Dual mode of embryonic development is highlighted by expression and function of Nasonia pair-rule genes
Figures
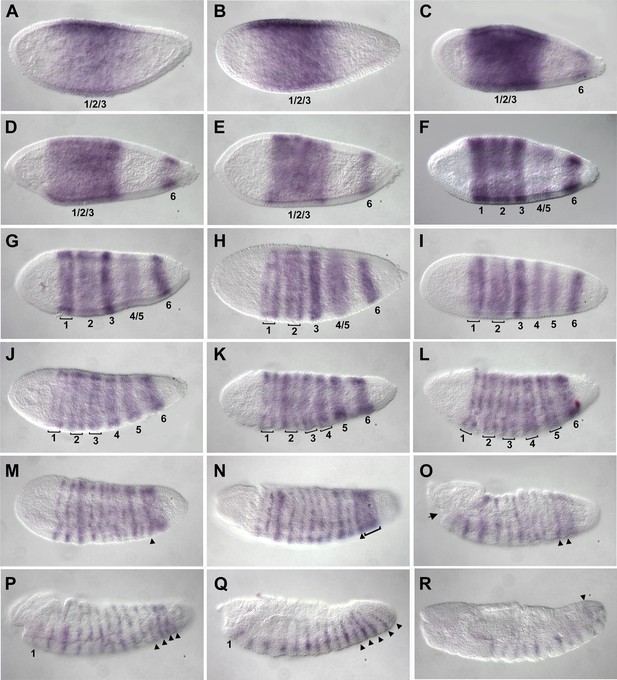
Summary of Nasonia eve mRNA expression.
Embryos are shown with anterior left and dorsal up. Nv eve is initially expressed in a broad domain (A and B), which sharpens as a posterior stripe becomes visible at around 4 hr after embryo laying (AEL) (C and D). The broad domain retracts anteriorly and gives rise to three apparently double-segment stripes (E and F). Between stripes 3 and posterior stripe 6, an additional double stripe precursor comes up at around 6 hr AEL (stripe 4/5; panels F and G) and this splits to form two double-segment stripes, ‘4’ and ‘5’ as double-segment stripes 1–3 split into two single-segment stripes each between 6 and 8 hr AEL (F–J). Stripes 4 and 5 also split to form single-segment stripes during early gastrulation, and stripe 6 broadens (K and L), giving rise to stripes that are visibly distinct during germ band extension in non-fluorescent staining by 10–12 hr AEL (M–R, arrowheads). There are a total of 16 single-segment stripes of Nv eve.
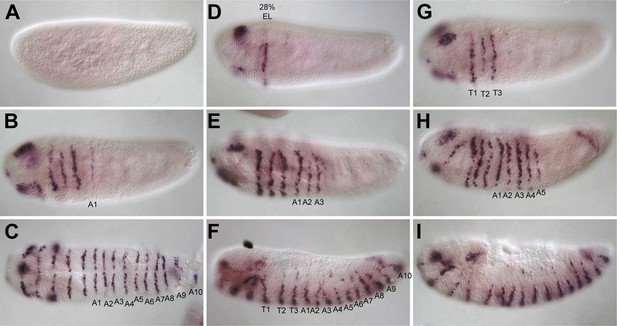
Nv wingless (Nv wg) mRNA expression in the embryo.
All embryos are shown with anterior to the left and dorsal up except as indicated. (A) Precellular blasoderm with no expression of Nv wg. (B) Cellular blastoderm embryo exhibiting head expression and one stripe of Nv wg. (C) Early gastrula embryo exhibiting Nv wg staining in three segmental stripes comprising thoracic segments. (D) Ventral view of early gastrula embryo with four segmental stripes, including one abdominal segment. (E) Germ band extension embryo with six segmental stripes of Nv wg expression. (F) Germ band extension embryo exhibiting eight segmental stripes, including three thoracic and five abdominal stripes, as well as additional head segmental stripes. (G) Ventral view of a fully extended germ band with the full complement of 16 segmental Nv wg stripes. (H) Lateral view of germ band retracting embryo with 16 segmental stripes, including three clear segmental stripes in the head. (I) Lateral view of dorsal closure embryo exhibiting fading segmental Nv wg staining in all segments.
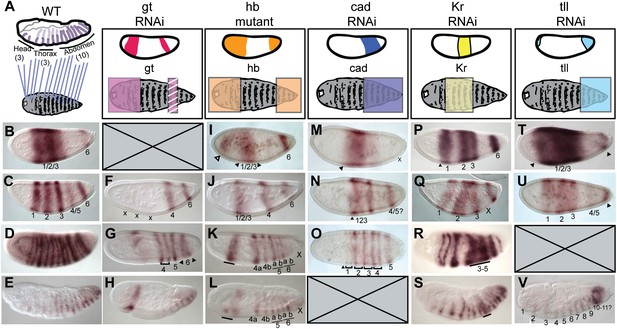
Nv eve epistasis with maternal and gap genes.
(A) Schematic representation of the germ-band-extended embryo, showing 16 single-segment stripes of Nv eve expression, and their segment counterparts in the patterned larval cuticle. Colored boxes cover the segments of the larval cuticle that are lost or fused in each RNAi background. All embryos are shown anterior left, dorsal up (except where indicated). Nv eve mRNA expression is shown in each embryo (B–G). Wild-type (WT) embryos are shown as staged controls for RNAi embryos. (B) WT early blastoderm embryo. (C) WT cellular blastoderm embryo. (D) WT early gastrula extension embryo. (E) WT germ-band-retracted embryo. (F–H) gt RNAi embryos stained for Nv eve mRNA expression. (F) Cellular blastoderm embryo with reduced Nv gt exhibits loss of anterior Nv eve stripes (x). (G) Nv gt RNAi embryo in early germ-band-extension exhibits loss of anterior Nv eve stripes and improper splitting of Nv eve stripe 5, as well as aberrant dorsal anterior expression of Nv eve. (H) Nv gt RNAi embryo at dorsal closure exhibits a stripe of Nv eve at the anterior, as well as a reduced number of posterior segmental Nv eve stripes. (I–L) Nv hb mutant embryos stained for Nv eve mRNA expression. (I) Early blastoderm Nv hb mutant embryos have a reduced central Nv eve domain (bounded by black arrowheads), and an ectopic anterior Nv eve stripe (white arrowhead). (J) Nv hb mutant cellular blastoderm embryo with a single anterior domain of Nv eve that has failed to resolve, and a single stripe 4 which exhibits delayed splitting. (K) Nv Hb mutant germ-band extension embryo with fused anterior domain (line) and 6 segmental stripes, representing derivatives of Nv eve stripes 4 and 5 and two derivatives of stripe 6; additional stripe 6 derivatives are absent (x). (L) hb mutant dorsal closure embryo exhibiting fused anterior domain (line) and the same number of derivatives as in (M), with more posterior segments missing (x). (M–O) Nv cad RNAi embryos stained for Nv eve mRNA expression. (M) Nv cad RNAi early blastoderm with reduced central Nv eve domain that is also posteriorly shifted (anterior boundary indicated by black arrowhead). (N) Nv cad RNAi cellular blastoderm embryo with posteriorly shifted (arrowhead), reduced Nv eve central domain, whose splitting is delayed. (O) Nv cad RNAi early gastrula embryo with posterior shift in Nv eve expression (black arrowhead). Four double-segment periodicity stripes are split into single-segment stripes and stripe 5 remains intact. (P–S) Nv Kr RNAi embryos stained for Nv eve mRNA expression. (P) Nv Kr RNAi precellular blastoderm embryo with aberrant Nv eve central domain resolution, where stripes 2–3 appear posteriorly shifted. (Q) Dorsolateral view of a Nv Kr RNAi embryo where stripes 2 and 3 are less refined than WT and 3 is posteriorly shifted. No stripe 4/5 expression is detected (X). (R) Nv Kr RNAi early gastrula embryo with aberrant stripe 2 splitting and aberrant resolution of stripes 3–5. (S) Moderately affected Nv Kr RNAi germ-band retraction embryo with fused segments in the middle of the embryo (line). (T–V) Nv tll RNAi embryos stained for Nv eve mRNA expression. (T) Nv tll RNAi early blastoderm embryo with expanded Nv eve expression domains toward both poles (arrowheads). (U) Nv tll RNAi precellular blastoderm embryo showing delayed resolution of Nv eve stripes 1–3 and Nv eve stripe 6 shifted to the extreme posterior pole of the embryo (arrowhead). (V) Nv tll RNAi dorsal closure embryo showing abnormal posterior Nv eve stripe formation.
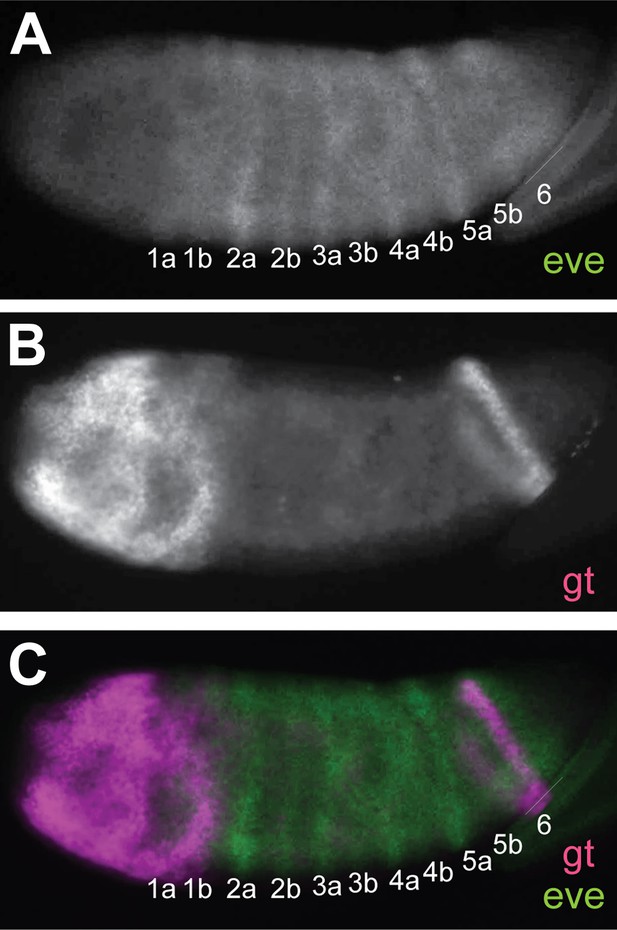
Nv eve/Nv gt double FISH in the embryo.
Lateral view of early Nasonia gastrula embryo double stained for Nv eve and Nv gt mRNA. (A) Nv eve mRNA. (B) Nv gt mRNA. (C) Merge of Nv eve and Nv gt channels. Note the position of the late posterior stripe of Nv gt relative to Nv eve stripe 6.
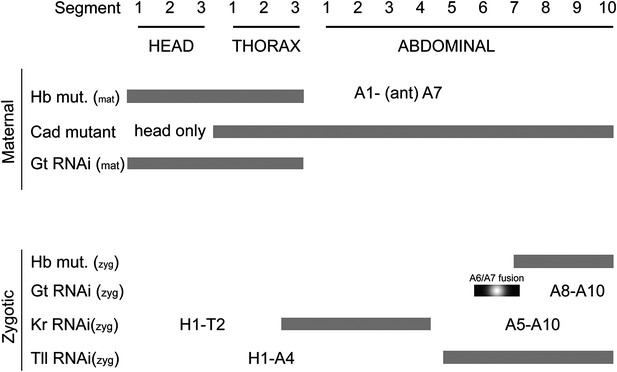
(A) Summary of maternal and gap gene loss of function phenotypes in Nasonia. On the vertical axis, genes affected by RNAi or genetic lesion are listed, and indicated as either maternal or zygotic phenotypes. Segments lost in each background are indicated as gray bars. Segments that remain in a given background are annotated alphanumerically.
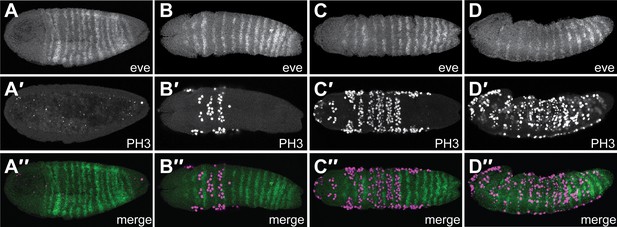
Nv eve expression and cell division appear to be coordinated.
Embryos co-stained for Nv eve mRNA using in situ hybridization and fluorescent detection, as well as for mitotic figures, using an antibody against phospho Histone H3. Embryos are shown with anterior left and dorsal up, except columns B and C, which are ventral views. (A–A″) An early gastrula embryo exhibiting 15 stripes of Nv eve, including five derivatives of stripe 6 (A), has no evident mitotic figures in the posterior domain of Nv eve stripe 6 differentiation (A′). (A″) Merge of panels A and A′. (B–D″) Timecourse series of wild-type embryos stained for Nv eve mRNA and phospho-Histone H3. (B–D). Top panels are Nv eve in situ alone, middle panels (B′–D′) are phospho-Histone H3 antibody staining, and bottom panels (B″–D″) are merge images of upper panels, showing localization of mitotic figures relative to Nv eve stripes.
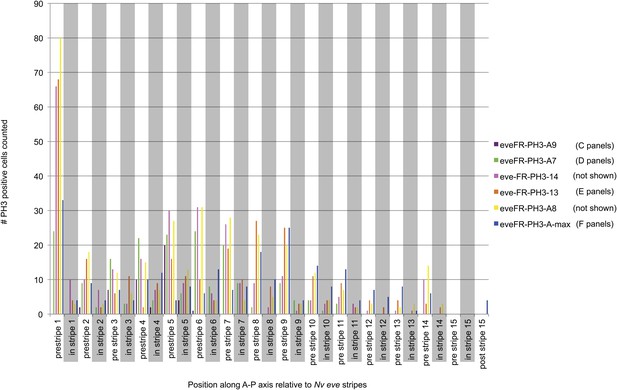
Quantification of PH3 positive cells relative to Nv eve stripes in the embryo.
Embryos stained for phosphorylated histone H3 (to label mitotic figures) and for Nv eve mRNA (as shown in Figure 3) were analyzed to quantify the number of mitotic cells occurring between stripes as compared to within eve stripes. Cell number counted is plotted on the Y axis as a function of position along the A–P axis of the embryo (as indicated by Nv eve stripe position), which is indicated on the X axis. Mitotic cells that fall partially on a stripe are counted as occurring in the stripe. Each unique color indicates one embryo quantified in this manner, and embryos corresponding to those shown in Figure 3 are also labeled according to their position in the figure.
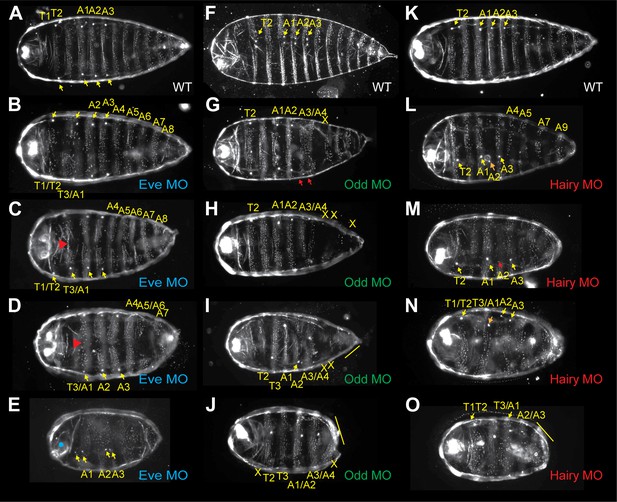
Morpholino knockdown of Nv eve, Nv hairy, and Nv odd results in embryo patterning defects.
First instar larval cuticles are shown with anterior left and generally ventral denticle patterns are shown. (A, F, K) Wild-type larval cuticles. Yellow arrows indicate spiracles present on segments T2, A1, A2 and A3. Bright anterior labral appendages are apparent at the extreme anterior of the larva. (B–E) Unhatched larvae from Nv eve morpholino (MO)-injected embryos, in order of increasing phenotype severity. Red arrowheads indicate loss of midline cuticle. Blue dot indicates head open defect. Yellow arrowheads indicate position of spiracles. (G–J) Unhatched larvae from Nv odd morpholino (MO)-injected embryos, in order of increasing phenotype severity. Yellow arrows indicate position of spiracles, red arrows indicate A3/A4 fusion. X indicates naked cuticle from segment loss. Yellow line indicates multi-segment fusion. (L–O) Unhatched larvae from Nv hairy morpholino (MO)-injected embryos, in order of increasing phenotype severity. Yellow arrowheads indicate position of spiracles. Red or orange arrowheads indicate aberrantly positioned or missing spiracles. Yellow line indicates segment fusion.
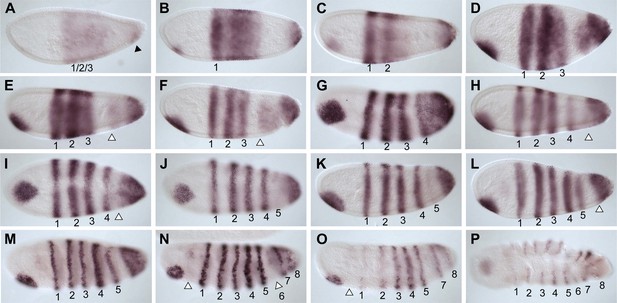
Summary of Nv odd-skipped mRNA expression.
Embryos are shown with anterior left and dorsal up, except where indicated. (A) Precellular blastoderm embryo showing early expression of Nv odd in a broad domain and a posterior cap with a slight clearing in between. (B) Precellular blastoderm embryo showing ventral head patch and darkened central broad domain and distinct posterior cap. (C) Precellular blastoderm embryo with sharpening pair-rule stripes and expanding posterior cap. (D) Precellular blastoderm embryo with dark ventral head patch and posterior cap, and expansion of expression between broad central domain and posterior domain. (E and F) Cellularizing blastoderm embryos with three double-segment periodicity stripes, and a continuous posterior domain of variable staining intensity. Arrowhead indicates boundary of faint expression, which prefigures position of double-segment stripe 4. (G) Ventral view of cellularizing embryo with three strong double-segment stripes, and a fourth stripe forming at the anterior boundary of a more uniformly staining posterior cap (arrowhead). (H) Cellularized blastoderm embryo with four distinct double-segment stripes and a receding posterior cap domain (arrowhead). (I) Ventral view of cellular blastoderm showing four strong double-segment stripes and receding posterior cap (arrowhead), whose anterior boundary prefigures the position of stripe 5. (J) Ventrolateral view of cellular blasoderm embryo showing early appearance of stripe 5 at the anterior boundary of receding posterior domain, whose staining intensity is now less uniform. (K) Cellular blastoderm embryo with five double-segment stripes of expression, a strong ventral head spot, and a reduced, uniform posterior cap. (L) Same as K, with five equivalently strong double-segment stripes. Arrowhead indicates slightly expanded posterior cap. (M) Early germ-band extension embryo with five double-segment periodicity stripes and two stripes becoming evident within the posterior cap. (N) Slightly later embryo than M, with 2 posterior cap stripes more clearly differentiated. (O) Slightly later embryo than N, with anterior stripes fading and posterior segments expanding. (P) Dorsal view, dorsal closure embryo exhibiting eight single-segment periodicity stripes.
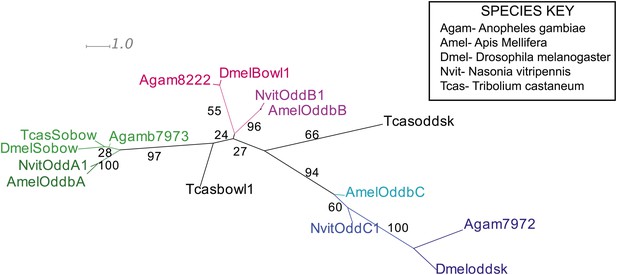
Phylogenetic analysis of odd-skipped.
Phylogenetic analysis of odd-skipped protein was performed using Maximum likelihood analysis with 1000-fold bootstrap support, using RAxML via the CIPRES portal (Copf et al., 2003; Nagy et al., 1994). The best scoring maximum likelihood tree is shown with bootstrap support values indicated adjacent to branch.

odd-skipped protein sequence alignment.
Predicted odd-skipped protein sequences from Nasonia were aligned to odd-skipped related proteins sequences from Drosophila melanogaster, Anopheles gambiae, Apis mellifera, and Tribolium castaneum using ClustalW using standard parameters. Full-length sequences were used for alignment.
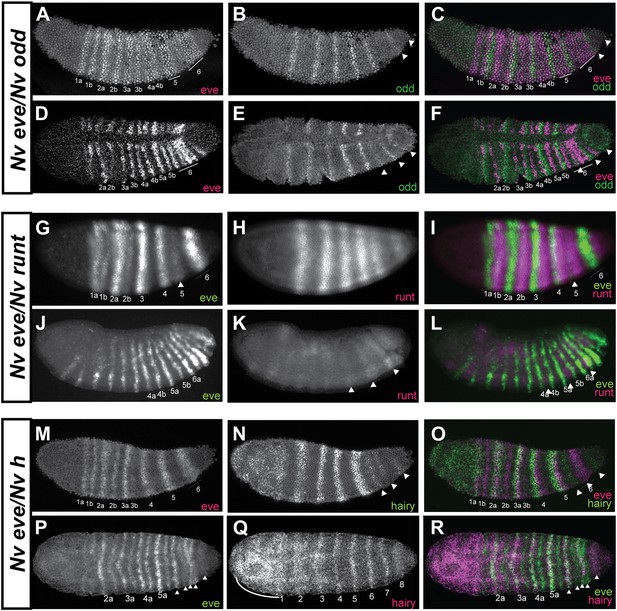
Phasing of Nasonia pair-rule genes in embryos using double fluorescent in situ hybridization.
(A) Lateral view of Nv eve expression in early gastrula embryo. (B) Nv odd expression alone in the same embryo. (C) Merge of Nv eve and Nv odd channels, illustrating their relative phasing. Nv eve mRNA is pseudo-colored pink, Nv odd is in green. Arrowheads indicate position of a posterior doublet of odd stripes. (D) Dorsolateral view of Nv eve in later gastrula embryo. (E) Nv odd expression alone in the same embryo. Arrowheads indicate position of posterior Nv odd stripes 6, 7 and 8. (F) Merge of Nv eve and Nv odd channels, illustrating their relative phasing. (G) Lateral view of Nv eve expression in blastoderm embryo. Arrowhead indicates position of Nv eve stripe 5. (H) Nv runt expression in the same blastoderm embryo. (I) Merge of Nv eve (green) and Nv runt (pink) channels, indicating relative phasing. (J) Lateral view of Nv eve expression in germ-band-extended embryo. Numbers indicate identity of Nv eve stripe. (K) Nv runt expression alone in the same embryo. Arrowheads indicate position of posterior primary Nv runt stripes. (L) Merge of Nv eve (green) and Nv runt (pink) channels, indicating relative phasing. Note that posterior Nv runt stripes, though faint, appear to be positioned posterior to odd-numbered Nv eve segmental stripes. (M) Lateral view of Nv eve expression in early gastrula embryo. Line indicates broadening stripe 6. (N) Nv hairy expression in the same gastrula embryo. Arrowheads indicate positions of three late forming posterior double-segment stripes. (O) Merge of Nv eve (pink) and Nv hairy (green) channels, indicating relative phasing. (P) Ventral view of gastrula embryo showing Nv eve expression alone. Arrowheads indicate positions of single-segment stripes derived from eve stripe 6. (Q) Nv hairy expression alone in the same gastrula embryo. Line indicates extended anterior domain continuous with stripe 1. (R) Merge of Nv eve (green) and Nv hairy (pink) channels, illustrating relative phasing.
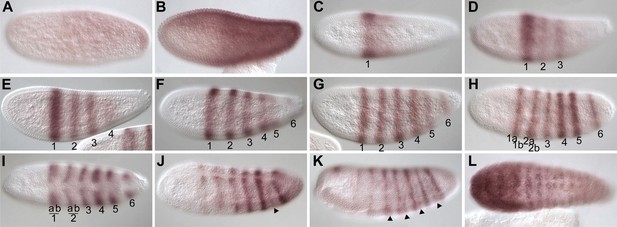
Summary of Nv runt mRNA expression.
Embryos are shown anterior left and ventral up unless indicated. (A) Early energid stage embryo with faint staining evident in posterior. (B) Early precellular blastoderm embryo exhibiting stronger, mostly ubiquitous Nv runt staining biased towards the posterior. (C) Precellular blastoderm embryo with strong stripe of Nv runt expression within larger, gap-type domain. Expression is absent from the rest of the embryo. (D) Precellular blastoderm embryo exhibiting three clear pair-rule stripes of Nv runt expression. (E) Cellular blastoderm embryo with four pair-rule stripes of Nv runt expression. (F) Early gastrula embryo exhibiting three pair-rule stripes, an expanding stripe 4/5 stripe domain and faint posterior stripe 6. (G) Early gastrula embryo exhibiting 6 pair-rule stripes of Nv runt expression. (H) Gastrula embryo exhibiting splitting of anterior stripes 1 and 2 (and possibly 3), and stronger expression of stripes 4 and 5, with expanded interstripe distance between stripes 5 and 6. (I) Dorsal view of gastrula embryo at approximately the same stage as in (H). (J) Dorsolateral view of germ-band extension embryo with fading anterior stripes, and an additional pair-rule stripe appearing between stripes 5 and 6 (arrowhead). (K) Germ-band extension embryo exhibiting seven pair-rule stripes of expression with segmental stripes appearing in posterior interstripes (arrowheads). (L) Ventral view of fully extended germ band exhibiting segmental expression in all segments, in addition to continuous expression in the ventral anterior of the embryo.
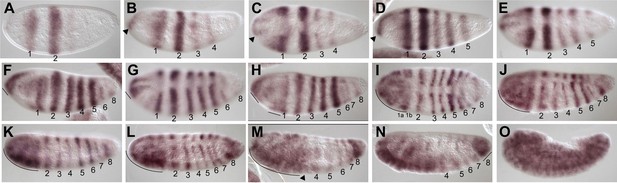
Summary of Nv hairy mRNA expression.
(A) Blastoderm embryo with two double-segment periodicity stripes of Nv hairy expression. Note that stripe 2 is broader and stronger than stripe 1. (B) Blastoderm embryo showing four double-segment periodicity stripes of expression plus an anterior accumulation of Nv hairy transcripts (arrowhead). (C) Dorsal view of embryo as in (B), illustrating the dorsal anterior expression (arrowhead) that is activated in the same pattern as the anterior domain of Nv tailless (Lynch et al., 2006). (D) Blastoderm embryo with strong anterior and dorsal anterior expression of Nv hairy and five pair-rule stripes. (E) Dorsal view of embryo as in (D) with increased dorsal anterior expression of Nv hairy, and the anterior spreading of expression from the anterior of double-segment pair-rule stripe 1. (F) Blastoderm embryo with expanding anterior domain (line), five double-segment ‘pair-rule’ stripes, and two additional stripes coming up. Note that the anterior domain between stripe 1 and the anterior pole is becoming more continuous in expression. (G) Dorsolateral view of embryo as in (F) highlighting the dorsal anterior expression. Stripe 2 is still wider than other stripes. Stripe 6 appears to be of single-segment periodicity. (H) Early gastrula embryo exhibiting a non-homogenous but largely continuous anterior cap of Nv hairy expression (that includes stripe 1). Four additional double-segment stripes and three single-segment stripes (two derived from stripe 6) are now evident. (I) Dorsal view of embryo slightly older than embryo in (H) showing the nearly continuous head domain, and the apparent splitting of stripe 1 within that domain. Double-segment stripes are thinning. (J) Dorsolateral view of extending germ-band embryo. Head domain is continuous (line). Stripes 1–7 have single-segment periodicity, are of non-uniform strength; stripe eight appears darker and broader. (K) Germ-band extending embryo with a continuous head domain (line) and eight discrete stripes. (L) Dorsolateral view of germ-band extending embryo. Stripe 8 is expanded into a wedge abutting the pole cells, and the anterior domain is expanding to include stripe 2. (M) Germ-band extension embryo with expanding anterior domain, that extends to include stripe 3 (arrowhead). Posterior domain is expanded. (N) Dorsolateral view of embryo as in (M) showing further expansion of posterior stripe 8 domain (line). (O) Germ-band-retracted embryo exhibiting ubiquitous staining with striated expression evident.
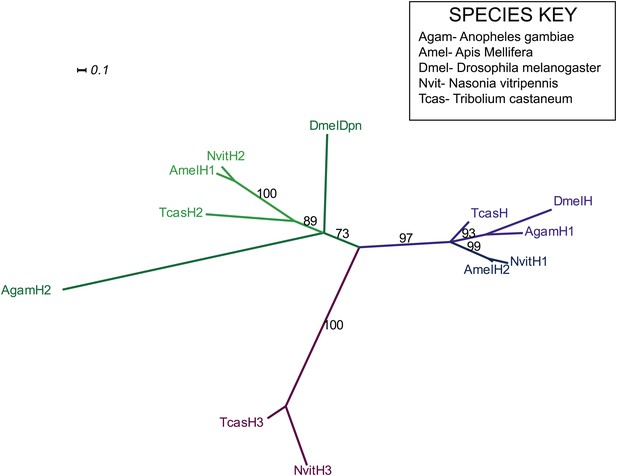
Phylogenetic analysis of hairy.
Phylogenetic analysis of hairy-related protein sequences was performed using Maximum likelihood analysis with 1000-fold bootstrap support, using RAxML via the CIPRES portal (Copf et al., 2003; Nagy et al., 1994). The best scoring maximum likelihood tree is shown with bootstrap support values indicated adjacent to branch.
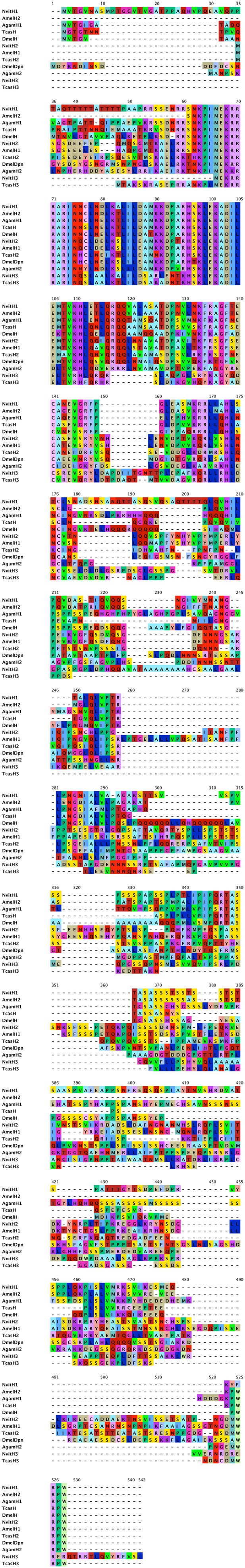
Hairy protein sequence alignment.
Predicted hairy-like protein sequences from Nasonia were aligned to hairy-like proteins from Drosophila melanogaster, Anopheles gambiae, Apis mellifera, and Tribolium castaneum using ClustalW. Full-length sequences were used for the alignment.
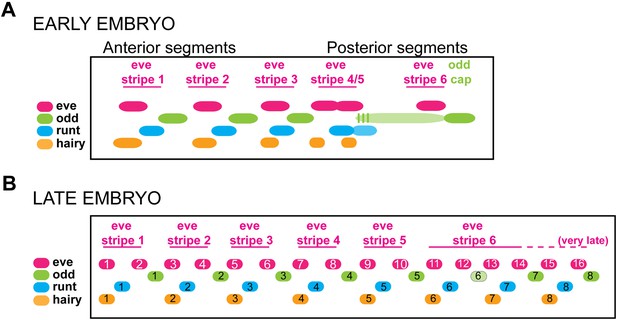
Summary model of pair-rule gene expression in the Nasonia embryo.
(A) Model of register of pair-rule gene expression in the early embryo. Nv eve and Nv odd stripes are totally complementary, whereas Nv runt stripes partly overlap each of these genes at their interface. Nv hairy stripes overlap Nv eve stripes toward the anterior of each double-segment periodicity stripe. Towards the posterior of the embryo, an extended domain of low-level Nv odd expression exhibits dynamic behavior over several nuclear cycles, and stripes 4/5 of Nv eve, Nv odd, and Nv runt each differentiate during this interval. Even more posteriorly, Nv eve stripe 6 lay anterior to a continuous Nv odd cap that extends to the posterior pole of the embryo. This region is set aside for segment specification and differentiation during germ-band extension. (B) Model of register of pair-rule gene expression in the germ-band extension (late) embryo. Single-segment periodicity stripes in the germ-band-extended embryo exhibit a variation upon early gene expression patterns. Nv eve single-segment stripes are interrupted by Nv runt and then Nv odd such that Nv runt stripes follow odd-numbered Nv eve stripes, and Nv odd stripes follow even-numbered Nv eve stripes. Each of 8 Nv hairy stripes overlaps odd-numbered Nv eve stripes that derive from the anterior of Nv eve pair-rule stripes. Additional expression of several of these genes in the ventral and head domains, which appears to rely on different regulatory logic, is not shown.





