Holocentromeres are dispersed point centromeres localized at transcription factor hotspots
Figures
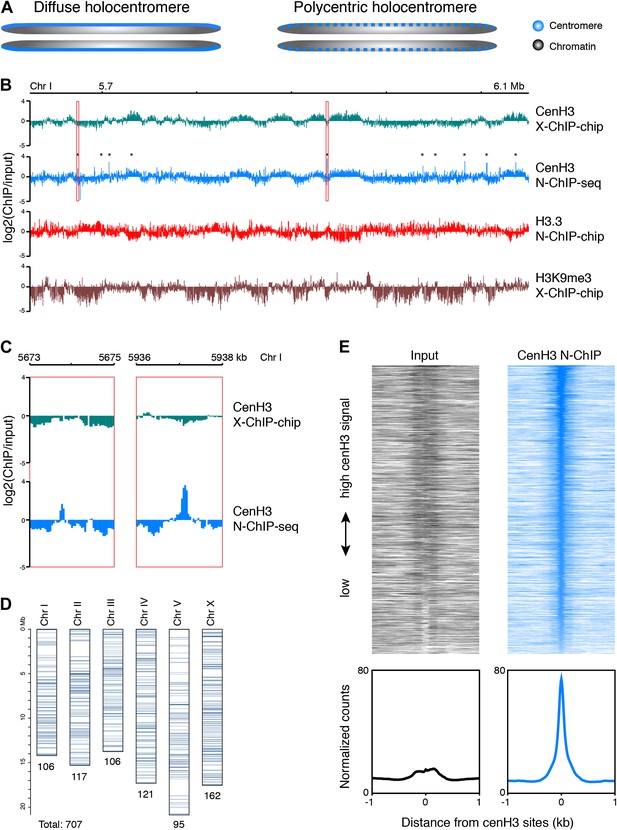
Genome-wide distribution of cenH3.
(A) Classic holocentromere models proposed by Schrader (Schrader, 1947): diffuse and polycentric holocentromeres. The diffuse model predicts full centromere coverage of the chromosomes. The polycentric model predicts discrete centromeric sites that together give the appearance of holocenticity. (B) Genome browser view of 525 kb on Chr I for cenH3 X-ChIP-chip (Gassmann et al., 2012), cenH3 N-ChIP-seq (this study), H3.3 N-ChIP-chip (Ooi et al., 2010) and H3K9me3 X-ChIP-chip (Liu et al., 2011), showing the close correspondance of cenH3 N-ChIP and X-ChIP signals for domains, but not peaks, and the positive correlation of cenH3 signal with H3K9me3 signal and the negative correlation of cenH3 signal with H3.3 signal. Log2 ratios of IP and input are shown to enable comparison between microarray and sequencing data. CenH3 peaks are marked by asterisks. (C) Genome browser view of cenH3 N-ChIP-seq (this study) and cenH3 X-ChIP-chip (Gassmann et al., 2012) at two representative cenH3 peaks marked in (B) with red boxes. (D) Number and genomic distribution of cenH3 peaks called per chromosome. See also Figure 1—source data 1. (E) Heatmaps and average plots of input and cenH3 N-ChIP signal within a 2-kb window around all 707 cenH3 peaks. Each line of the heatmaps represents an individual cenH3 site. The heatmaps are sorted from high to low cenH3 signal.
-
Figure 1—source data 1
Genomic coordinates of cenH3 peaks.
- https://doi.org/10.7554/eLife.02025.004
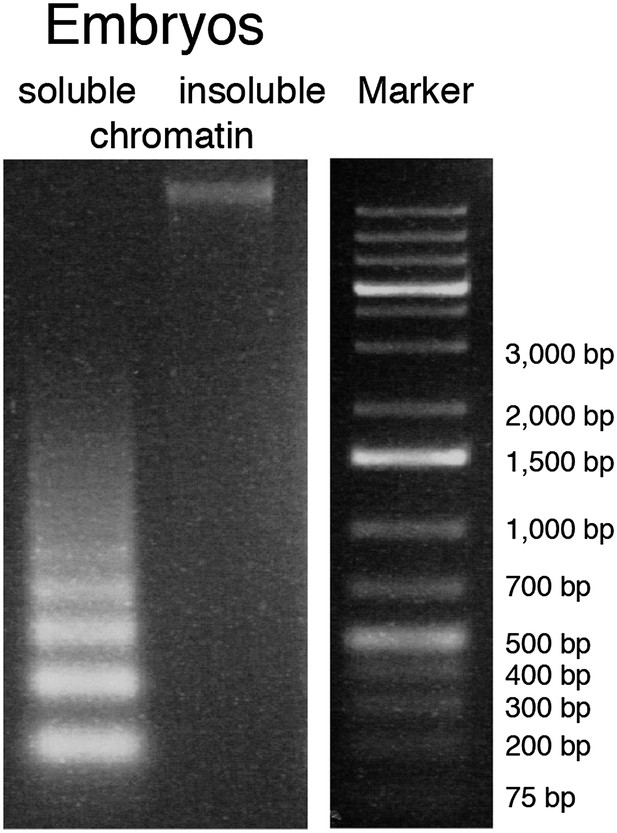
Solubilization of chromatin.
Agarose gel image of DNA isolated from soluble and insoluble chromatin after 2 min MNase digestion and needle extraction.
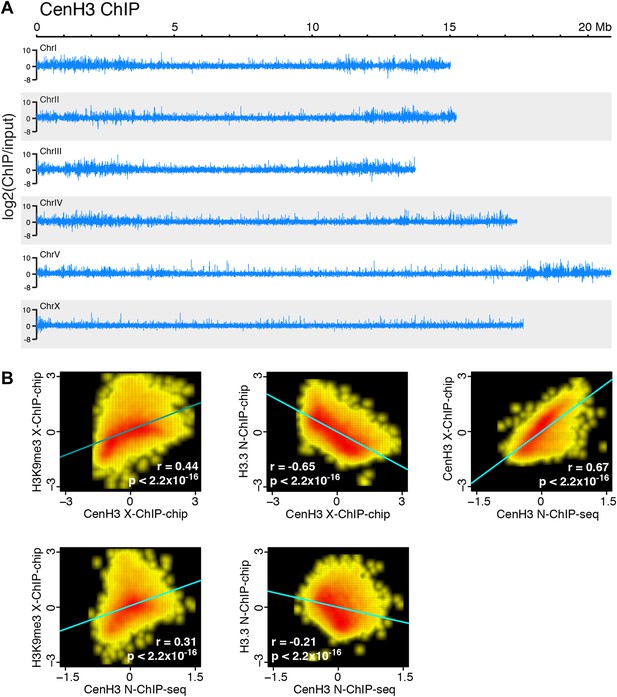
Comparison of cenH3 to H3K9me3 and H3.3 on a genome-wide scale.
(A) Genome browser view of cenH3 N-ChIP signal on a genome-wide scale. Log2 ratios of ChIP and input are shown. (B) Correlation density plots of cenH3 N-ChIP (this study) and cenH3 X-ChIP (Gassmann et al., 2012) with H3K9me3 X-ChIP (Liu et al., 2011) (left) and H3.3 N-ChIP (Ooi et al., 2010) (center). Correlation density plot of cenH3 N-ChIP with cenH3 X-ChIP (right). Average log2 ratios of ChIP over input in 10 kb-windows are plotted and used to calculate regression lines and correlation coefficients (r).

Analysis of cenH3 distribution and occupancy on a genome-wide scale.
(A) Distance between cenH3 peaks ranked by size for the entire genome (left) and for individual chromosomes (right). (B) Gene density at cenH3 sites. (C) Average cenH3 ChIP signal in the N-ChIP data (this study) and the X-ChIP data (Gassmann et al., 2012) within a 2-kb window around all 707 cenH3 sites. Average occupancy at domains defined in Gassmann et al. (Gassmann et al., 2012) is indicated by dashed lines for both data sets. (D) Box plots of fragment counts at cenH3 peaks, cenH3 domains, and regions between cenH3 domains for cenH3 N-ChIP data, illustrating that the highest signal genome-wide is found at cenH3 peaks. Differences of cenH3 ChIP and input samples are shown. The average cenH3 signal at each peak, domain, and region between domains was calculated, and the average values were plotted. N = 707 (peaks), 2865 (domains), 2868 (regions between domains). For cenH3 peaks, a 200-bp window around the summit of the peaks was used. Domains were defined in Gassmann et al. (Gassmann et al., 2012).
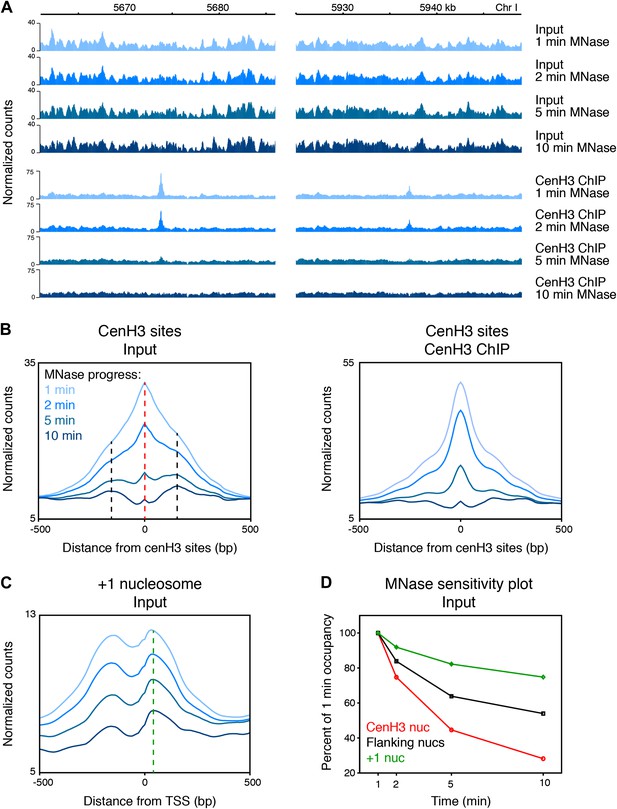
CenH3 peaks are especially MNase sensitive.
(A) Genome browser view of input chromatin and cenH3 ChIP signal within a 25-kb window surrounding two representative cenH3 peaks. Tracks for occupancy after 1 min, 2 min, 5 min and 10 min of MNase digestion are shown. (B) Average input signal (left) and cenH3 ChIP signal (right) within a 1-kb window around all 707 cenH3 sites after the indicated MNase digestion intervals. The dashed red line indicates the midpoint of the cenH3 nucleosome and the dashed black lines indicate the midpoints of the flanking nucleosomes. (C) Average input signal within a 1-kb window around 7043 transcriptional start sites (TSS) after the indicated MNase digestion intervals. The dashed green line indicates the midpoint of the +1 nucleosome. TSS were defined by Chen et al. (Chen et al., 2013). (D) MNase sensitivity plot for the cenH3 nucleosome and the flanking nucleosomes shown in (B) and the +1 nucleosome shown in (C). The occupancy of these nucleosomes at each MNase digestion time point was divided by the occupancy at the first time point. N = 707 (cenH3 nuc), 1414 (flanking nucs), 7043 (+1 nuc).
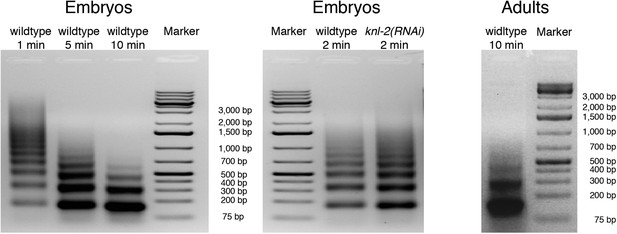
Progress of MNase digestion.
Agarose gel images of input DNA after MNase digestion of chromatin from wildtype and knl-2(RNAi) embryos and wildtype adults.
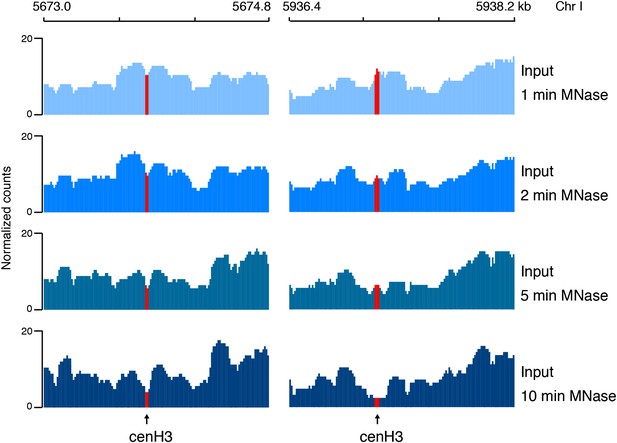
CenH3 sites are preferentially digested by MNase.
Genome browser view of input chromatin at two representative cenH3 sites after 1 min, 2 min, 5 min, and 10 min of MNase digestion, illustrating that the MNase-sensitivity of cenH3 nucleosomes is visible in the input chromatin. CenH3 sites are marked in red.
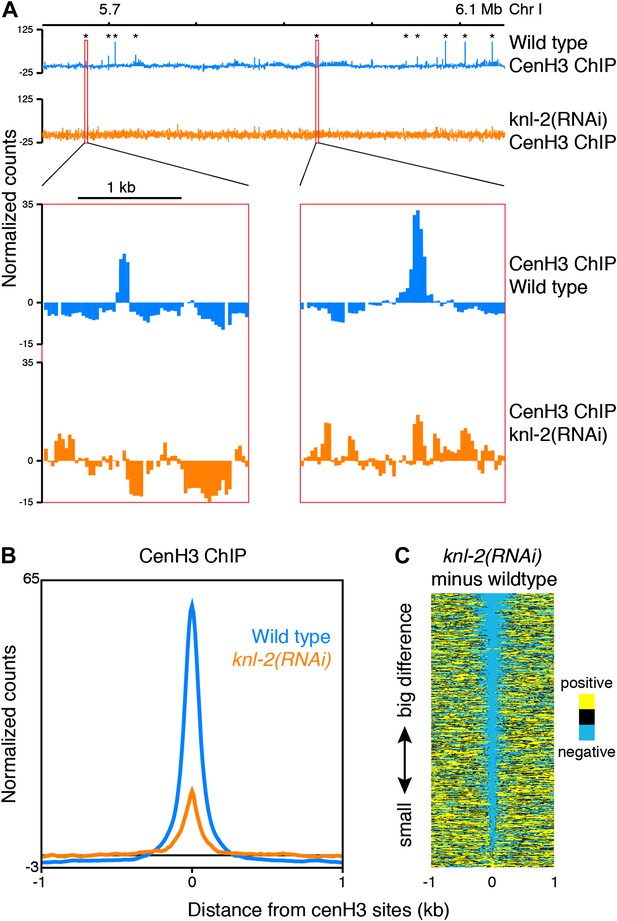
CenH3 peaks depend on cenH3 loading.
(A) Genome browser view of cenH3 ChIP in wildtype and knl-2(RNAi) embryos, with two enlarged cenH3 peaks. KNL-2 is required for cenH3 loading onto chromatin. Differences between ChIP and input are shown. CenH3 peaks are marked by asterisks. (B) Average cenH3 ChIP signal within a 2-kb window around all 707 cenH3 sites in wildtype and knl-2(RNAi) embryos. Differences between ChIP and input are plotted. (C) Heatmap of difference in cenH3 ChIP signal between wildtype and knl-2(RNAi). Each line of the heatmap represents an individual cenH3 site. The heatmap is sorted by decreasing difference between wildtype and knl-2(RNAi).
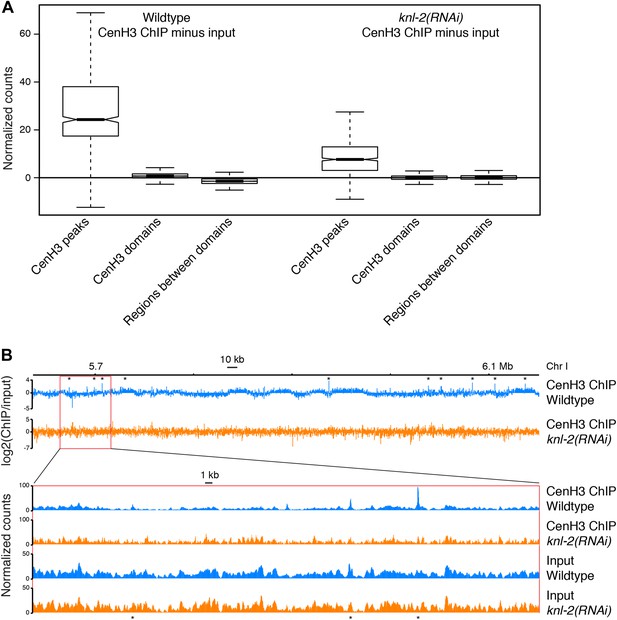
Comparison of wildtype and knl-2(RNAi) chromatin.
(A) Box plots of fragment counts at cenH3 peaks, cenH3 domains, and regions between cenH3 domains for cenH3 ChIP in wildtype and knl-2(RNAi) embryos. Differences of cenH3 ChIP and input samples are shown. The average cenH3 signal at each peak, domain, and region between domains was calculated, and the average values were plotted. N = 707 (peaks), 2865 (domains), 2868 (regions between domains). For cenH3 peaks, a 200-bp window around the summit of the peaks was used. Domains were defined in Gassmann et al. (Gassmann et al., 2012). (B) Genome browser view of cenH3 ChIP from wildtype and knl-2(RNAi) embryos. Log2 ratios of ChIP and input are plotted to visualize domains (top), and normalized counts are plotted for an enlarged region to show that the differences between the two samples are dominated by the ChIP, while input samples are highly similar. CenH3 peaks are marked by asterisks.
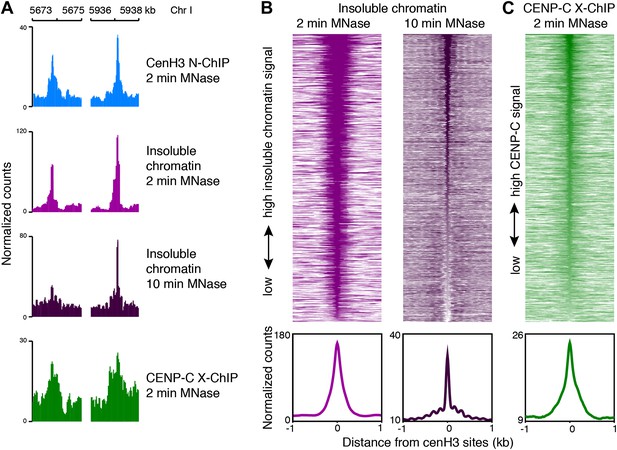
CenH3 sites are bound by the kinetochore.
(A) Genome browser view of cenH3 ChIP (native, 2 min MNase), insoluble chromatin (native, 2 min and 10 min MNase), and CENP-C ChIP (formaldehyde-crosslinked, 2 min MNase) signal at two representative cenH3 sites. (B and C) Heatmaps and average plots of insoluble chromatin signal after 2-min and 10-min MNase digestion (B) and CENP-C ChIP signal (C) within a 2-kb window around all 707 cenH3 sites. Each line of the heatmaps represents an individual cenH3 site. Heatmaps are sorted by decreasing signal.
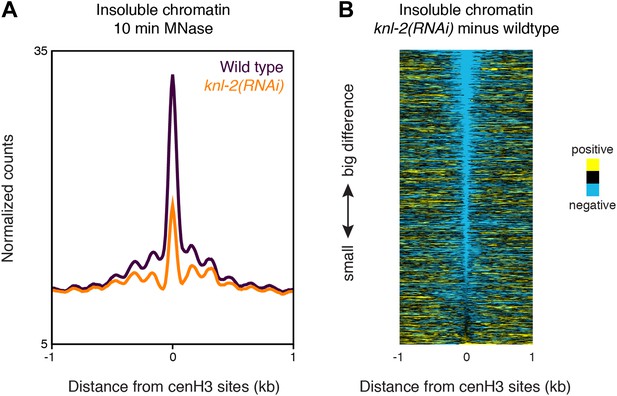
Insoluble chromatin at centromeric sites in knl-2(RNAi) embryos.
(A) Average insoluble chromatin signal after 10 min MNase digestion within a 2-kb window around all 707 cenH3 sites in wild-type and knl-2(RNAi) embryos. (B) Heatmap of the difference in insoluble chromatin signal between wildtype and knl-2(RNAi). Each line of the heatmap represents an individual cenH3 site. The heatmap is sorted by decreasing difference between wildtype and knl-2(RNAi).

Quantification of kinetochore occupancy.
(A) Box plots of fragment counts at cenH3 peaks, cenH3 domains, and regions between domains for insoluble chromatin (native), illustrating that the highest signal genome-wide is found at cenH3 peaks. (B) Box plots of fragment counts at cenH3 peaks, cenH3 domains, and regions between domains for CENP-C ChIP (formaldehyde-crosslinked), illustrating that the highest signal genome-wide is found at cenH3 peaks. Differences of CENP-C ChIP and input samples are shown. For (A) and (B), the average cenH3 signal at each peak, domain, and region between domains was calculated, and the average values were plotted. N = 707 (peaks), 2865 (domains), 2868 (regions between domains). For cenH3 peaks, a 200-bp window around the center of the high occupancy sites was used. Domains were defined in Gassmann et al. (Gassmann et al., 2012). (C) Normalized number of insoluble chromatin peaks coinciding with cenH3 peaks, cenH3 domains and regions between domains. (D) Normalized number of CENP-C peaks coinciding with cenH3 peaks, cenH3 domains and regions between domains. For (C) and (D), the number of insoluble chromatin peaks and CENP-C peaks overlapping with each group was divided by its genome coverage (141,400 bp for peaks, 42,722,880 bp for domains, 46,119,910 bp for rest of the genome) and then multiplied by 10 Mb.
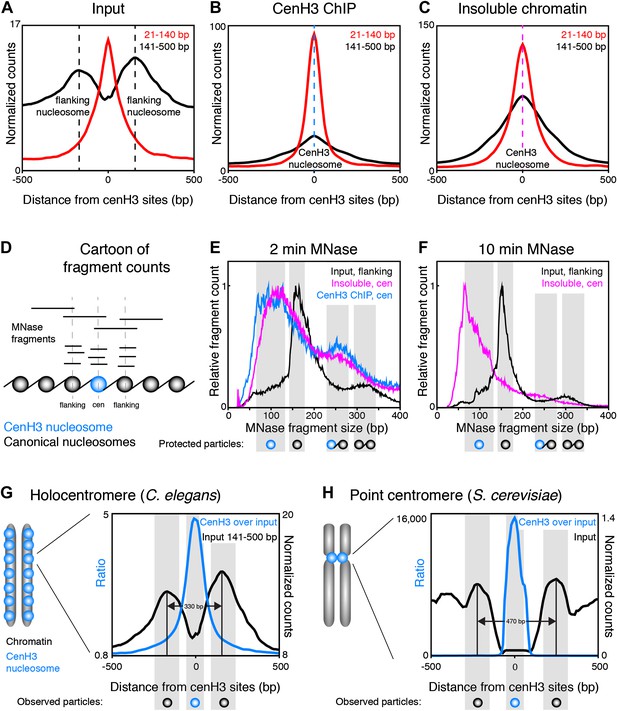
CenH3 nucleosomes protect small DNA fragments.
(A, B, C) Normalized fragment counts in input (A), cenH3 ChIP (B) and insoluble chromatin (C) samples at centromeric sites. MNase fragments were divided into nucleosomal (141–500 bp) and small (21–140 bp) size classes. Average signals within a 1-kb window around all 707 cenH3 sites are plotted. Dashed lines mark the centers of the flanking nucleosomes in (A) (black lines) or the centromeric nucleosome in (B) (blue line) and in (C) (magenta line). (D) Cartoon illustrating how MNase fragment size distributions shown in (E and F) were determined. Fragments that cross the center of the cenH3 nucleosome or of the flanking nucleosomes were counted. (E and F) MNase fragment size distribution after 2 min (E) and 10 min (F) of MNase digestion. Input fragments at flanking nucleosomes (black) and cenH3 ChIP fragments (blue) and insoluble chromatin fragments (magenta) at centromeric nucleosomes are shown. Cartoons of the protected particles are shown below each panel. (G and H) Comparison of worm holocentromere and budding yeast point centromere. (G) C. elegans holocentromere. Centromere model and cenH3 ChIP over input ratio (all size classes; left y-axis) and nucleosomal signal from input (141–500 bp; right y-axis). Average signals within a 1-kb window around all 707 cenH3 sites are shown. (H) Budding yeast point centromere. Centromere model and data from Krassovsky et al. (Krassovsky et al., 2012), cenH3 ChIP over input ratio (left y-axis) and input signal (right y-axis) from all 16 centromeres.

Comparison of input signal at centromeric sites in native and formaldehyde-crosslinked samples.
Nucleosome-size fragments at centromeric sites from native and formaldehyde-crosslinked input samples (top, MNase-seq, this study), and mononucleosome-size fragments from native and formaldehyde-crosslinked samples from the modENCODE database (http://intermine.modencode.org/), derived from microarray data (bottom).
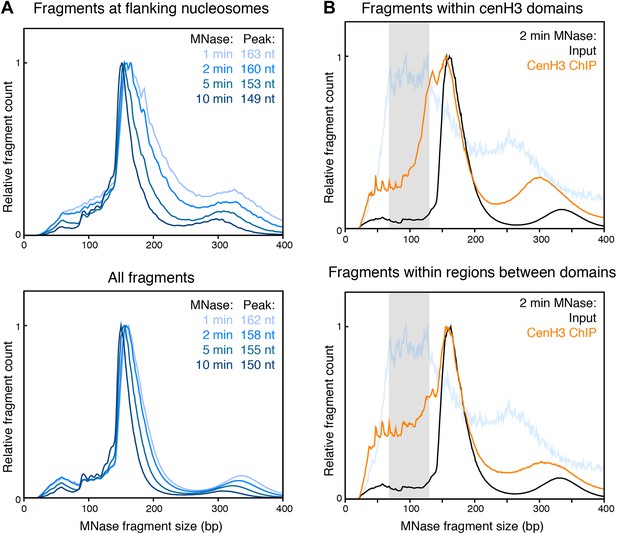
Fragment size distribution for different degrees of MNase digestions.
(A) MNase fragment size distribution of fragments crossing the dyad of the nucleosomes flanking the cenH3 peaks (top) or of all fragments genome wide (bottom). Fragment size distributions for the four indicated MNase digestion time points are shown. (B) MNase fragment size distribution of fragments within cenH3 domains (top) and regions between domains (bottom) for input and cenH3 ChIP after 2 min of MNase digestion. Domains were defined in Gassmann et al. (Gassmann et al., 2012). The MNase fragment size distribution of fragments crossing the center of the cenH3 nucleosome at cenH3 sites is shown in light blue for comparison, where the gray area marks the peak size interval.
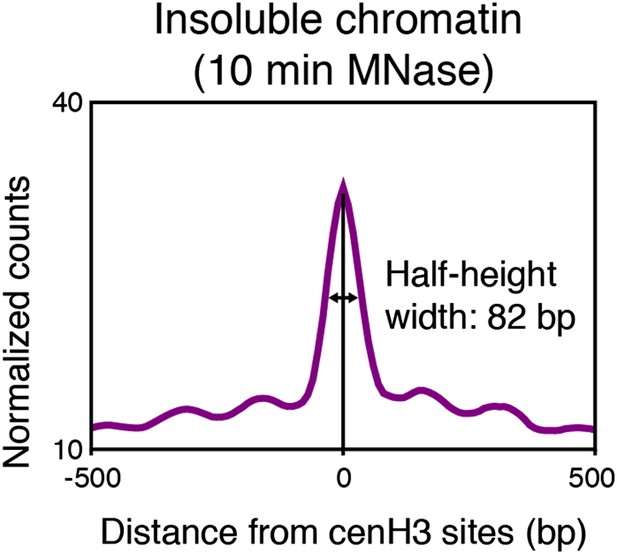
Particle size estimation from overdigested insoluble chromatin.
Half-height width of the average signal of insoluble chromatin at cenH3 peaks after 10 min MNase. Average signal within a 1-kb window around all 707 cenH3 sites is shown.
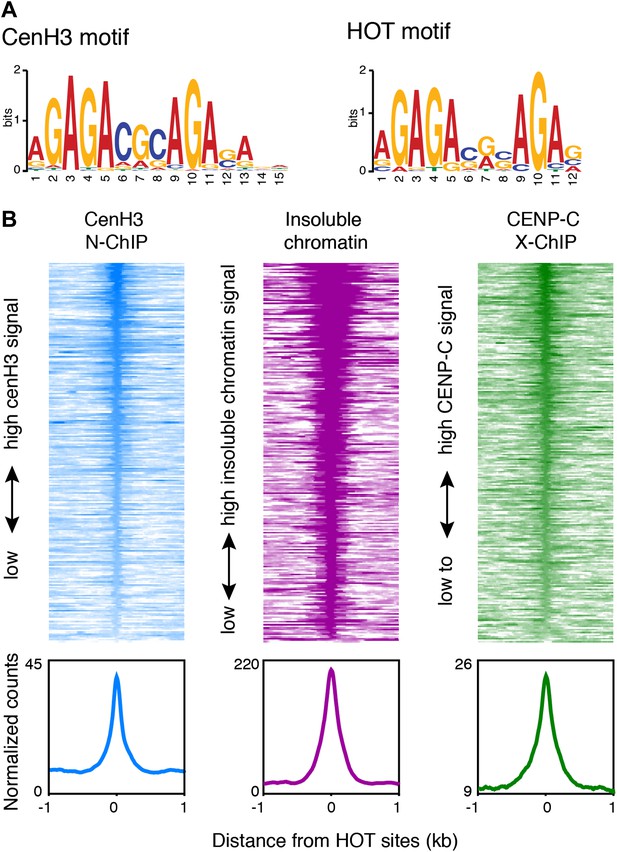
Centromeres coincide with transcription factor hotspots.
(A) MEME motif for the 80-bp cores of centromeric sites (left) and the 80-bp cores of high occupancy target (HOT) sites (right). (B) Heatmaps and average plots of cenH3 ChIP, insoluble chromatin and CENP-C ChIP signal within a 2-kb window around HOT sites, illustrating that HOT sites are highly occupied by cenH3, insoluble chromatin and CENP-C. Each line of the heatmaps represents an individual HOT site. Heatmaps are sorted by decreasing signal.
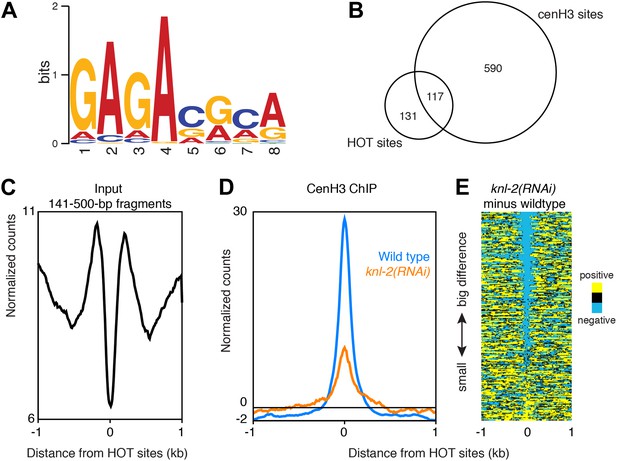
CenH3 site motif and characterization of HOT sites.
(A) MEME motif for 80-bp cores of all centromeric sites. (B) Venn diagram illustrating the overlap between cenH3 sites and HOT sites. Hypergeometric p=8.6 × 10−161. (C) Average signal of nucleosome-size input MNase fragments (141–500 bp) within a 2-kb window around all 248 HOT sites. (C) Average cenH3 ChIP signal within a 2-kb window around all 248 HOT sites in wildtype and knl-2(RNAi) embryos. Differences between ChIP and input are plotted. (D) Heatmap of difference in cenH3 ChIP signal between wildtype and knl-2(RNAi) at all 248 HOT sites. Each line of the heatmap represents an individual HOT site. The heatmap is sorted by decreasing difference between wildtype and knl-2(RNAi).
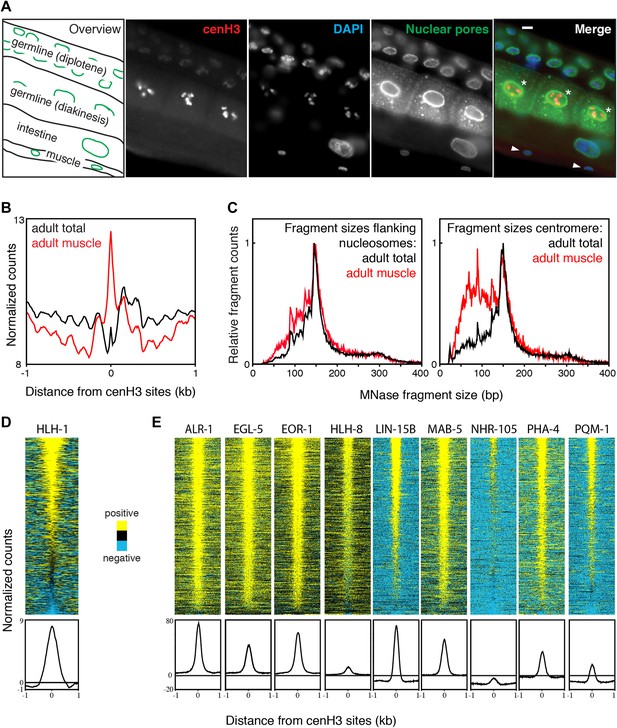
Transcription factor occupancy upon loss of cenH3 as cells exit the cell cycle.
(A) CenH3 in adult germline, intestine and muscle. Staining of a worm section with anti-cenH3, anti-NPP-9 (nuclear pores; staining control) and DAPI are shown. Germ cells in diakinesis are marked with asterisks, muscle cells with arrowheads in the merge. Scale bar is 3 µm. (B) Average MNase-seq signal within a 2-kb window around centromeric sites for adult total and adult muscle nuclei, illustrating that centromeric sites remain occupied in cenH3-depleted cells. MNase digestion >10 min. (C) MNase fragment size distribution at flanking nucleosomes and centromeric sites for total adult and adult muscle nuclei, illustrating that at least a sub-population of the particles occupying the centromeric sites are not canonical nucleosomes. (D) Heatmap and average plot of HLH-1 ChIP signal from adults within a 2-kb window around cenH3 sites. HLH-1 is a HOT site transcription factor. (E) Heatmaps and average plots of ChIP signal for another nine of the transcription factors used to define HOT sites within a 2-kb window around cenH3 sites, data from Gerstein et al. (Gerstein et al., 2010). All transcription factors in (E) were profiled in the third larval instar except ALR-1 (second larval instar) and PHA-4 (adults). Differences between ChIP and input are plotted in (D and E). Each line of the heatmaps represents an individual cenH3 site. Heatmaps are sorted by decreasing signal.
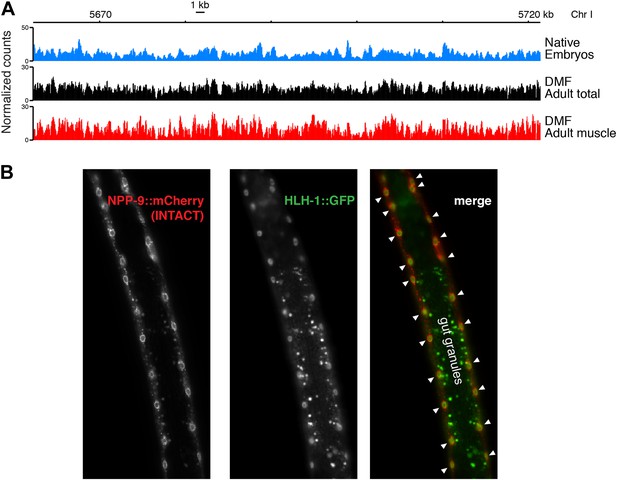
Comparison of chromatin from embryos and adults.
(A) Genome browser view of MNase-seq input signal from embryonic nuclei (native), total adult nuclei (dimethylformamide fixed), and adult muscle nuclei (dimethylformamide fixed). (B) Overlap of NPP-9::mCherry used for purifying muscle nuclei to obtain chromatin profiles (Steiner et al., 2012) and HLH-1::GFP, a muscle-specific HOT site transcription factor (Gerstein et al., 2010) in a live worm. Gut granules show auto-fluorescence in the green channel. Muscle nuclei are marked with arrowheads in the merge.





