Three-dimensional electron crystallography of protein microcrystals
Figures
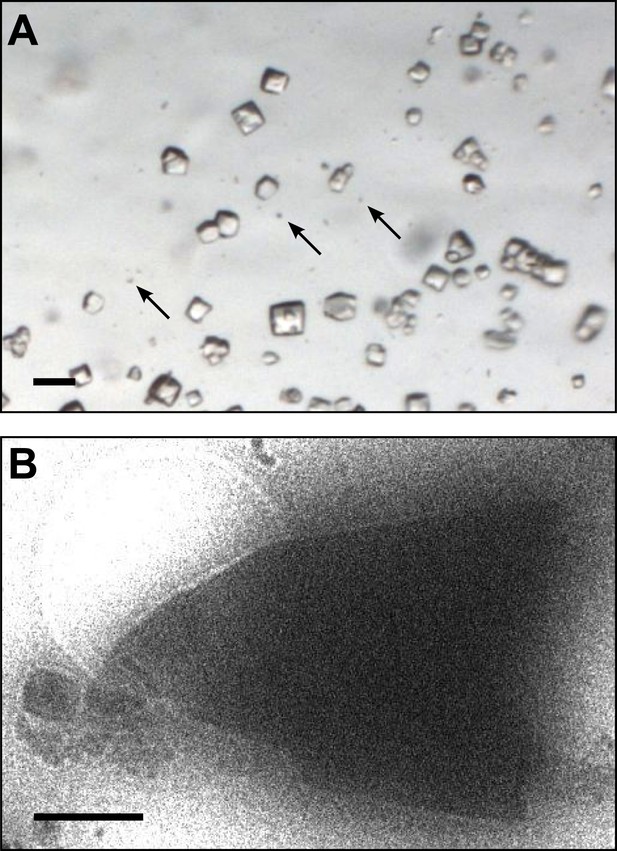
Images of lysozyme microrystals.
(A) Light micrograph showing lysozyme microcrystals (three examples indicated by arrows) in comparison with larger crystals of the size normally used for X-ray crystallography. Scale bar is 50 μm. (B) Lysozyme microcrystals visualized in over-focused diffraction mode on the cryo-EM prior to data collection. The length and width of the crystals varied from 2 to 6 µm with an estimated thickness of ∼0.5–1 µm. Scale bar is 1 µm.
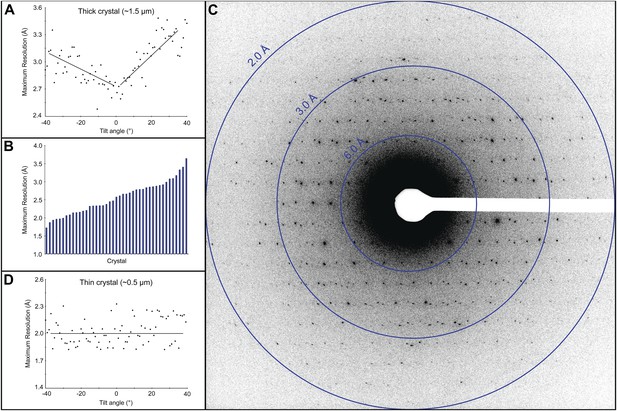
Resolution and data quality of lysozyme microcrystals.
(A) Analysis of the effects of crystal thickness on maximum resolution of observed reflections from thick crystals. The analysis shows adverse effects of crystal thickness on the obtainable resolution as large crystals are tilted. (B) For assessing the quality of our cryo preparations, diffraction data were obtained from 100 lysozyme microcrystals. 43/100 were thin crystals that showed reflections in the 2–4 Å range, with the best crystal in this set yielding data to ∼1.7 Å resolution. (C) An example of lysozyme diffraction data collected at 0.01e−/Å2/second and a 10 s exposure. The pattern shows strong and sharp spots surpassing 2 Å resolution. This diffraction pattern was processed with ImageJ and despeckled for ease of viewing. (D) Analysis of the effects of crystal thickness on maximum resolution of observed reflections from thin crystals. The small crystal shows a relatively constant maximum resolution that does not appear to be affected by crystal tilt.
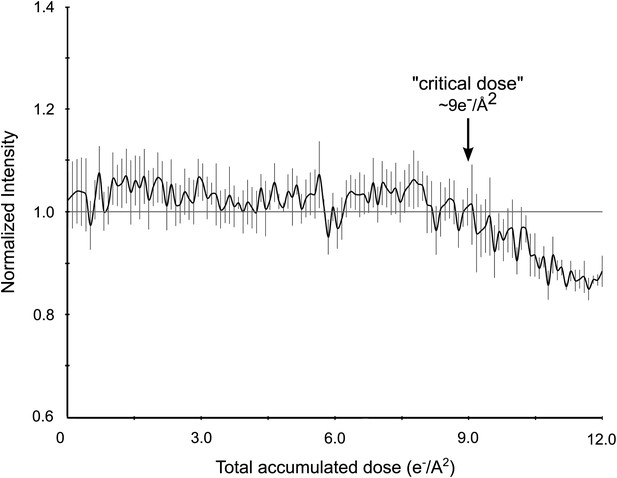
Effects of cumulative electron dose on diffraction data quality.
A single lysozyme microcrystal was subjected to 120 sequential exposures without tilting, each of a dose of ∼0.1 e−/Å2 for a total accumulated dose of ∼12 e−/Å2. Normalized intensity vs total accumulated dose for three diffraction spots observed over all 120 sequential frames was plotted. A decrease in diffraction intensity becomes apparent at a dosage of ∼9 e−/Å2 (‘critical dose’). Bars represent standard error of the mean.
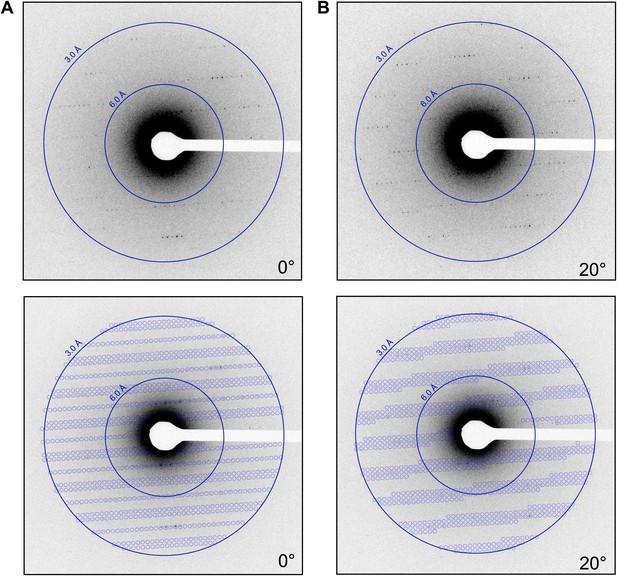
Prediction of reflections and indexing in the diffraction patterns.
(A and B) Two examples of diffraction patterns obtained from a single crystal at tilt angles of 0° and 20° respectively. Locations indicated by circles were predicted to contain diffraction spots by our spot prediction algorithm. Additional examples from the same crystal are presented in Video 2. The resolution limit was set at 2.9 Å resolution for this study.
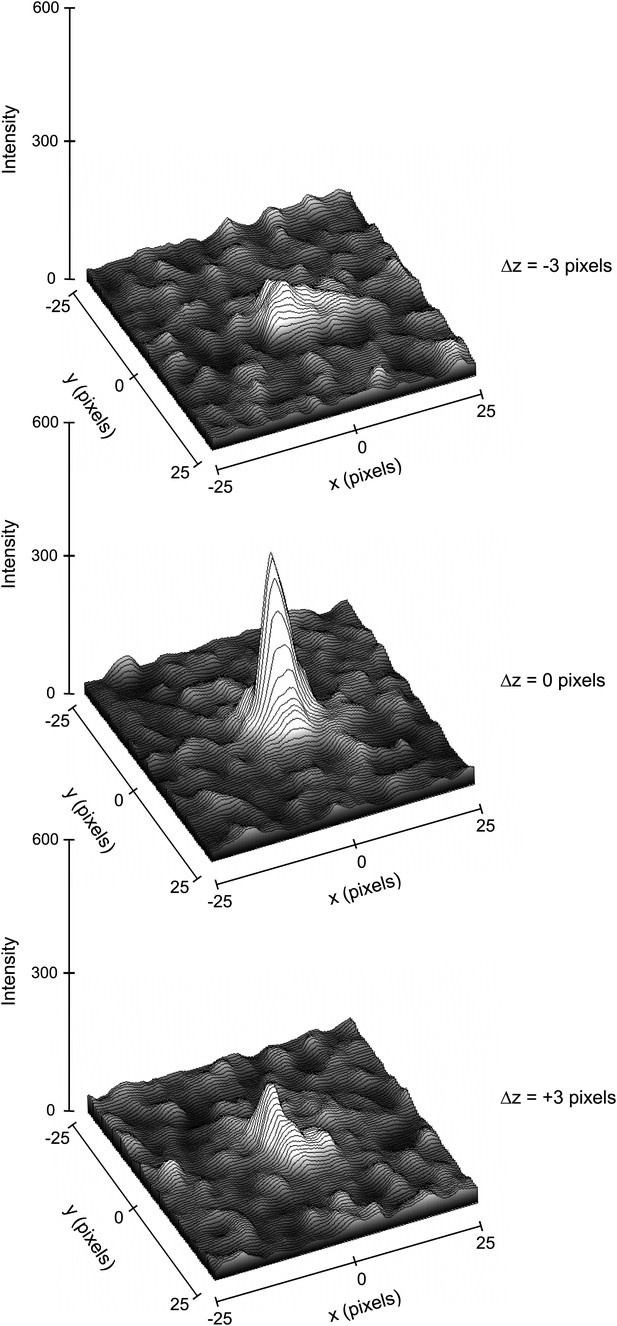
Three-dimensional profiles of the intensity of a single reflection over three consecutive diffraction patterns at −0.1°, 0°, and 0.1° degree tilts.
The plots show the approximate dimensions of the full reflection with a width (full width at half maximum height) of 3–5 pixels in the x, y, and z direction.
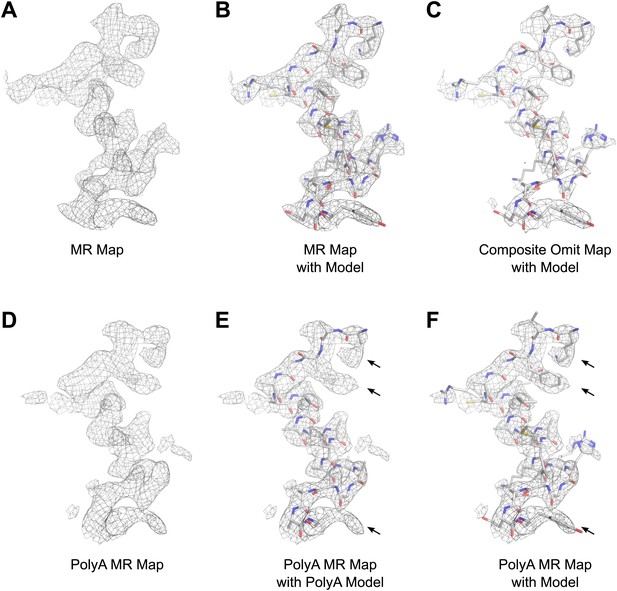
Results of phasing by molecular replacement prior to crystallographic refinement.
Molecular replacement was performed with both the full model of lysozyme (PDB 4AXT, top panels) as well as a poly-alanine model (bottom panels) and the resulting 2Fobs−Fcalc maps around residues 1–20 are shown. (A and B) The phases following molecular replacement with the full model were of good quality demonstrated by how well the density surrounding the model fits, even before any refinement is performed. (C) A composite-omit map calculated by omitting 5% at a time showed good agreement with the unrefined structure indicating the phases were not dominated by model bias. (D–F) As an additional test of model bias, phasing was done with a poly-alanine homology search model of lysozyme. The resulting 2Fobs−Fcalc map is of good quality (D) and shows density extending beyond the poly-alanine model (E and F, arrows). (F) The same density map as E but with the structure of lysozyme fit. Arrows in D and E show examples of clear side chain density from the poly-alanine map. All maps are contoured at 1.0σ.
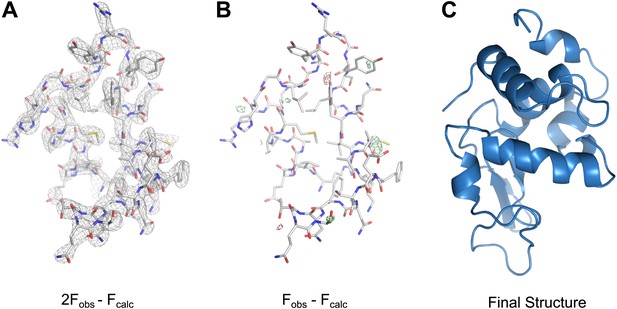
MicroED structure of lysozyme at 2.9 Å resolution.
(A) The 2Fobs−Fcalc (contoured at 1.5σ) map covers protein residues 5–45 of lysozyme. (B) Fobs−Fcalc difference map contoured at +3.0σ (green) and −3.0σ (red) for the same protein region. The map (A) shows well-defined density around the vast majority of side chains and the difference map (B) shows no large discrepancies between the observed data (Fobs) and the model (Fcalc). The final structure of lysozyme is shown in panel C and the complete three-dimensional map is presented in Video 3.
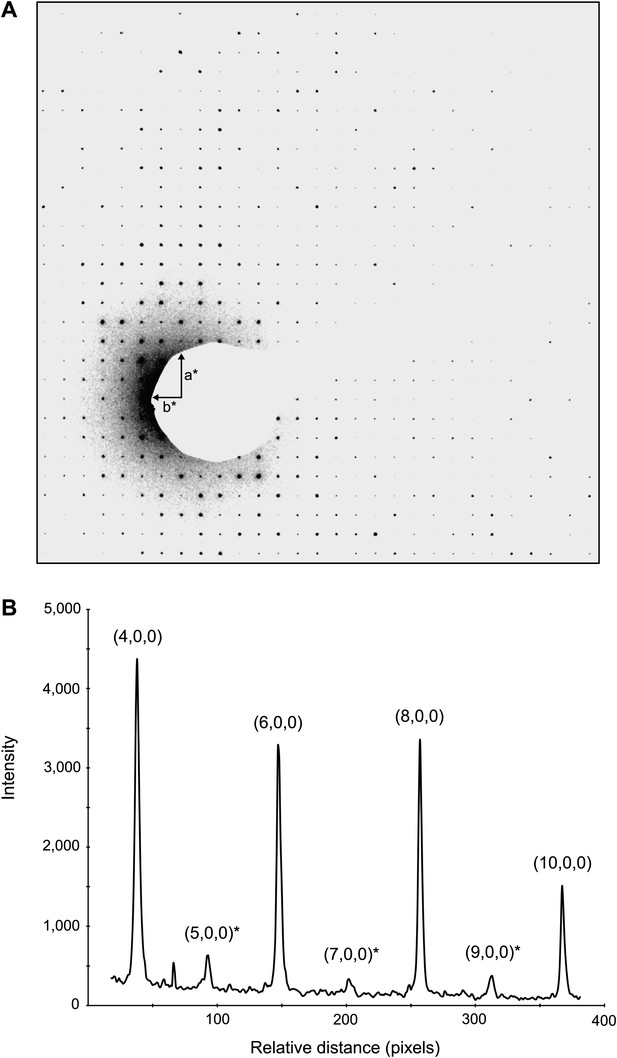
Dynamic scattering in lysozyme 3D crystals.
Intensity measurement along the a* axis of a raw diffraction pattern illustrating the relatively small contributions due to dynamic scattering. (A) Diffraction pattern from the major plane of a lysozyme crystal with visible intensity in the (2n+1,0,0) and (0,2n+1,0) Miller indices. (B) (2n+1, 0, 0) reflections (starred) are expected to be systematically absent and observed intensities at these indices are assumed to be the result of dynamic scattering. Image contrast was enhanced for clarity using ImageJ.
Videos
An example of a complete three-dimensional electron diffraction data set from a single lysozyme microcrystal.
In this example, diffraction patterns were recorded at 1° intervals from a single crystal, tilted over 47°. Cumulative dose was ∼5 e−/Å2 in this example.
An example of spot prediction in diffraction data from a single crystal.
Reflections predicted on representative diffraction patterns obtained from a single crystal tilted over 39° sampled every 2° in this video. Predictions were made to 2.9 Å resolution using our spot prediction algorithm.
Three-dimensional representation of merged intensity values.
2490 total unique reflections are present for an overall completeness of 92% at 2.9 Å resolution. Video begins with a* axis horizontal, b* axis vertical, and the c* axis normal to the image plane.
2Fobs−Fcalc density around the complete lysozyme model at 2.9 Å resolution (contoured at 1.5σ).
https://doi.org/10.7554/eLife.01345.014Tables
MicroED crystallographic data
| Data collection | |
| Excitation voltage | 200 kV |
| Electron source | Field emission gun |
| Wavelength (Å) | 0.025 |
| Total electron dose per crystal | ∼9 e−/Å2 |
| Number of patterns per crystal | 40–90 |
| No. crystals used | 3 |
| Total reflections to 2.9 Å | 84,889 |
| Data refinement | |
| Space group | P43212 |
| Unit cell dimensions | |
| a = b | 77 Å |
| c | 37 Å |
| α = β = γ | 90° |
| Resolution | 2.9–20.0 Å |
| Total unique reflections | 2490 |
| Reflections in working set | 2240 |
| Reflections in test set | 250 |
| Multiplicity* | 34 |
| Completeness (2.9–3.1) | 92% (57%) |
| Rwork/Rfree (%) | 25.5/27.8 |
| RMSD bonds | 0.051 Å |
| RMSD angles | 1.587° |
| Ramachandran (%)† (allowed, generous, disallowed) | 99.1; 0.9; 0 |
-
*
Multiplicity is defined as total measured reflections divided by number of unique reflections.
-
†
Statistics given by PROCHECK (Laskowski et al., 1993).
Results of model validation and bias tests
| Data set | Molecular replacement result | TFZ | Final Rfree (%)¶ |
|---|---|---|---|
| 1* | No solution | N/A | N/A |
| 2† | Solution** | 19.1 | 54.9 |
| 3‡ | No solution | N/A | N/A |
| 4§ | Solution | 12.6 | 35.2 |
| 5# | Solution | 14.7 | 27.8 |
-
*
Random intensities.
-
†
Shuffled Miller indices.
-
‡
Calmodulin replaced intensities.
-
§
Intensities ± 35%.
-
#
Original data.
-
¶
Final Rfree after a minimum of two cycles of refinement.
-
**
Solution was found; however, the space group was incorrect (P4121).
Models for molecular replacement validation
| Protein | PDB ID | Molecular weight (kDa) | Symmetry | Unit cell dimensions | MR solution |
|---|---|---|---|---|---|
| Hen Egg White Lysozyme* | 4AXT | 14.3 | P43212 | a = b = 78.24 Å | Yes |
| c = 37.47 Å | |||||
| α = β = γ = 90° | |||||
| T4 Lysozyme† | 2LZM | 18.7 | P3212 | a = b = 61.20 Å | No |
| c = 96.80 Å | |||||
| α = β = 90° | |||||
| γ = 120° | |||||
| Calmodulin‡ | 3CLN | 16.7 | P1 | a = 29.71 Å, | No |
| b = 53.79 Å, | |||||
| c = 24.99 Å | |||||
| α = 94.13°, | |||||
| β = 97.57°, | |||||
| γ = 89.46° | |||||
| Dodecin§ | 4B2J | 8.5 | F4132 | a = b = c = 142.90 Å | No |
| α = β = γ = 90° | |||||
| αA Crystallin# | 3L1E | 11.9 | P41212 | a = b = 56.22 Å, | No |
| c = 68.66 Å | |||||
| α = β = γ = 90° |
-
*
-
†
-
‡
-
§
-
#





