Histone supply regulates S phase timing and cell cycle progression
Figures
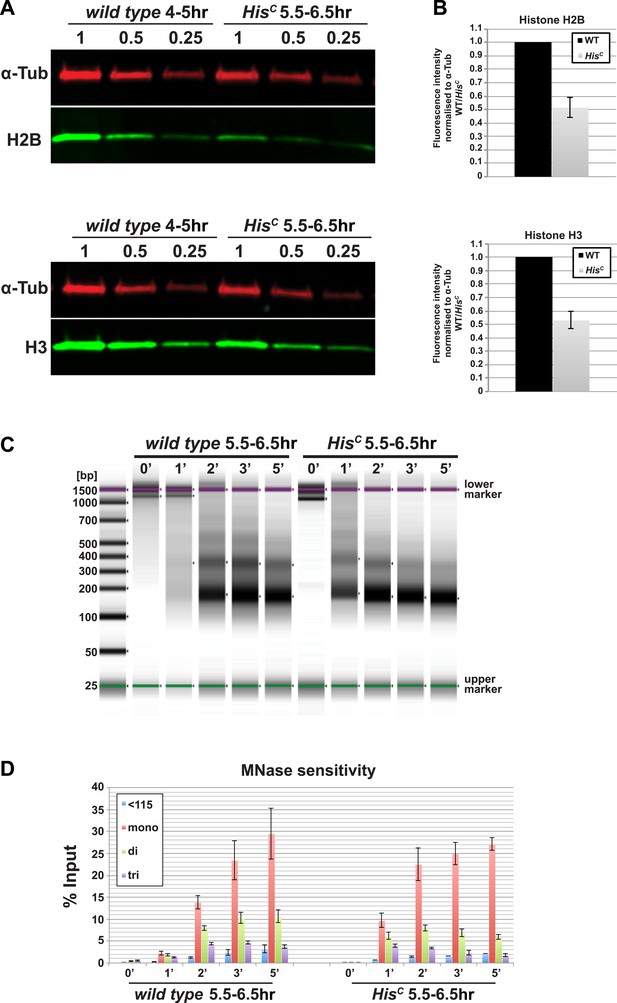
Nucleosome density is affected in HisC mutant cells.
(A) Fluorescent Western blot analysis for histone H2B and H3 (green) using wild type embryos at 4–5 hr and sorted HisC mutant embryos at 5.5–6.5 hr AEL, respectively. α-Tubulin (α-Tub; red) was used as loading control. A dilution series of each extract was loaded (1, 0.5, 0.25). (B) Quantification of Western blots as shown in (A). Fluorescence measurements of the histone signal was normalised to the α-Tubulin signal and the wild type (WT)/HisC ratio is shown. The histone protein content is ∼twofold reduced in HisC mutant embryos as compared to wild type. Mean values from three independent experiments are shown. Error bars indicate standard error. (C) Gel electrophoresis of time-course (0′–5′) microccocal nuclease (MNase) digestions using sorted HisC mutant and wild type sibling embryos at 5.5–6.5 hr after egg laying, respectively. (D) Quantification of MNase digestion experiments as shown in (C). HisC mutant chromatin is digested more rapidly into mononucleosomal DNA than control chromatin. Mean values from three independent experiments are shown. Error bars indicate standard error.
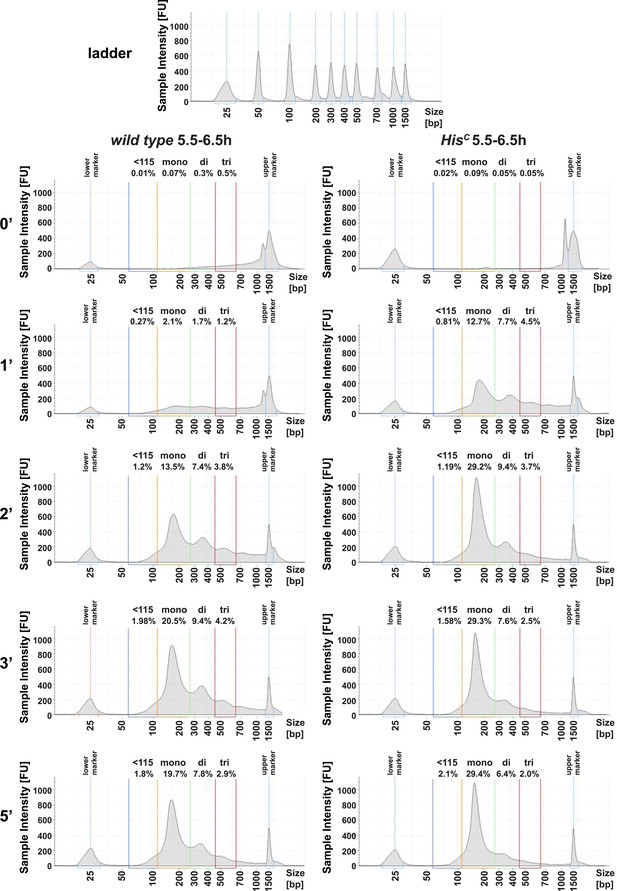
HisC mutant chromatin shows increased MNase sensitivity.
Quantification of the time-course microccocal nuclease (MNase) digestion experiment shown in Figure 1C using the Tapestation analysis software (Agilent Technologies). Shown are the individual electropherograms for each digestion (0′, 1′, 2′, 3′, 5′), which display the size and concentration of DNA. Regions for quantification were defined as follows: 60–115 bp (<115 bp), 115–270 bp (mononucleosomes), 270–455 bp (dinucleosomes), 455–650 bp (trinucleosomes). Numbers above regions indicate the concentration as % of input.
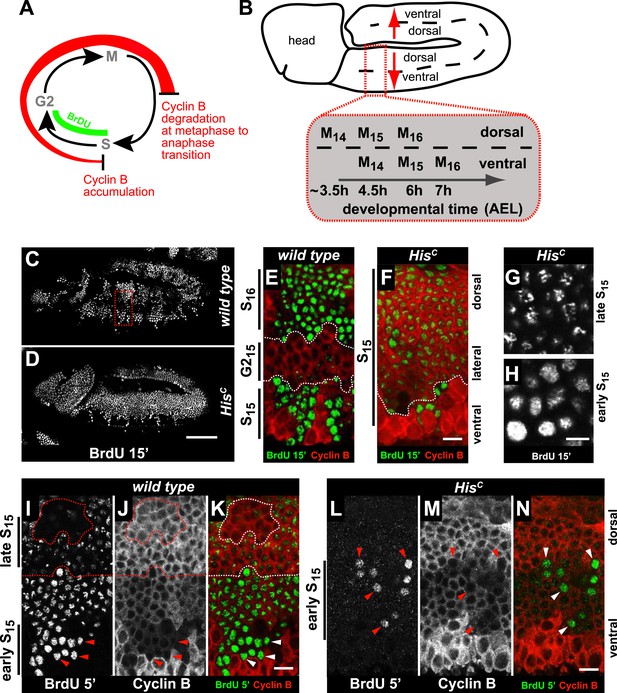
HisC mutant cells extend S phase and slow down DNA synthesis.
(A and B) Schematic models of Cyclin B accumulation and BrdU incorporation during the embryonic cell cycle (A) and its spatial control (B). AEL: after egg laying. (C–N) BrdU pulse labelling for 15 (C–H) or 5 min (I–N) and staining with antibodies against BrdU (green in merge) and Cyclin B (red in merge). (C and D) BrdU was detected in the epidermis of HisC mutant embryos indicating DNA replication but the pattern of replicating cells was distinct from wild type. (E and F) Magnifications of an epidermal region as shown for the wild type embryo (boxed area in C). (E) BrdU labelled wild type cells of the ventral epidermis in S15 (below dashed lines) and of the dorsal epidermis in S16 (above dashed lines). Cells in G215 were BrdU negative with high levels of Cyclin B and located in the lateral epidermis (between dashed lines). (F) In HisC mutant embryos, lateral and dorsal cells re-accumulated Cyclin B and were labelled for BrdU (above dashed line), indicating that mutant cells still replicated DNA during S15. (G and H) Punctate pattern of BrdU incorporation in cells that progressed into late S15 in the dorsal epidermis (G) and uniform incorporation pattern in cells of the ventral epidermis (H). (I–K) In wild type embryos replicating cells in early (ventral, below dashed line) and late S15 (dorsal, above dashed line) were detected after a 5 min BrdU labelling pulse. Some patches of dorsal cells completed S15 and did not incorporate BrdU (encircled). (L–N) In HisC mutant embryos BrdU was detected after a 5 min BrdU labelling pulse only in ventral cells that were in early S phase as indicated by low levels of Cyclin B (arrowheads). Dorsal up (E–N), scale bars: 100 µm (C and D), 10 µm (E–N).
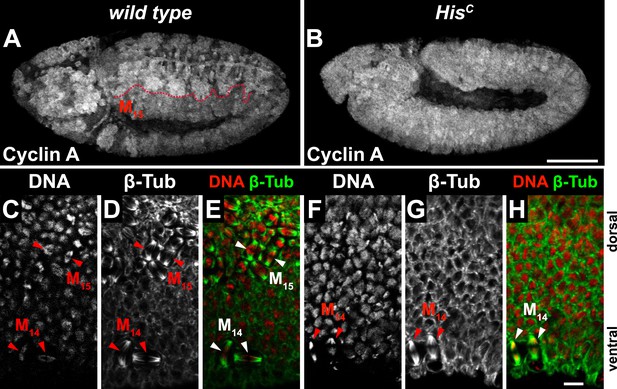
The cell cycle arrest of HisC mutant cells is before mitosis.
(A–H) Immunofluorescent staining with antibodies against Cyclin A (A and B) and β-tubulin (C–H). (A) Wild type embryos degraded Cyclin A in the dorsal epidermis during M15 (below dashed line). (B) HisC mutant embryos failed to degrade Cyclin A. (C–H) Magnifications of epidermal cells from embryos. (C–E) Wild type embryos show mitotic spindles in ventral epidermal cells at M14 and dorsal epidermal cells in M15 as visualised by β-tubulin staining (arrowheads, green in merge). (F–H) HisC mutant embryos show only mitotic spindles in the ventral epidermis during M14 (arrowheads) but not in the dorsal epidermis. Dorsal up in (C–H), scale bars: 100 µm (A and B), 10 µm (C–H).
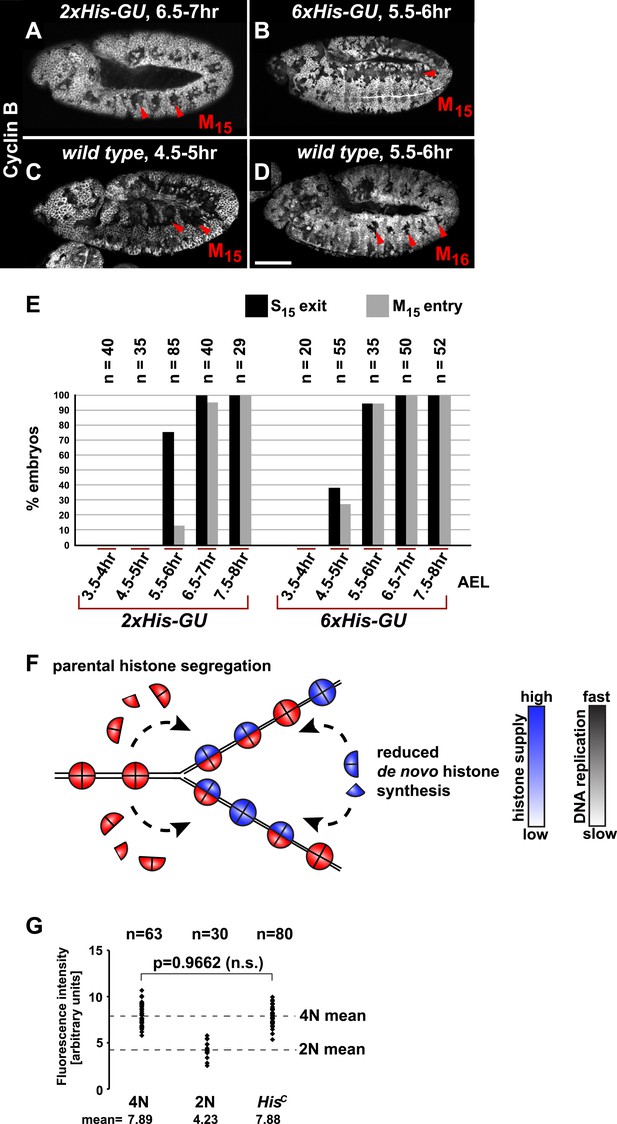
Histone availability determines the rate of S phase progression but is not required for completion of DNA replication.
(A–D) Immunofluorescent staining with antibodies against Cyclin B. (A) Cyclin B degradation at 6.5–7 hr AEL around the tracheal pits in 2xHis-GU embryos during M15 (arrowheads) shows that these embryos are lagging more than one cell cycle behind as compared to wild type (see also Figure 3—figure supplement 1). (B) Cyclin B degradation in dorsal cells of 6xHis-GU embryos at 5.5–6 hr AEL during M15 (arrowheads), showing that 6xHis-GU reduced the delay in cell cycle progression compared to 2xHis-GUs (see also Figure 3—figure supplement 2). (C) Cyclin B degradation in dorsal cells of wild type embryos at 4.5–5 hr AEL during M15 (arrowheads). (D) Cyclin B degradation at 5.5–6 hr AEL around the tracheal pits in wild type embryos during M16 (arrowheads). (E) BrdU pulse labelling for 15 min of 2xHis-GU and 6xHis-GU embryos at indicated developmental stages and staining with antibodies against BrdU and Cyclin B. Shown is the quantification of embryos that completed S15 (‘S15 exit’, based on lack of BrdU labelling) and progressed into M15 (‘M15 entry’, based on Cyclin B degradation), showing the interdependence of the number of histone genes and cell cycle progression. n: number of embryos. (F) Schematic model showing nucleosome assembly at the replication fork and its dependence on histone supply. (G) DNA quantification of single nuclei stained with DAPI. Wild type nuclei in G215 and early S16 defined 4N and 2N DNA content, respectively. HisC mutant nuclei show mean intensity value of 4N nuclei, suggesting that mutant cells completed genome duplication. n: number of nuclei, p: probability from Student's t test, n.s.: not significant. Scale bars: 100 µm (A–D).
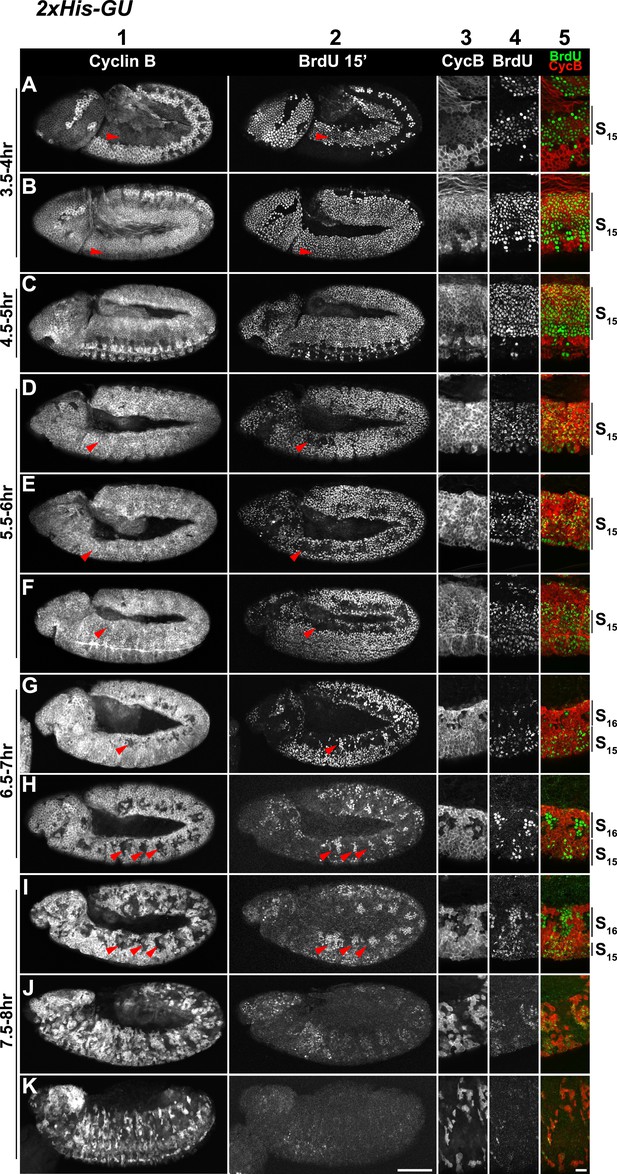
Cell cycle progression of HisC mutant embryos with two His-GUs (2xHis-GUs).
Shown are representative time-matched (as indicated on the left) embryos that were incubated with BrdU for 15′ immediately before fixation and staining with antibodies against Cyclin B and BrdU, respectively. BrdU incorporation visualises DNA replication and Cyclin B degradation is a marker for mitosis (Figure 1A). Columns 1–2 show the Cyclin B and BrdU staining in whole mount embryos. Columns 3–5 are magnifications of an epidermal region with single channel detections of Cyclin B (CycB), BrdU and the merge of both channels (BrdU in green, Cyclin B in red). Magnifications are oriented with dorsal side up. We previously described experiments showing the stereotyped cell cycle progression in wild type embryos (Günesdogan et al., 2010). AEL: after egg laying, scale bars: 100 µm in columns 1–2, 10 µm in columns 3–5. 3.5–4 hr AEL: (A) dorsal epidermal cells degraded Cyclin B, completed M14 and entered S15 as shown by the incorporation of BrdU (arrowheads). (B) Ventral cells underwent M14 and entered S15 (arrowheads). Accordingly, all cells re-accumulated Cyclin B and incorporated BrdU. 4.5–5 hr AEL: (C) all epidermal cells undergo S15 and thus show Cyclin B staining and incorporation of BrdU. At this stage, dorsal epidermal cells of wild type embryos already entered M15/S16 (Figure 2C). 5.5–6 hr AEL: (D–F) dorsal epidermal cells stopped incorporating BrdU and completed S15 (arrowheads). 6.5–7 hr AEL: (G and H) dorsal epidermal cells degraded Cyclin B and entered M15 (arrowheads). Some cells already went through M15 and showed BrdU incorporation, indicating that they had entered S16 (arrowheads, G). The division pattern in wild type embryos is highly stereotyped and controlled by developmental genes. In wild type embryos, cells around the tracheal pits, which are structures in the posterior of each parasegment, enter M16 first (Figure 2D). In 2xHis-GU embryos, M15 took place at a developmental stage, when wild type embryos progressed through or completed M16 in the epidermis. Interestingly, the M15 division pattern in 2xHis-GU embryos (H) resembled the M16 division pattern in wild type embryos (Figure 2D), suggesting that the developmental program proceeds normally in 2xHis-GU embryos. 7.5–8 hr AEL: (I) after completion of M15, cells entered S16 and showed BrdU incorporation (arrowheads). (J and K) During and after germ band retraction, most epidermal cells did not incorporate BrdU, indicating that they aborted S16. Wild type embryos at this developmental stage stop to proliferate after M16 by entering a G1/0 phase (Knoblich et al., 1994). 2xHis-GU embryos apparently also enter a G1/0 phase, enforced by the ongoing developmental program.
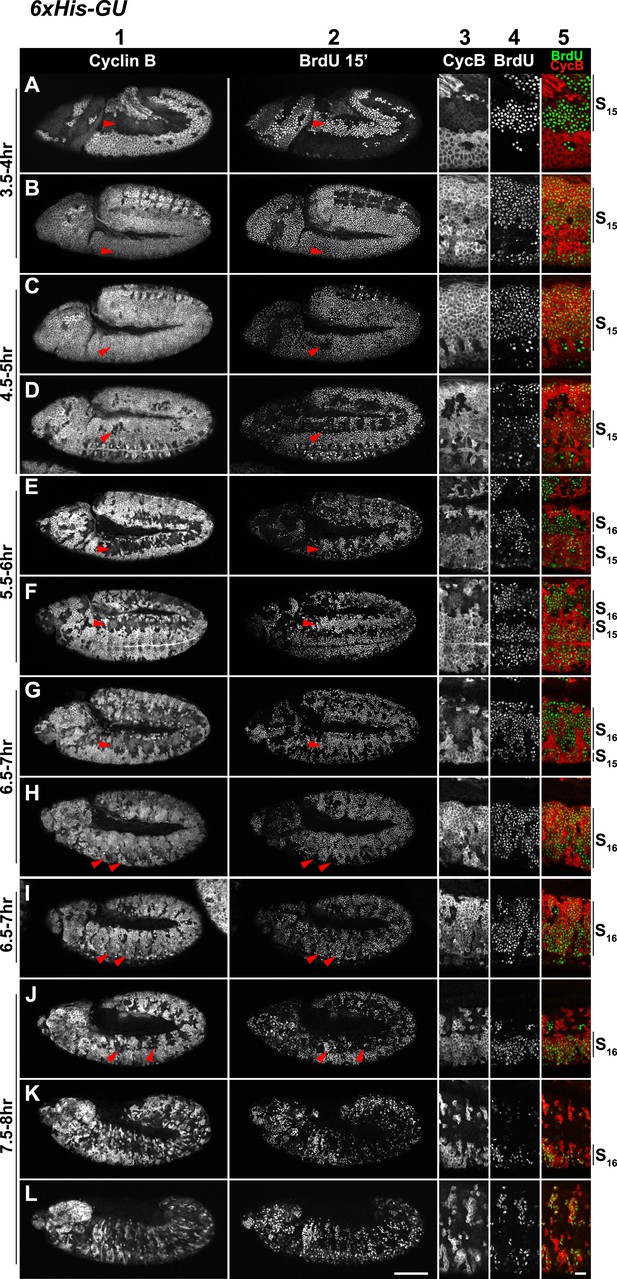
Cell cycle progression of HisC mutant embryos with six His-GUs (6xHis-GUs).
Shown are representative time-matched (as indicated on the left) embryos that were incubated with BrdU for 15′ immediately before fixation and staining with antibodies against Cyclin B and BrdU, respectively. BrdU incorporation visualises DNA replication and Cyclin B degradation is a marker for mitosis (Figure 1A). Columns 1–2 show the Cyclin B and BrdU staining pattern in whole mount embryos. Columns 3–5 are magnifications of an epidermal region with single channel detections of Cyclin B (CycB), BrdU and the merge of both channels (BrdU in green, Cyclin B in red). Magnifications are oriented with dorsal side up. We previously described experiments showing the stereotyped cell cycle progression in wild type embryos (Günesdogan et al., 2010). AEL: after egg laying, scale bars: 100 µm in columns 1–2, 10 µm in columns 3–5. 3.5–4 hr AEL: (A) dorsal epidermal cells degraded Cyclin B, completed M14, and entered S15 as shown by the incorporation of BrdU (arrowheads). (B) Ventral cells underwent M14 and entered S15 (arrowheads). Accordingly, all cells re-accumulated Cyclin B and incorporated BrdU. 4.5–5 hr AEL: (C and D) dorsal epidermal cells stopped incorporating BrdU and completed S15 (arrowheads, C). A few of these cells degraded Cyclin B and entered M15 (arrowheads, D). Thus, the delay during DNA replication in 6xHis-GU embryos is reduced as compared to 2xHis-GU embryos, which completed S15 at 5.5–6 hr AEL (Figure 2—figure supplement 1D–F). 5.5–6 hr AEL: (E and F) dorsal epidermal cells degraded Cyclin B, completed M15 and some cells already entered S16 as shown by BrdU incorporation (arrowheads). 6.5–7 hr AEL: (G–I) dorsal epidermal cells incorporated BrdU in S16 (arrowheads, G). Ventral epidermal cells degraded Cyclin B and entered M15 (arrowheads, H), followed by incorporation of BrdU in S16 (arrowheads, I). 7.5–8 hr AEL: (J–L) some dorsal epidermal cells degraded Cyclin B and went through M16 (arrowheads, J). During and after germ band retraction (K and L), most cells did not reaccumulate Cyclin B or incorporated BrdU, suggesting that they stopped proliferating, which is similar to wild type at this developmental stage (Günesdogan et al., 2010).
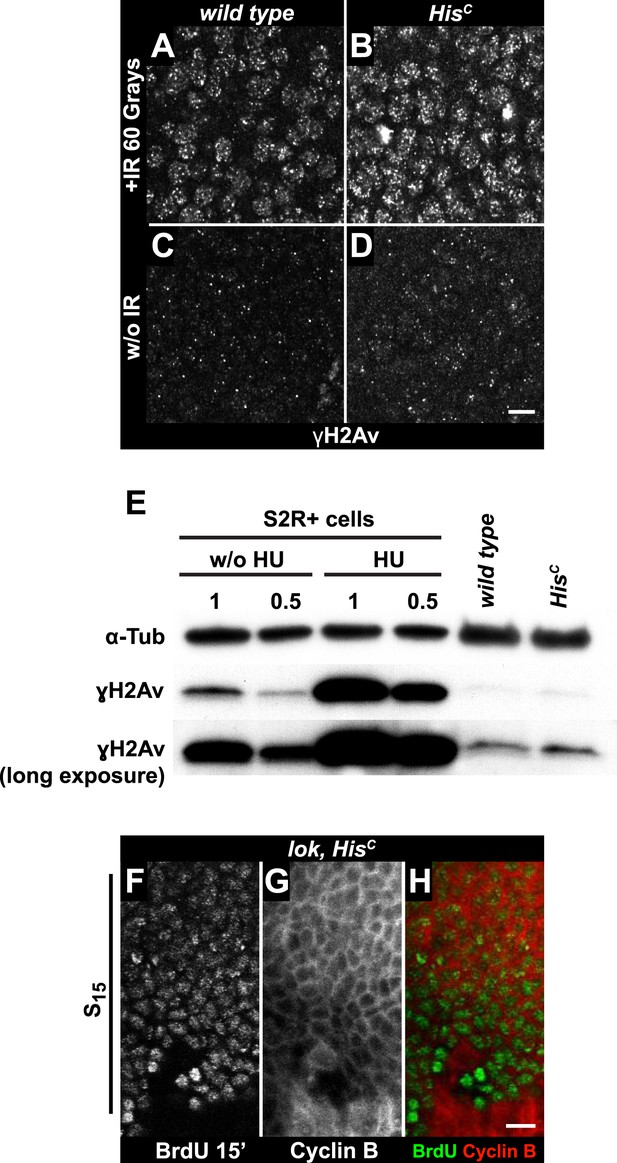
The cell cycle arrest of HisC mutant cells does not depend on the ATM/Chk2 DNA damage checkpoint.
(A and B) Wild type and HisC mutant cells responded to ionizing irradiation (IR) by phosphorylation of the variant histone H2Av, detected by a phosphospecific antibody against H2Av (γH2Av). (C and D) Without irradiation, HisC mutant cells did not show elevated γH2Av staining compared to wild type, indicating that mutant cells did not accumulate DNA damage. (E) Western blot for γH2Av using untreated (w/o HU) and HU-treated (HU) S2R+ cells as controls (two dilutions, 1 and 0.5) as well as HisC mutant embryos and wild type embryos at 5.5–6.5 hr AEL. α-Tubulin (α-Tub) was used as a loading control. HU treatment results in a significant increase of γH2Av, which was not observed in HisC mutant embryos. (F–H) BrdU pulse labelling for 15 min and staining with antibodies against BrdU and Cyclin B. lok, HisC double mutant embryos showed a similar phenotype as HisC mutant embryos (see Figure 1F). Dorsal up in (F–H), scale bars: 10 µm.
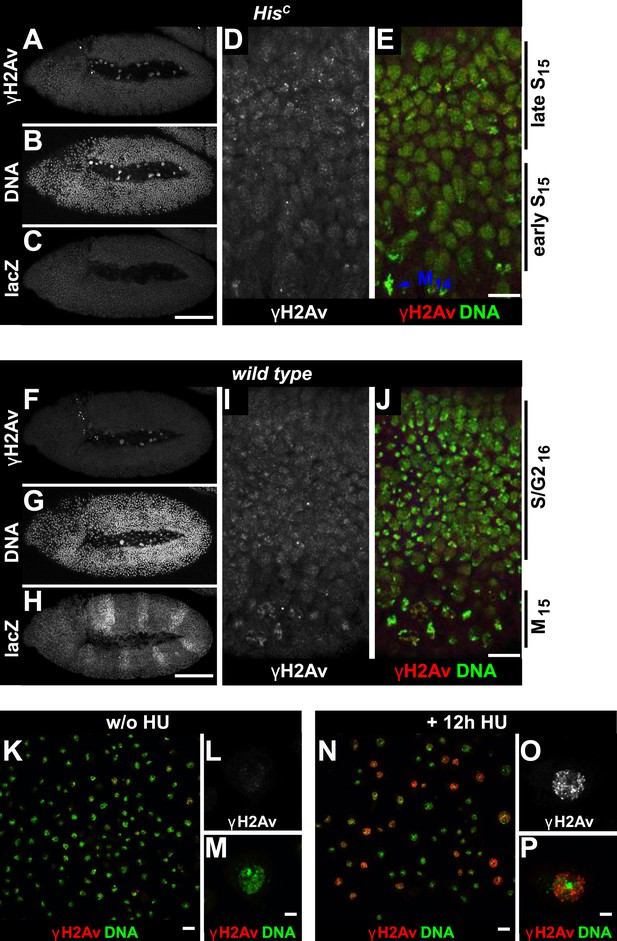
HisC mutant embryos show a moderate increase of γH2Av during S15 progression.
HisC mutant (A–E) and wild type sibling (F–J) embryos stained with antibodies against phosphorylated H2Av (γH2Av), against β-Galactosidase (lacZ) and with DAPI to detect DNA. (A–C) HisC mutant embryos were identified by the lack of lacZ staining. (D and E) Late S15 cells in the dorsal epidermis of HisC mutant embryos show a moderate increase of γH2Av as compared to ventral cells that are in early S15 or still in M14 (blue arrowhead). (F–H) Wild type sibling embryos of comparable age were identified based on lacZ staining. (I and J) In these embryos dorsal epidermal cells were already in S/G216. The increase in γH2Av staining was less pronounced than in HisC mutant embryos. (K–P) Untreated (w/o HU; K–M) and HU-treated (N–P) S2R+ cells were stained for γH2Av and DNA showing an increase of γH2Av signal in HU-treated cells. Dorsal up in (A–J), anterior to the left in (A–C, F–H), scale bars: 100 µm (A–C, F–H), 10 µm (D, E, I, J), 20 µm (K and N) and 5 µm (L, M, O, P).
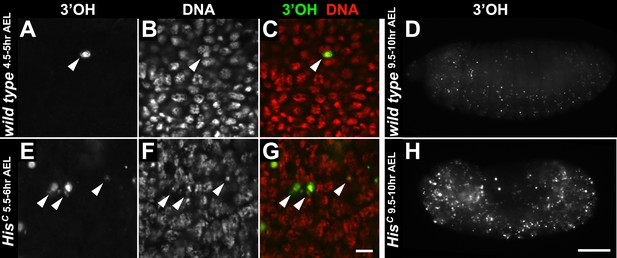
HisC mutant embryos do not show accumulation of DNA damage during early embryogenesis.
(A–H) TUNEL assays with whole mount embryos to detect free 3′OH groups of DNA, which indicate DNA damage. (A–C) Wild type embryos at 4.5–5 hr AEL contained a few apoptotic cells (arrowheads). In most nuclei, free 3′OH groups were not detected. (D) Whole mount wild type embryos at 9.5–10 hr AEL showed a few apoptotic cells. (E–G) HisC mutant embryos at 6.5–7 hr AEL also contained a few apoptotic cells (arrowheads). However, the level of TUNEL staining was not uniformly elevated in the mutant cells, indicating that there is no increase of DNA damage compared to wild type. (H) HisC mutant embryos die around 9.5–10 hr AEL and show an increased number of apoptotic cells. Scale bars: 10 µm (A–C), (E–G), 100 µm (D and H).
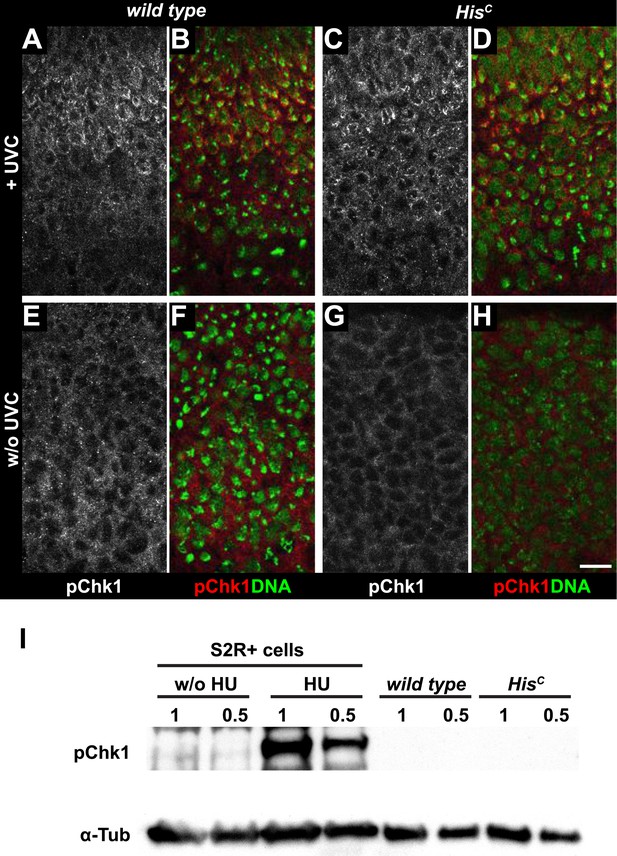
HisC mutant cells do not activate the ATR/Chk1 DNA damage checkpoint.
(A–D) Wild type and HisC mutant cells responded to UV irradiation (UVC) by phosphorylation of the Drosophila Chk1 ortholog GRP, detected by a phosphospecific antibody for Chk1 (pChk1). Embryos were counterstained for DNA to visualize nuclei (B and D). (E–H) Without irradiation, HisC mutant cells did not show elevated staining for pChk1 compared to wild type, indicating that mutant cells did not activate the ATR/Chk1 checkpoint. (I) Western blot detecting pGRP by an antibody to phosphorylated Chk1 (pChk1) and α−Tubulin (α-Tub). Extracts were prepared from SR2+ tissue culture cells that were either untreated (w/o HU) or treated with HU (HU). Embryos were either sorted wild type controls or HisC mutants. Two dilutions (1 and 0.5) of each sample were loaded. Dorsal up in (A–H), scale bars: 10 µm.
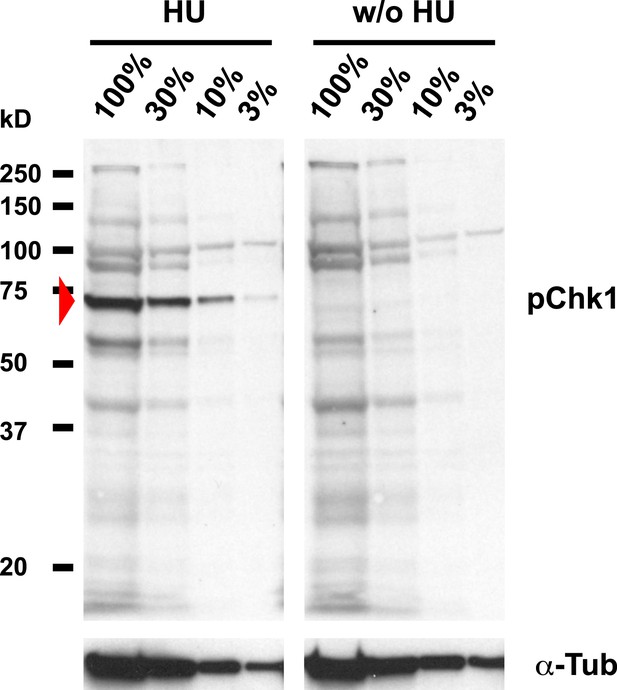
Detection of pGRP by a phosphospecific antibody for Chk1 by Western blotting.
Western blot detecting pGRP with an antibody against phosphorylated Chk1 (pChk1) and α−Tubulin (α-Tub). Extracts were prepared from SR2+ tissue culture cells that were either untreated (w/o HU) or treated with HU (HU). Serial dilutions of the respective extracts were loaded (100% to 3%). The apparent molecular weight of marker proteins is indicated to the left (kD). The anti-Chk1 phospho-S345 antibody detects background bands in extracts from HU treated and untreated S2R+ cells, but only one band is induced by HU (arrowhead). The molecular weight of the respective band is in the range of the expected molecular weight of GRP (58 kD).
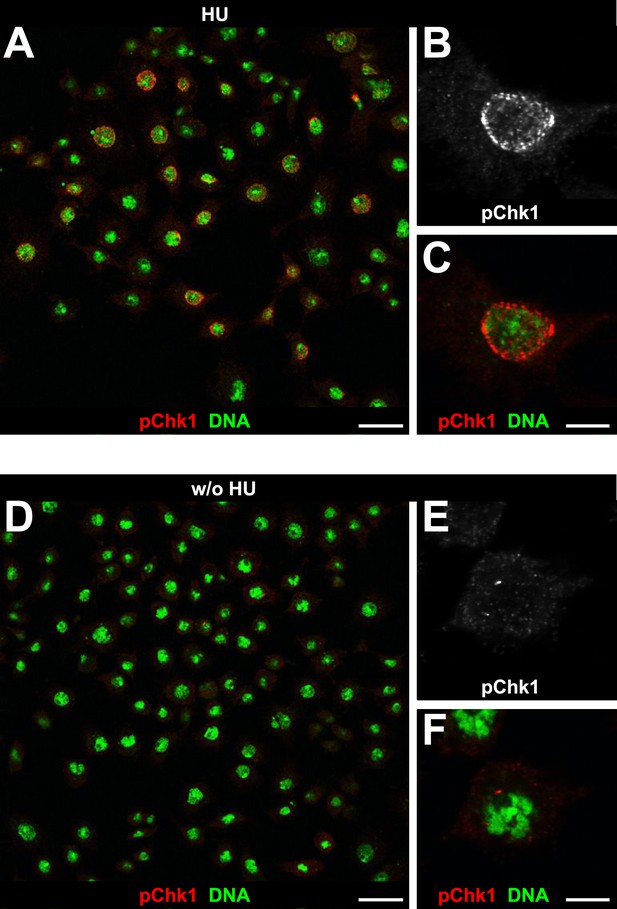
Detection of pGRP by a phosphospecific antibody for Chk1 by immunofluorescence.
S2R+ cells were either cultured in the presence of HU (HU) or without HU (w/o HU) and stained with an a phosphospecific antibody for Chk1 (pChk1) and with DAPI (DNA). (A) HU treatment induces a clear signal that is detected in S2R+ cells. HU induces replicative stress by depletion of intracellular dNTP pools. Therefore, only cells that were in S phase during the 12 hr treatment were susceptible to HU. (B and C) Single cell magnifications show that the signal detected by the anti-Chk1 phospho-S345 antibody localizes to the nucleus and resembles the focal staining pattern expected for phosphorylated and active GRP. (D) Untreated cells show only background signal for pChk1. (E and F) Single cell magnifications show that the residual background signal detected in untreated cells is mainly cytoplasmic. Scale bars: 20 µm (A and D), 5 µm (B, C, E, F).
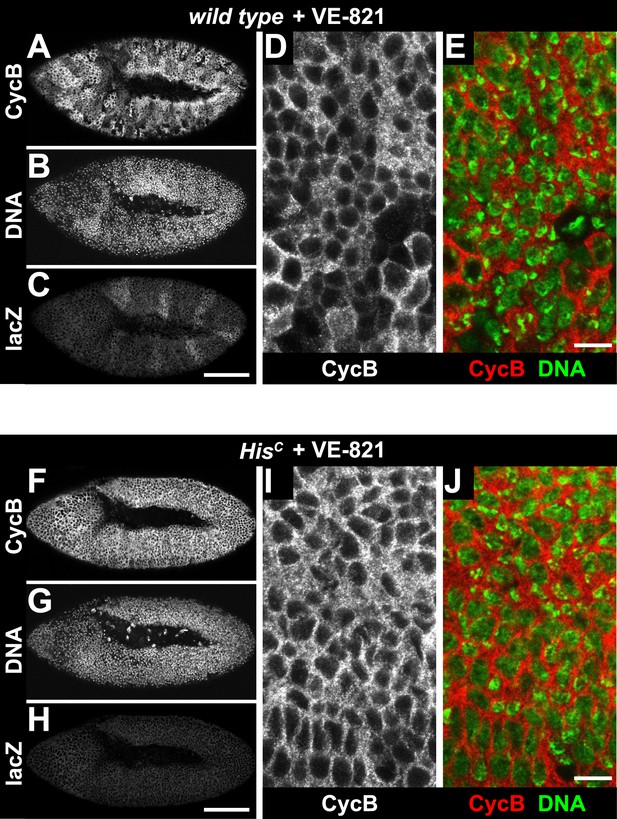
Treatment of HisC mutant embryos with the ATR inhibitor VE-821 does not release the cell cycle arrest.
Wild type sibling (A–E) and HisC mutant (F–J) embryos stained with DAPI (DNA) and antibodies against Cyclin B (CycB) and β-Galactosidase (lacZ). (A–C) Wild type sibling embryos were identified by lacZ staining. (D and E) Cyclin B degradation indicates that treatment with the VE-821 inhibitor did not block entry into mitosis. (F–H) HisC mutant embryos of comparable age were identified based on the lack of lacZ staining. (I and J) VE-821 treatment did not result in Cyclin B degradation in HisC mutant cells. Dorsal up in (A–J), anterior to the left in (A–C, F–H), scale bars: 100 µm (A–C, F–H) and 10 µm (D, E, I, J).
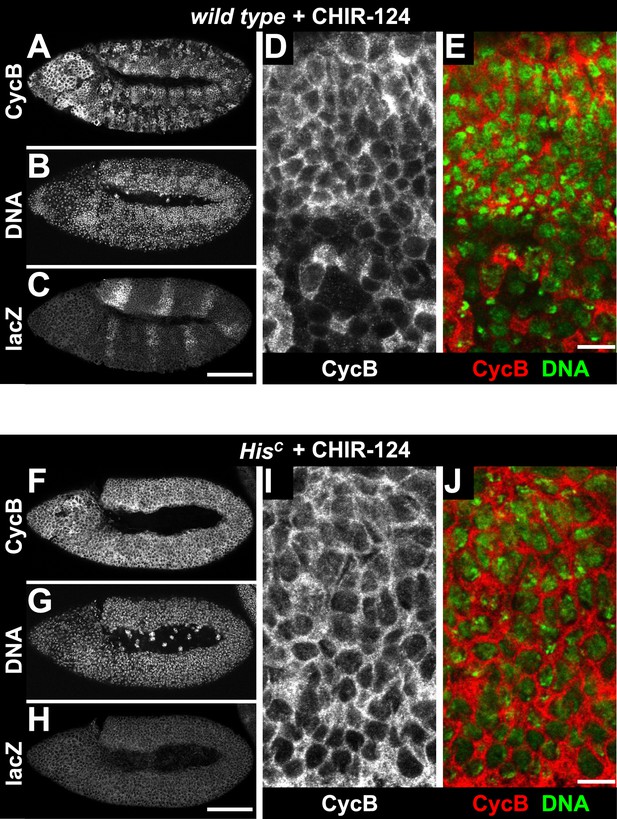
Treatment of HisC mutant embryos with the Chk1 inhibitor CHIR-124 does not release the cell cycle arrest.
Wild type sibling (A–E) and HisC mutant (F–J) embryos stained with DAPI (DNA) and antibodies against Cyclin B (CycB) and β-Galactosidase (lacZ). (A–C) Wild type sibling embryos were identified by lacZ staining. (D and E) Cyclin B degradation indicates that treatment with the CHIR-124 inhibitor did not block entry into mitosis. (F–H) HisC mutant embryos of comparable age were identified based on the lack of lacZ staining. (I and J) CHIR-124 treatment did not result in Cyclin B degradation in HisC mutant cells. Dorsal up in (A–J), anterior to the left in (A–C, F–H), scale bars: 100 µm (A–C, F–H) and 10 µm (D, E, I, J).
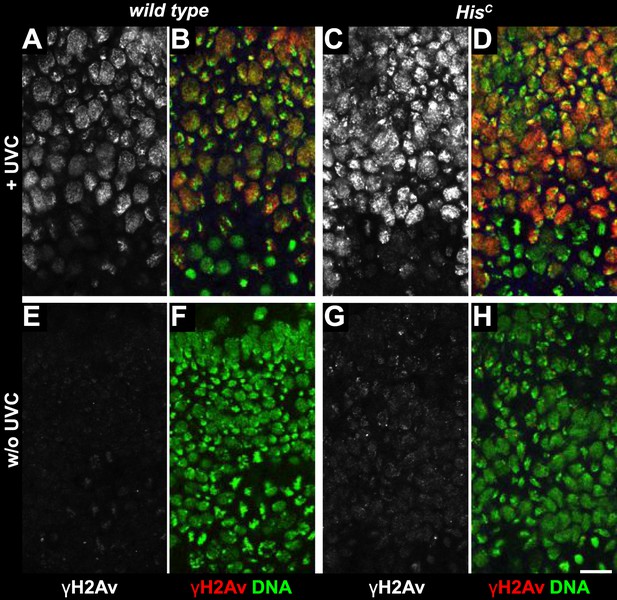
UV irradiation induces phosphorylation of H2Av.
Wild type sibling (A, B, E, F) and HisC mutant (C, D, G, H) embryos stained with DAPI (DNA) and antibodies against phosphorylated H2Av (γH2Av). (A–D) Wild type and HisC mutant cells responded to UV irradiation (UVC) by γH2Av. The lack of γH2Av in ventral cells that were still in G214 or in M14 indicates that the UVC treatment primarily affected cells that were in S15 at the time of irradiation or entered S15 during the recovery period after treatment (45 min). Notably, dorsal epidermal cells were either blocked or delayed in entry to M15 based on the absence of mitotic DNA structures. (E–H) Without irradiation, HisC mutant cells showed slightly elevated levels of γH2Av staining compared to cells from wild type siblings. Dorsal up in (A–H), scale bars: 10 µm.
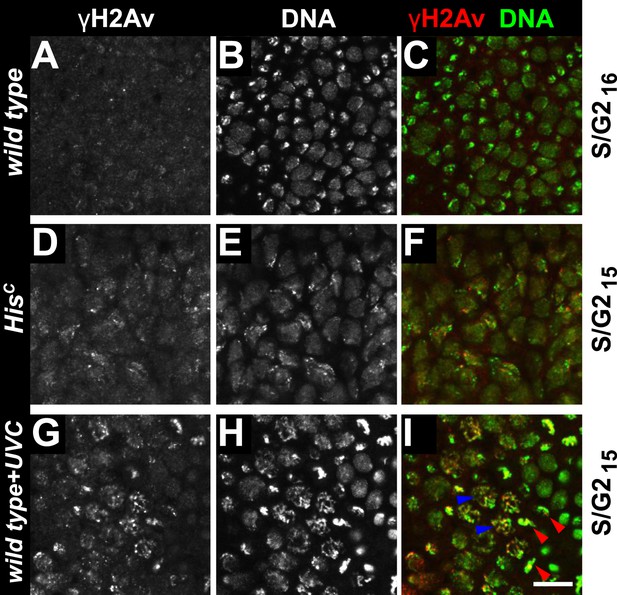
Mitotic entry with elevated γH2Av levels.
Wild type sibling (A–C, G–I) and HisC mutant (D–F) embryos stained with DAPI (DNA) and antibodies against phosphorylated H2Av (γH2Av). (A–C) Wild type cells that were in S16/G216 display background levels of γH2Av staining. (D–F) Compared to wild type cells, HisC mutant cells in late S15 show increased γH2Av staining. (G–I) UVC-irradiated wild type cells show increased γH2Av staining. These irradiated cells entered into M15 (blue arrowheads point at prophase structures, red arrowheads point at metaphase structures), indicating that they repaired DNA damage and were no longer arrested by the DNA damage checkpoints. The elevated γH2Av staining in HisC mutant cells compared to wild type cells is therefore unlikely to reflect significant DNA damage. Scale bar: 10 µm.
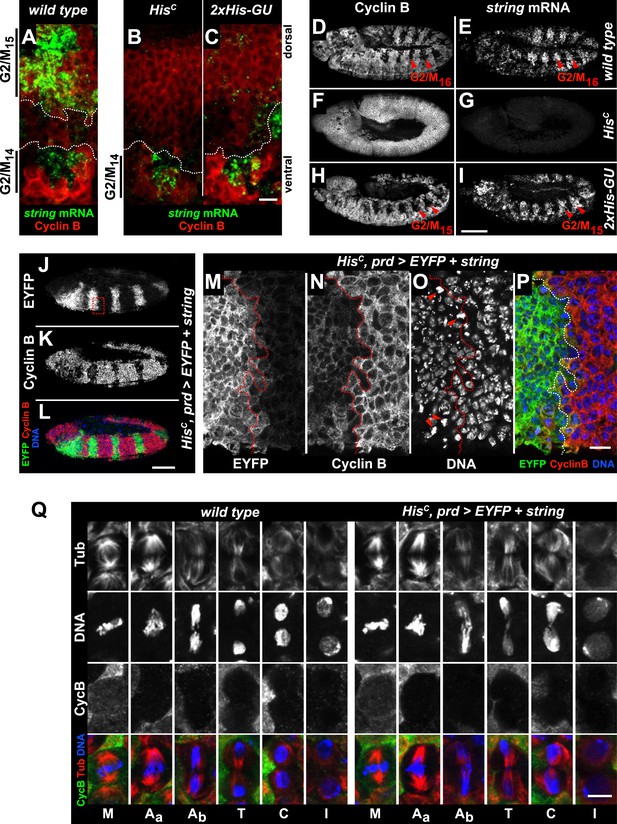
The cell cycle arrest of HisC mutant cells depends on string.
(A–I) string RNA in situ hybridisation (green in merge) and staining with an antibody against Cyclin B (red in merge). (A–C) Magnifications of epidermal cells from embryos at 4.5–5 hr AEL. (A) Wild type embryos upregulated string mRNA and accumulated Cyclin B in ventral epidermal cells during G214 (below dashed lines) and in dorsal epidermal cells during G215 (above dashed lines). In lateral cells with low levels of Cyclin B, string is not expressed during S15 (between dashed lines). (B) HisC mutant embryos failed to upregulate string in the dorsal epidermis (above dashed line). (C) 2xHis-GU embryos failed to upregulate string in the dorsal epidermis (above dashed line). (D–I) Whole mount embryos at 6.5–7 hr AEL. (D and E) Dorsal cells of wild type embryos progressed into G216, upregulated string mRNA and degraded Cyclin B during M16 (arrowheads). (F and G) HisC mutant embryos did not degrade Cyclin B and failed to accumulate string mRNA. (H and I) 2xHis-GU embryos accumulated string mRNA in G215 and degraded Cyclin B in M15 (arrowheads). (J–P) HisC mutant embryos expressing UAS-EYFP and UAS-string under the control of prd-GAL4 stained with antibodies against Cyclin B and EYFP. (M–P) Magnifications of epidermal cells from an embryo (boxed in J). (J–P) Epidermal cells within the EYFP, string expression domains degraded Cyclin B and entered M15/S16. Arrowheads in (O) show mitotic cells. (Q) Mitotic progression in wild type M15 and M15 of HisC mutant cells rescued by string expression visualized by staining for α-Tubulin (Tub), DNA, and Cyclin B (CycB). M: metaphase, Aa: anaphase-a, Ab: anaphase-b, T: telophase, C: cytokinesis, I: interphase. Dorsal up (A–C, M–P), scale bars 10 µm (A–C, M–P), 100 µm (D–L), 5 µm (Q).
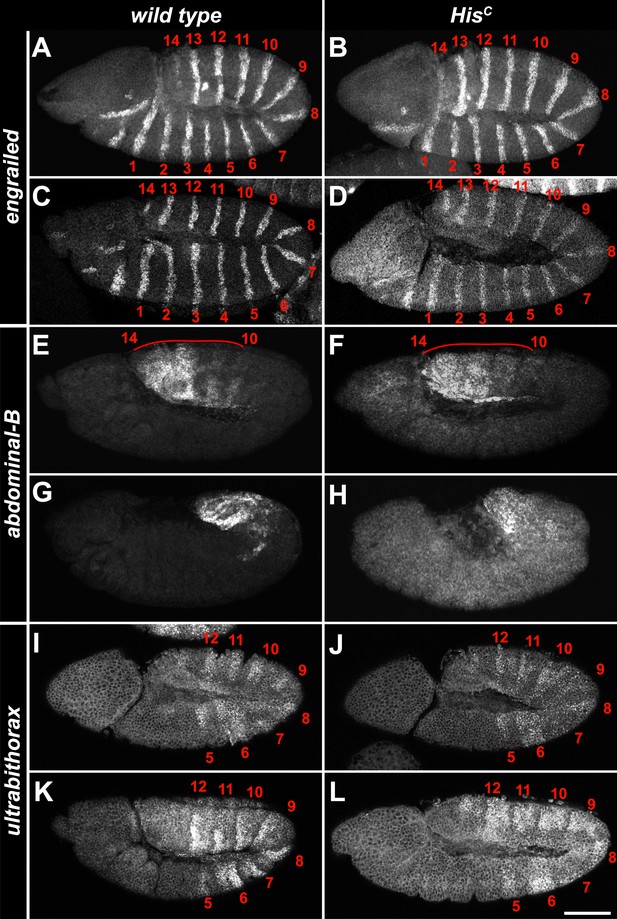
The expression pattern of developmental genes is normal in HisC mutant embryos.
(A–L) Fluorescent in situ hybridisations of whole mount embryos using probes for the segment polarity class gene engrailed (A–D), the homeotic genes Abdominal-B (E–H) and Ultrabithorax (I–L). (A–D) At embryonic stage 9, engrailed is expressed in 14 parasegmental stripes (1–14) in the trunk and in additional expression domains in the head (A). The same pattern was detected in HisC mutant embryos (B). In embryonic stage 10, engrailed is detectable in 14 parasegmental stripes in the trunk of wild type (C) and HisC mutant embryos (D). (E–H) Abdominal-B is expressed in the most posterior parasegments 10–14 at embryonic stage 10–11 (E). The same expression pattern was observed in HisC mutant embryos (F). During germ band retraction at stage 12 of embryogenesis, the expression patterns are still very similar between wild type (G) and mutant embryos (H). (I–L) Ultrabithorax is expressed in parasegments 5–12 at embryonic stage 9 (I). The same expression pattern was observed in HisC mutant embryos at stage 9 (J). Also, at stage 10–11 of embryogenesis, wild type (K) and mutant embryos (L) show a highly similar expression pattern. Scale bar: 100 µm.
Additional files
-
Source code 1
Source code of the ImageJ script used in this study to determine the volume and total fluorescence intensity of individual DAPI stained nuclei based on confocal z-stacks.
- https://doi.org/10.7554/eLife.02443.022





