Lhx1 maintains synchrony among circadian oscillator neurons of the SCN
Figures
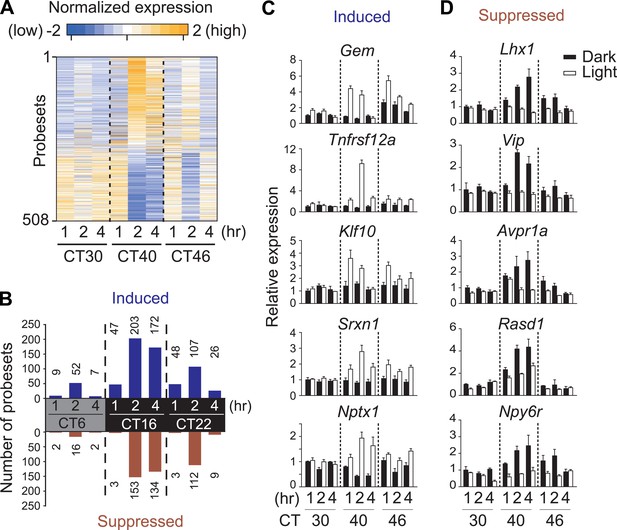
Light-regulated transcripts of the SCN.
(A) Heatmap rendering of light-regulated SCN transcripts. For each time point, fold change between respective light treated and dark control was plotted. (B) Circadian gating of light-modulated transcripts. Cutoffs of two fold were set for up-regulation (blue) or suppression (red) after light pulse, and the number of probesets that satisfy each cutoff was plotted for each point. Quantitative RT-PCR (qRT-PCR) expression confirmation of genes detected as light-regulated by microarray. Examples of genes (C) induced or (D) repressed by light pulses at three different time points. (Mean +s.e.m., n = 4).
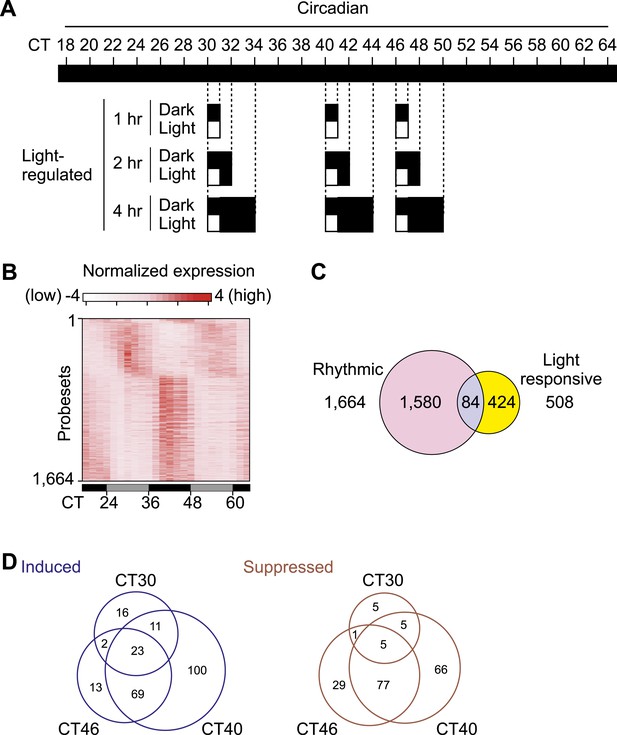
Transcriptional profiling of the mouse SCN.
(A) Sampling schedule for the collection of SCN. C57BL/6J male mice were entrained to 12 hr light:12 hr darkness for 2–3 weeks and transferred to constant darkness. From Circadian Time (CT) 18, 30 hr after lights off, four mice at each time point were collected every 2 hr in dark over two complete days till CT64. From CT30, CT40, or CT46, one group of mice was exposed to 1 hr light, while the control group was maintained in dark, then both groups stayed in dark after 1 hr. SCNs were collected 1, 2, or 4 hr after the beginning of 1 hr light pulse. (B) Heatmap rendering of circadianly expressed transcripts in the mouse SCN. Each horizontal line represents one probeset from MOE430 high density array. (C) Venn diagram for the overlap of light-regulated and cycling transcripts in the SCN. Numbers shown are for probesets. (D) Venn diagram of light induced and suppressed transcripts showing that the light pulse at CT16 that causes maximal phase shift also affects the expression of a large number of SCN transcripts.
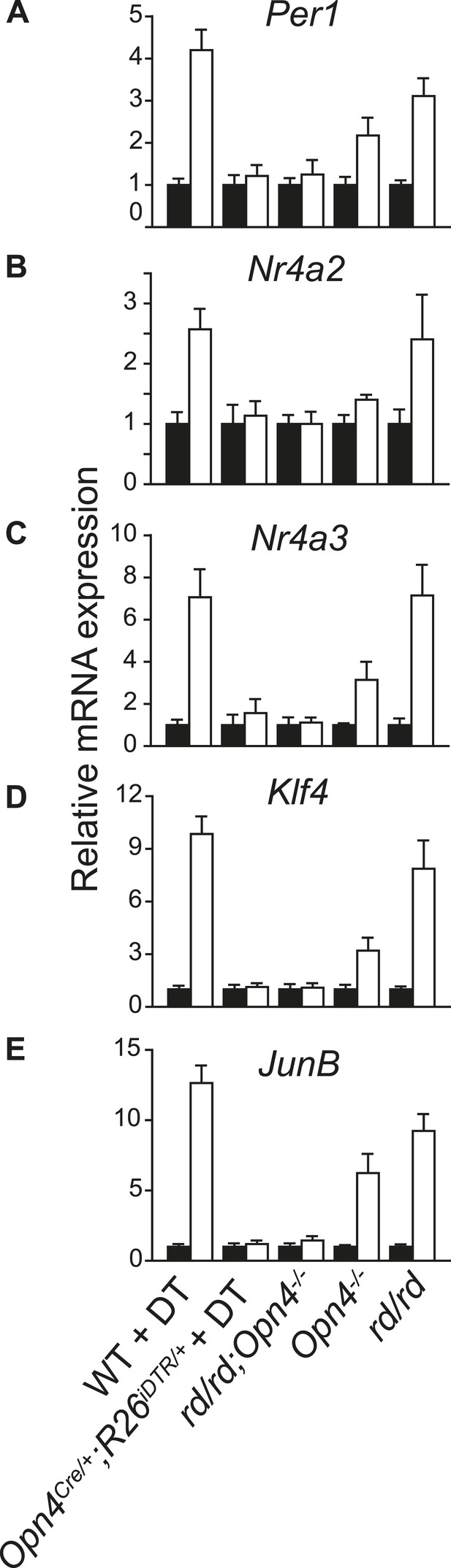
Light-induced changes in SCN gene expression correlate with the known effect of light on phase shift in different genetic models of light signaling.
qRT-PCR quantification of (A) Per1, (B) Nr4a2, (C) Nr4a3, (D) Klf4, and (E) JunB mRNA in the SCN of dark reared or 2 hr after a 1-hr light pulse delivered at CT16 are shown. (Mean + s.e.m., n = 4). The adult rd mice show outer retina degeneration, yet light resets their circadian clock as effectively as of the WT mice (Foster et al., 1991). Opn4−/− mice lack melanopsin and their circadian clock shows an attenuated light-induced phase shift (Panda et al., 2002; Ruby et al., 2002). Opn4−/−;rd mice lack rod, cone, and melanopsin photopigments and show no response to light (Panda et al. 2003). Opn4Cre/+;R26iDTR/+ mice treated with DT specifically lose melanopsin-expressing retinal ganglion cells and show no phase shifting effect of light (Hatori et al., 2008). All mice were dark reared for at least 7 days and their activity onset was used to calculate CT16.
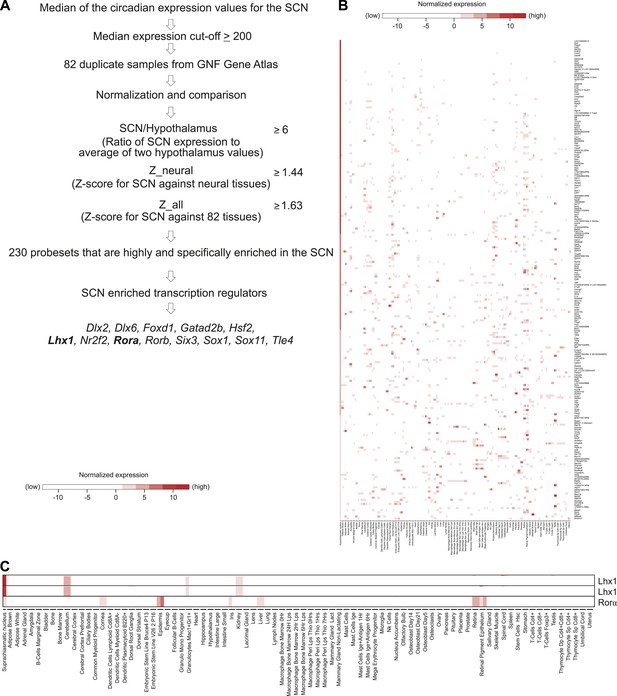
SCN enriched (not SCN-exclusive) transcripts.
(A) Criteria to find SCN-enriched genes among 83 mouse tissues revealed 230 probesets among which 13 were transcription factors. (B) Expression patterns of SCN-enriched transcripts in 83 mouse tissues. Except SCN, duplicate data sets were used for other 82 tissues. The value used for the SCN was the (normalized) median of all the circadian values (24 in total) for the given probeset. Affymetrix probeset IDs and raw data for each gene are shown in Supplementary file 3. (C) The SCN is the only tissue showing overlapping expression of Lhx1 and Rorα. Lhx1 (Affymetrix IDs 1421951_at and 1450428_at) and Rorα (1436325_at) in Figure 1—figure supplement 3C were extracted from Figure 1—figure supplement 3B.
Loss of Lhx1 expression in the SCN renders faster synchronization with change in LD regimes.
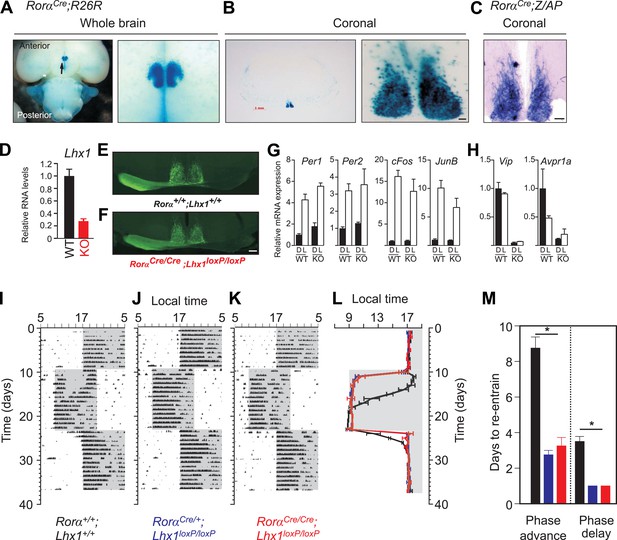
Enriched expression of a Rorα-driven marker in the SCN in RorαCre;R26R mice. (A) Ventral view of a whole brain (magnified view on the right) of adult RorαCre;R26R shows LacZ staining of the SCN. (B) Coronal section through the mid-SCN region (scale bar, 1 mm) and the magnified view of the SCN (scale bar, 100 µm) showing LacZ expression or (C) alkaline phosphatase expression in RorαCre;Z/AP mice. (D) qRT-PCR estimate of Lhx1 expression in the SCN (mean +s.e.m, n = 5). (E) Normal SCN innervation of the retinal ganglion cells in the WT mice as revealed by monocular injection of CTB-conjugated fluorescent marker is intact in (F) Lhx1SCN−KO mice. A 1 hr light pulse at CT16 causes (G) upregulation of light-induced genes (Per1, Per2, cFos, JunB), while (H) the light-suppressed transcripts (Lhx1, Vip, Avpr1a) in the WT SCN show reduced expression in the Lhx1SCN-KO mice. Mice were in DD for 2 days before the light pulse. Representative actograms of (I) RorαCre/Cre, (J) RorαCre/+;Lhx1loxP/loxP, and (K) RorαCre/Cre;Lhx1loxP/loxP mice subjected to 8 hr phase advance and 8 hr delay. (I) Average (+s.e.m., n = 5–8) activity onset and (K) average (+s.e.m.) number of days to re-entrain to advance or delay in light onset in three genotypes. Color codes in L and M correspond to the labels in I–K.
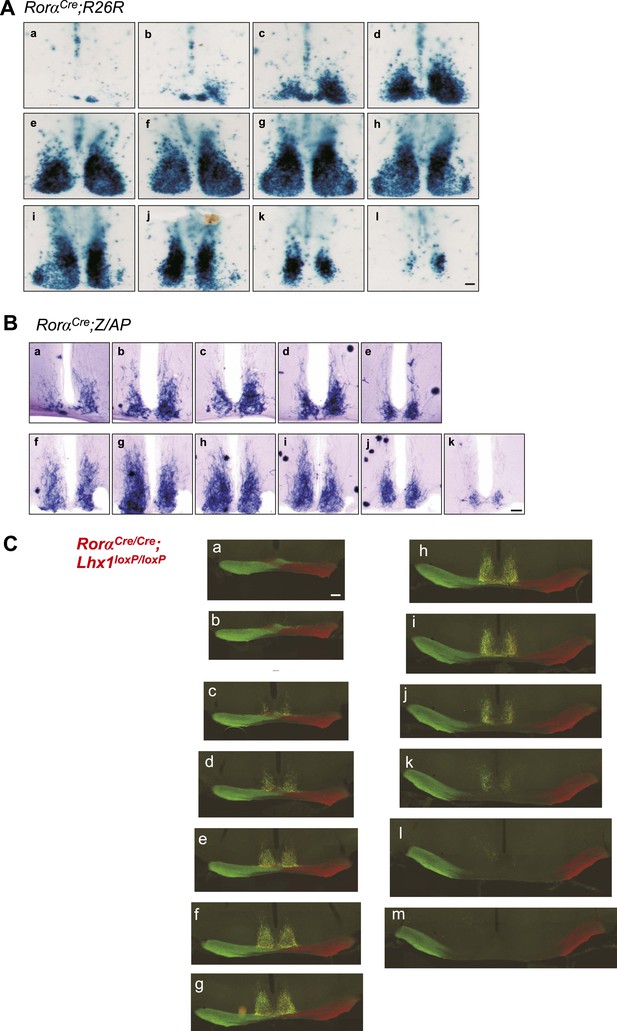
Histology of the adult SCN.
Serial coronal brain sections of adult. (A) RorαCre;R26R or (B) RorαCre;Z/AP mice showing LacZ or alkaline phosphatase staining in the SCN. Scale bar, 100 µm. (C) Serial coronal hypothalamic brain section of an adult RoraCre/Cre;Lhx1fl/fl mouse intra-ocularly injected with Cholera toxin B (CTB) conjugated Alexa Fluor 488 (green) or 594 (Red) showing normal innervation of the SCN.
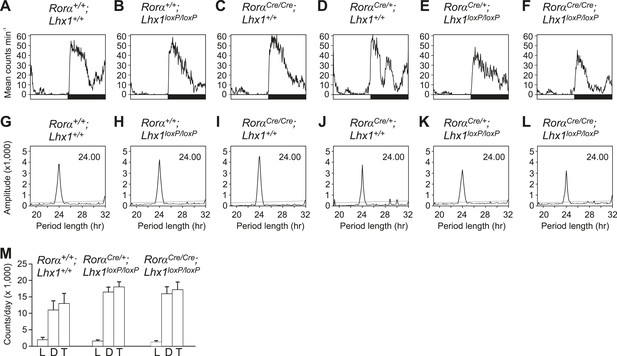
Activity profile under light-dark condition.
(A–F) Activity profiles and (G–L) Chi-squared periodograms of representative mice of indicated genotypes during LD cycles. The period length (H) is shown inside panels of (G–L). Respective actograms showing wheel-running activity during LD are shown in Figure 3A–F. (M) Quantitation of the amounts of RorαCre;Lhx1loxP wheel running activity. Activity counts in LD cycles (L; light, D; dark and T; total) were plotted. Error bars indicate standard error of the mean. Activity during light, activity during dark and the total daily activity among these three genotypes were not significantly different.
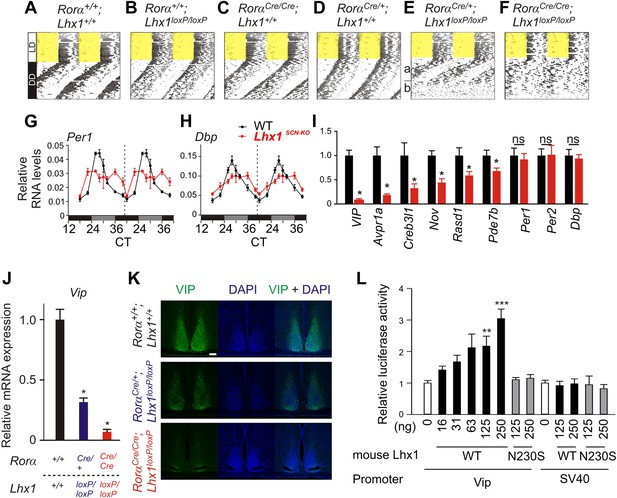
Lhx1 sustains normal circadian activity rhythms by regulating expression of synchronizing factors.
(A–F) Representative wheel running activity pattern over several days of LD followed by DD in wild type and mice lacking Lhx1 in the SCN. Double-plotted qRT-PCR quantification (average +s.e.m, n = 3–4 mice) of (G) Per1 and (H) Dbp in the SCN of DD adapted WT and Lhx1SCN−KO mice. (I) Average (+s.e.m. 8 time points every 3 hr over 24 hr) expression of several factors involved in intercellular communication or circadian clock in the SCN of dark adapted WT and Lhx1SCN−KO mice. (J) Average mRNA (+s.e.m., n = 3–6 mice, *p < 0.05) expression or (K) immunoreactivity of VIP is reduced in the SCN of Lhx1-deficient mice. (L) Transcriptional activation of mouse Vip promoter by mouse LHX1. pGL3-promoter vector was used as a control promoter vector. Values are mean +s.e.m, ANOVA **p<0.01, ***p<0.001 vs 0 ng (white bar).
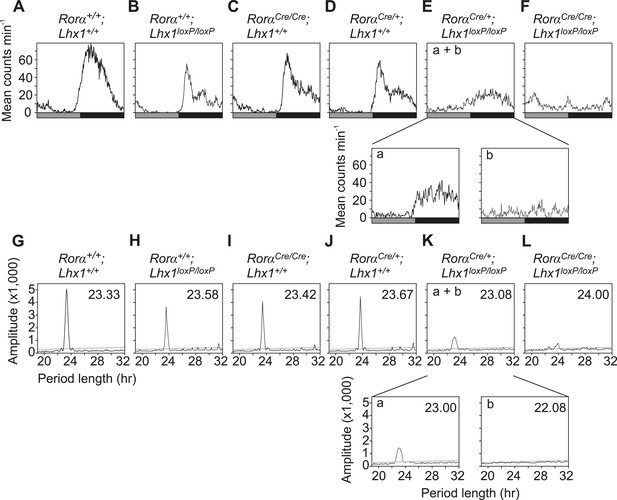
Activity profile under constant darkness.
(A–F) Activity profiles and (G–L) Chi-squared periodograms of representative mice of indicated genotypes during DD cycles. The period length (H) is shown inside panels of (G–L). Respective actograms showing wheel-running activity during DD are shown in Figure 3A–F. The insets in Ea and Eb show activity profile during the first 2 weeks in DD and last week of DD when the mice were rhythmic and arrhythmic respectively. Similarly, the insets in Ka and Kb show the respective chi-square periodogram. Average period lengths are shown in Table 1.
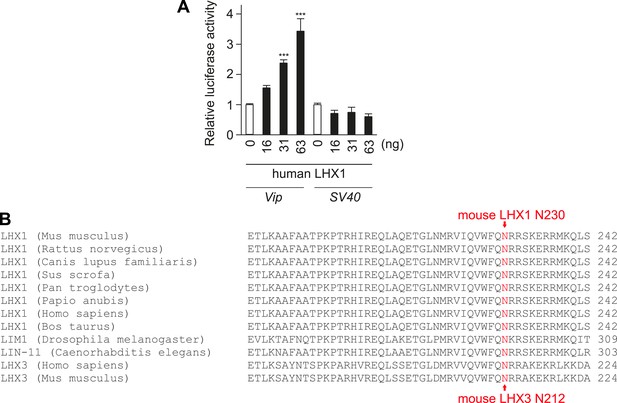
Lhx1 activates Vip transcription.
(A) Human Lhx1 activates Luciferase expression from Vip:Luc but not from SV40 promoter in a dose-dependent manner. (B) Amino acid sequence alignment of DNA binding region of LHX1 and LHX3 showing the N230 residue in mouse Lhx1 that is critical for normal transcriptional activation function of Lhx1.
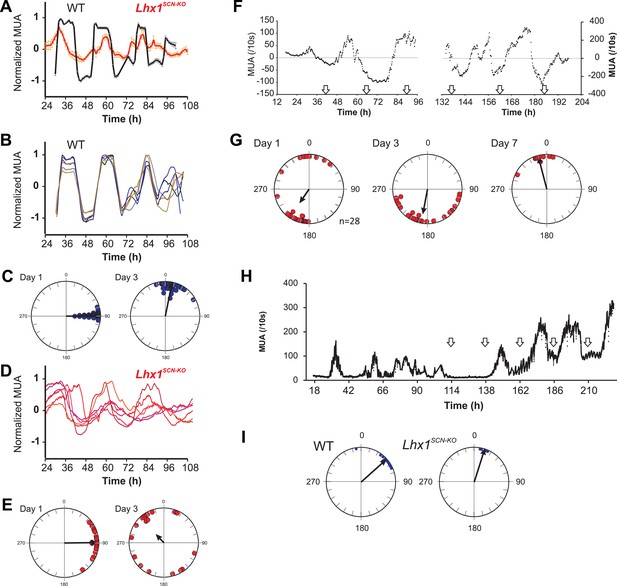
Lhx1 maintains synchrony among SCN neurons partly via VIP.
(A) Average (+s.e.m.) normalized multiunit activity (MUA) recorded from representative SCN slices of LD-adapted WT (n = 40, black) and Lhx1SCN−KO (n = 12, orange) mice. Data were binned every 60 min. Representative normalized MUAs and peak phase of activity from WT SCN (B and C, n = 40) and Lhx1SCN−KO mouse (D and E, n = 19). For C and E, left and right panels are respectively for days 1 and 3. (F) Average MUA of a DD adapted Lhx1SCN−KO SCN that received 1 hr perfusion of VIP daily for up to 7 days. (G) Peak phases of activity are gradually synchronized over 7 days. (H) Representative MUA from the SCN of an LD-adapted Lhx1SCN−KO mouse over several days. During the first 4 days, the activity dampened, which was rescued by daily application of VIP. Down arrows in F and H indicate the time of VIP application. (I) Peak phases of activity in WT and Lhx1SCN−KO SCN at the end of 7 days are shown.
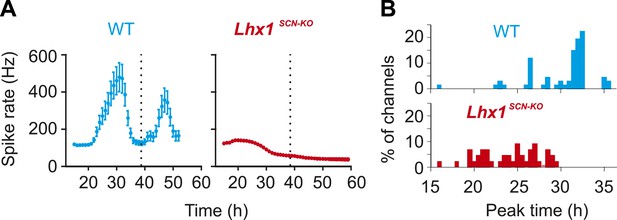
Normalized multi-unit activity recorded from DD adapted WT and Lhx1SCN-KO (mean ± SEM).
Peak time of multiunit activity from each channel shows relative synchrony in the WT mouse that is dispersed in the Lhx1SCN-KO mouse.
Tables
Circadian running wheel activity period length of various mouse strains under constant darkness (n = 7–22)
| Average (h) | SEM (h) | |
|---|---|---|
| Rora+/+;Lhx1+/+ | 23.78 | 0.23 |
| Rora+/+;Lhx1loxP/loxP | 23.64 | 0.05 |
| RoraCre/Cre;Lhx1+/+ | 23.40 | 0.14 |
| RoraCre/+;Lhx1+/+ | 23.86 | 0.07 |
| RoraCre/+;Lhx1loxP/loxP | 23.51 | 0.13 |
| RoraCre/Cre;Lhx1loxP/loxP | NA | NA |
-
Mice showing arrhythmic activity were excluded from the analysis. (NA = Not Applicable).
Additional files
-
Supplementary file 1
Probesets and corresponding genes showing circadian expression of mRNA abundance in the mouse adult SCN.
- https://doi.org/10.7554/eLife.03357.016
-
Supplementary file 2
(A) Transcripts showing >2 fold expression changes after 1 hr light pulse from CT6, CT16, and CT22. The dark controls for this data set were from Supplementary file 1. Therefore, for consistency in naming, the CT6, CT16, and CT22 samples are labeled as CT30, CT40, and CT 46. (B) Transcripts showing both light regulation and rhythmic expression. (C) Fold changes (ratio of light exposed SCN value to dark control SCN value) for light-modulated transcripts.
- https://doi.org/10.7554/eLife.03357.017
-
Supplementary file 3
SCN-enriched transcripts and their expression in 83 mouse tissues.
- https://doi.org/10.7554/eLife.03357.018





