The pseudo GTPase CENP-M drives human kinetochore assembly
Figures
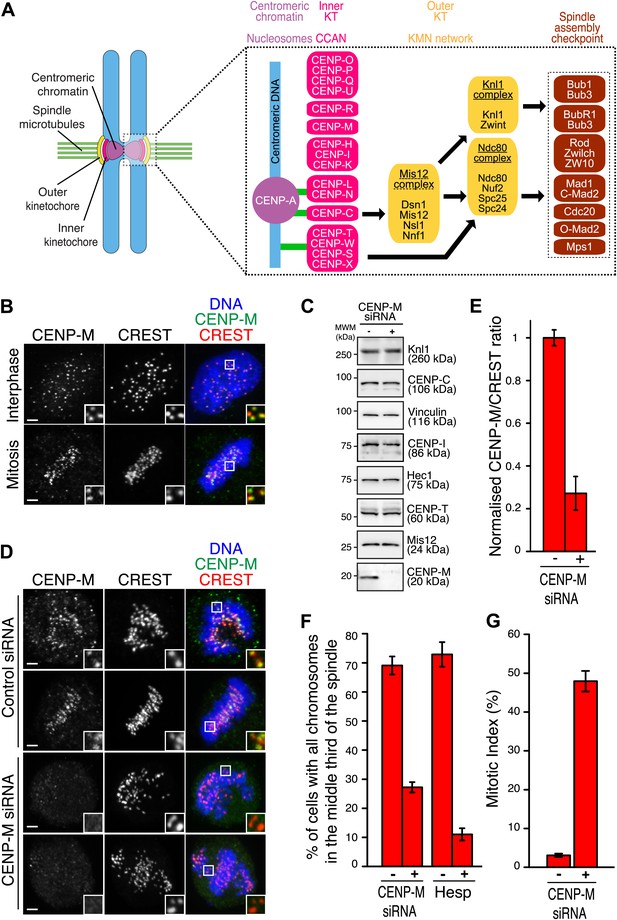
Ablation of CENP-M perturbs kinetochore function.
(A) Core kinetochore components. CENP-C and CENP-T/W may create independent connections between centromeres and outer kinetochores. Green lines indicate direct connections with centromeric DNA or chromatin. Black lines indicate recruitment dependencies. CENP-C binds directly to CENP-A and Mis12 complex (Przewloka et al., 2011; Screpanti et al., 2011; Kato et al., 2013). CENP-T, together with CENP-W, S and X, may form a nucleosome-like structure interacting directly with DNA and the Ndc80 complex (Hori et al., 2008a; Gascoigne et al., 2011; Schleiffer et al., 2012; Nishino et al., 2013, 2012). Sub-complexes of CCAN subunits were inferred from reconstitution or from similarity of depletion phenotypes (see main text). (B) Representative immunofluorescence (IF) images showing endogenous CENP-M localization to kinetochores of HeLa cells in both interphase and mitosis. Kinetochores were visualized with CREST sera and DNA stained with DAPI. Insets show a higher magnification of regions outlined by the white boxes. Scale bar = 2 µm. (C) Whole cell protein extracts from HeLa cells treated with specific siRNAs (showed in D) were run on SDS-PAGE and immunoblotted for the indicated kinetochore proteins. Vinculin was the loading control. MWM, molecular weight marker. (D) HeLa cells depleted for CENP-M display significant chromosome congression defects. Following fixation, cells treated with CENP-M siRNA were imaged for endogenous CENP-M, CREST and DNA (DAPI). Scale bars = 2 µm. (E) CENP-M kinetochore levels from the experiment in D. Quantifications are expressed as normalized CENP-M/CREST fluorescence intensity ratios. Graphs and bars indicate mean ± SEM. (F) Quantification of chromosome congression defects in D. As a positive control, cells treated with 500 nM Hesperadin were scored for alignment defects. (G) Quantification of the percentage of mitotic cells in the experiment in D.
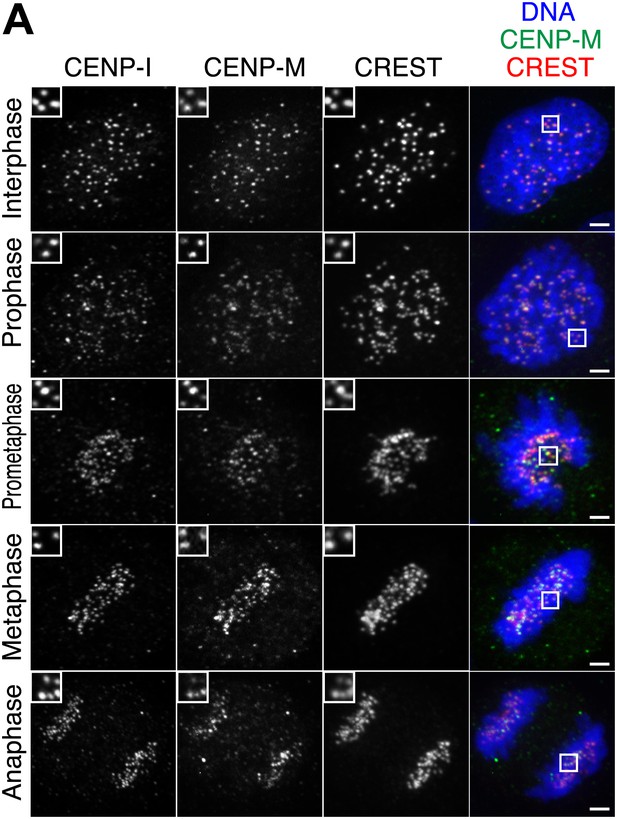
Additional localization data.
(A) Human CENP-M and CENP-I are constitutively associated with kinetochores during the cell cycle. Asynchronously growing HeLa cells were immunostained for endogenous CENP-M, endogenous CENP-I, and the centromeric marker CREST. DNA was stained with DAPI. The displayed interphase and metaphase cells are the same shown in Figure 1B but with the addition of the CENP-I staining (left panels). Insets show a higher magnification of the regions outlined by the white boxes. Scale bar = 2 µm.
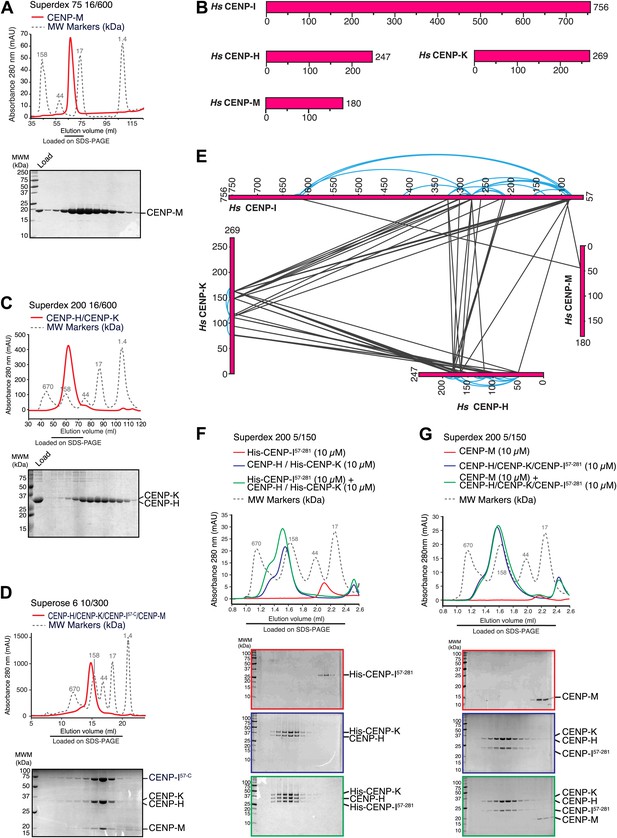
Characterization of the HIKM complex.
(A) SEC elution profile of CENP-M with associated SDS-PAGE separations of peak fractions indicated by the horizontal bar under the profile. CENP-M (∼20 kDa) elutes as expected for a monomeric species. (B) Schematic representation of the primary sequence of CENP-H, CENP-I, CENP-K, and CENP-M. (C) SEC elution profile and SDS-PAGE separation of the CENP-H/K complex. CENP-H/K forms a 1:1 dimer (∼61 kDa) (Figure 2—figure supplement 1, panel B) but elutes near the 158 kDa marker, indicative of an elongated complex. (D) SEC elution profile and SDS-PAGE separation of the CENP-HI57–CKM complex. CENP-HI57–CKM (∼159 kDa) elutes near the 158 kDa marker, suggesting that the complex contains a single copy of each subunit. (E) Summary of cross-links. Intra-molecular cross-links are shown in blue and outside the ideal perimeter designed by the four subunits of the complex. Inter-molecular cross-links are shown as black lines. (F) CENP-H/His-CENP-K complex and His-CENP-I57–281, both at 10 µM, form a stoichiometric complex as shown by co-elution from SEC runs and corresponding SDS-PAGE separations. (G) Lack of co-elution from SEC runs and SDS-PAGE analysis indicate that CENP-H/K/I57–281 complex and CENP-M do not bind.
-
Figure 2—source data 1
List of intra- and inter-molecular crosslinks of the CENP-HIKM complex.
- https://doi.org/10.7554/eLife.02978.006
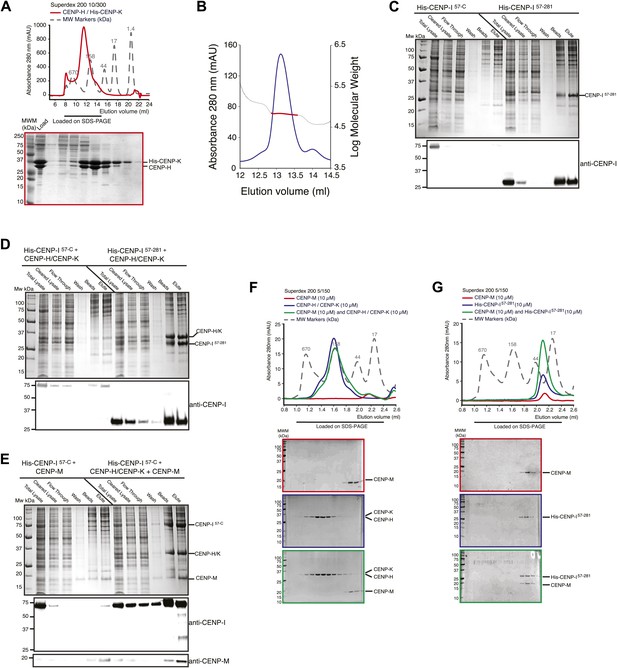
Additional biochemical characterization.
(A) SEC elution profile of the CENP-H/His-CENP-K complex. (B) Static light scattering analysis of the CENP-H/K complex indicates that the complex has a 1:1 stoichiometry and a molecular weight of ∼61 kDa. (C–E) Summary of expression or co-expression tests with the indicated CENP-I constructs. Only the co-expression of CENP-I57–C with both CENP-M and CENP-H-K (shown in panel E) was able to promote full solubilisation and stabilization of CENP-I57–C. Co-expression with only CENP-H/K or CENP-M was insufficient. (F) SEC analysis shows that CENP-M does not interact with CENP-H/K complex. (G) CENP-M does not interact with His-CENP-I57–281.
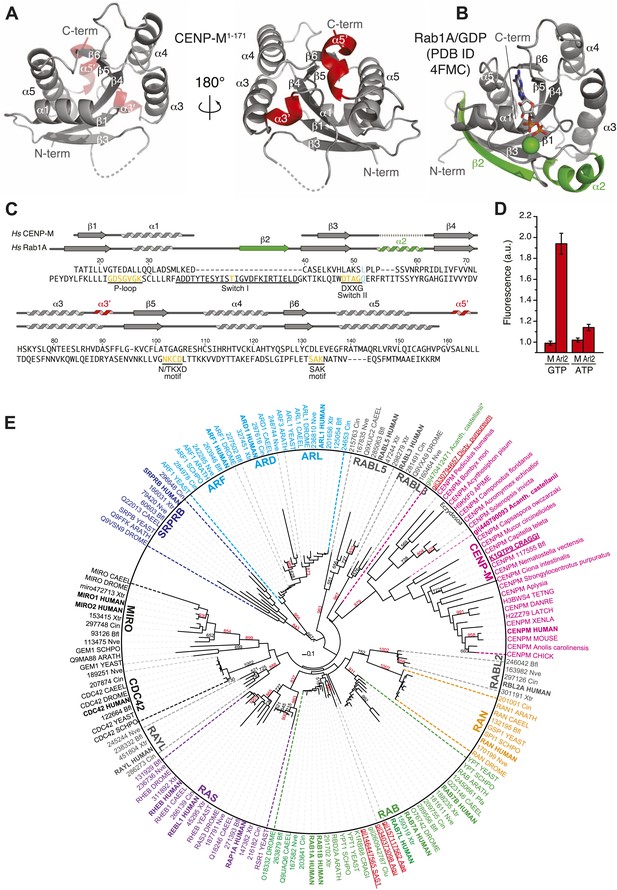
A small G-protein fold in CENP-M.
(A) Cartoon model of CENP-M1–171 in two orientations. (B) Cartoon model of Rab1A/GDP (PDB ID 4FMC). (C) Sequence alignment based on the structural superposition of CENP-M1–171 with Rab1A. Conserved elements of small G proteins are in yellow. A conserved residue involved in catalysis and targeted by activating mutations in Ras is in light blue. Secondary structure elements of CENP-M1–171 not present in Rab1A/GDP are in red, while those present in Rab1A/GDP but not in CENP-M1–171 are in green. (D) Experiments with N-methylanthraniloyl (MANT) derivatives of GTP and ATP. Binding of MANT-GTP or MANT-ATP to the small GTPase Arl2 (‘Arl2’) or CENP-M (‘M’) was monitored at an emission wavelength of 440 nm (See Figure 3—figure supplement 1, panel D). The histogram shows the time-averaged fluorescence value after addition of the indicated proteins to a solution of the indicated MANT nucleotides, normalized against the time-averaged value prior to protein addition. Only the addition of Arl2 to MANT-GTP (or MANT-GDP, see Figure 3—figure supplement 1, panel D) gave a clear increase in signal indicative of a physical interaction of the MANT nucleotide with Arl2. (E) Unrooted maximum likelihood tree of 157 sequences. Shown are members of classical small GTPase families in many species, covering a wide range of evolutionary points. Gray names are reclassified families (Rojas et al., 2012). Bold names are human sequences. CENP-M sequences are pink. Uppercase indicates Uniprot code for proteins. 3-code labels are: Nve (Nematostella vectensis), Bfl (Branchiostoma floridae), Xtr (Xenopus tropicalis), and Cin (Ciona intestinalis). Numbers on the left of 3-code labels are accession numbers corresponding to the DOE Joint Genome Institute (JGI) database. Red and underlined names are Genbank gi identifiers found in iterative hmmer searches against non-redundant database. Acanth. castellanii indicates Acantanthamoeba castellanii, where * indicates that this entry is annotated as a RAS protein. Dicty. purpureum is Dictyostelium purpureum, Aae is Aedes aegypti, Aqu is Amphimedon queenslandica (sponge), and Clu is Clavispora lusitaniae (fungi). Red numbers within the tree indicate number of trees corresponding to more than 80% of statistical support for a given group, whereas black indicates values below 80%. Only representative numbers have been shown for clarity.
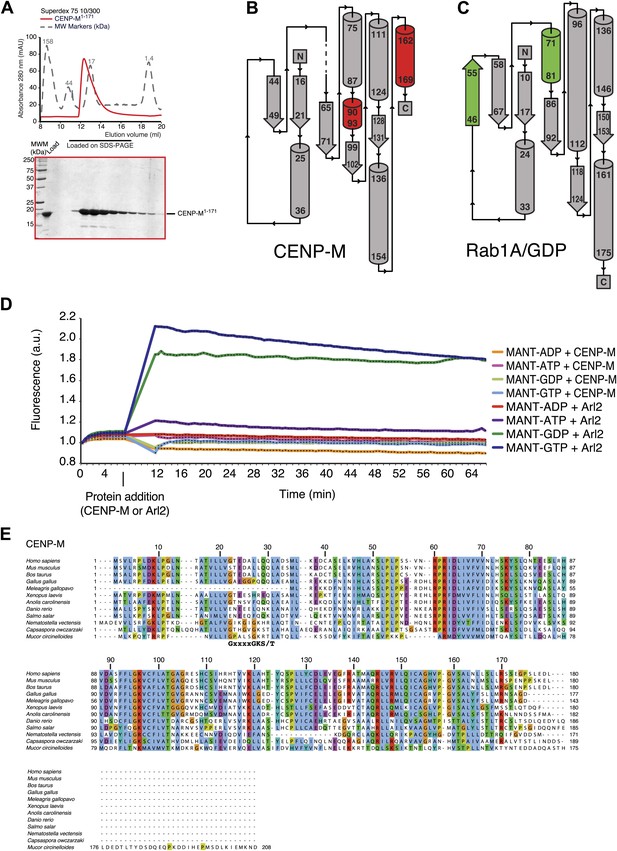
Additional analyses of CENP-M (related to Figure 3).
(A) SEC elution profile of CENP-M1–171. The protein (∼19 kDa) elutes as expected for a monomer. (B) Topology diagram of CENP-M. (C) Topology diagram of the small GTPase Rab1A. (D) Experiments with N-methylanthraniloyl (MANT) derivatives of GTP and ATP (Hiratsuka, 1983) demonstrate that CENP-M does not bind to GTP, GDP, ATP or ADP. The small GTPase Arl2 was used both as positive and as negative control. (E) Alignment of the sequences of 12 distant CENP-M orthologs.
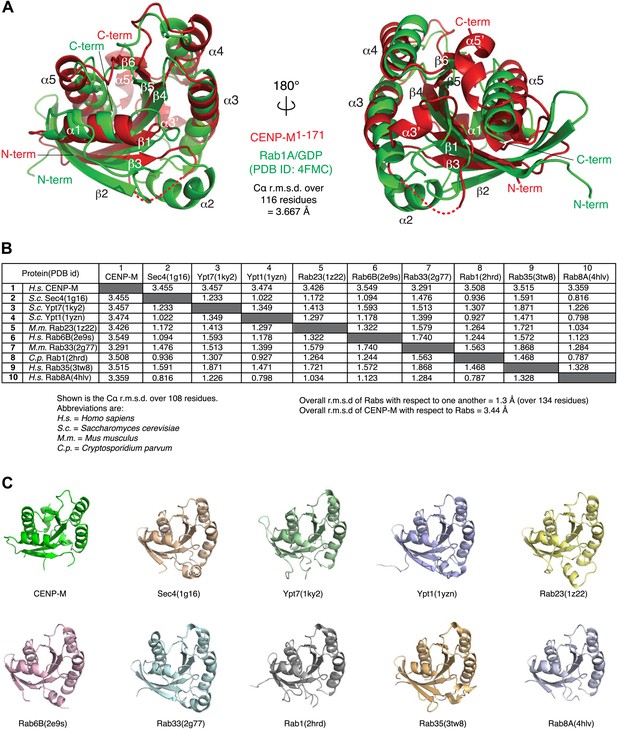
Comparison of CENP-M with Rab-like GTPases.
(A) The coordinates of CENP-M (red) and Rab1A (green; PDB code 4FMC) were superposed using the PDBeFold server (http://www.ebi.ac.uk/msd-srv/ssm/cgi-bin/ssmserver; Krissinel and Henrick, 2004) and visualized using Pymol. (B) Table showing the root mean square deviation in the coordinates of Cα atoms after superposition of CENP-M with the indicated models of Rab-family GTPases (PDB ID codes are reported). The superpositions were carried out using PDBeFOLD. (C) Cartoon models of CENP-M and the indicated Rab-family GTPases viewed with the same orientation.
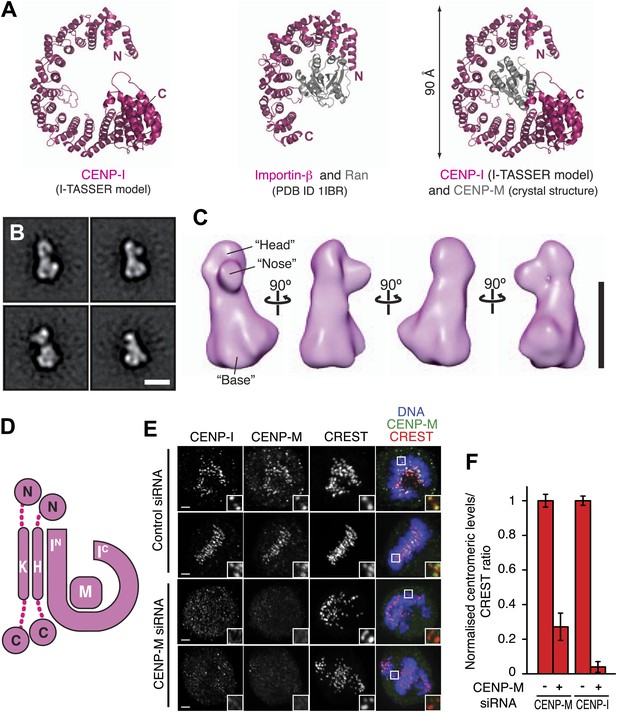
Structural organization of the HIKM complex.
(A) Cartoon representation of the CENP-I model generated by program I-TASSER (left), of the Importin-β/Ran complex (middle), and of a hypothetical structure between CENP-I and CENP-M modeled on the Importin-β/Ran complex (right). A scoring function (C-score) associated with I-TASSER models estimates accuracy of structure predictions. C-score is typically in a range from −5 to 2, where a higher score reflects a model of better quality. Both false positive and false negative rates are estimated to be below 0.1 when a C-score >−1.5 is displayed (Zhang, 2008). The CENP-I model is associated with a C-score of −1. (B) Representative class averages of the negatively stained HIKM complex. Figure 4—figure supplement 3 shows the complete set of class averages. Scale bar = 10 nm. (C) A 3D reconstruction of HIKM complex from negatively stained particles at ∼22 Å resolution. Scale bar = 10 nm. (D) Summary of interactions in the CENP-HIKM complex. The central regions of CENP-H and CENP-K may form an extended parallel interaction, possibly through an α-helical arrangement, which interacts more or less co-linearly with the N-terminal region of CENP-I (IN). Additional globular domains may be present at the N- and C-termini of CENP-H and CENP-K. The entire sequence of CENP-I may fold as a helical solenoid. CENP-M does not interact with CENP-H/K and may bind near the concave surface of the predicted CENP-I solenoid, becoming largely buried. (E) siRNA depletion of endogenous CENP-M abrogates CENP-I kinetochore localization in HeLa cells. Representative cells displayed here are the same shown in Figure 1D, but with addition of CENP-I staining (left panels). Insets display a higher magnification of regions outlined by white boxes. Scale bars = 2 µm. (F) CENP-M and CENP-I kinetochore levels from the experiment illustrated in E. Quantification for CENP-M kinetochore levels are the same shown in Figure 1D and were performed as previously described. Graphs and bars indicate mean ± SEM.
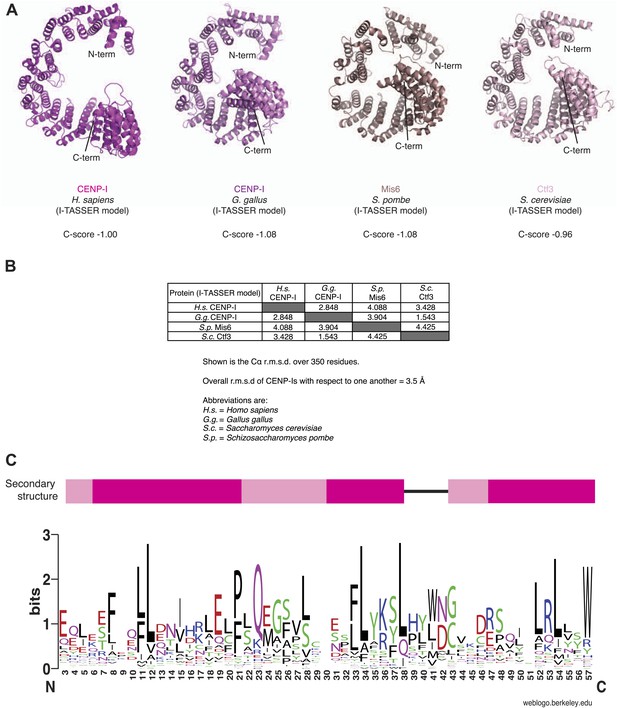
Structural predictions on CENP-I orthologs.
(A) The sequences of human CENP-I plus three orthologues were submitted to the I-TASSER server (http://zhanglab.ccmb.med.umich.edu/I-TASSER/). The best models from each search were superposed on the human model and displayed. The C-score is shown for each model. (B) Summary of the superposition for each homology model. Importantly, while similar to one another, the models are also sufficiently different, as a result of different templates being used for homology modeling. (C) Analysis of α-helical repeat elements. Repeat elements within the human CENP-I sequence were identified by the RADAR server (http://www.ebi.ac.uk/Tools/pfa/radar/) (Heger and Holm, 2000). RADAR also detected repeat elements in the equivalent region for chicken CENP-I, Mis6, and Ctf3. From a multiple sequence alignment of 25 CENP-I orthologs, the three repeat regions were all aligned against each other. The multiple sequence alignment was then submitted to the WebLogo server (http://weblogo.berkeley.edu/logo.cgi) (Crooks et al., 2004) to generate a consensus sequence. The secondary structure for the repeats, based on the homology models, is shown above. Consistent α-helical elements are shown in pink; variable elements are shown in light pink; unstructured regions are represented as a black line. The repeat is rather divergent, and indeed is sometimes comprised of three rather than two α-helices.
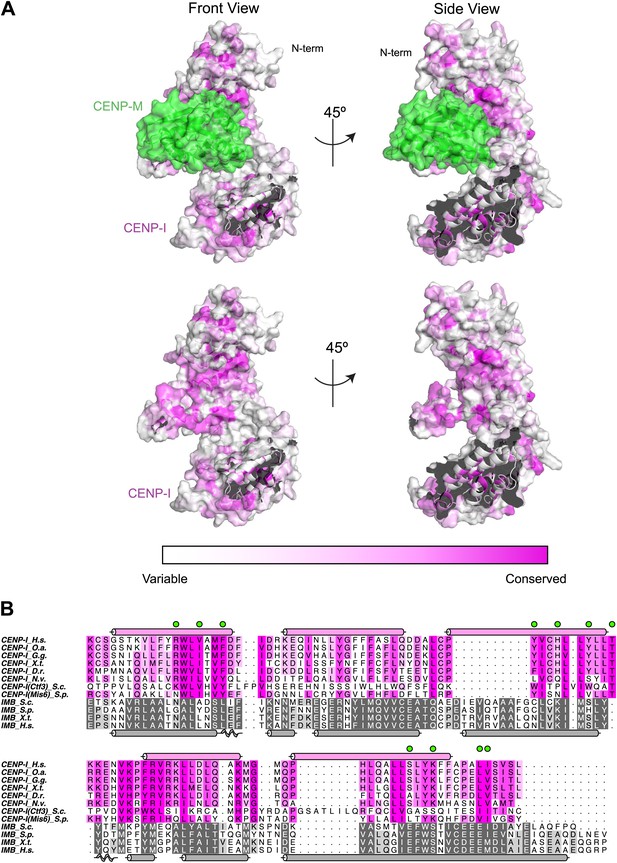
Conservation mapped on the CENP-I model.
(A) Surface representation of the CENP-M/I model (as shown in Figure 4). The C-terminal part of the CENP-I model has been removed to show the concave surface with clarity. Residues are colored from white to magenta according to conservation within 23 different CENP-I orthologues. (B) A representative sequence alignment of eight CENP-I sequences from different organisms is shown, with a focus on the region that, according to the model, may be involved in CENP-M binding. Surface residues predicted to be in contact with CENP-M are highlighted with green circles. A comparison of four importin-β (ImpB) orthologues is shown to demonstrate that while many residues are conserved between CENP-I and ImpB (consistent with a similar fold) those residues that might contact CENP-M are divergent (consistent with binding a different ligand). Species abbreviations are as follows H.s., Homo sapiens; O.s., Ornithorhynchus anatinus; G.g., Gallus gallus; X.t., Xenopus tropicalis; D.r., Danio rerio; N.v.; Nematostella vectensis; S.c., Saccharomyces cerevisiae S288C; S.p., Schizosaccharomyces pombe.
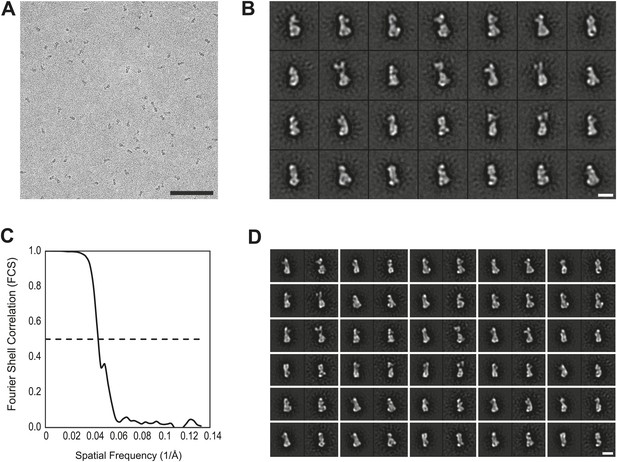
EM analysis.
(A) Representative electron micrograph area of the negatively stained CENP-HIKM complex. Scale bar = 100 nm. (B) Collection of class averages of the CENP-HIKM complex derived from a data set of 5958 single particles. Selected classes are shown in Figure 4B. Scale bar = 10 nm. (C) Fourier shell correlation (FSC) curves of the negative stain reconstruction of the CENP-HIKM complex. The resolution was estimated by the FSC 0.5 criterion to be 22 Å. (D) Reprojections of the 3D reconstruction paired with their corresponding class averages. Scale bar = 10 nm.
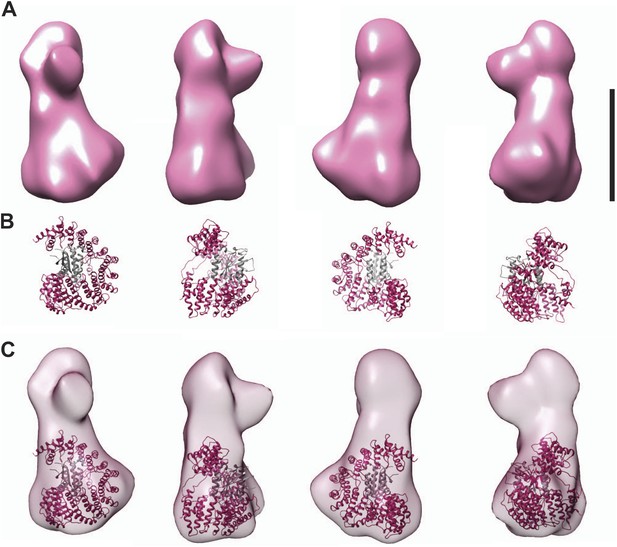
Fitting the CENP-I/M model in the EM density.
(A) A 3D reconstruction of HIKM complex from negatively stained particles at ∼22 Å resolution (already shown in Figure 4C). Scale bar = 10 nm. (B) Four orientations of the CENP-I/M model built by I-TASSER (see Figure 4A, right). (C) Tentative manual fitting of the CENP-I/M model into the EM density. The CENP-I/M model fits snugly in the ‘base’ density, leaving empty space in the ‘head’ and ‘nose’ domains, which is therefore predicted to host CENP-H/K.
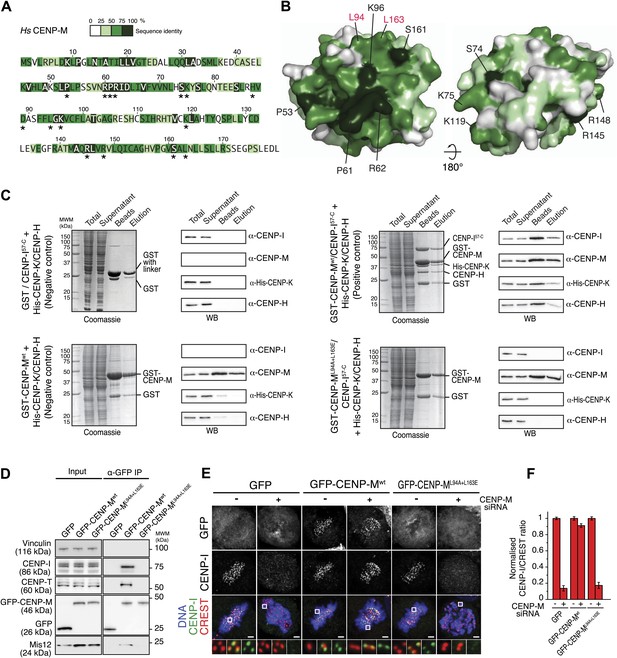
CENP-M residues required for kinetochore targeting of CENP-I.
(A) The CENP-M alignment identifies highly conserved residues (based on alignment in Figure 3—figure supplement 1, panel D), a subset of which (asterisks) is exposed at the surface of CENP-M. (B) Position of conserved residues shown in A on two opposite faces of the CENP-M surface. (C) After insect cell co-expression of indicated proteins, affinity purification with GST-CENP-M (if present) led to isolation of associated proteins shown, after SDS-PAGE separation, in the ‘beads’ fraction. Material eluted from beads was collected and shown in lanes labeled ‘elution’. Co-expression of all four CENP-HI57–CKM subunits (positive control) is necessary for identification of the CENP-HI57–CKM complex on beads and in elution fraction. CENP-ML94A–L163E fails to assemble CENP-HI57–CKM despite the expression of CENP-H, CENP-I57–C and CENP-K. (D) GFP-CENP-Mwt, but not GFP-CENP-ML94A+L163E, co-immunoprecipitates CENP-I, CENP-T and Mis12 from mitotic cells. Panels represent the α-GFP co-immunoprecipitation analysis of protein extracts obtained from mitotic HeLa Flp-In T-REx cells stably expressing GFP, GFP-CENP-Mwt or GFP-CENP-ML94A+L163E from an inducible promoter. Total protein extracts (Input) and immunoprecipitates (α-GFP IP) were run on SDS-PAGE and subjected to WB with indicated antibodies. Vinculin was used as a loading control. (E) Representative images of HeLa Flp-In T-REx cells treated with siRNA for endogenous CENP-M and expressing the indicated siRNA-resistant GFP-CENP-M fusions. Expression of GFP-CENP-Mwt, but not of GFP-CENP-ML94A+L163E, rescues chromosome alignment defects and loss of CENP-I kinetochore localization observed upon depletion of endogenous CENP-M. Following fixation, cells were immunostained and imaged for GFP, CENP-I, CREST and DNA (DAPI). Insets show a higher magnification of regions outlined by white boxes. Scale bars = 2 µm. (F) Quantification, for experiment in E, of the CENP-I kinetochore levels normalized to CREST kinetochore signal. Graphs and bars indicate mean ± SEM. See ‘Materials and methods’ section for details on quantification.
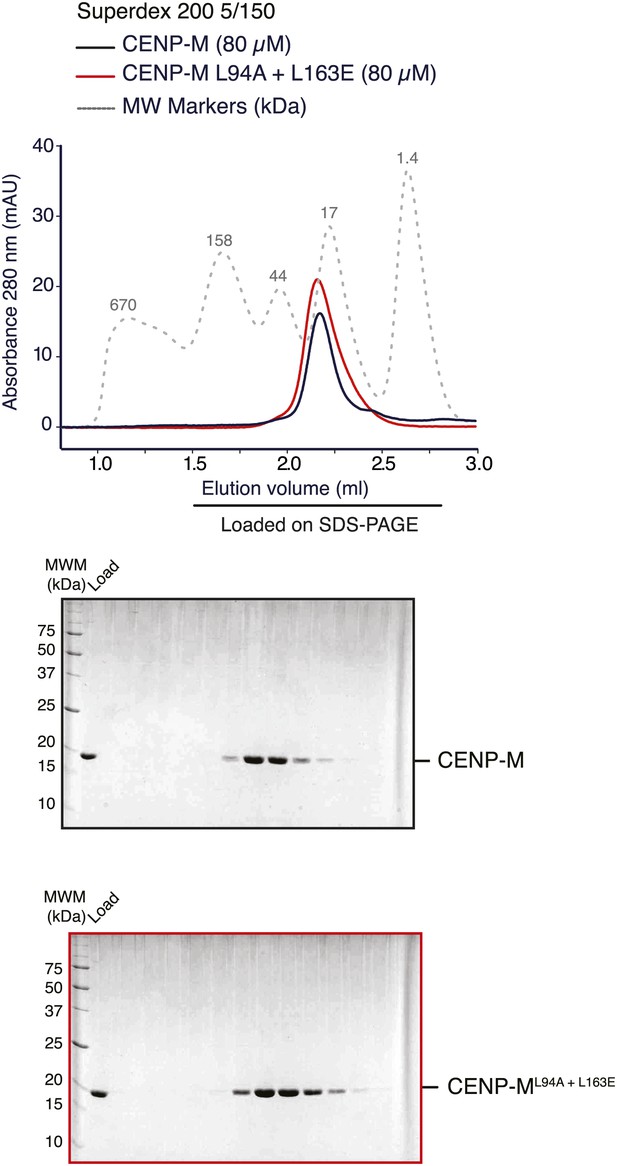
Stability of CENP-M mutant.
SEC runs and corresponding SDS-PAGE of CENP-Mwt or CENP-ML94A+L163E purified to homogeneity after expression in E. coli indicate essentially identical elution profiles, suggesting that the point mutations do not affect the stability of the CENP-M mutant.
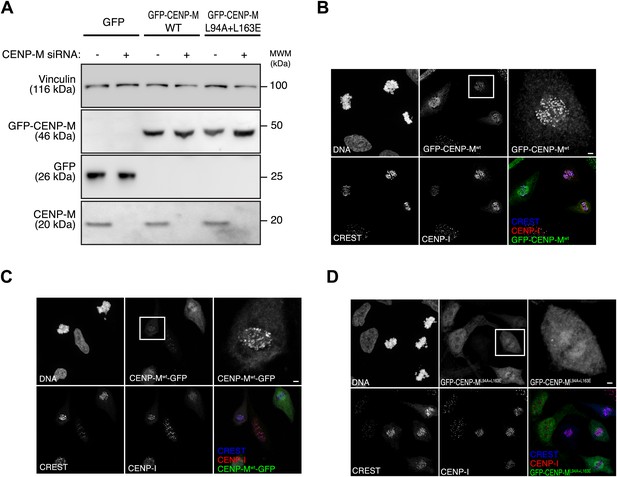
Inducible expression and localization of CENP-M.
(A) HeLa Flp-In T-REx cells expressing siRNA resistant GFP-CENP-M fusions and treated with siRNA specific for endogenous CENP-M show significantly reduced levels of the target protein. The effect of CENP-M siRNA treatment on the cellular levels of CENP-M, GFP, and GFP-CENP-M fusions was monitored by Western blotting. Protein extracts from the indicated conditions were run on SDS-PAGE and immunoblotted for the indicated proteins. Vinculin was used as a loading control. (B–D) Representative images of the cellular localization of CENP-M fusions to GFP. HeLa Flp-In T-REx cells expressing GFP fused to CENP-Mwt (either N-terminally [B] or C-terminally [C]) or CENP-ML94A+L163E (D) were analysed for the localization of CENP-I and the GFP fusions. Cultures were enriched for G2 cells by inhibiting Cdk1 with RO-3306 (Vassilev et al., 2006). G2 cells were then released into mitosis with a washout of the inhibitor in pre-warmed media, fixed after 1 hr and imaged for GFP, endogenous CENP-I, CREST and DNA (DAPI). Insets show a higher magnification of the regions outlined by the white boxes. Scale bar represents 2 µm.
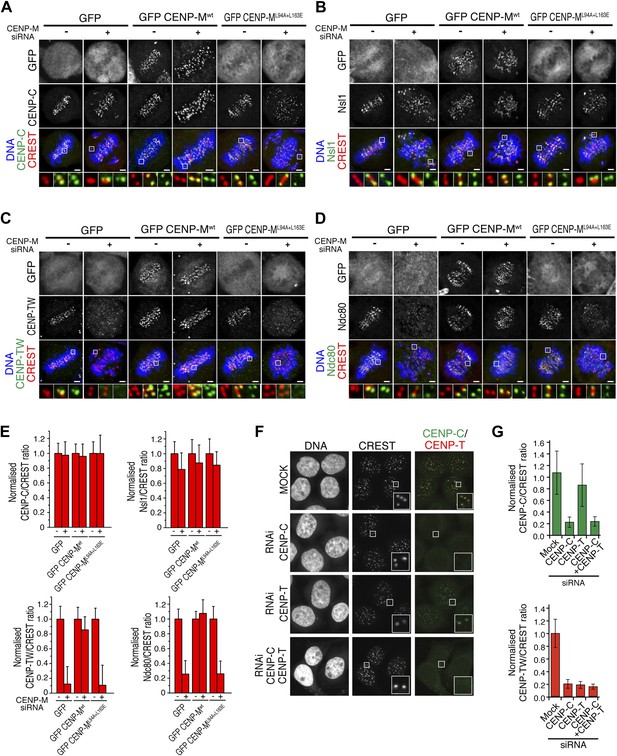
Significance of the CENP-M/CENP-I interaction for kinetochore assembly.
(A–D) Representative images of the localization of kinetochore proteins in HeLa Flp-In T-REx cells treated with siRNA for endogenous CENP-M and expressing the indicated siRNA-resistant GFP-CENP-M fusions. Scale bars = 2 µm. (E) Quantification, for experiments A–D, of the kinetochore levels of the indicated proteins normalized to CREST. Graphs and bars indicate mean ± SEM. (F) Depletion of CENP-C abrogates kinetochore accumulation of CENP-T/W. Representative images of HeLa cells treated with siRNA for CENP-C or CENP-T and arrested in G2 with the Cdk1 inhibitor RO-3306 (‘Materials and methods’). Following fixation, cells were immunostained for CENP-C, CENP-T/W and CREST. DNA was stained with DAPI. Scale bars = 10 µm. (G) Quantification, for experiment F, of the kinetochore levels of the indicated proteins normalized to CREST kinetochore signal. Graphs and bars indicate mean ± SEM.
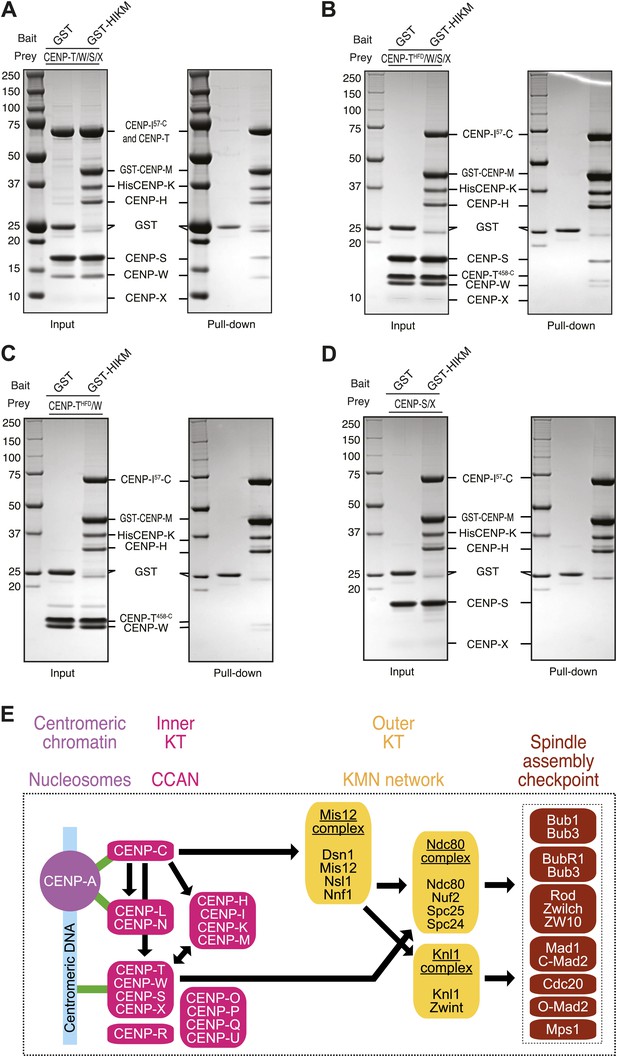
Direct interaction of HIKM complex with the CENP-T/W complex.
(A–D) GST or GST-HIKM baits were immobilized on beads and incubated with (A) CENP-T/W/S/X complex, (B) CENP-T452–C/W/S/X complex, (C) CENP-T452–C/W, or CENP-S/X. For each sample, both the input and the solid phase bound material (indicated as ‘pull-down’) are shown after separation by SDS-PAGE and staining with Coomassie brilliant blue. Note that full-length CENP-T and CENP-I57–C migrated indistinguishably. (E) Model of kinetochore assembly supported by our analysis. CENP-C and possibly CENP-N/L interact directly with the CENP-A nucleosome. The presence of CENP-C at the centromere is essential for the recruitment of CENP-T/W and CENP-HIKM complex. CENP-T/W and CENP-HIKM complex are co-dependent and interact physically with each other.
Tables
List of potential binding interactions of CENP-M tested with purified proteins and complexes
| CENP-M incubation with: | Binding: |
|---|---|
| H3-containing mononucleosomes (DNA 601–167 bp) | NO |
| CENP-A-containing mononucleosomes (DNA 601–167 bp) | NO |
| CENP-C constructs (1–544, 509–760, 631–C-terminus) | NO |
| CENP-L/CENP–N complex | NO |
| CENP-H/CENP–K complex, CENP-I57–281, CENP-H/CENP-K/CENP-I57–281 complex | NO |
| CENP-O/CENP-P/CENP-Q/CENP–U complex, CENP-O/CENP-P and CENP-Q/CENP-U sub-complexes | NO |
| CENP-R | NO |
| CENP-T/CENP-W/CENP-S/CENP-X complex, CENP-T/CENP-W sub-complex (phosphorylated by Cdk1 or not), CENP-S/CENP-X sub-complex | NO |
| Mis12 complex | NO |
| Ndc80 complex | NO |
| Knl12000–2311 | NO |
| Zwint | NO |
| KMN network | NO |
| Microtubules | NO |
Data collection, phasing and refinement statistics
| Native | Derivative | |
|---|---|---|
| Data collection | ||
| Beamline | ESRF ID14-4 | SLS X06DA (PXIII) |
| Spacegroup | P3 | P3 |
| Unit cell parameters (Å, °) | a = b = 104.50, c = 33.59 | a = b = 104.03, c = 33.56 |
| α = β = 90, γ = 120 | α = β = 90, γ = 120 | |
| Wavelength (Å) | 0.91970 | 0.97942 |
| Resolution limits (Å) | 52.25–1.49 (1.54–1.49)* | 31.45–2.00 (2.06–2.00)* |
| Reflections observed/unique | 607786/64606 | 419456/27271 |
| Completeness (%) | 98.3 (96.3)* | 99.9 (98.9)* |
| Rsym†(%) | 5.6 (36.1)* | 9.4 (75.2)* |
| <I>/<σI> | 26.3 (6.4)* | 24.1 (3.9)* |
| Redundancy | 9.4 (8.8)* | 15.4 (13.9)* |
| SAD phasing | ||
| BAYES-CC | 49.1 ± 18.5 | |
| Se sites found/expected | 5/6 | |
| FOM before solvent flattening and density modification | 0.35 | |
| FOM after solvent flattening and density modification | 0.69 | |
| Refinement | ||
| Resolution limits (Å) | 52.25–1.49 (1.52–1.49)* | |
| Reflections for Rcryst/for Rfree | 59806/4800 | |
| Rcryst‡(%) | 12.4 (20.6)* | |
| Rfree‡(%) | 16.4 (23.0)* | |
| No. of protein atoms/water atoms | 2307/297 | |
| Average B factor protein atoms/water atoms (Å2) | 21.67/35.34 | |
| RMSD bond lengths (Å) | 0.005 | |
| RMSD bond angles (°) | 0.854 | |
| Twin fraction (operator −h, −k, l) | 0.49 | |
| Ramachandran Plot Statistics§ | ||
| Favoured region (%) | 99.3 | |
| Outliers (%) | 0.0 | |
-
BAYES-CC: Bayesian estimate of the correlation coefficient (CC) between the experimental map and an ideal map, reported as CC * 100 ± 2 standard deviations.
-
FOM: figure of merit.
-
RMSD: root mean square deviation.
-
*
Values in parentheses refer to the highest resolution shell.
-
†
Rsym = ΣhΣi|Ih,i − <Ih>|/ΣhΣi Ih,i.
-
‡
Rcryst and Rfree = Σ|Fobs − Fcalc|/Σ Fobs; Rfree calculated for a 7.4% subset of reflections not used in the refinement.
-
§
Calculated using MOLPROBITY within the PHENIX suite.





