p50-associated COX-2 extragenic RNA (PACER) activates COX-2 gene expression by occluding repressive NF-κB complexes
Figures
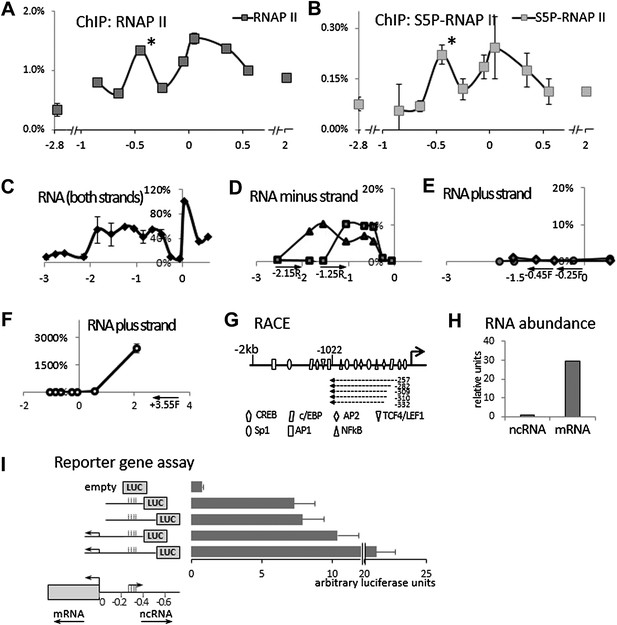
Identification of an antisense long non-coding RNA in the upstream region of the COX-2 gene.
(A and B) ChiP experiments were performed with chromatin extracts from HMEC cells with antibodies directed against RNAP II or phospho-serine 5 modified RNAP II, respectively, and analyzed by qPCR. Each histogram represents one qPCR data point (amplicon). Non-COX-2 promoter RNAP II peak is marked with an asterisk. X-axis scale is relative to COX-2 transcription start site. (C) Transcript abundance in the COX-2 locus in HMECs was measured by RT-qPCR. cDNA was primed with random oligos and therefore provide no information on strand specificity. Each diamond represents one RT-qPCR data point (amplicon). Y-axis values are relative to the signal obtained with the 5′ UTR amplicon. (D through F) Strand specific RT-qPCR. cDNA synthesis was performed with two sense oligos (m2R and jR) measuring antisense transcription or three antisense oligos (jF, eF, and control −3.55F) measuring sense transcription. (G) A schematic representation of lncRNA ends mapped with RACE. (H) Relative abundance of COX-2 mRNA and non-coding RNA. (I) A reporter gene assay used to measure the strength of the ncRNA promoter. Luciferase-only construct (empty) or constructs containing indicated fragments of COX-2 upstream region were transfected into 293T cells and luciferase activities were measured 48 hr post-transfection. Activity units reflect the ratios between Firefly and control Renilla luciferases.
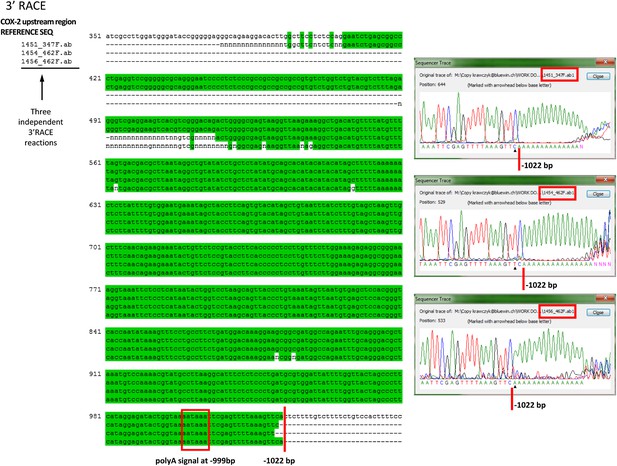
Mapping the 3′ end of the non-coding RNA transcript within the COX-2 upstream region by RACE.
Sequencing of three independent clones after amplification of 3′ ends. Oligos located at position −312 were used for initial RACE step, and resulting clones were sequenced with oligos at −347 or −462. Sequencer traces are shown on the right. Green shading indicates identical sequences.
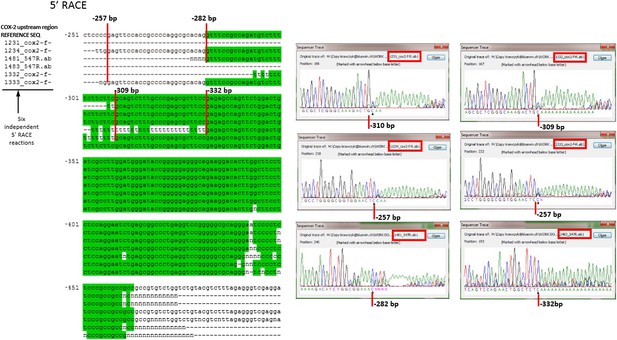
Mapping the 5′ end of the non-coding RNA transcript within the COX-2 upstream region by RACE.
Sequencing of six independent clones to map 5′ ends of the lncRNA. Nested RACE was performed using oligos at positions −900 and −700, and sequencing with oligos at −547 or −500 (f oligo).

Entire sequence of PACER, corresponding to the longest 5′ variant mapping to nucleotides −257 to −1022.
All positions relative to transcription start site of COX-2 mRNA.
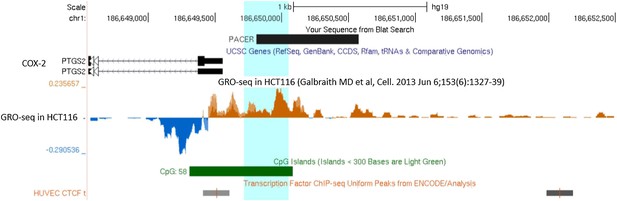
GRO-seq data supports the existence of a long non-coding RNA species within COX-2 upstream region.
A picture from UCSC genome browser (genome.ucsc.edu) showing COX-2 (marked PTGS2) region from 1 kb downstream (left) up to 3 kb upstream (right) of COX-2 transcription start site. PACER is depicted as mapped by our RACE experiments. Sense run-on transcription is shown below the horizontal line in blue, and antisense transcription in green above the line. Position of CTCF ChIP peaks from ENCODE study in HUVEC cells and location of CpG island are shown at the bottom.
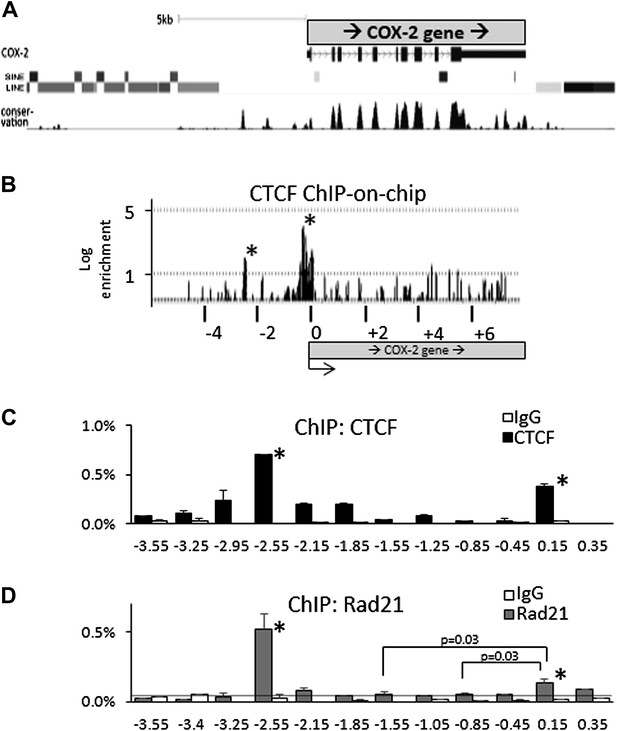
CTCF/cohesin complexes bind to two sites within the COX-2 locus, encompassing the promoters of mRNA and lncRNA.
(A) Genomic neighborhood of the human COX-2 gene. Note the repeat-DNA-free domain extending up to approximately −3 kb upstream of the COX-2 promoter. (B) ChIP-on-chip analysis of CTCF binding within the COX-2 genomic domain in human mammary epithelial cells (HMECs). Two CTCF peaks are evident at approximately −2.5 kb and the proximal promoter. Numerous other loci on the array served as negative controls (Figure 2—figure supplement 1). Results are represented as log2ratios between hybridization signals obtained with CTCF-ChIP and input DNA samples. Each vertical line represents one probe on the array. X-axis kb scale is relative to the COX-2 transcription start site. Significant signals are marked by asterisks. (C and D) Mapping of CTCF and cohesin binding in HMECs analyzed by ChIP-qPCR primer-walking. Amplicon coordinates are relative to the COX-2 transcription start-site.
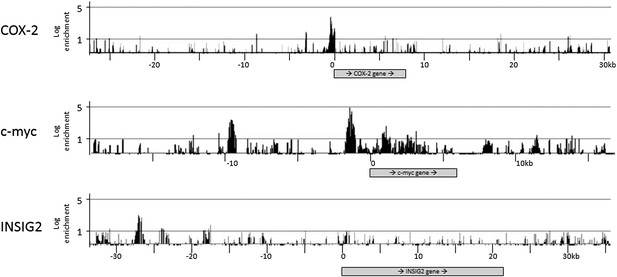
Selected results of CTCF ChIP-on-chip experiments.
The experiments were performed in duplicates with essentially identical results. COX-2 is located on chr1, c-myc on chr8 and INSIG2 on chr2.
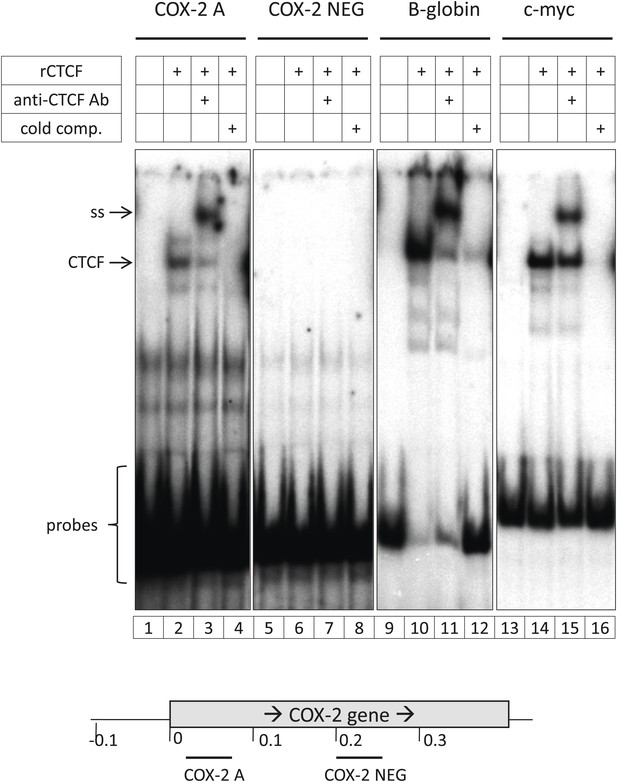
Identification and characterization of the 5’ UTR CTCF binding site in the COX-2 gene.
(A). EMSA assays were used to map the promoter-proximal CTCF binding site. Purified recombinant CTCF was incubated with oligonucleotide probes in the COX-2 region (lanes 1–4 and 5–8) B-globin LCR site IV (lanes 9–12) and c-myc (lanes 13–16). Lanes 1, 5, 9, and 13 served as probe-only controls. Anti-CTCF antibodies were added in lanes 3, 7, 11, and 15 to produce supershifts (ss). B-globin fragment was used as cold competitor in lanes 4, 8, 12, and 16.
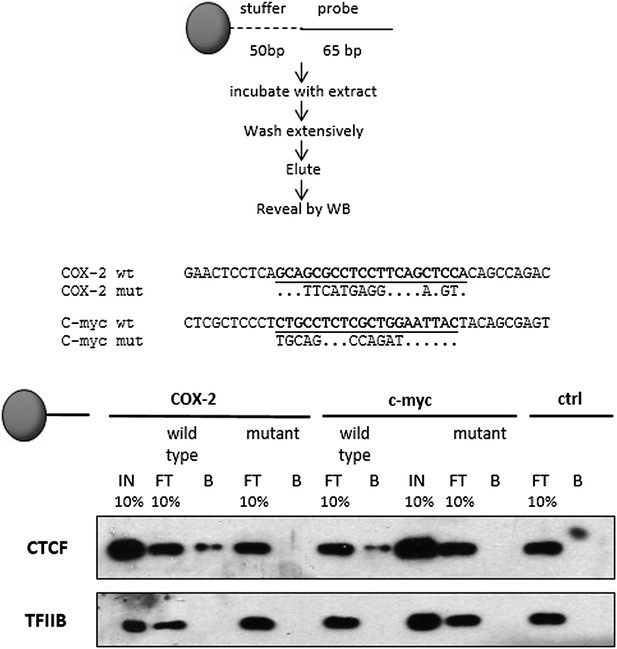
Mutations in predicted CTCF sites abolish CTCF recruitment to DNA templates.
Biochemical recruitment assays using extracts prepared from cells transfected with CTCF constructs were performed with wild-type or mutant COX-2 (lanes 1–5) or c-myc (lanes 6–10) templates. A template which did not contain recognizable CTCF sites served as a negative control (lanes 11–12). Wild type and mutant sequences are shown at the bottom. IN, input; FT, flowthrough; B, bead bound DNA-associated fraction.
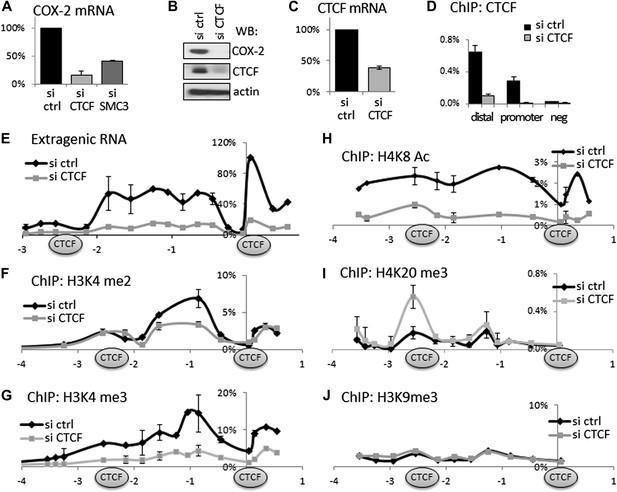
CTCF and cohesin maintain COX-2 mRNA and PACER lncRNA expression by demarcating a chromatin domain that is characterized by decreased H4K20 trimethylation, increased H3K4 di- and tri- methylation and increased histone acetylation.
(A) siRNA-mediated knockdown of CTCF or a cohesin subunit SMC3 reduces COX-2 mRNA levels in HMECs. RT-qPCR was performed with RNA prepared from HMECs 72 hr after transfection with the indicated siRNAs. Signals were normalized with 18S rRNA, TBP and GAPDH genes. (B) Western blot analysis of COX-2 and CTCF levels upon siRNA-mediated knockdown of CTCF. (C) CTCF mRNA levels in control and siRNA-CTCF transfected HMECs. (D) Binding of CTCF to the COX-2 promoter and distal sites were analyzed by ChIP-qPCR in control and CTCF knockdown cells. (E) Transcription across the COX-2 locus was measured by RT-qPCR in control (black diamonds) or siCTCF-transfected HMECs (grey squares). Signals were normalized as above and re-normalized using genomic DNA. (F–J) Levels of activation-associated (H3K4me2 and me3, H4K8Ac) and repression-associated (H4K20me3, H3K9me3) histone modifications were measured using ChIP-qPCR in HMECs transfected with control siRNAs (black lines) or siRNA against CTCF (grey lines). Each diamond or square denotes a qPCR amplicon. CTCF/cohesin binding sites are marked with oval shapes. X-axis scale is relative to the COX-2 transcription start site.
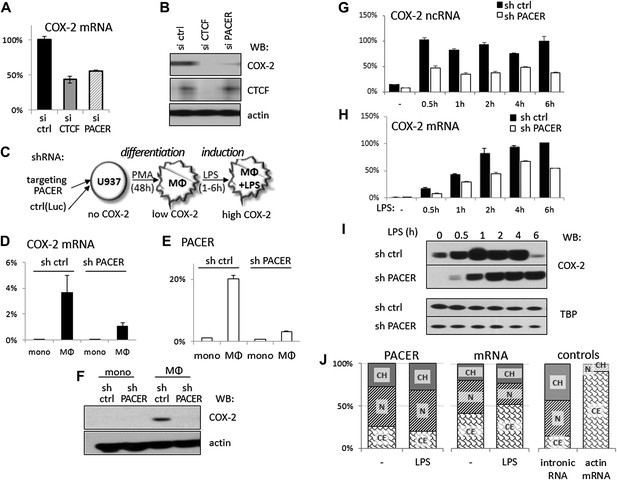
The antisense PACER lncRNA is a positive regulator of COX-2 expression in human mammary epithelial cells and in monocyte-derived macrophages before and after LPS stimulation.
Levels of COX-2 mRNA (A) and protein (B) were measured by RT-qPCR and Western blotting in control-scrambled siRNA, siRNA-CTCF, and siRNA-lncRNA transfected HMECs 72 hr post transfection. (C) Schematic representation of the monocyte-macrophage system used in this study. The human monocyte cell line, U937, was differentiated to macrophages with PMA and induced to express high levels of COX-2 by LPS stimulation. (D and E) COX-2 mRNA (D) and PACER (E) levels were measured by RT-qPCR in U937 lines carrying stably integrated control shRNA or shRNA-PACER expressing constructs before and after differentiation into macrophages. Values were normalized using 18S rRNA, TBP, and ß-globin genes. (F) COX-2 protein expression analyzed by Western blotting in the same experiment. (G and H) COX-2 ncRNA and mRNA levels were measured in a time course of LPS stimulation in control U937 (black bars) and PACER knockdown U937 macrophages (white bars). Values are relative to maximal COX-2 expression in control U937 cells after 6 hr LPS stimulation. (I) COX-2 protein expression analyzed by Western blotting in the same experiment. TBP served as a control. (J) Subcellular localization of COX-2 mRNA, PACER lncRNA and controls: nuclear intronic RNA and exclusively cytoplasmic actin mRNA. Relative levels in each fraction were measured by RT-qPCR and plotted such that they add up to 100%. CH, chromatin-bound fraction, N, nucleoplasm, CE, cytoplasm.
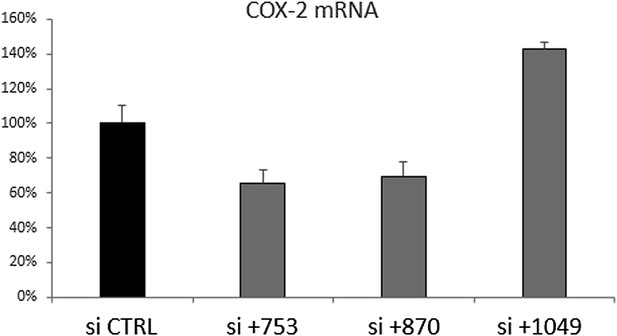
Knockdown of COX-2 mRNA with siRNA targeting PACER (si+753 and si+870) or positioned outside PACER (+1049).
Numbers are relative to COX-2 transcription start site, PACER extends from position +257 to +1022. Scrambled sequence was used as CTRL. Knockdown was performed by transient transfection of siRNAs into HMEC cells. RNA was harvested 3 days post transfection.
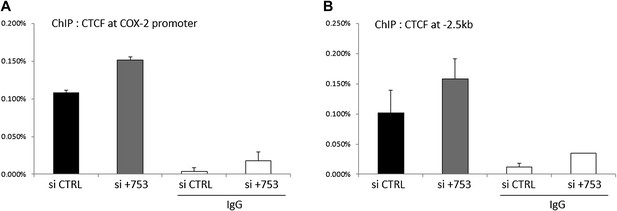
Binding of CTCF to the COX-2 promoter upon PACER knock-down in HMECs.
Binding of CTCF to the COX-2 promoter (A) and distal site (−2.5 kb, B) was assessed by ChIP.
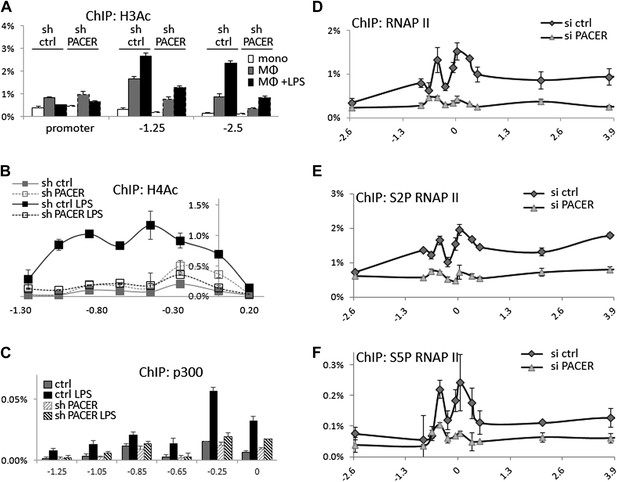
PACER facilitates recruitment of p300 HAT and RNAP II pre-initiation complexes to increase histone acetylation and induce COX-2 transcription.
(A) ChIP-qPCR was used to measure levels of histone H3 acetylation in control U937 and sh-lncRNA U937 monocytes (white bars), macrophages (grey-bars) and LPS-stimulated macrophages (black bars) at the COX-2 promoter, −1.25 kb upstream and −2.5 kb upstream of the COX-2 transcription start site. (B) ChIP-qPCR analysis of histone H4 acetylation across the COX-2 upstream region in control and sh-PACER U937 macrophages before and after LPS stimulation. (C) Association of p300 with the COX-2 upstream region was analyzed by ChIP-qPCR in control (filled squares) and sh-PACER (open squares) U937 macrophages before and after LPS stimulation. (D–F) Association of bulk RNAP II (D), S2-phosphorylated (E) and S5-phosphorylated (F) RNAP II across the COX-2 locus analyzed by ChIP-qPCR in control HMECs (diamonds) or PACER-knockdown (triangles) HMECs. Association of RNAP II with the COX-2 locus in control or siRNA-PACER transfected HMECs was assayed using ChIP-qPCR 72 hr post transfection. X-axis scale is relative to the COX-2 transcriptional start site.
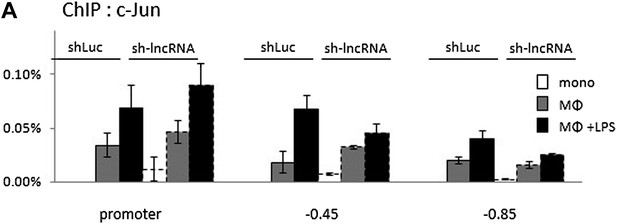
Binding of c-Jun to the COX-2 region upon PACER knock-down in U937 monocytes, macrophages, and LPS-stimulated macrophages analyzed by ChIP.
https://doi.org/10.7554/eLife.01776.017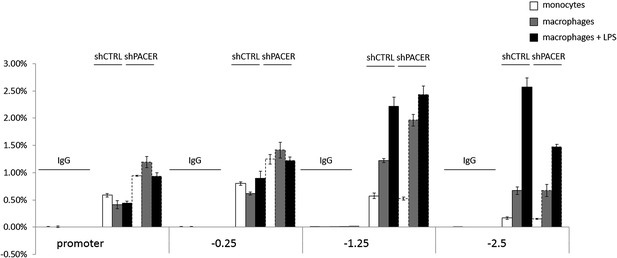
Levels of H3K4 dimethylation at COX-2 region upon PACER knock-down in U937 monocytes, macrophages, and LPS-stimulated macrophages analyzed by ChIP.
https://doi.org/10.7554/eLife.01776.018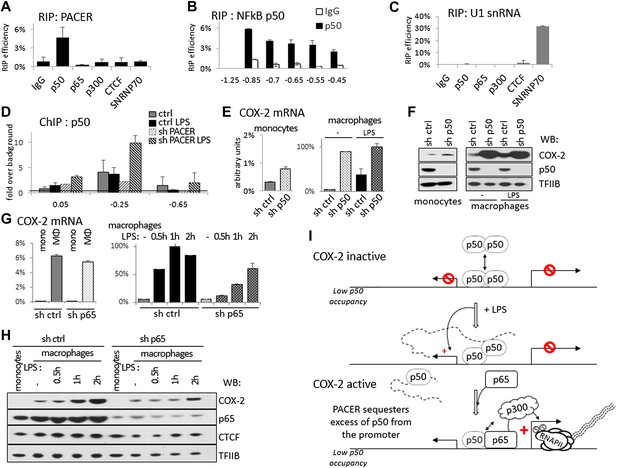
PACER controls COX-2 expression through binding of the repressive NF-κB subunit p50.
(A) Direct association of p50, p65/RelA, p300, and CTCF with PACER analyzed by RNA immunoprecipitation (RIP) in U937 macrophages. Primers for amplicon −0.85 kb were used. Signal is detected only in p50 immunoprecipitates. (B) Direct association of p50 with PACER analyzed by RNA immunoprecipitation (RIP) in U937 macrophages. Sites −0.45 kb through −0.85 kb lie within the lncRNA, while site −1.25 kb is outside and shows no detectable signal. IgG immunoprecipitation is shown as a control. (C) Control RIP experiments using the same antibodies as in (A) to analyze association with U1 snRNA. (D) Association of a small NF-κB subunit p50 with the COX-2 control region was analyzed by ChIP-qPCR in control and sh-lncRNA U937 macrophages. Binding is evident at the −0.25 kb site. (E and F) Expression of COX-2 mRNA (E) and protein (F) analyzed in control U937 monocytes and in mock and LPS-stimulated U937 macrophages and upon p50 knockdown. (G and H) Expression of COX-2 mRNA (G) and protein (H) analyzed in control U937 monocytes and in mock and LPS-stimulated U937 macrophages and upon p65 knockdown (I). A model of PACER-controlled COX-2 activation involving restricting promoter interaction of repressive p50 to facilitate recruitment of p300 HAT and RNAP II pre-initiation complexes to activate mRNA transcription.
Additional files
-
Supplementary file 1
Sequences of oligonucleotides used in this study
- https://doi.org/10.7554/eLife.01776.020





