The metal transporter ZIP13 supplies iron into the secretory pathway in Drosophila melanogaster
Figures
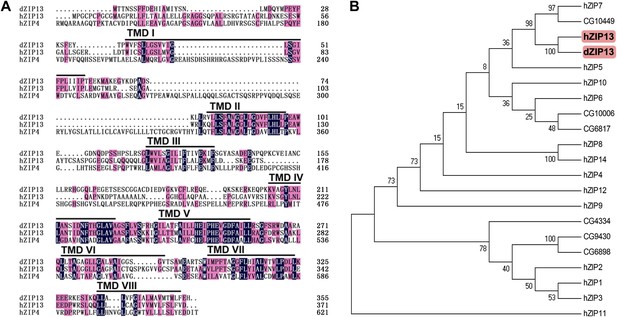
Sequence analysis of Drosophila ZIP13.
(A) Alignment of Drosophila ZIP13 (dZIP13, the top), human ZIP13 (hZIP13, the middle), and human ZIP4 (hZIP4, the bottom) proteins. Amino acid sequences for hZIP13, hZIP4, and dZIP13 (CG7816) were obtained from GenBank and aligned by HMHMM software. Black and pink shadings indicate respectively identical and conservative amino acids. The eight putative transmembrane (TM) regions are underlined and denoted as ‘TM I’ through ‘TM VIII’. (B) Phylogenetic tree analysis of human and putative Drosophila ZIP family members. The tree was generated using ClustalX version 1.81 and displayed with TreeView. Bootstrap probabilities for major clusters are shown by percentages. Accession numbers are listed for other Drosophila ZIPs used for the alignment.
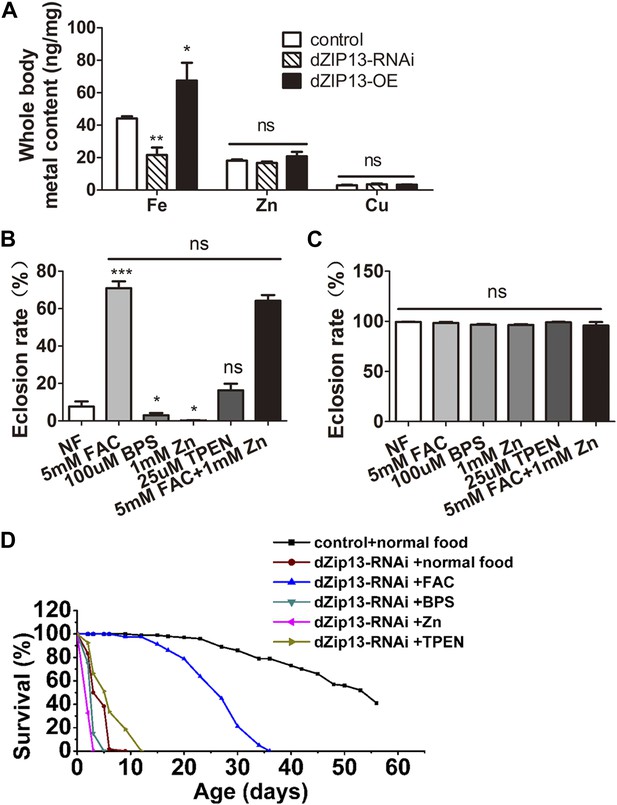
dZIP13-RNAi flies display iron-rescuable defects.
(A) Body metal contents when dZIP13 expression was modulated. Shown are flies with modulated dZIP13 expression in the midgut (NP3084 as the Gal 4 driver). A significant decrease in the whole body iron content, but not that of zinc or copper, was observed in dZIP13-RNAi flies, while dZIP13 overexpression led to an iron increase. Values represent three independent measurements and are normalized to the dry body weights; data are presented as means + SEM; n = 3 or 6. *p<0.05, **p<0.01; two-tailed Student's t test. (B) The eclosion rate of ubiquitously-RNA-interferenced-dZIP13 (Da > dZIP13-RNAi) larvae could be rescued by dietary iron supplementation. Da-Gal4 was crossed to wild-type or dZIP13-RNAi flies on juice-agar plates. Newly hatched progeny were transferred to normal food, or food supplemented with ZnCl2, TEPN, FAC, or BPS. Percentages of flies that eclosed to adults were counted; n = 6 or 8. (C) A control showing that the same amount of zinc, iron, or chelators supplemented in the food had no effect on the eclosion rate of wild-type Drosophila. (D) The shortened lifespan of Da > dZIP13-RNAi adults was partially rescued by dietary iron supplementation but not zinc. Percentages of flies that eclosed to adults were counted; n = 5.
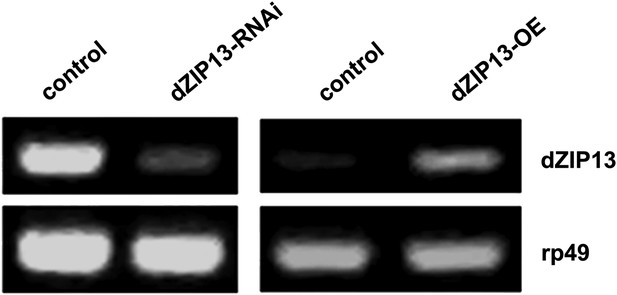
RT-PCR analysis of efficacy of dZIP13 knockdown or overexpression.
RT-PCR analysis of dZIP13 mRNA abundance in third instar larvae. rp49 was used as the loading control. Da-GAL4 was used as the expression driver.
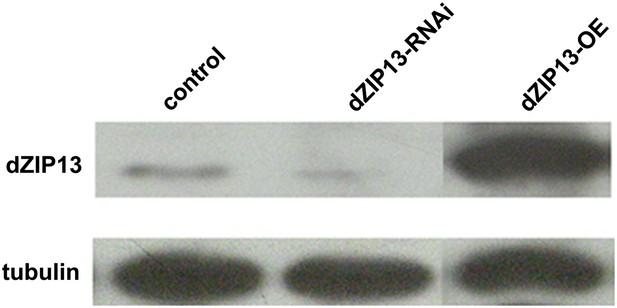
Western blot analysis of efficacy of dZIP13 knockdown or overexpression.
Western blot showing that the RNAi used in this study suppressed dZIP13 protein to a significantly reduced level. Tubulin was used as the loading control. Da-GAL4 was used as the expression driver.
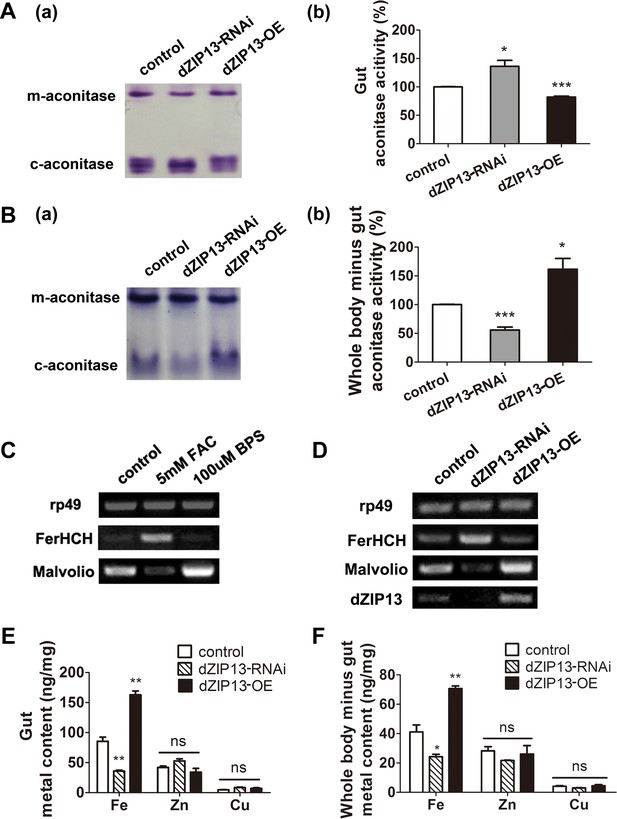
dZIP13 knockdown led to iron deficiency in the body but not the cytosol of gut cells.
(A) Cytosolic aconitase activity was increased in the gut of NP3084>dZIP13-RNAi larvae and decreased in NP3084>dZIP13-OE larvae, suggesting respectively iron elevation and iron deficiency in the cytoplasm. Panel (b) was quantitative measurement of (a). Results are presented as mean + SEM relative activity; n = 3. *p<0.05, **p<0.01, ***p<0.001; two-tailed Student's t test. (B) Cytosolic aconitase activity was decreased in the whole body minus gut (body parts other than the gut) of NP3084>dZIP13-RNAi larvae and increased in NP3084>dZIP13-OE larvae. Panel (b) was quantitative measurement of (a). Results are presented as mean + SEM relative activity; n = 3. *p<0.05, **p<0.01, ***p<0.001; two-tailed Student's t test. (C) RT-PCR analysis of iron homeostasis genes of normal flies in response to iron changes (feeding with iron or chelator). RNA was made from third instar larvae midguts. rp49 was used as the loading control. (D) RT-PCR analysis of iron homeostasis genes in the midgut of dZIP13 RNAi or OE third instar larvae. Expression is driven by the midgut driver NP3084. (E) An analysis of metal contents in the gut when dZIP13 expression was modulated. Shown are metal levels from fly larvae with modulated dZIP13 expression in the midgut (NP3084 as the Gal 4 driver). A significant decrease in the gut iron, but not zinc or copper, was observed in dZIP13-RNAi flies; dZIP13 overexpression led to an iron increase. Results are presented as mean + SEM relative activity; n = 3. *p<0.05, **p<0.01, ***p<0.001; two-tailed Student's t test. (F) An analysis of metal contents in the whole-body-minus-gut parts when dZIP13 expression was modulated in the larval midgut (NP3084 as the Gal 4 driver). A significant decrease in the whole-body-minus-gut iron, but not zinc or copper, was observed in dZIP13-RNAi flies; dZIP13 overexpression led to an iron increase. Results are presented as mean + SEM relative activity; n = 3. *p<0.05, **p<0.01, ***p<0.001; two-tailed Student's t test.
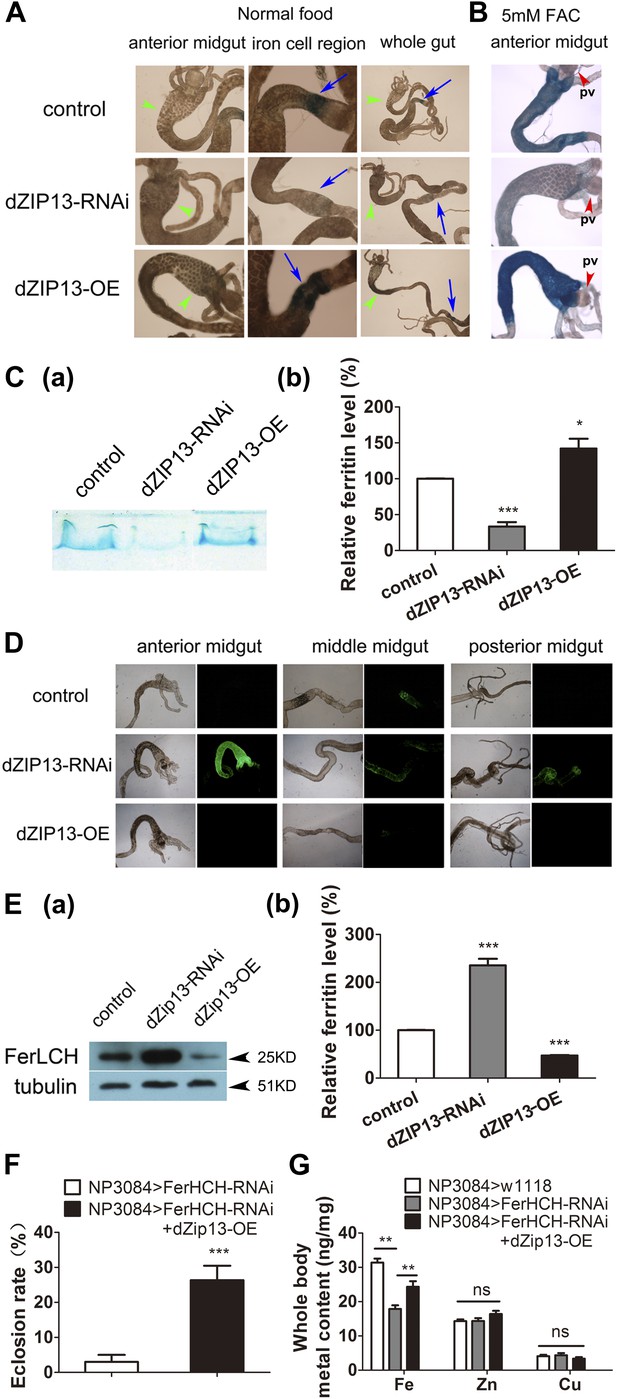
dZIP13 knockdown results in reduced ferritin iron loading in the gut.
(A) Staining of ferric iron in the larval gut. The staining in the midgut constriction and ectopic ferric staining in the anterior midgut are noted separately by arrows (blue) and arrow heads (green). The anterior midgut of NP3084>dZIP13-OE fly larvae deposited obviously a higher amount of iron than the control. Ferric iron significantly accumulated in the iron cell region of NP3084>dZIP13-OE while NP3084>dZIP13-RNAi adults showed almost no iron staining. Shown are representative images and in bright and dark fields. More images are shown in Figure 4—figure supplement 1. (B) Staining of ferric iron in the anterior midgut of iron-fed larvae. The anterior midgut follows the preceding distinct proventriculus (pv, red arrowheads). (C) Staining of ferric iron (bound to ferritin) on native PAGE. Same amounts of total protein extracts from control and NP3084>dZIP13-RNAi or NP3084>dZIP13-OE larval guts were loaded. The gel was directly stained with Prussian blue staining solution. For an intact gel image, see Figure 4—figure supplement 2. Panel (b) was quantitative measurement of (a). n = 3. *p<0.05, ***p<0.001; two-tailed Student's t test. (D) Ferritin expression in the gut. An obviously higher amount of ferritin was expressed in the gut of NP3084>dZIP13-RNAi larvae. The expression of ferritin was indicated with a protein trap line Fer1HCHG188, which tags the endogenous Fer1HCH through an N-terminal GFP fusion. Shown are representative results and more images are shown in Figure 4—figure supplement 3. (E) Western blot of ferritin of NP3084>dZIP13-RNAi and NP3084>dZIP13-OE larvae. Anti-ferritin light chain antibody was used. Tubulin was used as a loading control. For an intact gel image, see Figure 4—figure supplement 4. Panel (b) was quantitative measurement of (a). n = 3. ***p<0.001; two-tailed Student's t test. (F) Eclosion rescue of ferritin-RNAi by dZIP13. The eclosion rate of gut-specific (NP3084) ferritin-RNAi flies was rescued from <5% to ∼30% by dZIP13 overexpression. Newly hatched progeny were transferred to normal food, and percentages of flies that eclosed to adults were counted. n = 6. ***p<0.001; two-tailed Student's t test. (G) Rescue of iron deficiency of ferritin-RNAi by dZIP13. The reduced body iron content in gut-specific ferritin-knockdown larvae was also partially rescued by dZIP13 overexpression. n = 6. **p<0.01; two-tailed Student's t test.
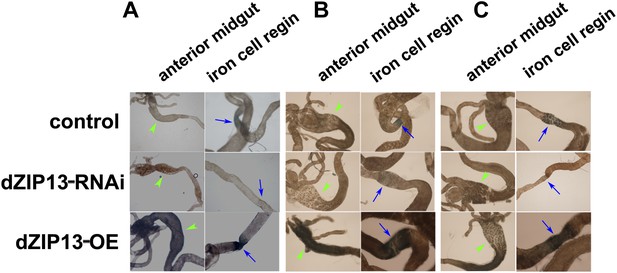
Staining of ferric iron in the larval gut.
The ferric staining in the midgut constriction and the ectopic ferric staining in the anterior midgut are noted separately by arrows (blue) and arrow heads (green). The anterior midgut of NP3084>dZIP13-OE fly larvae deposited an obviously higher amount of iron than the control. Ferric iron accumulated prominently in the iron cell region of NP3084>dZIP13-OE while NP3084>dZIP13-RNAi larvae showed almost no iron staining. (A), (B) and (C) are results from three independent experiments.
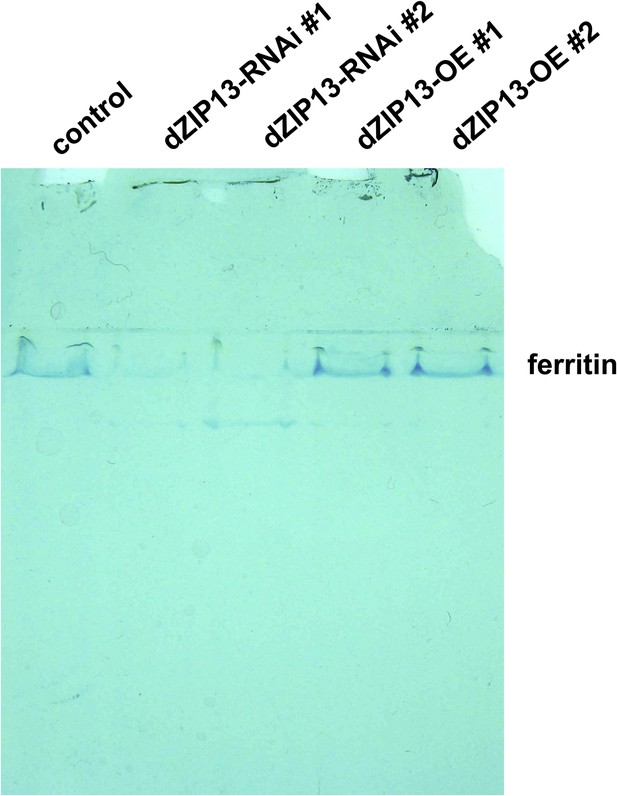
Staining of ferric iron (bound to ferritin) on native PAGE.
Same amounts of total protein extracted from control and NP3084>dZIP13-RNAi or NP3084>dZIP13-OE larvae guts were loaded. The gel was stained with Prussian blue staining solution.
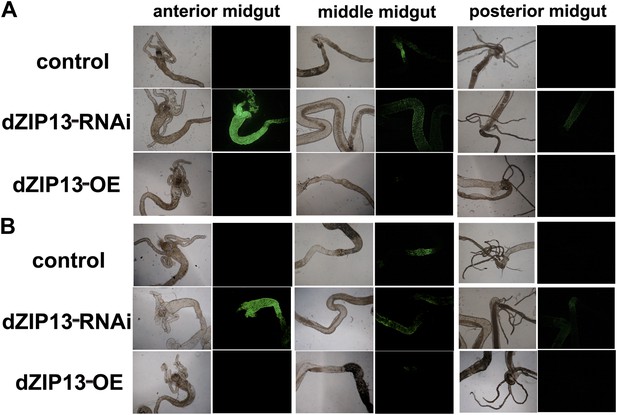
A significantly higher amount of ferritin was detected in the gut of NP3084>dZIP13-RNAi fly larvae.
The expression of ferritin was indicated with a protein trap line Fer1HCHG188 expressing an N-terminal GFP-tagged Fer1HCH protein. Shown are images from the bright and dark fields. (A) and (B) are results from two independent experiments.
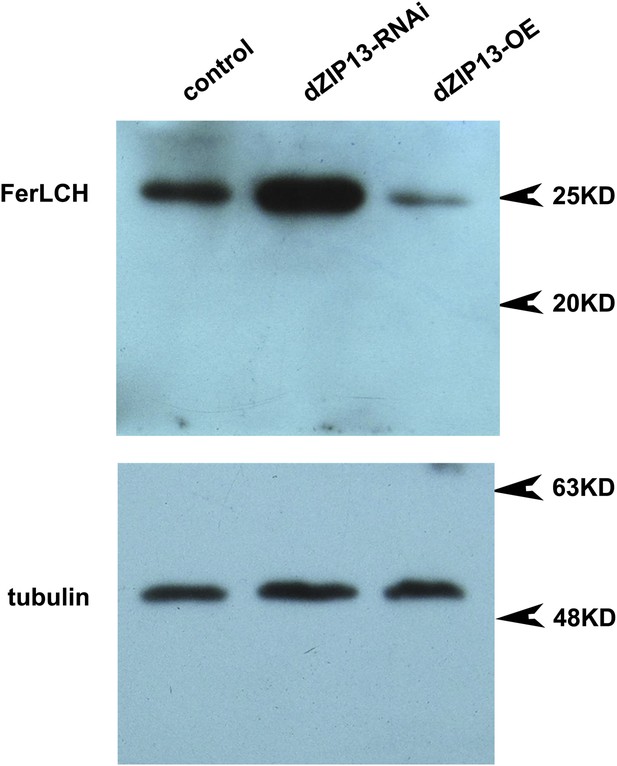
Western blot of ferritin of NP3084>dZIP13-RNAi and NP3084>dZIP13-OE larvae.
Anti-ferritin light chain antibody was used. Tubulin was used as the loading control.
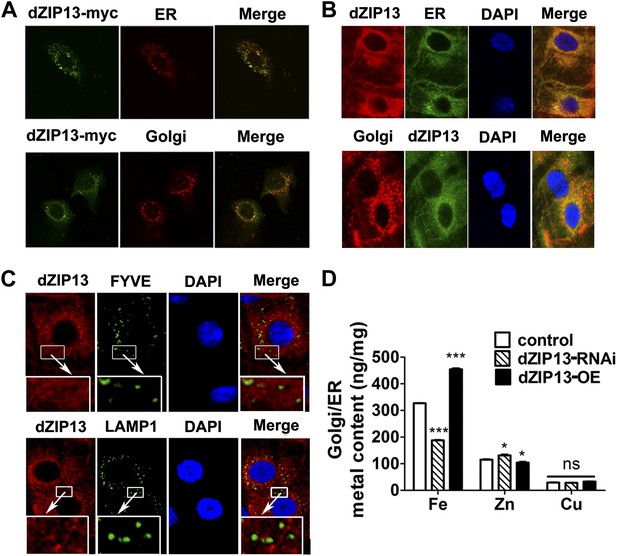
dZIP13 is located to ER/Golgi and involved in their iron regulation.
(A) The localization of dZIP13 in Caco2 cells. dZIP13-myc was detected by myc antibody. This immunofluorescence staining showed that dZIP13 partially co-localizes with the ER/Golgi in Caco2 cells. (B) The localization of dZIP13 in Drosophila midgut epithelial cells. dZIP13 was detected directly by dZIP13 antibody. These images indicated dZIP13 partially co-localizes with ER/Golgi in Drosophila gut cells. (C) dZip13 does not co-localize well with endosome markers. The localizations of dZIP13 and endosomes in Drosophila midgut epithelial cells were shown. dZIP13 was detected directly with dZIP13 antibody. Lysosome-associated membrane protein 1 (LAMP1) fused with GFP was used to indicate lysosomes and late endosomes; FYVE was used to mark the early endosomes. (D) Less iron in ER/Golgi from dZIP13-RNAi larvae and more in that of dZIP13-OE larvae. Copper was not affected while zinc contents were marginally different. n = 3. *p<0.05, ***p<0.001; 2-tailed Student's t test.
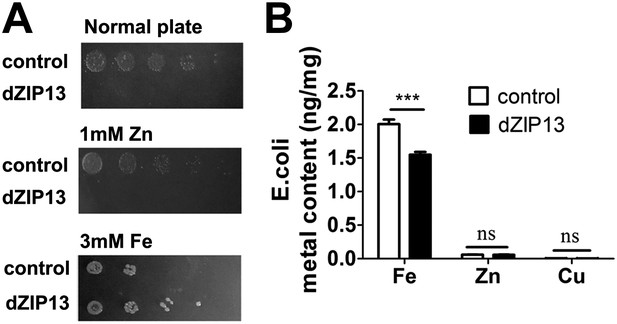
Heterologous dZIP13 expression renders the growth of E. coli iron-dependent and iron-resistant.
(A) E. coli expressing dZIP13 required iron addition to grow and was more resistant to iron excess. (B) Less iron content was detected in E. coli expressing dZIP13 while zinc and copper were not much affected. n = 3. ***p<0.001; two-tailed Student's t test.
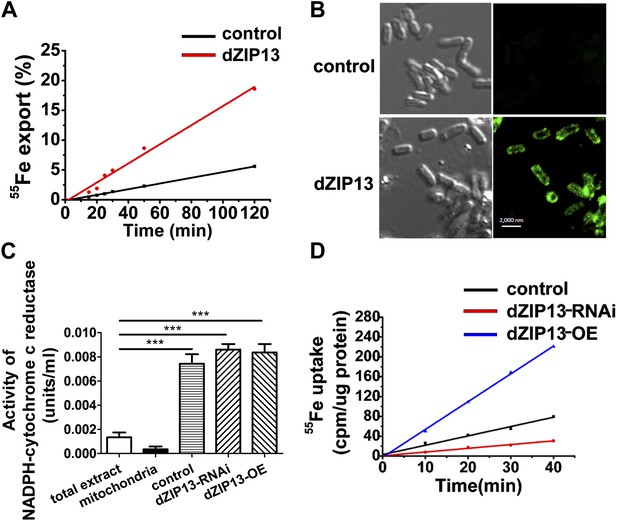
Iron radioisotope transport indicates dZIP13 as an iron exporter.
(A) A time-course of iron released from dZIP13-expressing E. coli cells. dZIP13 expression in E. coli significantly increased the iron efflux rate than the vector control. (B) Imunofluoresence of E. coli expressing dZIP13 indicated that at least some dZIP13 was located on the membrane of E. coli. (C) NADPH-cytochrome c reductase activity assay showed that the ER–Golgi preparation was indeed enriched with a large amount of ER/Golgi. (D) A time-course of iron uptake into the ER/Golgi sample. The rate of iron uptake was much higher in dZIP13-OE and lower in dZIP13-RNAi ER/Golgi samples.
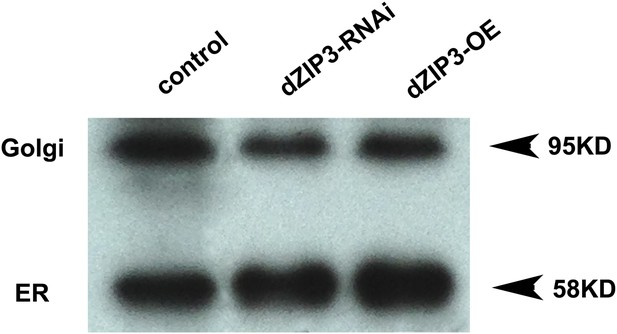
Western blot showing that the ER/Golgi samples purified contain both ER and Golgi.
Golgi and ER were detected with anti-GM130 and anti-PDI antibodies respectively.
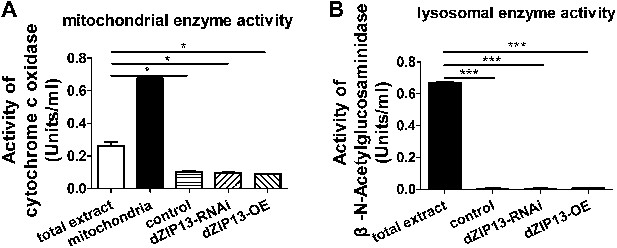
Purity of the ER/Golgi samples isolated from Drosophila.
(A) Cytochrome c oxidase assay indicated that the purified samples still contained some residual mitochondrial parts. (B) β-N-Acetylglucosaminidase (NAG) activity assay showed that the samples were essentially free of lysomal contamination.
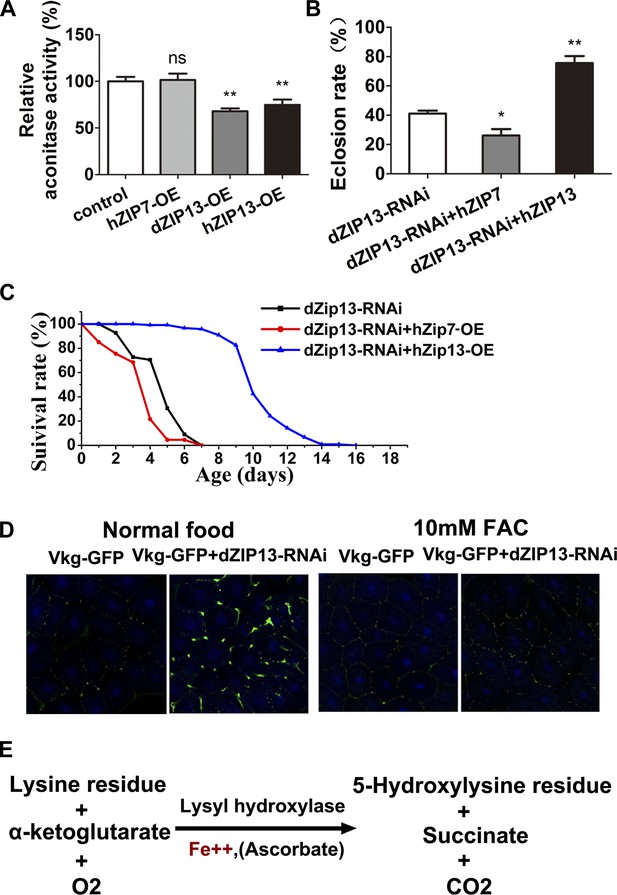
dZIP13 is functionally analogous to hZIP13.
(A) Decreased aconitase activity was observed after either dZIP13 or hZIP13 overexpression, but not a closely related zinc transporter hZIP7. n = 6. **p<0.01; two-tailed Student's t test. (B) The eclosion defect of dZIP13-RNAi flies could be significantly rescued by hZIP13 expression. Eclosion was rescued from ∼40% to ∼75% by hZIP13. n = 6. *p<0.05, **p<0.01; two-tailed Student's t test. (C) The shortened lifespan of dZIP13-RNAi was also rescued by hZIP13. (D) dZIP13-RNAi flies exhibited iron-rescuable collagen defects. Shown are images of fat body cells in VkgG454/+ control larvae and dZIP13-RNAi larvae, cultured on normal food and food with 10 mM FAC. Green: Vkg-GFP, blue: DAPI. (E) The lysyl hydroxylation reaction. The reaction depends on the presence of ferrous iron.

Two conserved amino acids D and H in the fourth transmembrane domain of other ZIPs are switched in ZIP13.
Shown here are the fourth transmembrane domain and adjacent sequences of ZIP13 and several other ZIPs. 1–3 are ZIP13 sequences from different organisms. 4–6 are several other representative ZIPs. Predicted TM4 segments are underlined. The highlighted two amino acids D and H are conserved in all other closely related ZIPs while their positions are switched in ZIP13s. Numbers in parenthesis indicate the starting/ending positions of the ZIPs shown. 1. ZIP13 (Danio rerio); 2. ZIP13 (CG7816) (Drosophila melanogaster); 3. ZIP13 (Homo sapiens); 4. ZIP7 (Homo sapiens); 5. ZIP8 (Homo sapiens); 6. ZIP14 (Homo sapiens).
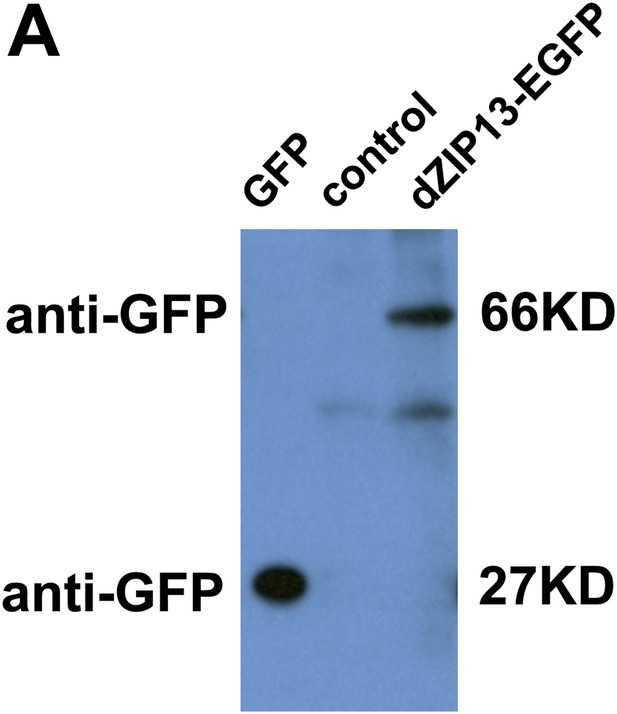
Western blot of dZIP13-EGFP larval extract for EGFP to asses EGFP fusion protein stability. Anti-GFP antibody was used. The GFP lane is the extract from Da>GFP flies (4ug). The control lane is normal fly sample (no EGFP expression) (80ug). Little free EGFP was detected in dZIP13-EGFP sample (80ug). An extra band below that of dZIP13-EGFP is likely a background signal because it is also detected in normal fly sample (control lane).
Tables
Drosophila used in this study.
| Drosophila | Descriptions | Origin |
|---|---|---|
| Da-Gal4 (#8641) | Ubiquitous Gal4 | Bloomington Drosophila Stock Center |
| NP3084 (#113094) | Expresses Gal4 in salivary glands, gastric caecae, and whole midgut in third instar larvae | Genetic Resource Center at the Kyoto Institute of Technology (DGRC) |
| Vkg-GFP (#G00454) | Carries a GFP fused to viking | Flytrap |
| Fer1HCHG188/TM3(#G00188) | Carries a GFP fused to Fer1HCH (ferritin 1heavy-chain homolog) | Flytrap |
| dZIP13-RNAi (#1364) | CG7816 RNAi line | Vienna Drosophila RNAi Center |
| dZIP13-OE | CG7816 over-expression line | This study |
| FerHCH-RNAi (#12925) | Fer1HCH RNAi line | Vienna Drosophila RNAi Center |
| hZIP7-OE | Human ZIP7 over-expression line | This study |
| hZIP13-OE | Human ZIP13 over-expression line | This study |
| Golgi marker | Carries a RFP fused to Rho1 | This study |
| ER marker (#ZCL1503) | Carries a GFP fused to PDI | (Morin et al., 2001) |
| Early endosome marker (#39695) | Carries a GFP fused to FYVE | Bloomington Drosophila Stock Center |
| Late endosome marker (#42714) | Carries a GFP fused to LAMP | Bloomington Drosophila Stock Center |
| dZIP13 mutant (#18595) | CG7816 mutant line | Bloomington Drosophila Stock Center |





