Distribution of dN/dS for orphan genes.
Most orphans have dN/dS lower than 1, consistent with the hypothesis of purifying selections acting on these genes.
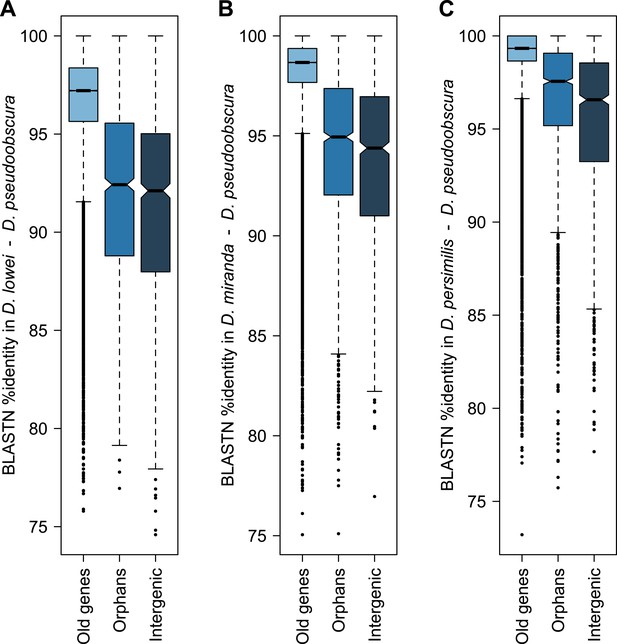
Conservation of orphans in the obscura group.
Sequence similarity of old genes, orphans and random intergenic region obtained from BLASTing D. pseudoobscura genes against the genomes of D. lowei (A), D. miranda (B) and D. persimilis (C). Orphans are significantly more conserved than random intergenic regions in D. lowei (Mann–Whitney test, p=0.00857), D. miranda (Mann–Whitney test, p<0.00034) and D. persimilis (Mann–Whitney test, p<2.8 × 10−13). These results are consistent with purifying selection acting on orphans.
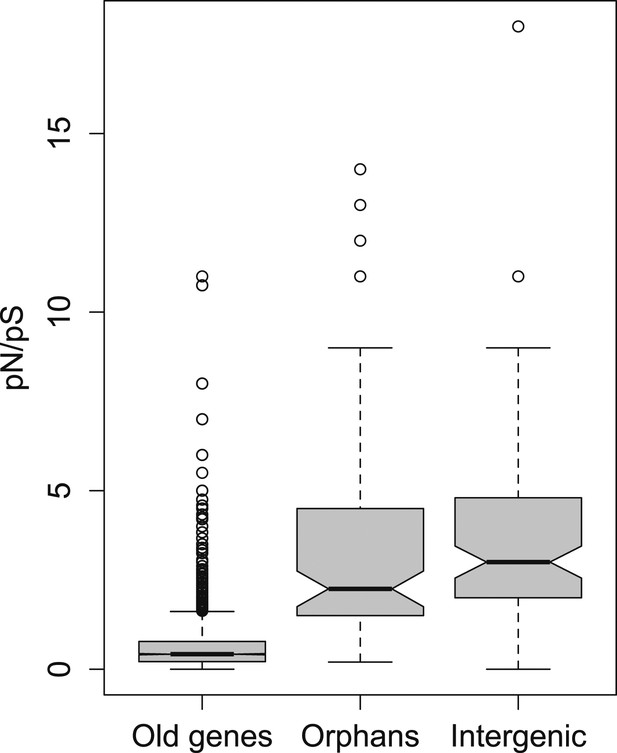
pN/pS for old genes, orphans, and intergenic regions.
Orphans show a pN/pS intermediate between old genes and intergenic regions. Nevertheless, pN/pS is significantly smaller for orphans compared to intergenic regions (Mann–Whitney test, p<1.0 × 10−15), indicating coding purifying selection acting on orphans.
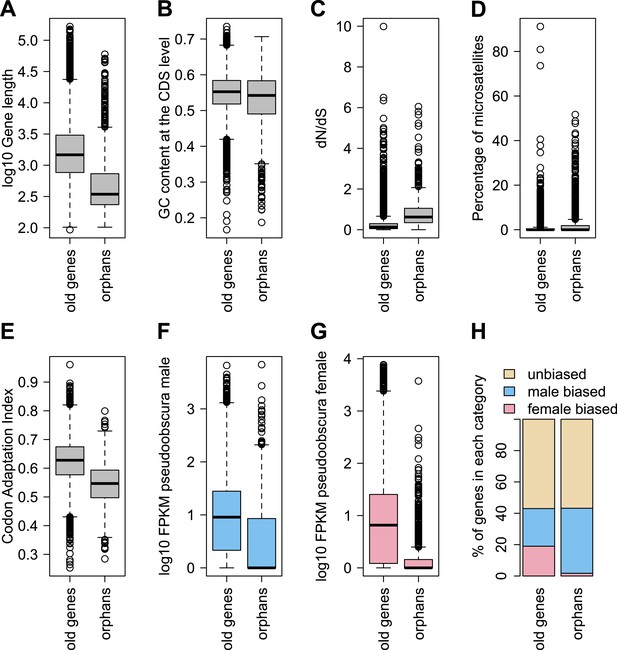
Comparison of orphans and genes conserved among 10 Drosophila species outside of the obscura group.
Orphans differ from old genes in various features: (A) gene length (B) GC content, (C) dN/dS (D) percentage of microsatellites in coding sequence (E) Codon Adaptation Index (F) gene expression level in D. pseudoobscura males (G) gene expression level in D. pseudoobscura females (H) sex-biased expression. Orphans are shorter (Mann–Whitney test, p<1.0 × 10−15), have lower GC content (Mann–Whitney test, p=3.9 × 10−7), lower codon usage bias (Mann–Whitney test, p<1.0 × 10−15), lower expression (Mann–Whitney test, p<1.0 × 10−15), higher proportion of microsatellites (Mann–Whitney test, p=1.8 × 10−4) and higher dN/dS (Mann–Whitney test, p<1.0 × 10−15) compared to old genes. Moreover, orphans are more enriched in male-biased genes compared to old genes (χ2-test, p<1.0 × 10−15).
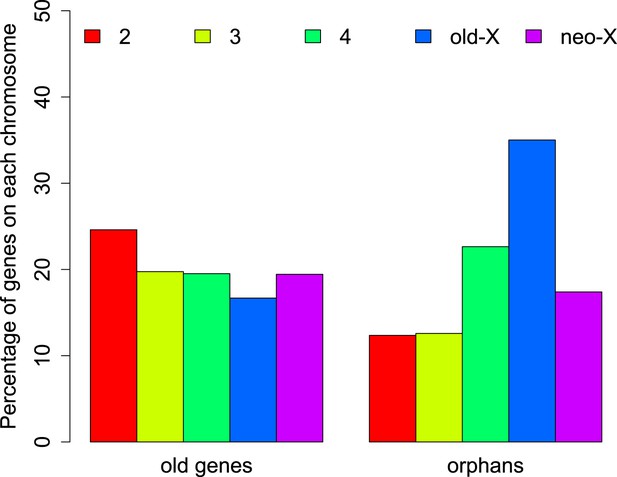
Chromosomal distribution of old genes and orphan genes.
Orphans are overrepresented on the old-X. The number of orphan genes on the neo-X (XR) is significantly lower than on the old-X (XL) (χ2-test, p<1.0 × 10−15).
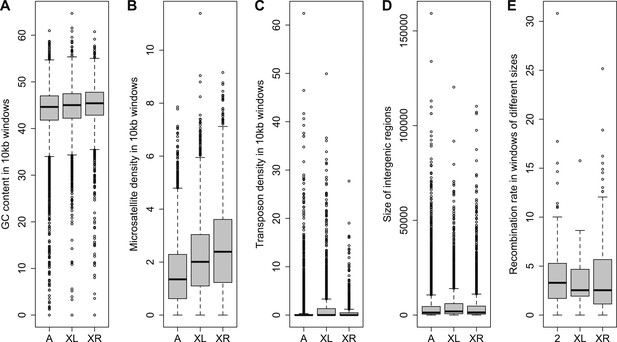
Comparison of genomic features among autosomes, old-X and neo-X.
(A) GC content in 100 kb windows, (B) Microsatellite density in 100 kb windows, (C) Transposon density in 100-kb windows, (D) Length of intergenic regions, (E) Recombination rate. GC content is significantly greater on the neo-X compared to old-X for 10 kb windows (Mann–Whitney test, p=0.00020), but not for 100 kb windows (Mann–Whitney test, p=0.1092). Microsatellite density is significantly higher on the neo-X for both windows of 10 kb (Mann–Whitney test, p=1.9 × 10−12) and 100 kb (Mann–Whitney test, p=0.00025). Transposon density is significantly lower on the neo-X for both windows of 10 kb (Mann–Whitney test, p<1.0 × 10−15) and 100 kb (Mann–Whitney test, p=4.6 × 10−12). Intergenic regions are significantly shorter on the neo-X compared to the old-X (Mann–Whitney test, p=7.4 × 10−9). Recombination rate does not differ significantly between old-X and neo-X (Mann–Whitney test, p=0.629).
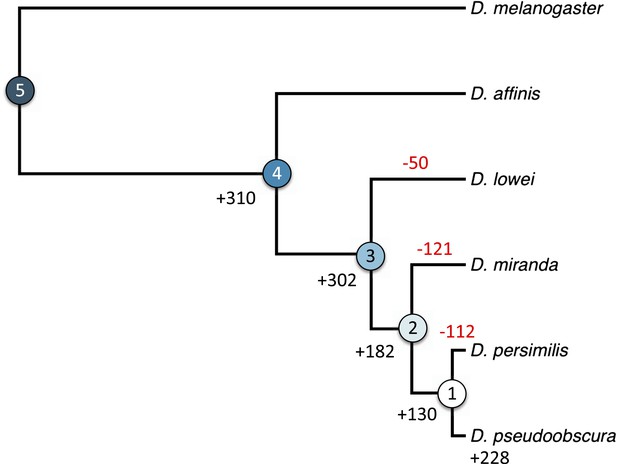
Orphan gain and losses in the Drosophila obscura group.
Schematic phylogenetic tree of the Drosophila obscura group species according to Beckenbach et al. (1993) with D. melanogaster as outgroup. Genes conserved between D. pseudoobscura and 10 non-obscura Drosophila species correspond to age class 5 (old genes). For each age class the number of gene gains is shown in black. Orphans lost at a given branch are indicated in red. Note that losses at internal branches cannot be calculated, since all the orphans are present in D. pseudoobscura. Losses in D. affinis cannot be unambiguously assigned due to the absence of an additional obscura outgroup.
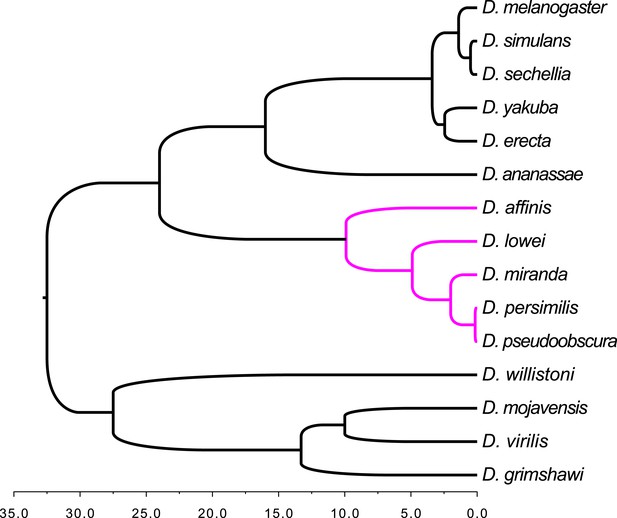
Schematic tree of the Drosophila species analyzed in this study.
The tree includes the 12 Drosophila species from FlyBase (Clark et al., 2007) plus three additional members of the obscura group (D. affinis, D. lowei, and D. miranda). The obscura group is highlighted in magenta. The species corresponding to the black subtrees were used as outgroups in the orphan detection pipeline (see ‘Materials and methods’). Divergence times for the 12 Drosophila species are taken from Table 3 in Obbard et al. (2012) (estimates based on mutation rate); for D. affinis and D. miranda from Gao et al. (2007); for D. lowei from Beckenbach et al. (1993).
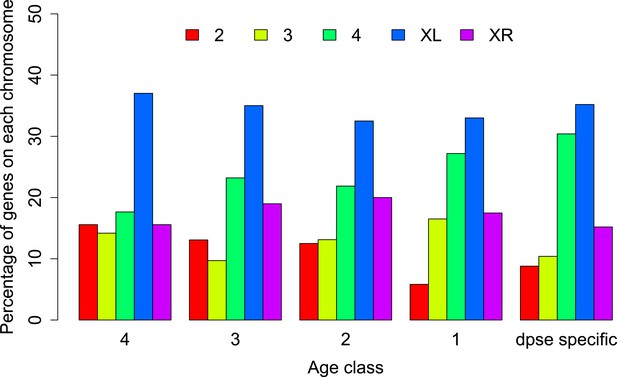
Chromosomal distribution of orphans of different age classes.
In each age class orphans are underrepresented on the neo-X (XR) compared to old-X (XL) (Age class 4: χ2-test, p=6.3 × 10−9; age class 3: χ2-test, p=4.4 × 10−5; age class 2: χ2-test, p=0.00590; age class 1: χ2-test, p=0.00876; D. pseudoobscura specific: χ2-test, p=0.00030).
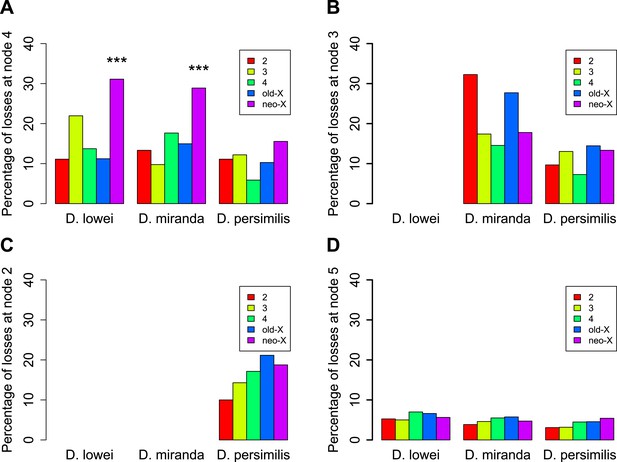
Orphans predating the XL-XR fusion are preferentially lost on the neo-X.
For three terminal branches (D. lowei, D. miranda, and D. persimilis) the fraction of lost genes for each age class is shown. Each autosome and both X-chromosome arms are shown in different color. At node 4, where the neo-X originated, we observed the highest rate of orphan pseudogenization on the neo-X (A). Notably, this effect is not seen for younger orphans (B and C) neither for old genes (D).
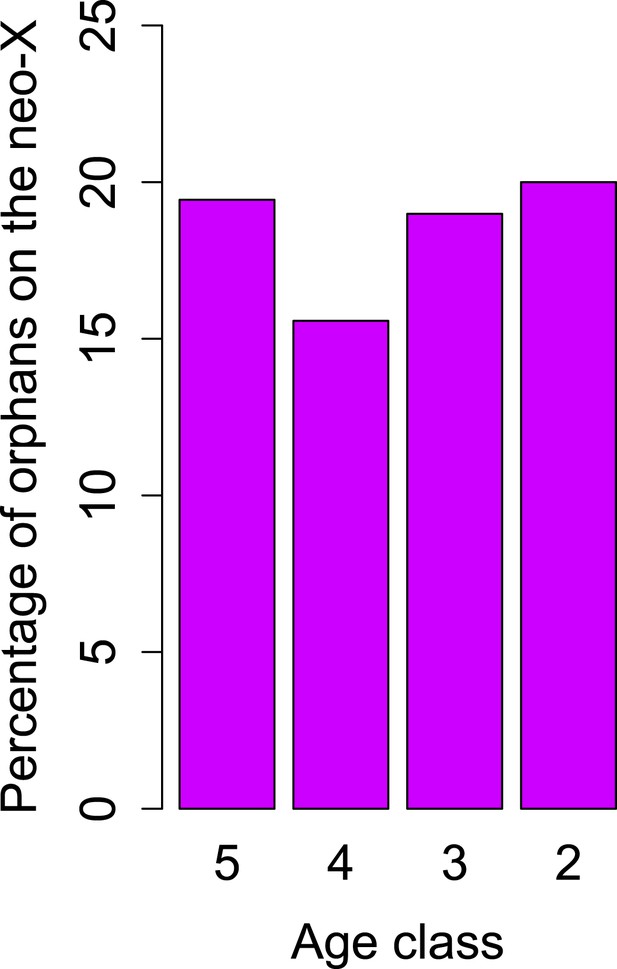
No change in orphan gain on the neo-X chromosome.
The percentage of orphan genes on the neo-X chromosome remains constant through time (indicated by age classes).
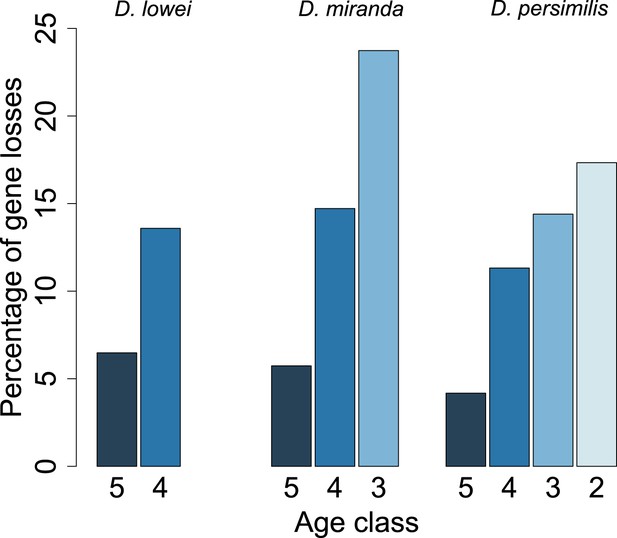
Young orphan genes are more likely to be lost.
The barplot shows the fraction of orphans that has acquired a frameshift or premature stop codon (i.e., lost function). For D. lowei, D. miranda, and D. persimilis, the fraction of lost orphans is shown for different age classes. Orphans are more likely to be lost than old genes. Both the D. miranda and D. persimilis lineage show that younger orphans are more likely to lose function than older ones.
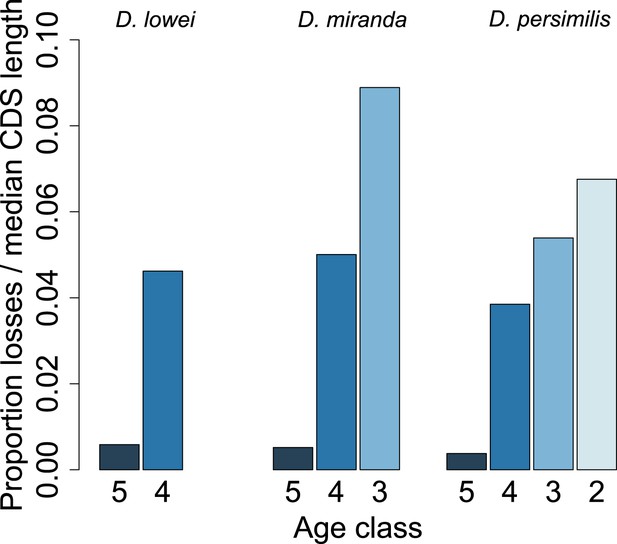
Young orphan genes are more likely to be lost: accounting for CDS length.
To test if the short CDS of orphans affects the pattern that young orphans are more likely to lose function, we normalized the percentage of losses by the median CDS length of genes at that node.
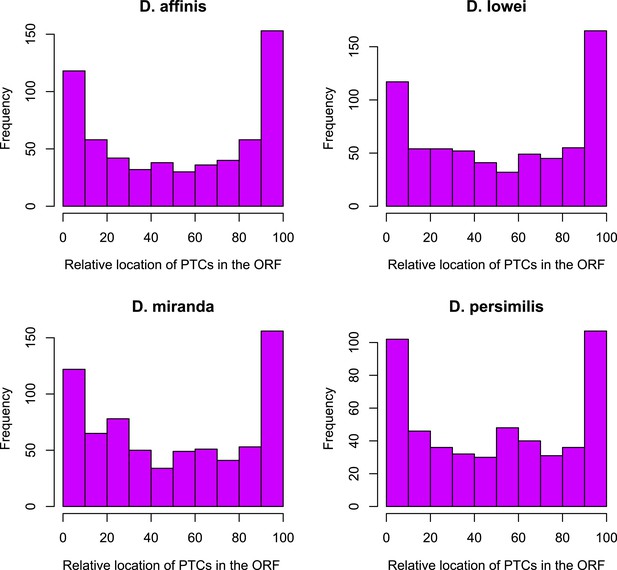
Distribution of premature stop codons (PTCs) along the ORF for all genes containing PTCs.
PTCs are enriched at the beginning and at the end of the ORF in each species.
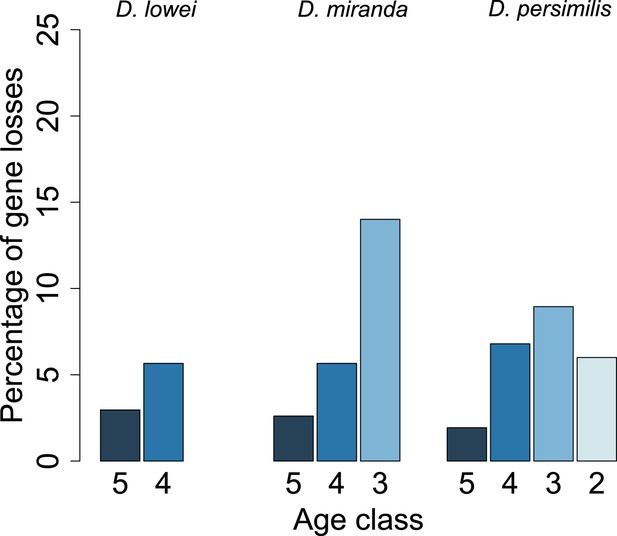
Young orphan genes are more likely to be lost: considering only frameshifts and premature stop codons occurring in the first half of the ORF.
We repeated the analysis shown in Figure 10 by considering only frameshifts and premature stop codons occurring in the first half of the ORF to define a conservative set of pseudogenes, since disrupting mutations occurring at the end of the ORF are likely to have little impact on the gene function.
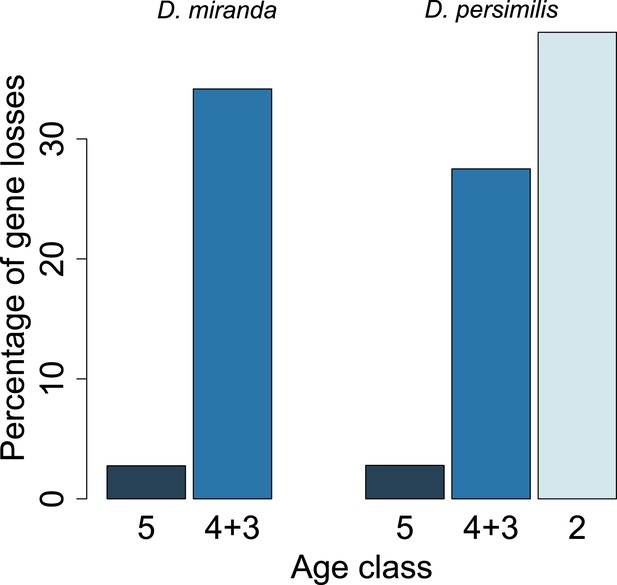
Young orphan genes are more likely to be lost: the conservative set of orthologs.
We repeated the analysis shown in Figure 10 by restricting it to orthologs for which at least one flanking gene is identified in the same contig (see ‘Annotation of the obscura species’). Due to the substantially reduced number of orphans in the older age classes, we combined age class 3 and 4.
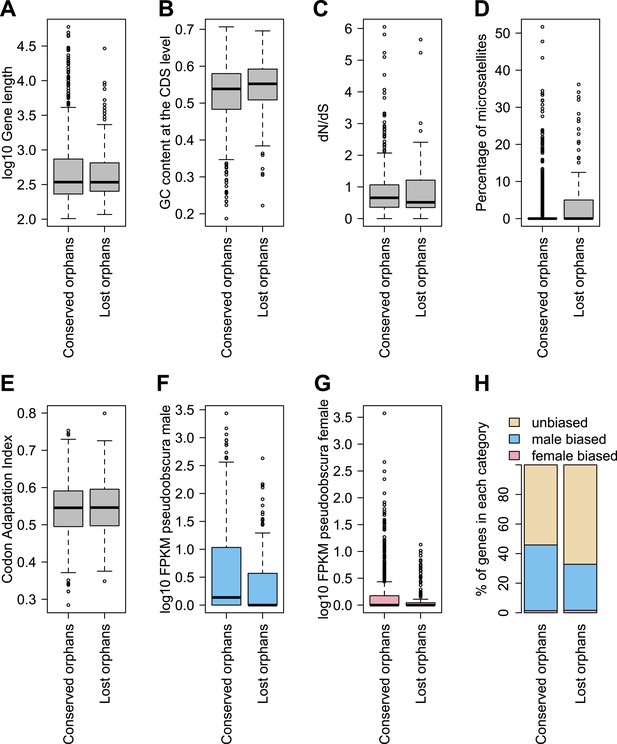
Features of conserved orphans vs lost orphans measured in D. pseudoobscura.
(A) Gene length (B) GC content, (C) dN/dS (D) percentage of microsatellites in coding sequence (E) Codon Adaptation Index (F) gene expression levels in D. pseudoobscura males (G) gene expression levels in D. pseudoobscura females (H) sex-biased expression. Gene length (Mann–Whitney test, p=0.7235) and evolutionary rates (Mann–Whitney test, p=0.5835) are not significantly different between conserved and lost orphans. Lost orphans have higher GC content (Mann–Whitney test, p=0.00325), lower expression in D. pseudoobscura males (Mann–Whitney test, p=0.00012) and females (Mann–Whitney test, p=0.00230) and a higher microsatellite content (Mann–Whitney test, p=0.00049) compared to conserved orphans. Lost orphans are enriched in unbiased genes compared to conserved orphans (χ2-test, p=0.02611).
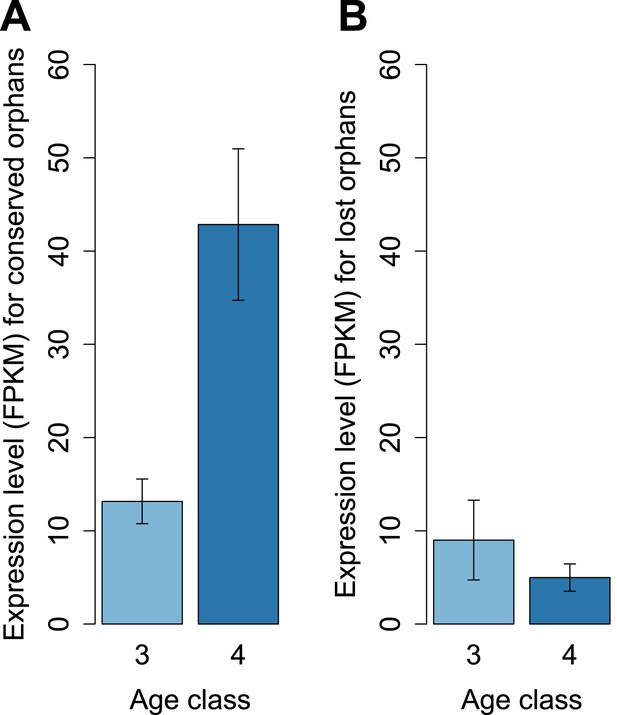
Conserved and lost orphans differ in their gene expression pattern.
Expression intensity and sex bias in D. miranda for orphans conserved in all the obscura species (conserved orphans) vs orphans that pseudogenized in D. lowei and/or D. persimilis (lost orphans). Expression is calculated in males for orphans of age classes 3 and 4. Expression level increases with age for conserved orphans (A), while it decreases for lost orphans (B).
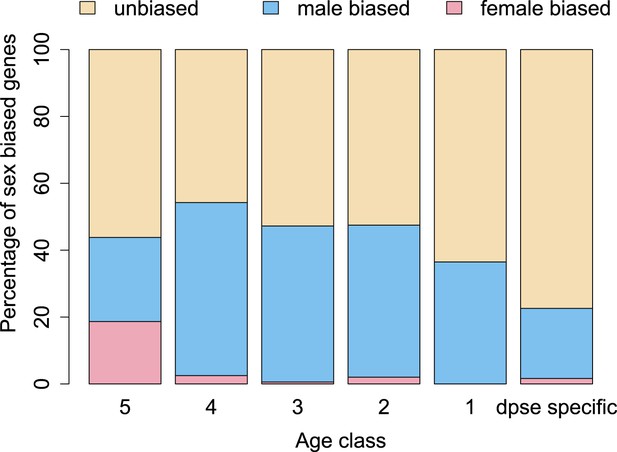
The proportion of male-biased orphans increases with age.
Sex-biased expression was measured in D. pseudoobscura for orphans belonging to different age classes and for old genes (age class 5).
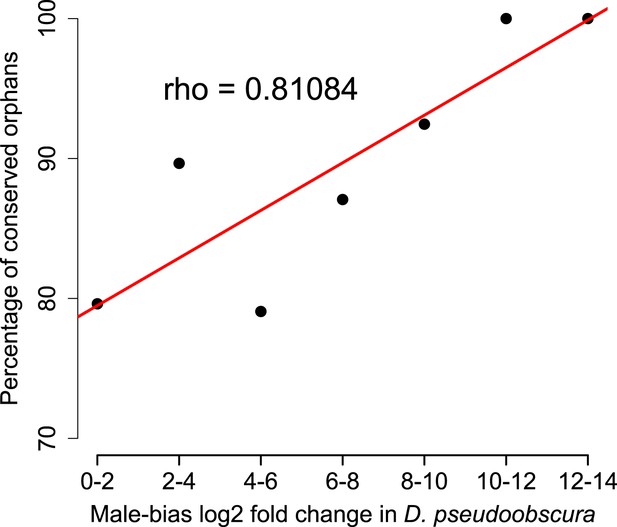
Conservation of orphans is correlated with male-biased gene expression.
Orphans with male-biased gene expression in D. pseudoobscura were grouped into classes according to expression bias strength. The fraction of conserved orphans in each bin shows a significant positive correlation with expression bias (Spearman’s rho = 0.811, p=0.02692). This correlation suggests that orphans with a more pronounced male-biased expression tend to persist longer than less male-biased orphans. No similar trend was seen for female-biased orphans (Spearman’s rho = 0.78262, p=0.1176).
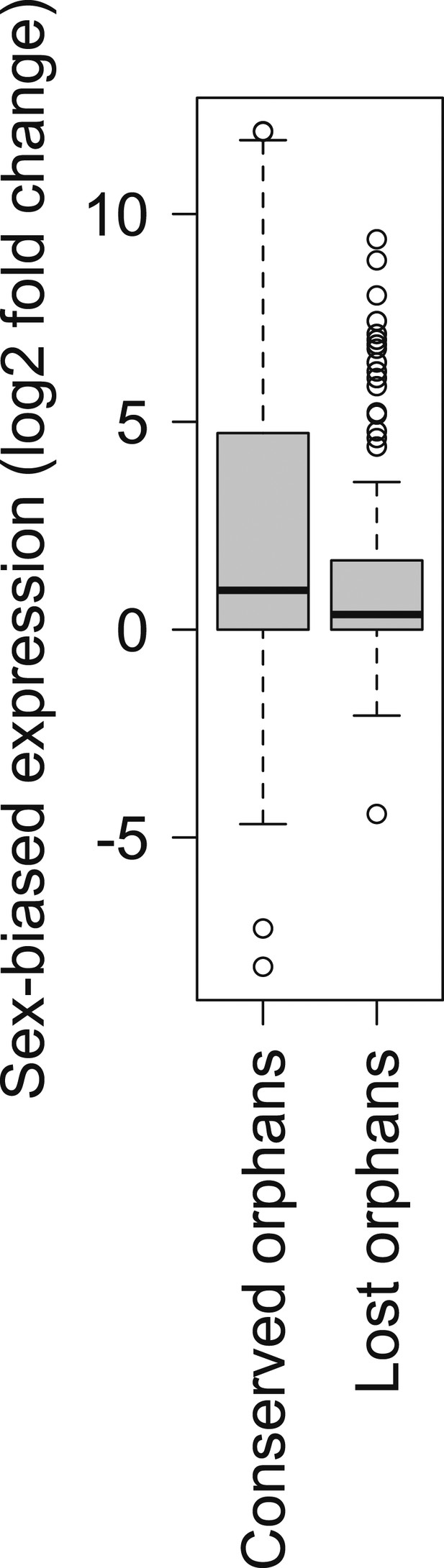
Comparison of strength of sex-biased gene expression for conserved and lost orphans in D. miranda.
A sex-biased gene expression larger than zero indicates a higher gene expression intensity in males than in females (male-biased gene expression). Conserved orphans have significantly higher male-biased expression than lost orphans (Mann–Whitney test, p=0.03158).
Tables
De novo assembly statistics
| D. affinis | D. lowei | D. persimilis | |
|---|---|---|---|
| Number of contigs | 28,946 | 106,465 | 17,387 |
| N75 | 9,478 | 1,218 | 10,359 |
| N50 | 25,160 | 3,230 | 24,172 |
| N25 | 49,062 | 7,357 | 49,047 |
| Minimum length | 121 | 162 | 147 |
| Maximum length | 216,903 | 87,164 | 204,742 |
| Average length | 5,183 | 1,388 | 7,736 |
| Total bp | 150,030,247 | 147,756,871 | 134,501,523 |
| Average coverage | 51 X | 92 X | 44 X |
-
The D. miranda genome was available at NCBI, thus no de novo assembly was made for this species.
Orthology annotation statistics
| D. affinis | D. lowei | D. miranda | D. persimilis | |
|---|---|---|---|---|
| Total genes | 14,287 | 14,952 | 15,282 | 14,995 |
| Genes with frameshifts/PTC* | 1,233 | 1,266 | 1,171 | 898 |
| Mean number of genes per contig | 3.4 | 1.6 | – | 3.4 |
| Median number of genes per contig | 2 | 1 | – | 2 |
| Maximum number of genes per contig | 35 | 24 | – | 37 |
-
*
PTC = Premature termination codons.





