tRNA synthetase counteracts c-Myc to develop functional vasculature
Figures
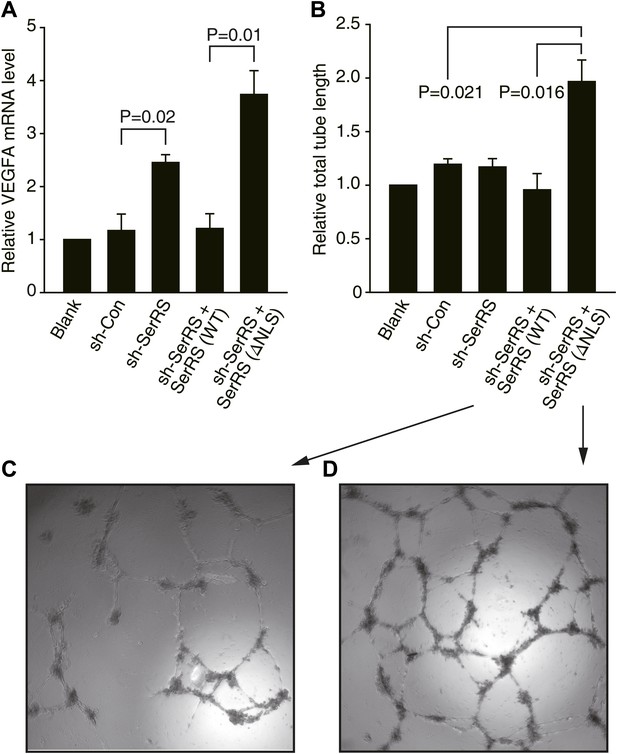
Nuclear SerRS suppresses VEGFA expression and angiogenesis.
(A) VEGFA mRNA levels as detected by real-time RT-qPCR in HUVECs infected with lentiviral plasmids expressing nonspecific control shRNA (sh-Con), SerRS-specific shRNA (sh-SerRS), or sh-SerRS and wild type (WT) or NLS-deleted (ΔNLS) SerRS simultaneously. Values are means ± SEM (n = 3). (B) Endothelial tube formation assay to show that excluding SerRS from the nucleus promotes angiogenesis. Values are means ± SEM (n = 3). (C and D) Representative images of the tubular network formed by HUVECs expressing WT and ΔNLS SerRS, respectively.
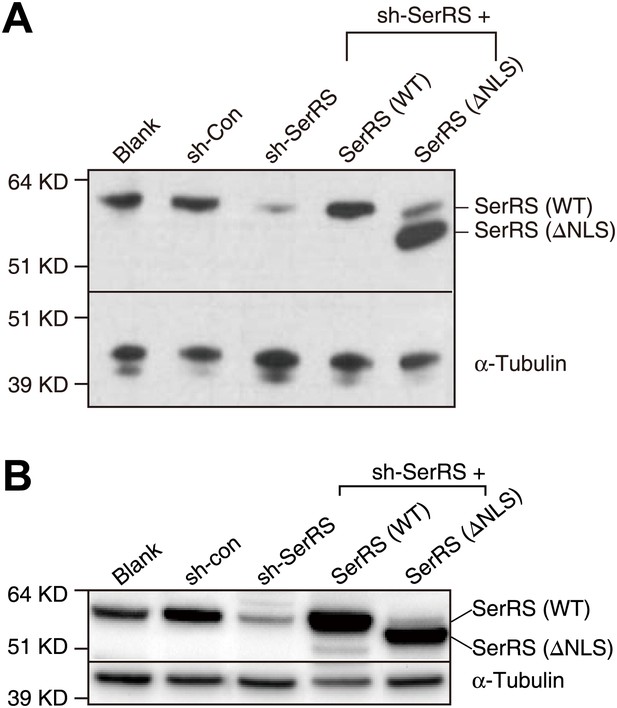
Manipulations of the expression of SerRS in HUVEC and HEK 293 cells.
HUVEC (A) and HEK 293 (B) cells were infected with lentiviruses expressing nonspecific control shRNA (sh-Con), shRNA targeting 3′-UTR of the endogenous SerRS gene (sh-SerRS), or sh-SerRS with WT or NLS-deleted (ΔNLS) SerRS simultaneously. The expression levels were measure 48 hr post-transfection by Western blot analyses.
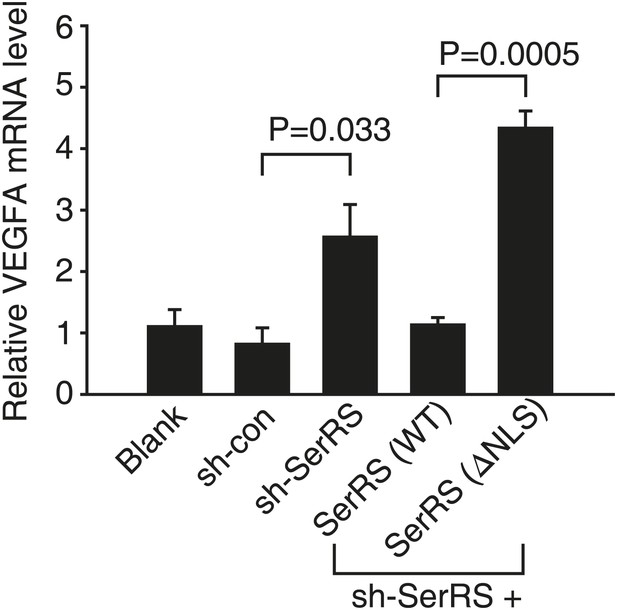
Nuclear SerRS suppresses VEGFA expression in HEK 293 cells.
VEGFA mRNA levels were detected by real-time RT-PCR in HEK 293 cells expressing indicated molecules. Values are means ± SEM (n = 3).
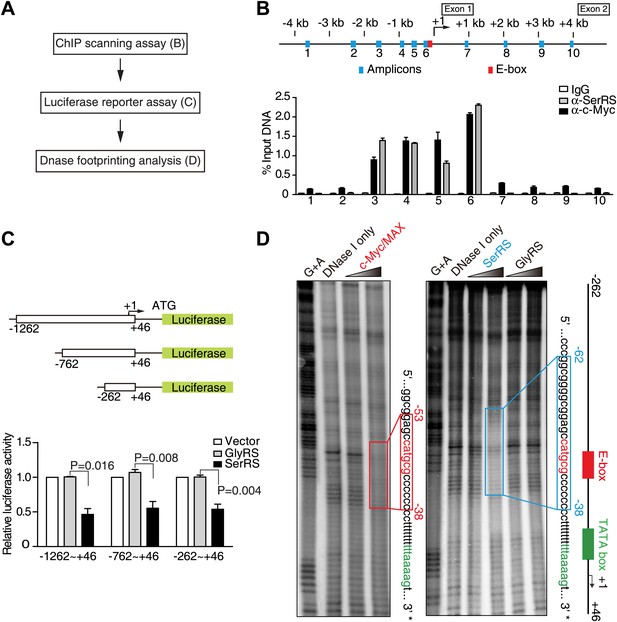
Identification of SerRS and c-Myc binding sites on the VEGFA promoter.
(A) Flow chart of consecutive methods used for determining the SerRS binding site. (B) Chromatin immunoprecipitation (ChIP) scanning assay to probe the SerRS and c-Myc binding sites. The promoter region of the VEGFA gene scanned by 10 amplicons is shown on the top. The amounts of DNA immunoprecipitated by anti-SerRS or anti-c-Myc antibodies or by control IgG from HEK 293 cell lysates were measured by real-time quantitative PCR at each amplicon. The results are represented as percentages of the total input of the chromatin DNA and shown as means ± SEM (n = 3). (C) Luciferase assay to confirm the repressive activity of SerRS and narrow down the SerRS binding site on the VEGFA promoter. Three different lengths of the VEGFA promoter were used to drive luciferase expressions in HEK 293 cells transfected with plasmids expressing SerRS, GlyRS or empty vector. The normalized luciferase activities are shown as mean ± SEM (n = 3). (D) In vitro DNase I footprint assay to identify the SerRS binding site. A 308-bp DNA fragment (−262 ∼ +46 on the VEGFA promoter) radiolabeled at the 3′ end was incubated with purified recombinant c-Myc/MAX (1:1 molar ratio), SerRS or GlyRS each at 1 or 5 µM, and then subjected to DNase I digestion. The regions protected by c-Myc/MAX and by SerRS are indicated in red and blue boxes, respectively.
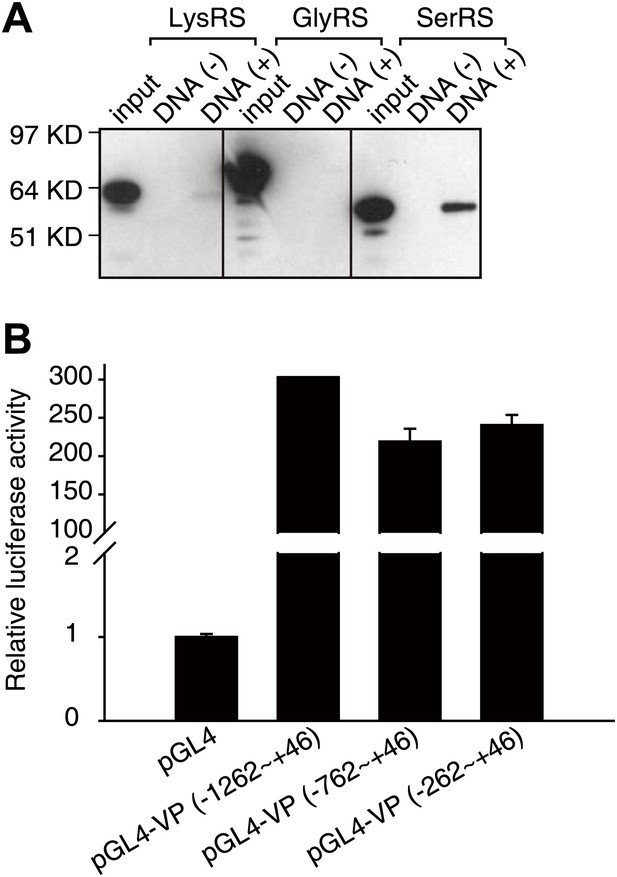
Identification of the interaction between SerRS and DNA.
(A) Human SerRS, but not LysRS or GlyRS, can bind to DNA. Purified, His6-tagged recombinant human LysRS, GlyRS, and SerRS proteins were incubated with calf thymus genomic DNA linked cellulose beads or empty beads. After wash, the beads were examined with Western blot analysis using anti-His antibody to detect protein binding. (B) Background luciferase activities driven by three different lengths of the VEGFA promoter. The promoter regions were inserted in front of the firefly luciferase reporter gene. The constructs (together with the Renilla luciferase control reporter vector for transfection efficiency control) were transfected into HEK 293 cells. The luciferase activities were measured 24-hr post-transfection, and the results are shown as relative firefly luciferase activities after normalized to Renilla luciferase activities and as means ± SEM (n = 3).
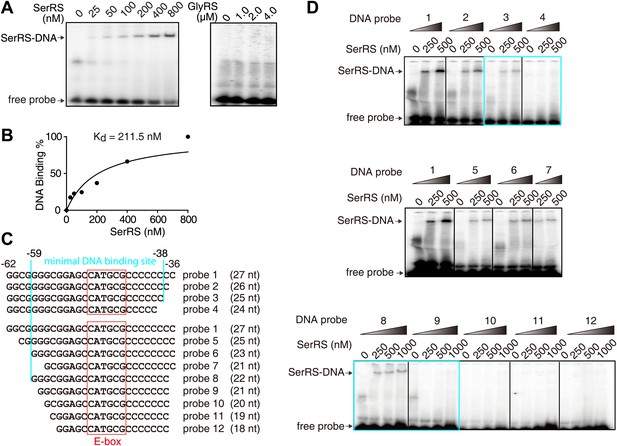
Characterization of the interaction between SerRS and DNA.
(A and B) In vitro EMSA assay to determine the binding affinity between SerRS and the 27-bp DNA. The 27-bp DNA fragment containing SerRS binding site on the VEGFA promoter (−62 ∼ −36) were labeled by 32P at the 5′ end, and then incubated with purified SerRS or GlyRS at indicated concentrations. The SerRS–DNA complex was followed by electrophoresis on native acrylamide gels. (C and D) EMSA to determine the minimal SerRS binding site on the VEGFA promoter. Truncations of the DNA from either end weakened the SerRS–DNA interaction. Purified recombinant SerRS protein was used at the indicated concentrations.
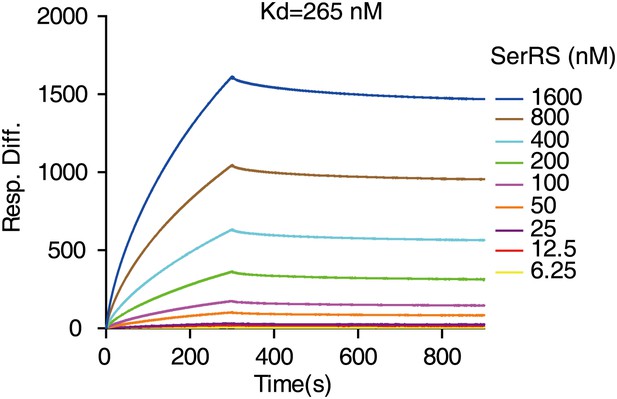
Determination of the binding affinity between SerRS and DNA by SPR.
The binding affinity of SerRS with the 27-bp DNA from the VEGFA promoter as measured by surface plasmon resonance (SPR) analysis.
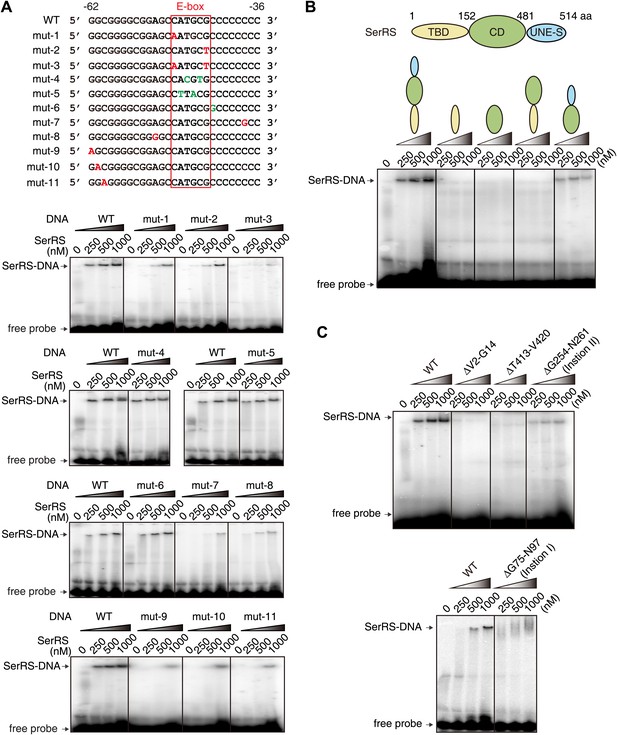
Further characterization of the interaction between SerRS and DNA.
(A) EMSA assay to probe the DNA sequence specificity for interacting with SerRS. DNA mutations that do or do not impact SerRS binding are colored in red and green, respectively. (B) Domain mapping analysis and EMSA assay to reveal multiple DNA binding sites on SerRS. TBD: tRNA binding domain; CD: catalytic domain; UNE-S: C-terminal appended domain unique to vertebrates. (C) Deletion mutagenesis to further define DNA binding sites on SerRS. Deletion of either insertion I, insertion II, motif V2-G14, or loop T413-V420 greatly weakens or abolishes the DNA interaction.
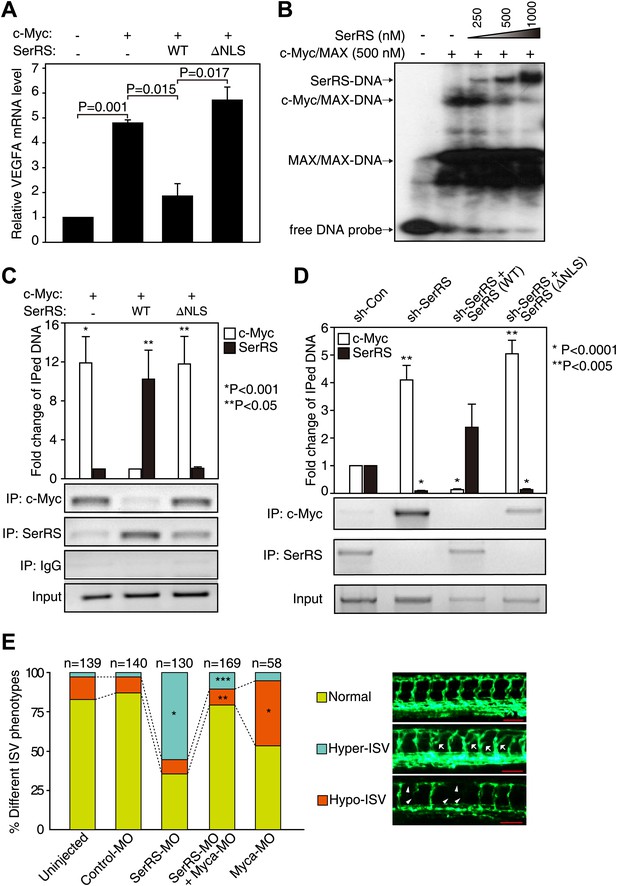
Competition between SerRS and c-Myc for DNA binding and their opposing effect in vascular development.
(A) Competition between c-Myc and SerRS on VEGFA expression. HEK 293 cells were transfected with c-Myc alone or c-Myc with WT or ΔNLS SerRS. The mRNA levels of VEGFA were determined by RT-PCR. Values are shown as means ± SEM (n = 3). (B) Competition between c-Myc/MAX and SerRS for DNA binding in vitro as examined by EMSA. The 27-bp DNA was radio-labeled and incubated with purified recombinant c-Myc/MAX together with purified recombinant SerRS at indicated concentrations. The protein–DNA complexes were followed by electrophoresis on a native acrylamide gel. (C) Competition between ectopically expressed SerRS and c-Myc for DNA binding on the VEGFA promoter in HEK 293 cells as examined by ChIP. HEK 293 cells were co-transfected with plasmids expressing c-Myc and WT or ΔNLS SerRS or empty vector (−) 24 hr prior to ChIP analysis. The amounts of DNA immunoprecipitated by anti-SerRS or anti-c-Myc antibodies or by control IgG from HEK 293 cell lysates were measured by PCR using a primer set targeting the VEGFA promoter. The normalized results (top panel) are represented as fold change of immunoprecipitated DNA by anti-SerRS vs anti-c-Myc and are shown as means ± SEM (n = 3, *p<0.001, **p<0.05). The bottom panel shows representative gel images. (D) Competition between endogenously expressed SerRS and c-Myc for DNA binding on the VEGFA promoter in HUVECs. HUVECs were infected to express the indicated molecules 48 hr prior to ChIP analysis. The same ChIP experiment and data analysis were performed as described in (C). *p<0.0001, **p<0.005. (E) Opposing effect of SerRS and c-Myc in zebrafish vascular development and their mutual phenotypic rescue. The percentage of Tg(Fli1a:GFP) zebrafish embryos showing different ISV phenotypes at 3 days post fertilization after the injection of morpholinos targeting SerRS (SerRS-MO), Myca (Myca-MO), or a control morpholino (Control-MO) are illustrated. Scale bars represent 0.125 mm. *p<0.0001 vs Control-MO, **p<0.0001 vs Myca-MO, ***p<0.0001 vs SerRS-MO. Control-MO was added to SerRS-MO or Myca-MO experiments in order to maintain a constant level of total morpholinos in each experiment.
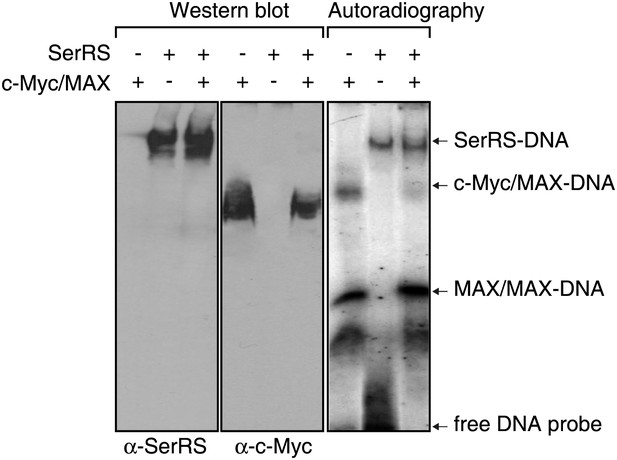
SerRS and c-Myc/MAX do not simultaneously bind to the DNA.
Purified recombinant SerRS (500 nM) and c-Myc/MAX (500 nM each) proteins were mixed as indicated and incubated with cold or 32P-labeled 27-bp DNA fragment on the VEGFA promoter (−62 ∼ −36). The mixtures were subjected to electrophoresis in 6% native polyacrylamide gel and the protein–DNA complexes were detected by autoradiography and Western blot analysis using anti-SerRS and anti-c-Myc antibodies, respectively.
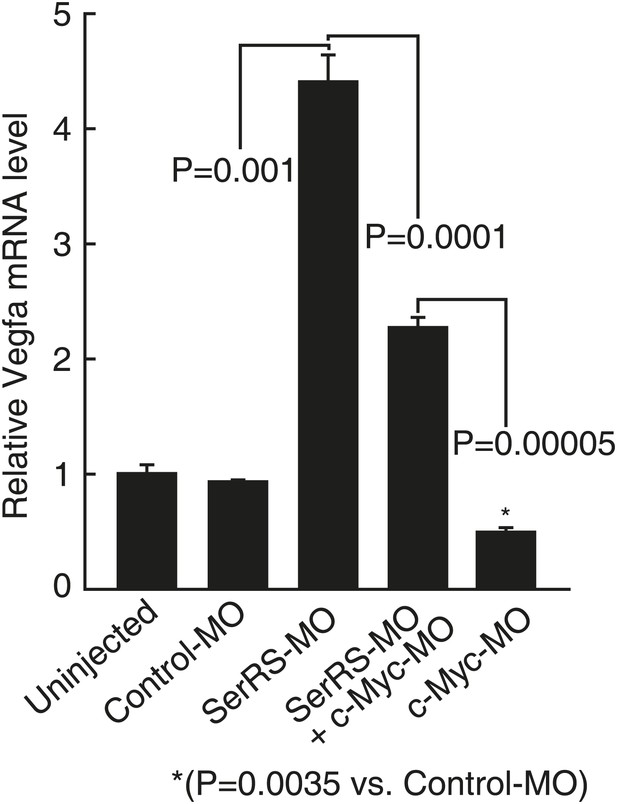
Effect of knocking down Myca or SerRS on Vegfa expression in zebrafish.
The effects of injecting the indicated morpholinos on Vegfa expression in zebrafish embryos (3 dpf) as measured by real-time RT-PCR. Data are shown as means ± SEM (n = 10–15).
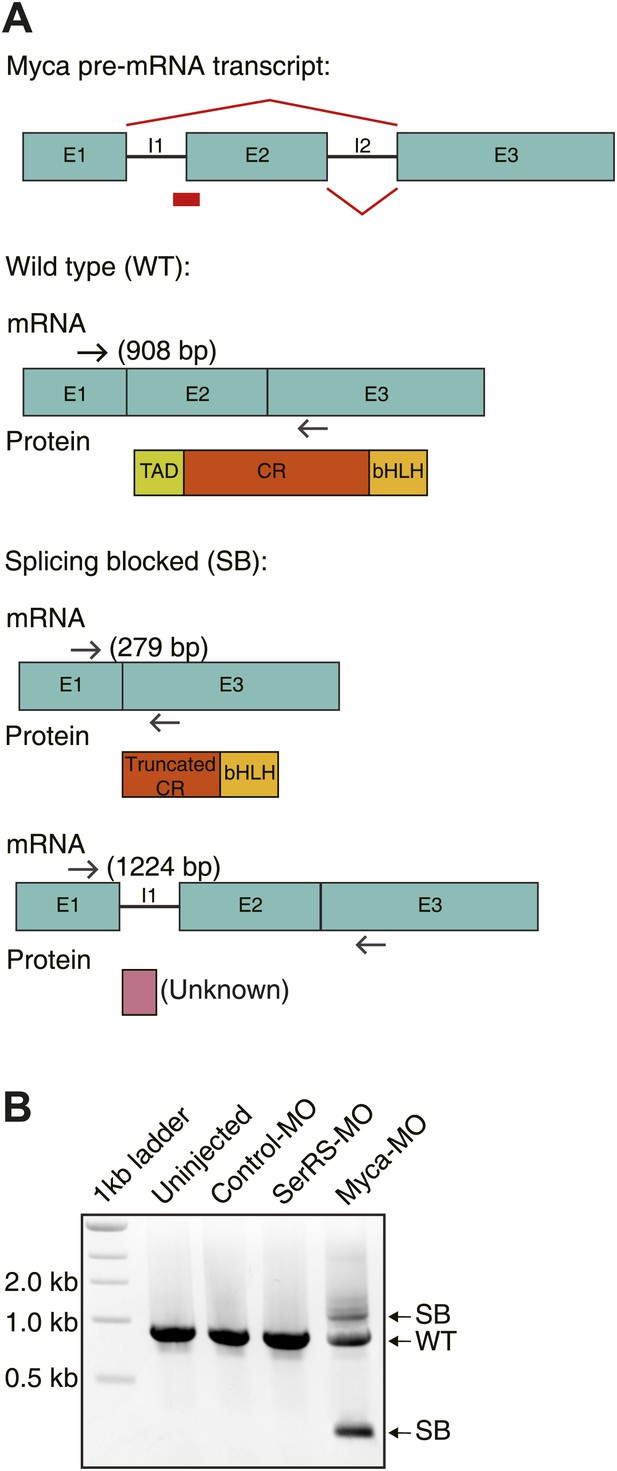
Design and efficiency of the antisense morpholino against Myca.
(A) The design of Myca-MO and the expected mRNA/protein products. The Myca-MO targets the intron1-exon2 splicing site of Myca precursor mRNA to potentially generate two alternatively spliced mRNA products (SB) that would be translated into non-functional proteins. (B) RT-PCR analysis confirming the dramatic decrease of WT Myca mature mRNA and the expected alternative splicing variants (SB) as the result of injecting Myca-MO.
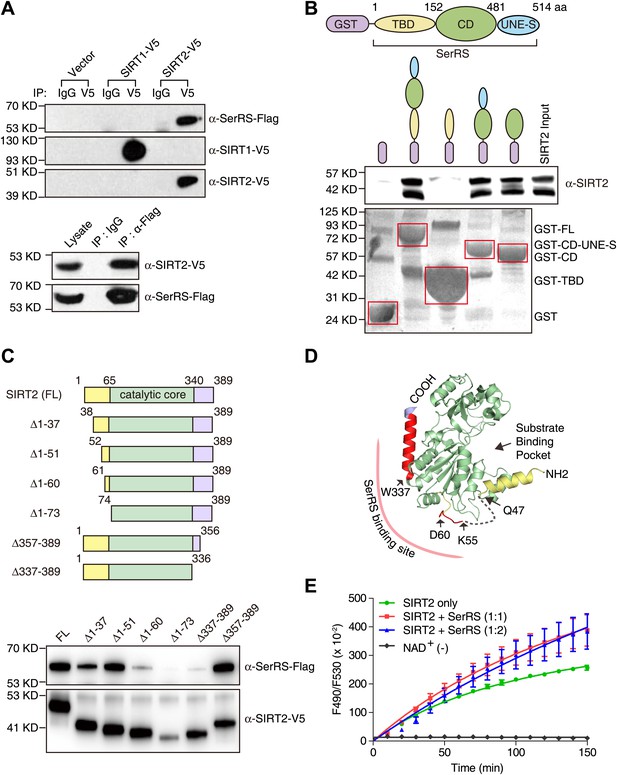
Demonstration and characterization of SerRS/SIRT2 interaction.
(A) SerRS specifically interacts with SIRT2 but not SIRT1. HEK 293 cells were co-transfected with plasmids expressing Flag-tagged SerRS and V5-tagged SIRT1 or SIRT2. Cell lysate was immunoprecipitated with anti-V5 (top panel), anti-Flag (bottom panel) antibodies or control IgG. The experiment was followed by Western blot analysis to detect the interaction between SerRS and SIRT1/SIRT2 using anti-Flag and anti-V5 antibodies. (B) GST-pull down assay to show that SerRS/SIRT2 interaction is direct and that the interaction is mediated by the catalytic domain of SerRS. Full-length SerRS or its domain fragments were fused with GST at N-termini to pull down purified His-tagged SIRT2. SIRT2 was detected by Western blot analysis using anti-His6 antibody, and the GST fusion proteins attached on the Glutathione-Sepharose beads were analyzed using ponceau S staining. TBD: tRNA-binding domain; CD: catalytic domain; UNE-S: C-terminal appended domain. (C) Mapping study to identify the SerRS binding sites on SIRT2. V5-tagged full-length SIRT2 or its truncated fragments was co-transfected with Flag-tagged SerRS into HEK 293 cells. SIRT2 proteins were immunoprecipitated with anti-V5 antibody and the SIRT2-bound SerRS proteins were detected by Western blot using anti-Flag antibody. (D) Illustration of the SerRS binding sites on the structure of SIRT2. Two SerRS binding sites (Gly52-Asp60, Trp337-Ser356) are highlighted in red. The catalytic domain of SIRT2 is in green, while the partially disordered N- and C-terminal regions are in yellow and purple, respectively. The gray dash line represents a disordered internal region. (E) Effect of SerRS on SIRT2 deacetylation activity. Recombinant human SIRT2 (1 µM) were incubated with purified SerRS (concentration measured as monomer) at the indicated ratios. The deacetylase activities of SIRT2 were measured by using a substrate peptide with one end coupled to a fluorophore and the other end to a quencher. An internal acetylated lysine residue serves as the substrate of SIRT2, and the deacetylation allows the peptide to be cleaved by a lysylendopeptidase to release the fluorophore from the quencher to emit fluorescence. Therefore, the SIRT2 acitivity was measured by monitoring the fluororescence intensity (excitation at 490 nm and emission at 530 nm). A reaction without NAD+ (NAD+ [−]) was performed as a negative control.
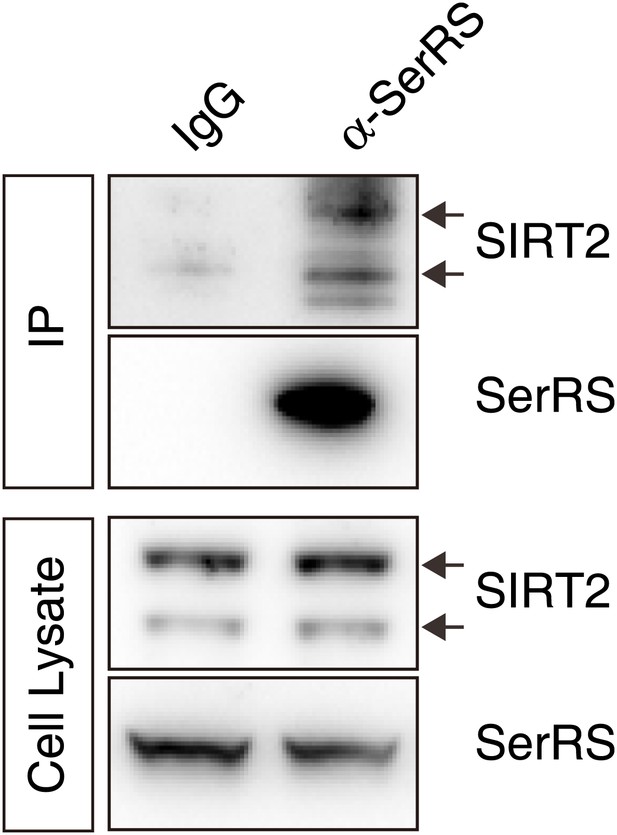
Endogenous interaction between SerRS and SIRT2.
Approximately 107 HEK 293 cells were lysed and the supernatant were immunoprecipicated with mouse monoclonal anti-SerRS antibody or the control mouse IgG. The co-immunoprecipitated proteins were analyzed by Western blot analysis using anti-SerRS and anti-SIRT2 antibodies, respectively.
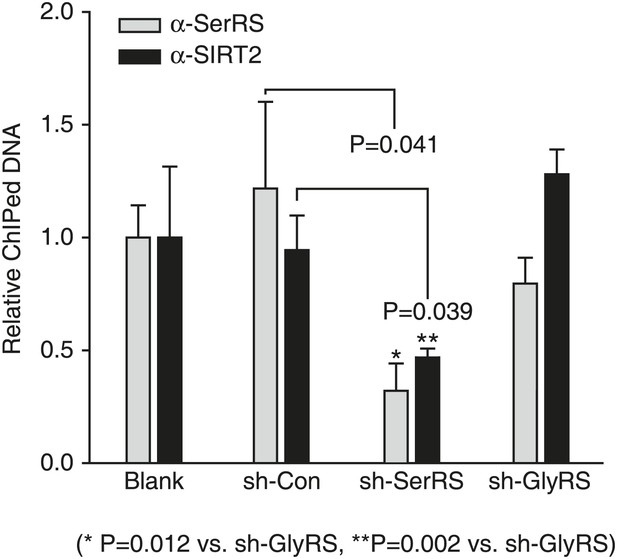
SerRS recruits SIRT2 to the VEGFA promoter.
ChIP assay to detect the impact of SerRS and GlyRS (as a control) on SIRT2 binding to the VEGFA promoter. HEK 293 cells were transfected with plasmids expressing the indicated shRNA for knocking down the expression of SerRS or GlyRS and subjected to ChIP analysis 48 hr post-transfection. The normalized results are represented as fold change of immunoprecipitated DNA by anti-SerRS vs anti-SIRT2 and are shown as means ± SEM (n = 3).
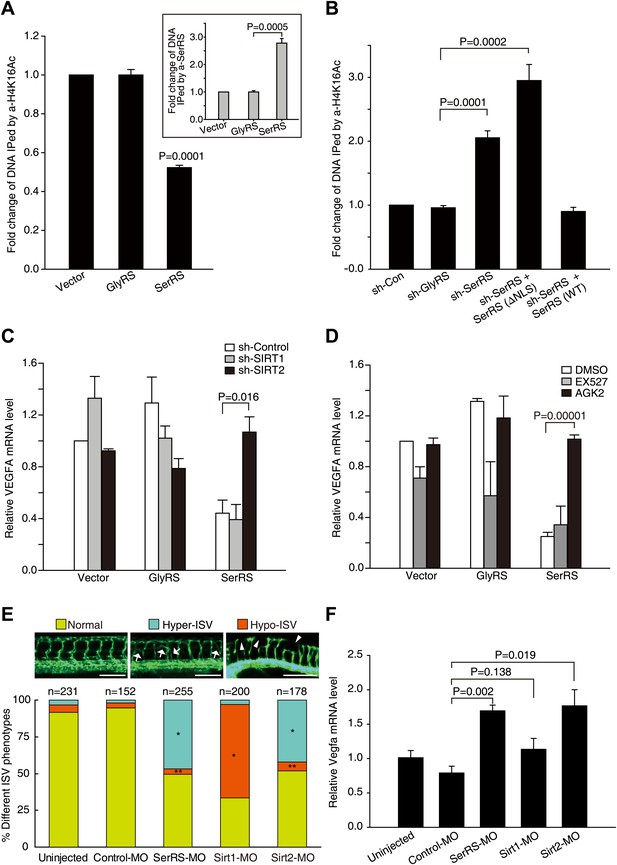
SerRS recruits SIRT2 to epigenetically silence VEGFA expression.
(A) ChIP assay to show that overexpression of SerRS reduces histone H4 acetylation level on the VEGFA promoter. HEK 293 cells were transfected with plasmids expressing SerRS, GlyRS or empty vector. The cell lysates were subjected to local ChIP analysis using anti-H4K16Ac (acetylated H4 at K16), anti-H4 (total), or anti-SerRS antibodies and a primer set targeting the VEGFA promoter. The amounts of DNA immunoprecipitated by anti-H4K16Ac antibody were normalized to those by anti-H4 antibody prior to fold change calculation. Inset: the normalized amounts of DNA immunoprecipitated by anti-SerRS. All data were shown as means ± SEM (n = 3). (B) ChIP assay to show that knock down of SerRS expression or exclusion of SerRS from the nucleus increases histone H4 acetylation level on the VEGFA promoter. HEK 293 cells were transfected with plasmids expressing the indicated molecules and subjected to local ChIP analysis as described above. As a control, GlyRS expression was knocked down but had no effect on H4 acetylation. (C) Effect of SIRT2 expression on the transcriptional repressor activity of SerRS as measured by VEGFA expression. HEK 293 cells were co-transfected with plasmids expressing shRNAs targeting SIRT1, SIRT2 or control shRNA and plasmids expressing SerRS, GlyRS or empty vector for 36 hr. The VEGFA expression levels were determined by using real-time RT-qPCR and are shown as means ± SEM (n = 3). (D) Effect of SIRT2-specific inhibitor on the transcriptional repressor activity of SerRS as measured by VEGFA expression. HEK 293 cells were transfected with plasmids expressing SerRS, GlyRS or empty vector. SIRT2-specific inhibitor AGK2 (10 µM, final concentration) or SIRT1-specific inhibitor EX-527 (1 µm, final concentration) or solvent alone (DMSO) was added to the cell culture media 2 hr post-transfection. VEGFA expression levels were measured 24 hr post-transfection by using real-time RT-qPCR and are shown as means ± SEM (n = 3). (E) Functional correlation between SerRS and SIRT2 in zebrafish. The percentage of Tg(Fli1a:EGFP) zebrafish embryos showing different ISV phenotypes at 3 days post fertilization after the injection of morpholinos targeting SerRS (SerRS-MO), Sirt1 (Sirt1-MO), Sirt2 (Sirt2-MO), or a control morpholino (Control-MO) are illustrated. Scale bars represent 0.25 mm. *p<0.0001 vs Control-MO, **p>0.1 vs Control-MO. (F) The effects of knocking down SerRS, Sirt2, or Sirt1 in zebrafish on Vegfa expression were examined by real-time RT-qPCR at 1 day post fertilization after injection of morpholinos as indicated. Data are shown as means ± SEM (n = 10–15).
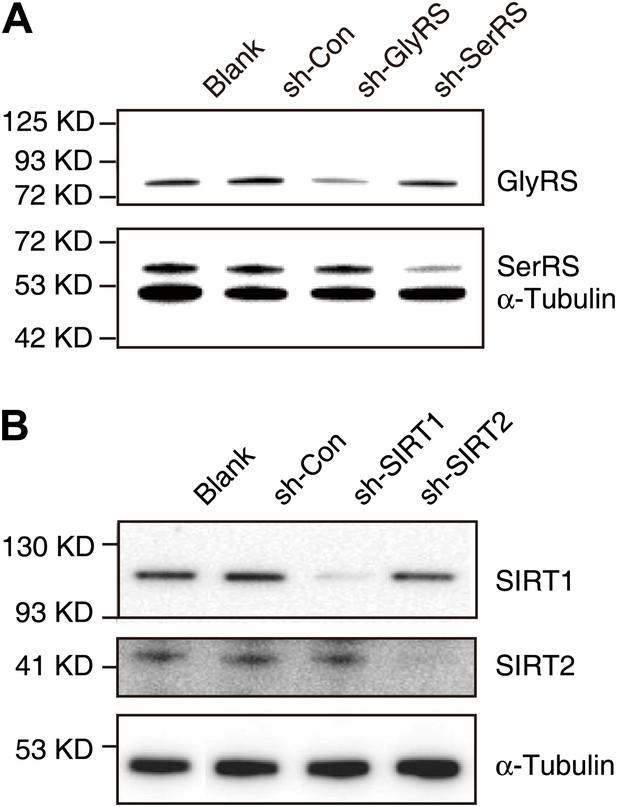
Knock-down efficiencies of shRNAs targeting GlyRS, SerRS, SIRT1, and SIRT2.
(A) Western blot analysis to confirm the efficiency of shRNAs in knocking down endogenous GlyRS (sh-GlyRS) or SerRS (sh-SerRS) expression in HEK 293 cells. (B) Western blot analysis to confirm the efficiency of shRNAs in knocking down endogenous SIRT1 (sh-SIRT1) or SIRT2 (sh-SIRT2) expression in HEK 293 cells.
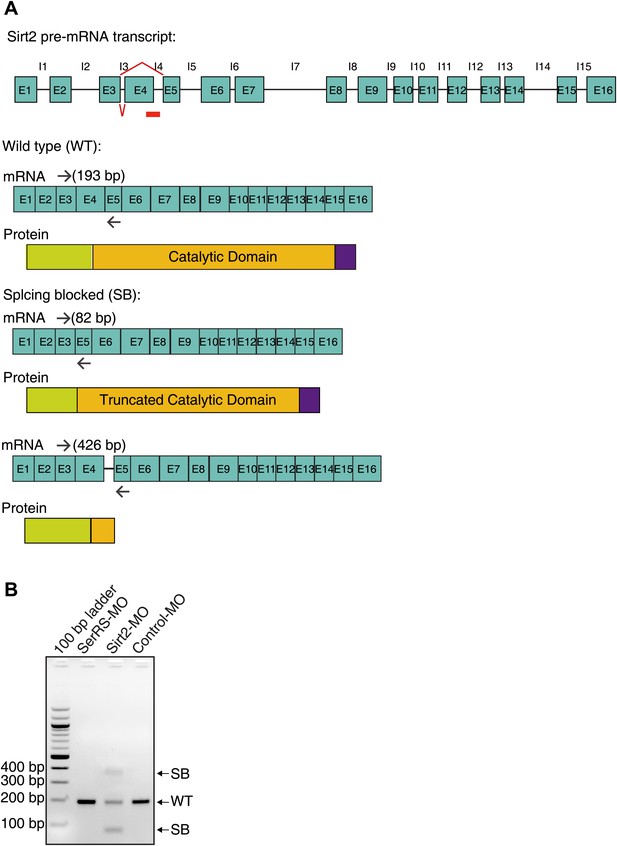
Design and efficiency of the antisense morpholino against Sirt2.
(A) The design of Sirt2-MO and the expected mRNA/protein products. The Sirt2-MO targets the exon4-intron4 splicing site of Sirt2 precursor mRNA to potentially generate two alternatively spliced mRNA products (SB) that would be translated into non-functional proteins. (B) RT-PCR analysis confirming the dramatic decrease of WT Sirt2 mature mRNA and the expected alternative splicing variants (SB) as the result of injecting Sirt2-MO.
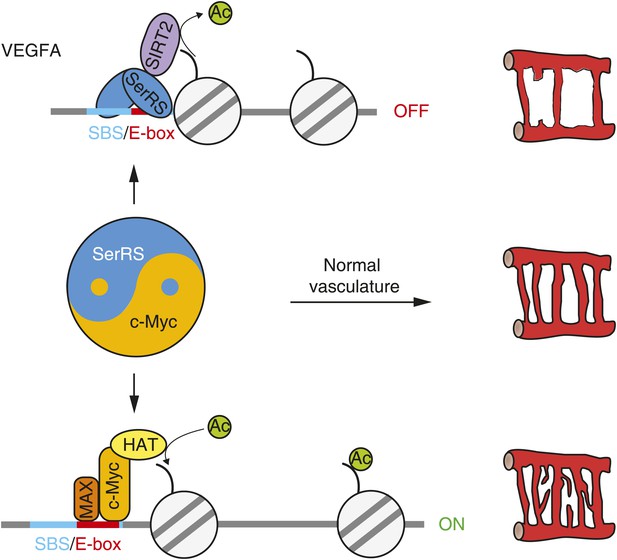
The ‘Yin-Yang’ relationship of SerRS and c-Myc in vascular development.
Nuclear SerRS binds to the VEGFA promoter at the identified SerRS binding site (SBS) and recruits the SIRT2 histone deacetylase to condense the local chromatin to shut down VEGFA expression. These tandem actions of SerRS symmetrically offset the VEGFA-promoting actions of c-Myc to maintain a delicate balance for the development of a functional vasculature.





