A molecular mechanism of mitotic centrosome assembly in Drosophila
Figures
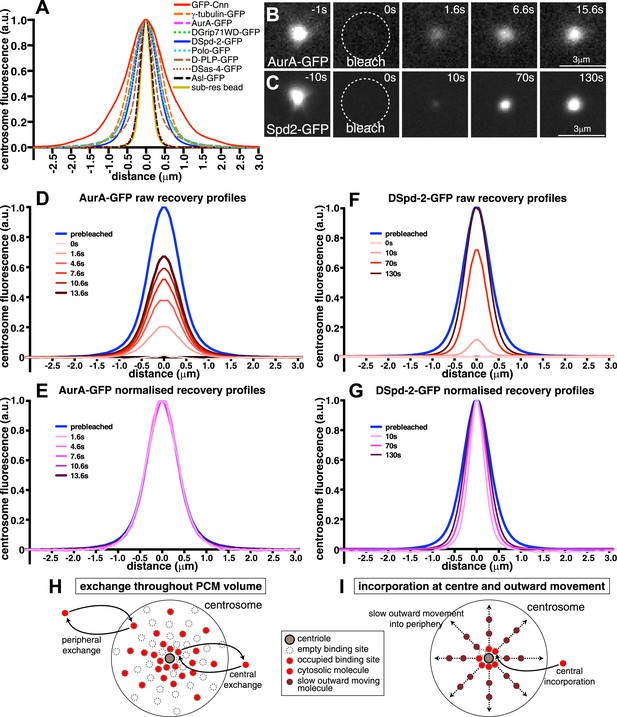
Centrosomal DSpd-2 displays an unusual dynamic behaviour.
(A) Graph shows the centrosomal fluorescence intensity profiles of various centrosomal proteins, along with the profile of 170 nm sub-resolution beads. (B and C) Images show the FRAP behaviour of AurA-GFP (B) and DSpd-2-GFP (C); time before and after photobleaching (t = 0) is indicated. Note how AurA-GFP fluorescence appears to recover evenly throughout the region it originally occupied, whereas DSpd-2-GFP fluorescence appears to initially recover only in the centre of the PCM and then spread outward. (D–G) Quantification of the recovery dynamics of AurA-GFP (D and E) and DSpd-2-GFP (F and G). Graphs show the average fluorescence intensity profile of at least 10 centrosomes at the selected time-points after photobleaching: (D) and (F) show the pre-bleached profiles (blue lines) and successive ‘raw’ recovery profiles (various shades of red), whereas (E) and (G) show the pre-bleached profiles and successive normalized recovery profiles (various shades of pink/purple—normalized so that their peak intensity is equal to the peak intensity of the pre-bleached profile). The normalized recovery curves of AurA-GFP are essentially identical to the pre-bleached profile at all time-points (E), while for DSpd-2-GFP they are initially narrower and spread outward over time (G). (H and I) Schematics illustrate the dynamic behaviour of AurA-GFP (and most other PCM components) (H) and DSpd-2-GFP (and GFP-Cnn) (I). Cytoplasmic AurA-GFP molecules exchange with binding sites spread throughout the PCM (H), whereas DSpd-2-GFP molecules are recruited by binding sites located close to the centrioles; once released from these binding sites the molecules spread slowly outward into the more peripheral regions of the PCM (I). See also Figure 1—figure supplements 1 and 2, and Video 1.
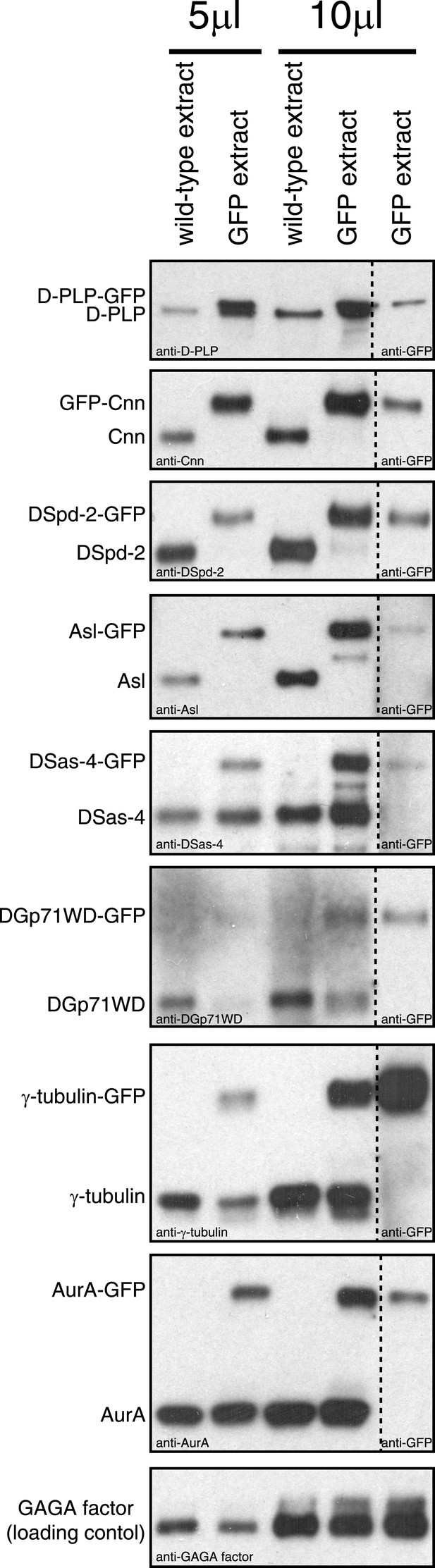
An analysis of the expression levels of several GFP-tagged centrosomal proteins.
Western blots compare the levels of various centrosomal proteins (as indicated) from wild-type embryo extracts (lanes 1 and 3) or from extracts of embryos expressing a GFP-fused version of the protein (lanes 2, 4, and 5). Lanes 1 to 4 were probed with antibodies raised against the centrosomal protein in question; lane 5 was probed with anti-GFP antibodies. Lanes 1 and 2 were loaded with 5 μl of extract; lanes 3 to 5 were loaded with 10 μl of extract. The bottom panel shows a Western blot for GAGA factor, which acts as a loading control (all blots were controlled in this way, but only one example is shown here). GFP-Cnn, DSpd-2-GFP, and Asl-GFP were expressed in a null mutant background; the other GFP-fusions were expressed in a wild-type background. Note how GFP-Cnn and D-PLP-GFP appear to be expressed at slightly higher levels than the endogenous protein, but that the other fusion proteins are expressed either at similar or at slightly lower levels than the endogenous protein. We previously showed that increasing the cytoplasmic concentration of GFP-Cnn increases the rate at which it is incorporated into the PCM, but does not change the inside out incorporation behaviour (Conduit et al., 2010). Note also that GFP-Cnn and DSpd-2-GFP are overexpressed and underexpressed, respectively, but both display a similar dynamic behaviour (Figure 1F,G, Figure 1—figure supplement 2M,N), again suggesting that the expression level of a GFP-fusion protein does not significantly affect the mode in which it is incorporated into the PCM. We have not been able to analyze the relative expression level of Polo-GFP, as we were unable to get anti-Polo antibodies to work on Western blots.
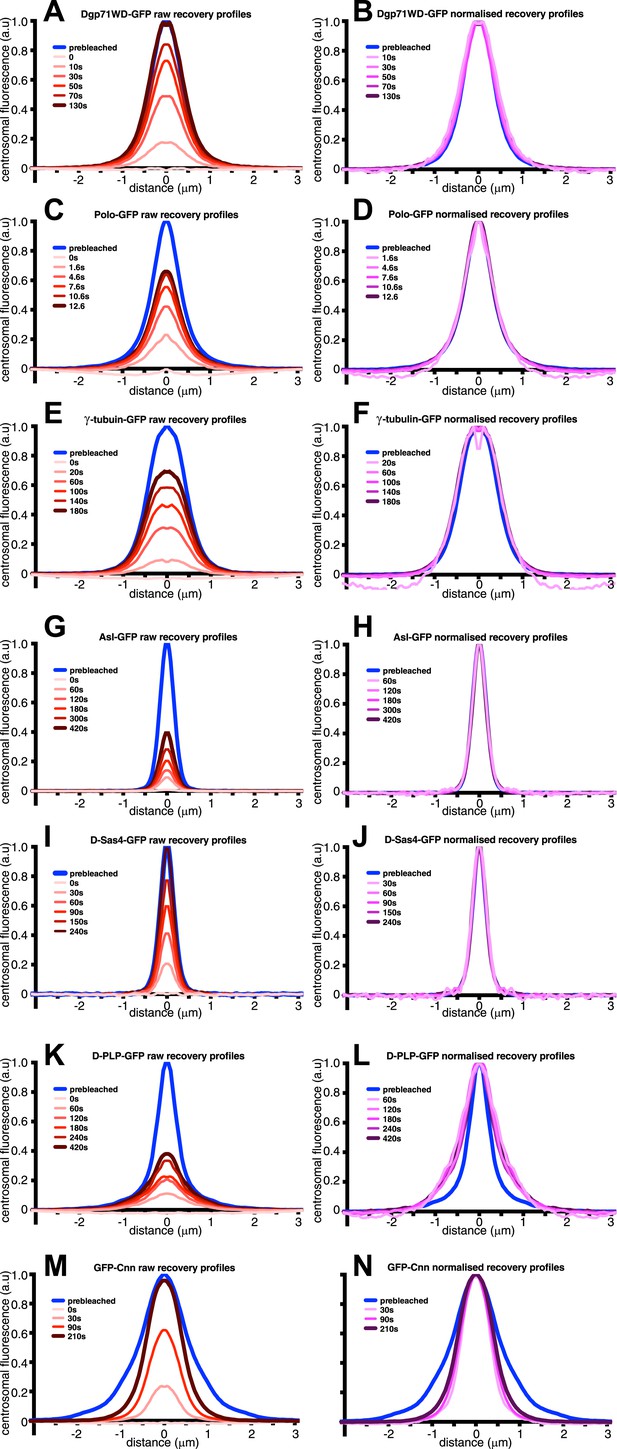
An analysis of the dynamic behaviour of several GFP-tagged centrosomal proteins.
Quantification of the FRAP recovery dynamics of DGp71WD-GFP (A and B), Polo-GFP (C and D), γ-tubulin-GFP (E and F), Asl-GFP (G and H), DSas-4-GFP (I and J), D-PLP-GFP (K and L), and GFP-Cnn (M and N). Graphs show the average fluorescence intensity profile of ≥10 centrosomes (‘Materials and methods’) at selected time-points after photobleaching (t = 0). Graphs on the left of each panel display pre-bleached profiles (blue lines) and ‘raw’ recovery profiles (various shades of red lines). Graphs on the right of each panel display pre-bleached profiles and recovery profiles (various shades of purple lines) that have all been normalized so that their peak intensity is equal to the peak intensity of the pre-bleach profile. Note how the normalized recovery profiles of DGp71WD-GFP (B), Polo-GFP (D), and γ-tubulin-GFP (F) are very similar to their pre-bleached profiles, indicating that they are all incorporated evenly throughout the PCM domain they originally occupied. This is also true for Asl-GFP (H) and DSas-4-GFP (J), but these proteins have the same distribution as sub-resolution beads (Figure 1A), so their true distribution cannot be resolved on this microscope system. The recovery dynamics of D-PLP-GFP (K and L) are complicated by the presence of a slow-exchanging centriole fraction and a fast-exchanging PCM fraction (Martinez-Campos et al., 2004), which cannot be properly distinguished at this resolution. It appears that the slow recovery rate of the centriole fraction compared to the PCM fraction causes the normalized recovery profiles to be wider than the pre-bleached profile. It can be seen, however, that the PCM fraction of D-PLP shows no sign of outward movement through the PCM (L). The normalized recovery profiles of GFP-Cnn (N) are narrower than the pre-bleached profile and spread outwards over time, reflecting the fact that Cnn molecules are initially incorporated only in the centre of the PCM and then move slowly outward (Conduit et al., 2014).
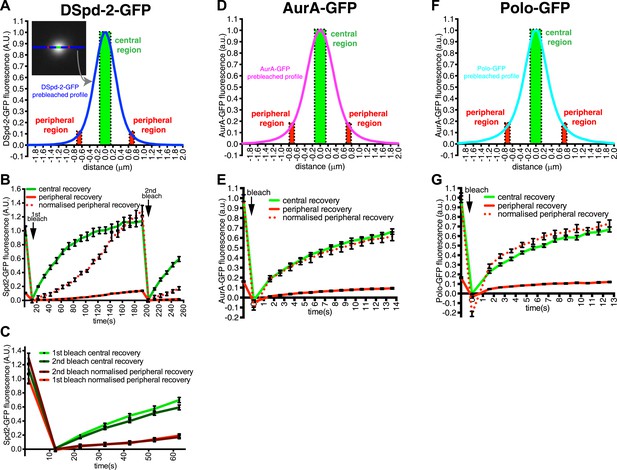
DSpd-2-GFP molecules spread away from the centrioles.
(A) Graph displays the pre-bleached profile of DSpd-2-GFP at centrosomes in Drosophila embryos (blue line, average of 10 centrosomes). Boxes highlight the central region of the PCM (green box) and peripheral regions of the PCM (red boxes) that were analyzed in a FRAP experiment. (B) Graph displays the average fluorescence intensity through time in the centre (green line) and periphery (red and dotted red lines) of the PCM after photobleaching. Arrows indicate times of photobleaching. The dotted red line represents the peripheral recovery after it has been normalized so that its initial pre-bleached value is equal to the initial pre-bleached value of the central recovery curve. Note how the central DSpd-2-GFP fluorescence recovery exhibits a typical logarithmic-shaped FRAP curve, with a fast initial rate that slows over time. In contrast, peripheral DSpd-2-GFP fluorescence recovery exhibits a very unusual behaviour as it is initially slow and then speeds up over time. This strongly suggests that DSpd-2-GFP molecules move from the centre to the periphery of the PCM (see main text). (C) Graph compares the initial recovery kinetics in the central (green lines) or peripheral (red lines) regions of the PCM after a first (light lines) and then second (dark lines) photobleaching event. The second photobleaching event took place when the rate of recovery in the centre was slow and the rate of recovery in the periphery was fast (see t = 200 s in B). Note that after the second bleaching event the central recovery returned to its original fast rate and the peripheral recovery returned to its original slow rate, showing that the increasing rate of recovery in the periphery was not due to peripheral binding sites gradually exchanging faster over time. (D–G) Graphs display the pre-bleached profiles (D and F) and recovery kinetics (E and G) of AurA-GFP (D and E) and Polo-GFP (F and G) at centrosomes in embryos (average of 11 centrosomes) in the same format as shown for DSpd-2-GFP (A and B). Note how the peripheral recovery curves of AurA-GFP (red and dotted red lines in E) and Polo-GFP (red and dotted red lines in G) both have a similar shape to their central recovery curves, indicating that the unusual peripheral kinetics of DSpd-2-GFP (red and dotted red lines in B) are not observed with other proteins that have a similar distribution around the centrioles. Error bars = SEM.
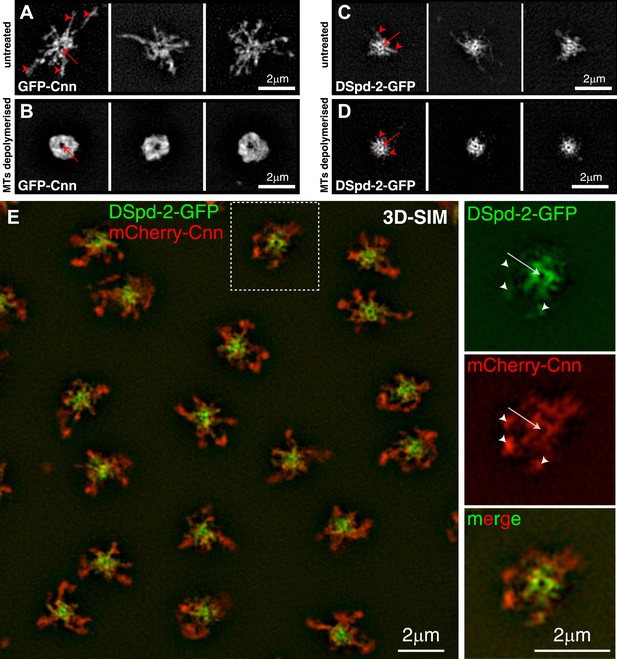
DSpd-2-GFP appears to form scaffold-like structure around the centrioles that partially co-localizes with the Cnn scaffold.
(A–D) 3D-SIM images of centrosomes from embryos expressing either GFP-Cnn (A and B) or DSpd-2-GFP (C and D) where the MTs are either present (A and C) or have been depolymerized by colchicine injection (B and D). (A) In untreated embryos, large projections of GFP-Cnn extend outwards (red arrowheads) from a central hollow (red arrow), which presumably contains the mother centriole. (B) After MT depolymerisation, the GFP-Cnn scaffold collapses into a largely amorphous structure; presumably, the molecular detail of the scaffold cannot be resolved even at this high resolution. The slightly larger central ‘hollow’ in the GFP-Cnn signal (red arrow) likely reflects the ability of the Cnn molecules to move a short distance away from the centre of the centrosome in the absence of microtubules (Conduit et al., 2014); these molecules then get ‘trapped’ in the more peripheral regions of the PCM, as they cannot efficiently leave the centrosome in the absence of MTs (Conduit et al., 2014). (C) In untreated embryos, DSpd-2-GFP appears as a series of spoke-like projections (red arrowheads) that extend away from a central ring (red arrow), which presumably surrounds the mother centriole; some of these projections weakly extend into the peripheral PCM. (D) After MT de-polymerisation, DSpd-2-GFP retains a large degree of its structure: there is a clear central ring (red arrow) with several spoke-like projections extending outwards (red arrowheads); these projections, however, no longer appear to extend into the more peripheral regions of the PCM. (E) Two-colour 3D-SIM images of centrosomes in untreated embryos co-expressing DSpd-2-GFP (green) and mCherry-Cnn (red). The panels on the right are enlargements of the boxed centrosome in the panel on the left; note the clear hollow in the centre of the centrosome (arrows), and how the mCherry-Cnn signal extends further away from the centrioles than the DSpd-2-GFP signal, although weak DSpd-2-GFP fluorescence can often be observed in the same region as the more peripheral mCherry-Cnn (arrowheads). See also Figure 3—figure supplement 1.
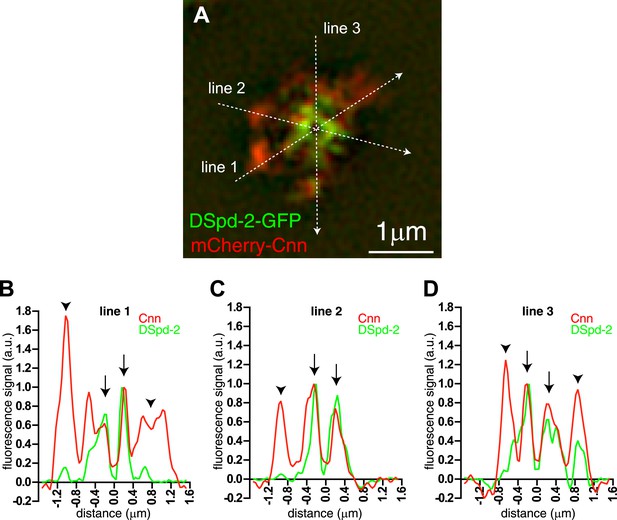
An approximate quantification of mCherry-Cnn and DSpd-2-GFP co-localisation.
The mCherry-Cnn (red) and DSpd-2-GFP (green) fluorescence intensity along the three lines that run through a 3D-SIM image of a centrosome in (A) are plotted on the graphs in (B), (C), and (D). Green lines represent the fluorescent intensity of DSpd-2-GFP, and red lines represent the fluorescent intensity of mCherry-Cnn. The graphs have been normalized so that the peak value in the central region of the PCM is equal to 1. Arrows within the graphs indicate the central regions of the centrosome where mCherry-Cnn and DSpd-2-GFP extensively co-localize. Arrowheads indicate the more peripheral regions of the centrosome that contain high levels of mCherry-Cnn fluorescence but only low levels of DSpd-2-GFP fluorescence.
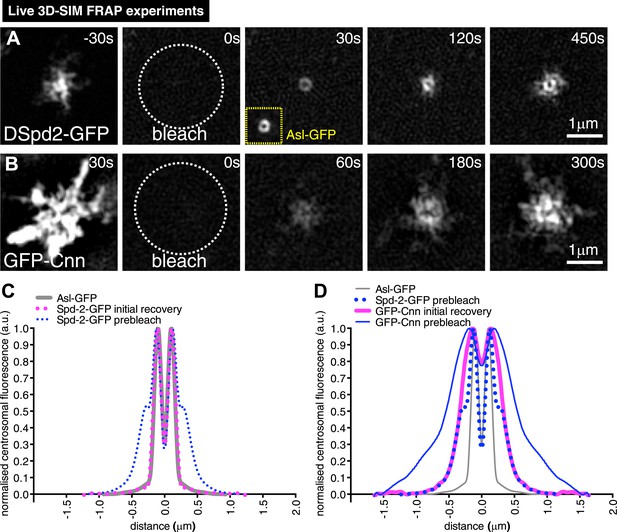
3D-SIM FRAP analysis of DSpd-2-GFP and GFP-Cnn behaviour at centrosomes.
(A and B) 3D-SIM images show the dynamic behaviour of DSpd-2-GFP (A) and GFP-Cnn (B) at centrosomes in living Drosophila embryos after FRAP; time before and after photobleaching (t = 0) is indicated. (A) DSpd-2-GFP fluorescence initially recovers in a toroid shape around the centriole (A, t = 30 s), which has similar dimensions to unbleached Asl-GFP (yellow inset). The protein then moves slowly outwards, forming dynamic projections that spread away from the centriole. (B) GFP-Cnn fluorescence initially recovers in a broader region around the centrioles (t = 60 s in B), which has similar dimensions to the pre-bleached DSpd-2-GFP signal (t = −30 s in A). (C) Graph compares the average prebleached (dotted blue line) and initial recovery (dotted pink line) profiles of DSpd-2-GFP to the average unbleached profile of Asl-GFP (grey line); all profiles were normalized so that their peak value is equal to 1. Note how the initial DSpd-2-GFP recovery profile is essentially identical to the un-bleached Asl-GFP profile. (D) Graph compares the average pre-bleached (solid blue line) and initial recovery (solid pink line) profiles of GFP-Cnn, to the average pre-bleached profile of DSpd-2-GFP (dotted blue line), and the average unbleached profile of Asl-GFP (grey line); all profiles were normalized so that their peak value is equal to 1. Note how the initial GFP-Cnn recovery profile is very similar (particularly in the more peripheral regions), to the pre-bleached DSpd-2-GFP profile, but is quite distinct from the pre-bleached Asl-GFP profile. See also Video 2.
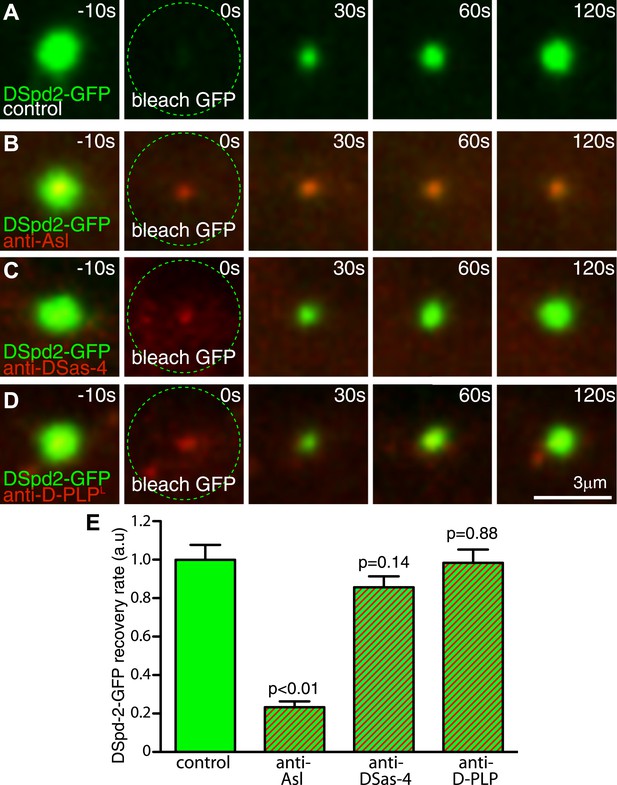
Inhibiting Asl function strongly perturbs DSpd-2 incorporation into the PCM.
Images (A–D) show results from FRAP experiments monitoring how DSpd-2-GFP (green) incorporation into the PCM is affected by inhibiting the function of various centriole-associated components with injected Texas-red-labelled antibodies (red—as indicated). The antibodies bind to their cognate protein at centrosomes close to the injection site (B–D), but not to those far from the injection site (A), which therefore act as internal controls. (E) Graph displaying the initial recovery rate of DSpd-2-GFP fluorescence at centrosomes bound with antibodies (as indicated below the graph) relative to the centrosomes not bound by antibodies (control). The rate of recovery was calculated by measuring the gradient of the initial linear phase of recovery that occurred over the first 60 s after photobleaching. Note how anti-Asl antibodies reduce the rate of DSpd-2-GFP incorporation into the PCM by ∼75%, but that anti-DSas-4 or anti-D-PLP antibodies have little or no effect. Error bars = standard error. See also Video 3.
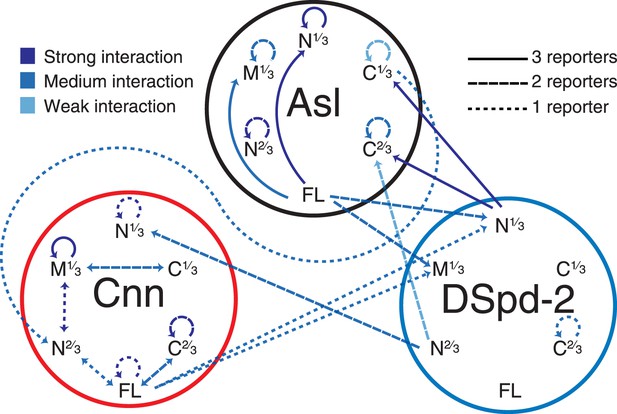
A yeast-two-hybrid analysis examining the interactions between Asl, DSpd-2, and Cnn.
A schematic summary of the yeast two-hybrid interactions observed between Asl, DSpd-2, and Cnn. The shades of the lines indicate the strength of the interactions observed: strong (dark blue), medium (blue), and weak (light blue). The characteristics of the lines indicate the number of different positive reporter assays for each interaction: solid line (3/3 assays), large dashed line (2/3 assays), and small dashed line (1/3 assays). Arrows point from bait to prey fragments—double-headed arrows indicate that the interaction scored positive with either fragment as bait or prey. See Figure 6—source data 1.
-
Figure 6—source data 1
A yeast-2-hybrid analysis testing the interactions between various Asl, DSpd-2, and Cnn fragments.
(Related to Figure 6) A yeast-2-hybrid analysis was carried out using various bait and prey fragments of each protein (as indicated in columns A and B). Three different reporters were tested—His (columns C–F), Ade (columns G and H), and LacZ (column I). Interaction levels are indicated as strong, medium, weak, or none. All combinations of baits and preys were tested, but only those that scored positive in at least one assay are shown.
- https://doi.org/10.7554/eLife.03399.015
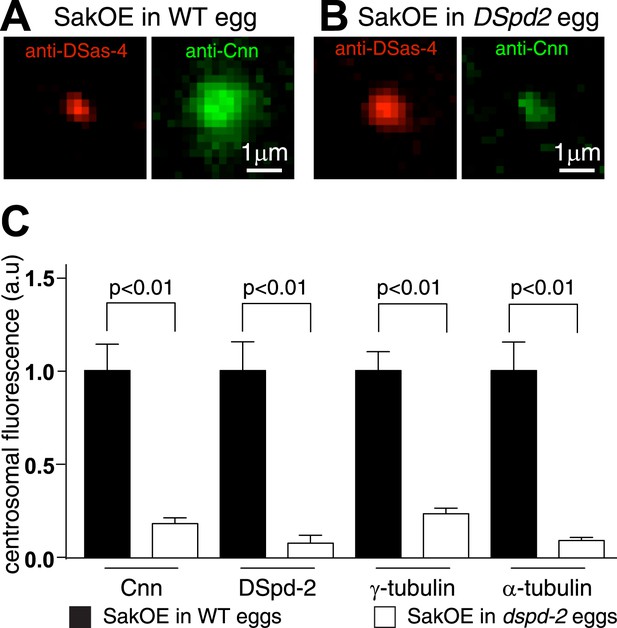
The centrosomal levels of Cnn are strongly reduced in eggs lacking DSpd-2.
(A and B) Images show centrosomes co-stained with DSas-4 antibodies (red) and Cnn antibodies (green) in fixed eggs that either contained (A) or lacked (B) endogenous DSpd-2. As embryos lacking DSpd-2 fail in pronuclear fusion (Dix and Raff, 2007) (and so fail to develop), we induced the de novo formation of centrosomes in WT and DSpd-2 mutant eggs by over-expressing Sak kinase (Peel et al., 2007; Rodrigues-Martins et al., 2007). The centrosomal levels of Cnn are dramatically reduced in the absence of DSpd-2. (C) The graph quantifying the centrosomal levels of Cnn, DSpd-2, γ-tubulin, and α-tubulin in Sak over-expressing eggs that either contained (black bars) or lacked (white bars) endogenous DSpd-2. The centrosomal levels of Cnn, γ-tubulin, and α-tubulin are dramatically reduced in the absence of DSpd-2.
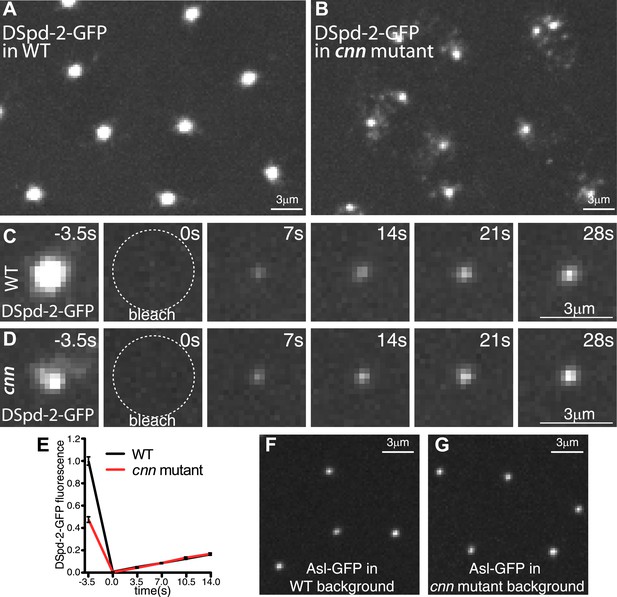

Cnn helps maintain DSpd-2 in the PCM.
(A and B) Images show the localization of DSpd-2-GFP at centrosomes in either WT (A) or cnn mutant (B) embryos. (C and D) Images show the initial dynamic behaviour of DSpd-2-GFP at centrosomes in either WT (C) or cnn mutant (D) embryos; time before and after photobleaching (t = 0 s) is indicated. (E) Quantification of DSpd-2-GFP fluorescence recovery at centrosomes in either WT (black line) or cnn mutant (red line) embryos. The initial rate of DSpd-2-GFP fluorescence recovery is very similar in both WT and cnn mutant embryos, revealing that the initial incorporation of DSpd-2-GFP into the PCM is not dependent on Cnn. (F and G) Images show the localization of Asl-GFP at centrosomes in living embryos in the presence (F) or absence (G) of Cnn. The ability of Asl to localize efficiently in the absence of Cnn presumably explains why DSpd-2 can still be recruited to centrioles at normal rates in the absence of Cnn. Error bars = standard error. See Video 4.
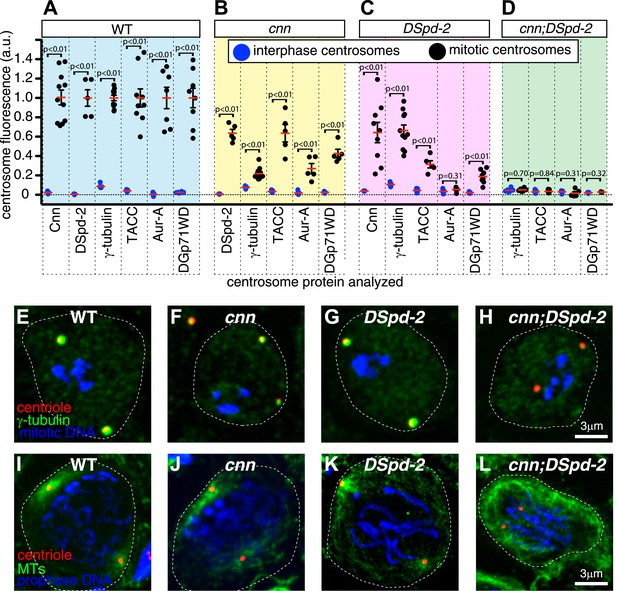
Cnn and DSpd-2 cooperate to recruit the mitotic PCM.
(A–D) Graphs show the average fluorescence intensities of interphase (blue dots) and mitotic (black dots) centrosomes from either WT (A), cnn mutant (B), dspd-2 mutant (C), or cnn;dspd-2 double mutant (D) larval brain cells stained for various centrosomal proteins (as indicated below graphs). Each data-point represents the average centrosome value from one brain. The horizontal red bars indicate the average value of all the brains. All the PCM proteins are still partially recruited to centrosomes in the absence of Cnn or DSpd-2 (with the possible exception of Aurora A, which does not appear to be recruited in the absence of DSpd-2). The mitotic PCM levels do not rise above interphase levels in the absence of both Cnn and DSpd-2, indicating that centrosome maturation has been abolished. (E–L) Images show typical mitotic cells from either WT (E and I), cnn (F and J), dspd-2 (G and K), or cnn;dspd-2 double mutant (H and L) larval brain cells stained for the centriole marker Asl (red), mitotic DNA (phospho-histone H3, blue), and either the PCM marker γ-tubulin (green, E–H) or MTs (I–L, green). Error bars = SEM. See also Figure 9—figure supplements 1 and 2.
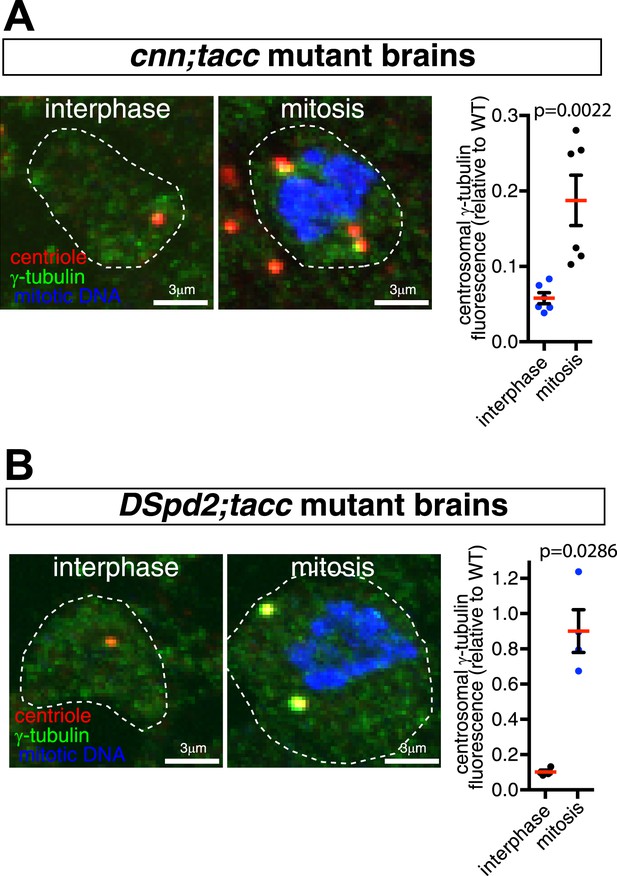
Centrosome maturation is not abolished in cnn;tacc or dspd-2;tacc double mutants.
(A and B) Images and associated graphs show mitotic and interphase larval brain cells stained for γ-tubulin (green), the centriole marker Asl (red), and mitotic DNA (phospho-histone H3, blue) from either cnn;tacc double mutant (A) or dspd-2;tacc double mutant larvae (B). Graphs display the average fluorescent intensities of mitotic and interphase centrosomes (relative to a WT mitotic value of 1) stained for γ-tubulin. Each data-point represents the average centrosome value from one brain. Note how centrosomes still partially mature in each mutant combination, as the levels of γ-tubulin centrosomal fluorescence are significantly higher in mitotic cells than in interphase cells.
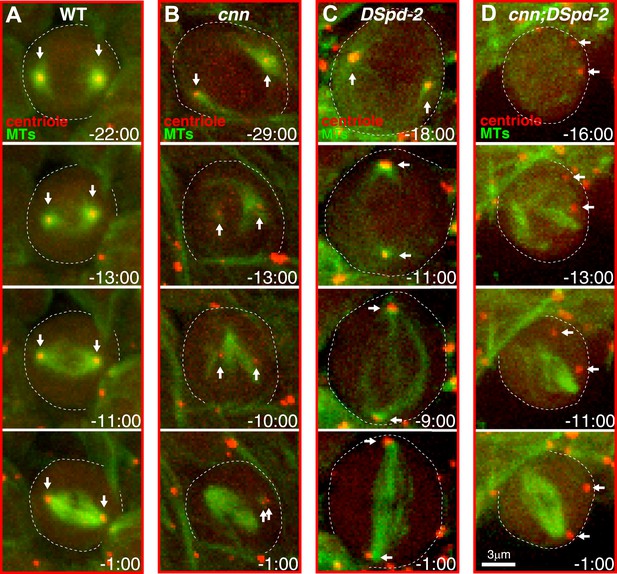
Centrosomes in cnn;dspd-2 double mutants fail to organise MTs.
Images show selected time-points from videos of either WT (A), cnn (B), dspd-2 (C), or cnn;dspd-2 double mutant (D) brain cells expressing the MT marker Jupiter-mCherry (pseudo-coloured green) and the centriole marker GFP-PACT (pseudo-coloured red); time before and after anaphase onset (t = 0 s) is indicated. Arrows indicate the position of the centrioles. The centrioles in WT cells, or cnn or dspd-2 single mutant cells can organize MT asters, but no centriole-associated MTs can be detected in the cnn;dspd-2 double mutants.
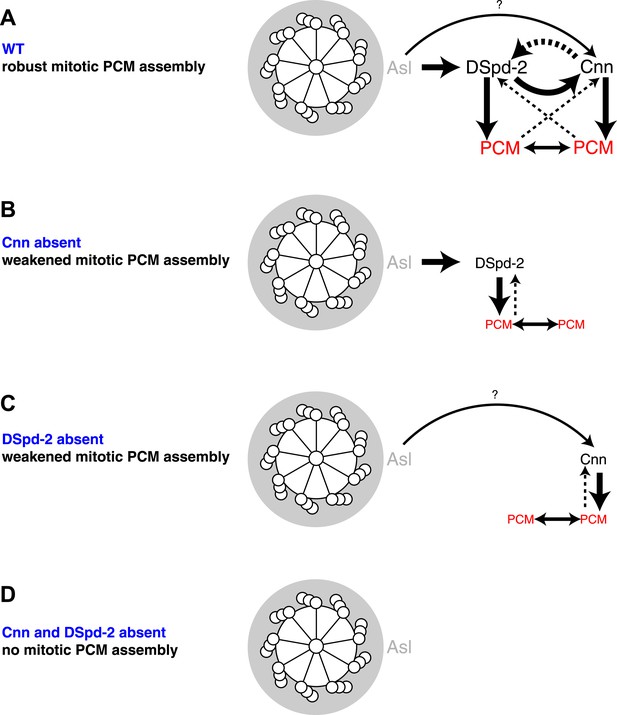
A model for mitotic PCM assembly in flies.
Schematics illustrate a putative pathway of mitotic PCM assembly in a WT cell (A), or in cells lacking either Cnn (B), DSpd-2 (C), or both Cnn and DSpd-2 (D). A top view of the mother centriole is shown surrounded by a layer of Asl (grey); solid arrows represent recruiting interactions, dotted arrows represent maintaining interactions. Arrow thickness reflects the relative strength of the recruitment or maintenance, and the size of the text reflects the amount of protein localized at centrosomes. In WT cells (A), Asl has an important role in recruiting DSpd-2 to centrosomes, which in turn has an important role in recruiting Cnn; Cnn then has an important role in maintaining DSpd-2 at centrosomes. Thus, a positive feedback loop is generated where increasing amounts of DSpd-2 can recruit increasing amounts of Cnn, which can then maintain increasing amounts of DSpd-2. DSpd-2 and Cnn both independently recruit other PCM components (red), which themselves help support the PCM structure and can recruit further PCM components. In the absence of Cnn (B), Asl can still recruit DSpd-2 normally, but DSpd-2 cannot efficiently accumulate around the centrioles. The reduced levels of DSpd-2 recruit reduced levels of PCM. In the absence of DSpd-2 (C), an alternative pathway recruits reduced levels of Cnn. This pathway most likely involves Asl (as indicated here), as inhibiting Asl reduces the rate of Cnn incorporation into the PCM (Conduit et al., 2010) and Asl and Cnn appear to weakly interact in a Y2H analysis (Figure 6); other pathways, however, could also be involved. The reduced levels of Cnn recruit reduced levels of PCM. In the absence of Cnn and DSpd-2 (D), no mitotic PCM can be assembled.
Videos
DSpd-2-GFP molecules are initially incorporated into the centre of the PCM.
(related to Figure 1). All Videos shown here are maximum intensity projections of image stacks. These videos illustrate the dynamic behaviour of AurA-GFP (A–C) or DSpd-2-GFP (D–F) at centrosomes in Drosophila embryos. Time before and after photobleaching (t = 0 s) is shown at the top right of (C) and (F). The graphs in (B) and (E) are the line profiles representing the distribution of the AurA-GFP and the DSpd-2-GFP centrosomal fluorescence, respectively, through time: the blue lines represent the pre-bleached profiles, and the red lines represent the recovering profiles. Note how the AurA-GFP fluorescence signal appears to recover evenly throughout the PCM domain it originally occupied, whereas the DSpd-2-GFP fluorescence signal appears to initially recover in the centre of the PCM and then spreads outwards. This is most clearly seen in (C) and (F) where the recovery profiles at all time-points have been normalized so that their peak intensities are equal to 1 (purple lines). Note how the normalized AurA-GFP recovery profiles all closely overlap with the pre-bleached profile, whereas the normalized DSpd-2-GFP recovery profiles are all narrower than the pre-bleached profile and spread out over time.
Super-resolution 3D-SIM FRAP experiments reveal the differences between how DSpd-2 and Cnn are incorporated into the PCM.
(Related to Figure 4). All Videos shown here are maximum intensity projections of image stacks. Live cell SD-SIM illustrating the dynamic behaviour of DSpd-2-GFP (A) and GFP-Cnn (B) at centrosomes in Drosophila embryos. The employed OMX Blaze 3D-SIM system enables sub-diffraction live cell imaging at high frame rates with ∼two-fold better xy- and z-resolution compared to conventional microscopy. Time before and after photobleaching (t = 0 s) is shown at the top right of each panel. Note how, prior to photobleaching, GFP-Cnn has a broader distribution that DSpd-2-GFP within the PCM. Immediately after photobleaching, DSpd-2-GFP fluorescence recovers in the shape of a toroid around the centriole, supporting our conclusion that Asl, which is distributed as a toroid around the centriole, is the major recruiter of DSpd-2 to centrosomes. In contrast, GFP-Cnn fluorescence recovers in a broader region around the centrioles, supporting our conclusion that DSpd-2 is the major recruiter of Cnn to centrosomes.
The rate of DSpd-2-GFP incorporation into the PCM is reduced in embryos injected with anti-Asl antibodies.
(related to Figure 5). All Videos shown here are maximum intensity projections of image stacks. These videos show the fluorescence recovery of DSpd-2-GFP (green) at centrosomes in embryos that were injected with fluorescently labelled antibodies (red) against Asl (A, B), DSas-4 (C), or D-PLP (D). Time before and after photobleaching (t = 0 s) is shown at the top right of (D). An example of a centrosome in an embryo injected with anti-Asl antibodies that was located far from the injection site and so received a low concentration of the antibody is shown in (A); these centrosomes acted as internal controls. Panels (B–D) show examples of centrosomes located close to the injection sites of antibodies against Asl (B), DSas-4 (C), and D-PLP (D); these centrosomes received a high concentration of each antibody. Note that DSpd-2GFP fluorescence recovered at a much slower rate at centrosomes that bound anti-Asl, whereas DSpd-2GFP fluorescence recovered at near normal rates at centrosomes that bound anti-DSas-4 or anti-D-PLP antibodies.
The centrosomal localization of DSpd-2-GFP is perturbed in the absence of Cnn.
(related to Figure 8). All Videos shown here are maximum intensity projections of image stacks. These videos illustrate the dynamic behaviour of DSpd-2-GFP at centrosomes in embryos where Cnn is present (A) or where Cnn is absent (B). Note how in the absence of Cnn, DSpd-2-GFP cannot properly spread out through the PCM and a haze of DSpd-2-GFP fluorescence, including small particles of DSpd-2-GFP, appears to be rapidly lost from the centrosomes.
Centrosomal MTOC activity is only abolished in cells lacking both Cnn and DSpd-2.
(Related to Figure 9). All Videos shown here are maximum intensity projections of image stacks. These videos show the distribution of the centriole marker GFP-PACT (pseudo-coloured red) and the MT marker Jupiter-mCherry (pseudo-coloured green) in either WT (A), cnn (B), dspd-2 (C), or cnn;dspd-2 mutant neuroblasts as the cells progress through mitosis. Time before and after anaphase onset (t = 0 s) is shown at the top right of each panel. Note how the centrosomal MT asters can be observed in WT (A), cnn (B), and dspd-2 (C) mutant neuroblasts, and these MT asters appear to contribute to spindle assembly. No centrosomal MT asters, however, can be observed in cnn;dspd-2 double mutant neuroblasts and the spindle appears to form independently of centrosomes (D).





