Overcoming myelosuppression due to synthetic lethal toxicity for FLT3-targeted acute myeloid leukemia therapy
Figures
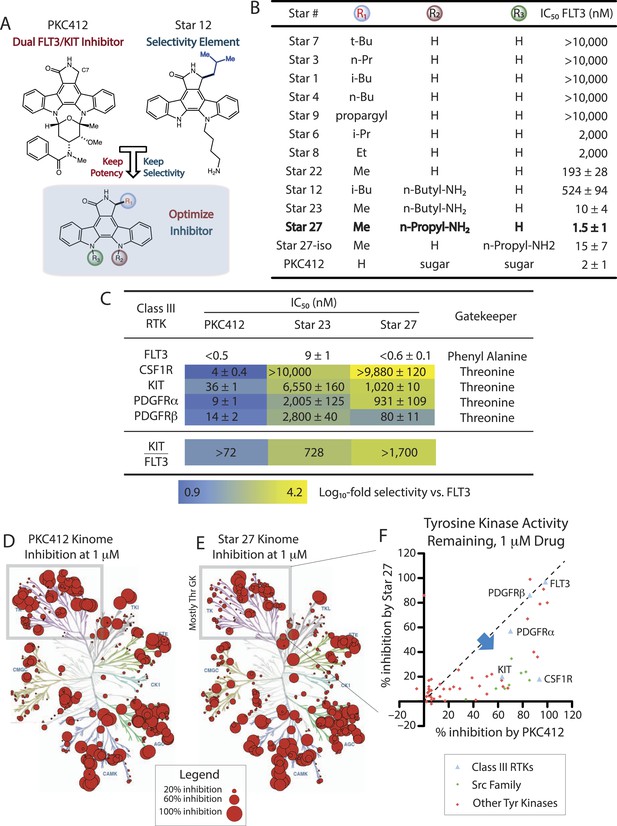
Purified kinase assays comparing SAR of staralogs.
(A) PKC412 (N-benzoylstaurosporine) potently inhibits all Class III/PDGFR family hematopoietic stem cell kinases. Staurosporine analog (staralog) Star 12, with large alkyl C7 (R1) substituent is a weak inhibitor of FLT3 but suggested more selectivity for Class III RTKs. (B) Table shows purified kinase in vitro assays testing SAR showing optimal potency at R1 = methyl; rigid potency window found with Star 27 with n-propyl-NH2 at R2 but not R3. Star ## ordered by increasing potency and numbering consistent with prior usage (Lopez et al., 2013). Error ranges represent standard error of the mean (SEM) and are the result of at least three independent measurements, each in triplicate. (C) IC50 values of PKC412 and lead compounds Star 23 and 27 against the hematopoietically relevant five Class III RTKs showing the optimal selectivity achieved for Star 27. Log10-scale heat map highlights IC50 ratios relative to FLT3, indicating progression from PKC412 having lower selectivity towards Star 27 having high selectivity. (D) Dendrogram showing single point inhibition for 319 kinases for PKC412. Each value represents the average of two experiments ± SEM (performed by RBC). (E) Dendrogram showing single point inhibition for 319 kinases for Star 27. (F) Graph of Tyr kinases. y-axis = potency by single point for Star 27 as a function of the corresponding potency with PKC412.
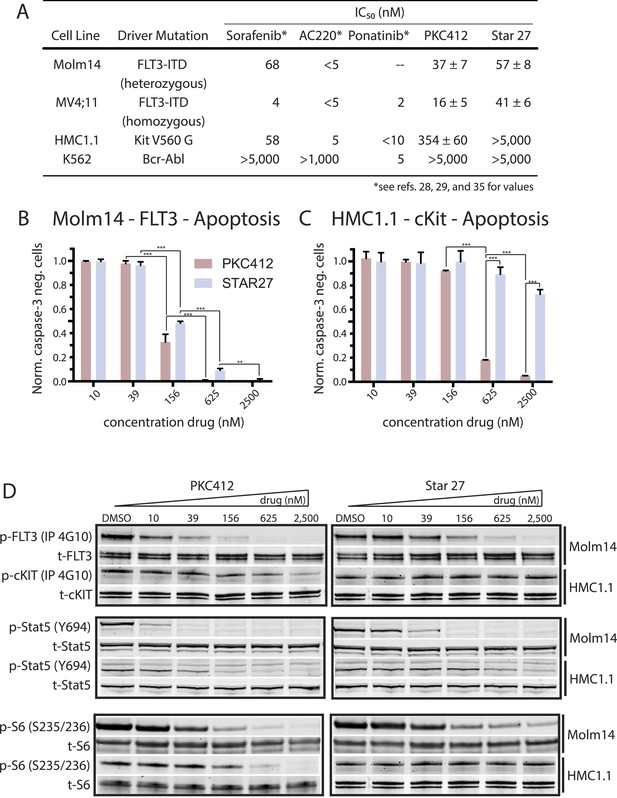
Cellular proliferation, apoptosis, and biochemical validation of Star 27’s potency against FLT3 mutants, but not in the KIT context.
(A) Table of cellular IC50 values for leading experimental clinical therapies (Sorafenib, AC220, Ponatinib, and PKC412) and Star 27 against a panel of AML-relevant human-derived (MV4;11 and molm14) cell lines and toxicity controls (HMC1.1 and K562) (adapted* from Guo et al., 2007; Kampa-Shittenhelm et al., 2013; Smith et al., 2013). Star 27 shows a similar order of magnitude potency against Molm14 and MV4;11 compared to PKC412. Relevant therapeutic windows between PKC412 and Star 27: an over fivefold increase in selectivity for HMC1.1/MV4;11 cells and an 11-fold increase for HMC1.1/Molm14 is observed for Star 27 over PKC412. Replicates shown are the result of at least three attempts, each in triplicate, and error ranges represent the standard error of the mean. (B) Normalized caspase-3 negative cells plotted against escalating drug dosage. Star 27 shows a similar degree of apoptosis to PKC412 in Molm14 cells. (C) In KIT-addicted HMC1.1 cells PKC412 exhibits potent toxicity while Star 27 shows very little up to 2,500 nM. Results performed in triplicate with two biological replicates. **p < 0.01; ***p < 0.001. (D) PKC412 and Star 27 have a similar degree of inhibition against p-FLT3 in Molm14 cells as well as a similar degree for p-STAT5 and p-S6 (for p-ERK, p-AKT, and p-MEK data not shown). In the HMC1.1 cells, by contrast, PKC412 and Star 27 show a larger difference in p-KIT inhibition.
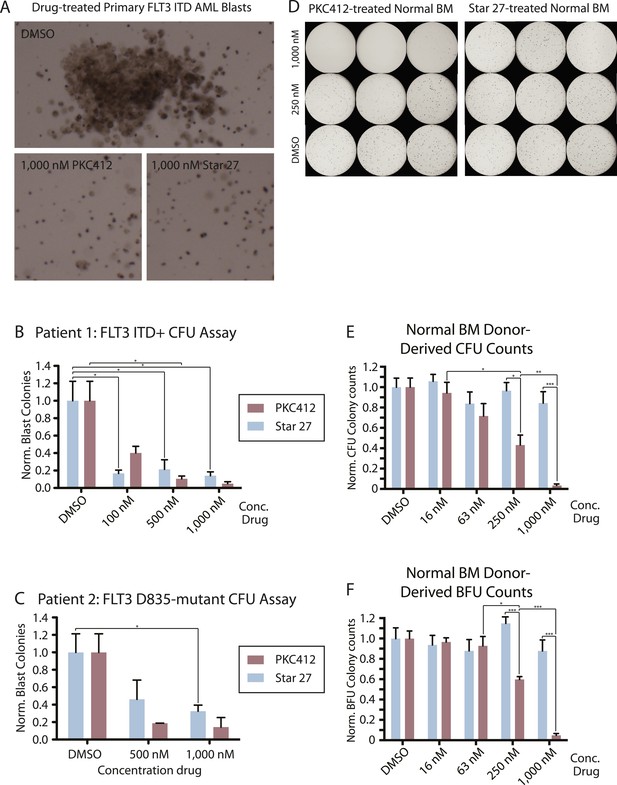
Colony forming assays comparing effect of Star 27 and PKC412 on normal donor-derived stimulated peripheral blood (SPB), normal bone marrow (BM) growth, and malignant blast reduction.
(A, B, C) Colony forming assays comparing effects of Star 27 and PKC412 on primary patient AML circulating blast growth in methylcellulose. Colonies consist primarily of leukemic blasts of various hematopoietic identities. BFU colonies are not shown. Colony numbers scored individually for each replicate 3 cm plate. (A) Images showing leukemic blast morphology in DMSO-treated plates and representative images in 1,000 nM-treated plates for both drugs as indicated. (B) Primary patient FLT3-ITD+ AML circulating blasts. Raw colony numbers range from ca. 20 in PKC412 or Star 27-treated cells at 1,000 nM to ca. 1,200 for CFUs in DMSO-treated plates. Concentrations shown were applied in triplicate to each concentration for two different cell densities, the larger (2.5 × 10^5 cells/ml) showing adequate cell growth and colony formation for statistically significant counts, the averages of those trails then calculated for standard error of the mean. *p < 0.05. (C) Primary patient FLT3 D835Y AML circulating blasts. Raw colony numbers range from ca. 25 for PKC412 (or 50 for Star 27) at 1,000 nM to ca. 120 for DMSO-treated plates. Conditions repeated in duplicate or higher replicates. Colonies scored individually for the highest cell density tested (2.5 × 10^5 cells/ml). Similar to primary ITD patient samples, colonies consist mostly of poly-hematopoietic leukemic blasts. *p < 0.05. (D, E, F) Normal BM and SPB colony data. (D) Images of colonies grown and derived from normal BM (images of SPB not shown) in methylcellulose. Raw colony numbers for BM range from zero in most PKC412-treated replicates at 1,000 nM to ca. 400 for BFUs and ca. 300 for CFUs in all Star 27-treated conditions. Non-magnified differences are particularly noticeable between the 1,000 nM and 250 nM dosages. (E and F) Concentrations shown were applied in triplicate to three normal SPB and one normal BM donors, the averages of those trials then calculated for standard error of the mean (full data not shown). Colony counts for DMSO ranged from the low 100's to 400's. Graphs show Star 27 having no effect on hematopoiesis up to 1,000 nM while PKC412 eliminates most normal hematopoietic colony forming potential at 1 mM on colony forming units (CFU) and blood forming units (BFU), respectively. CFU-GEMM colonies were counted as one CFU plus one BFU. *p < 0.05; ***p < 0.001.
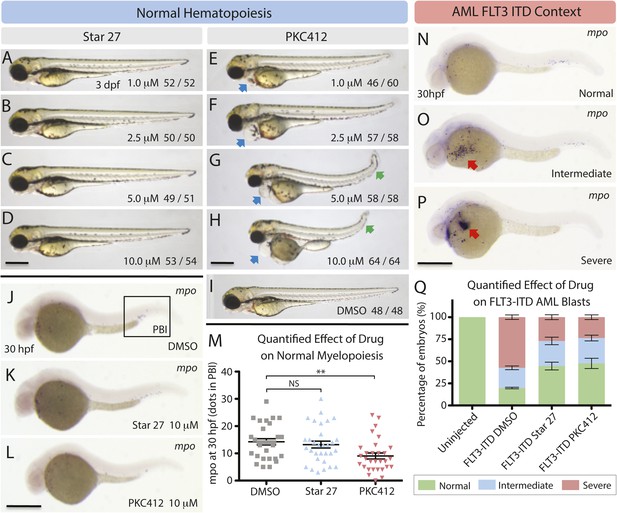
Effects of Star 27 on D. rario (zebrafish) WT morphology and myelopoiesis, and FLT3-ITD AML context.
Effect of Star 27 on zebrafish normal myelopoiesis and the FLT3/ITD-induced myeloid cells expansion. (A–D) The effect of Star 27 on zebrafish embryonic development at 3 dpf, showing no noticeable morphological defects up to 10 μM. (E–I) The effect of PKC412 on zebrafish embryonic development at 3 dpf, showing substantial pericardial defects beginning at 500 nM (data not shown), and tail curvature and length defects beginning at 1 μM and 2.5 μM, respectively vs. vehicle. (J, K, M) The effect of 10 μM Star 27 treatment on mpo+ myeloid cells development in the posterior blood island (PBI) at 30 hpf, showing no statistically significant change. (J, L, M) The effect of 10 μM PKC412 treatment on mpo+ myeloid cells development in the PBI at 30 hpf, showing a statistically significant granulogenesis/myelosuppression. (N–P) Three categories of mpo transcription (N, normal; O, intermediate; P, severe) were defined based on the WISH results (three experiments). (Q) The rescue effect of 10 μM Star 27 treatment on FLT3/ITD-induced mpo+ myeloid cells expansion at 30 hpf, showing rescue of normal phenotype approaching that seen for AC220 (He et al., 2014). Scale bar equals 500 μm. Blue arrows indicate the pericardial edema, green arrows indicate tail shortening and curving, and red arrows indicate the FLT3-ITD AML mpo+ myeloid cells expansion. PKC412 treatment on FLT3/ITD-induced mpo+ myeloid cells expansion at 30 hpf, showing an efficaciousness similar to Star 27, consistent with Figures 1–3. For all experiments: Zebrafish embryos were collected and kept in standard E3 medium at 28°C. Different concentrations of either drug were added to the E3 medium from 6 hr post fertilization (hpf) to 3 days post fertilization (dpf). Embryos treated with DMSO or 10 μM drug from 6 to 30 hpf were collected for mpo whole mount in situ hybridization (WISH) analysis. 80 ng plasmid DNA containing FLT3/ITD sequence was microinjected into one-cell stage embryos, and the uninjected embryos were used as control. FLT3/ITD-injected embryos were treated with DMSO or 10 μM drug from 6 to 30 hpf. Embryos were collected at 30 hpf for mpo WISH analysis. **p <0.01.
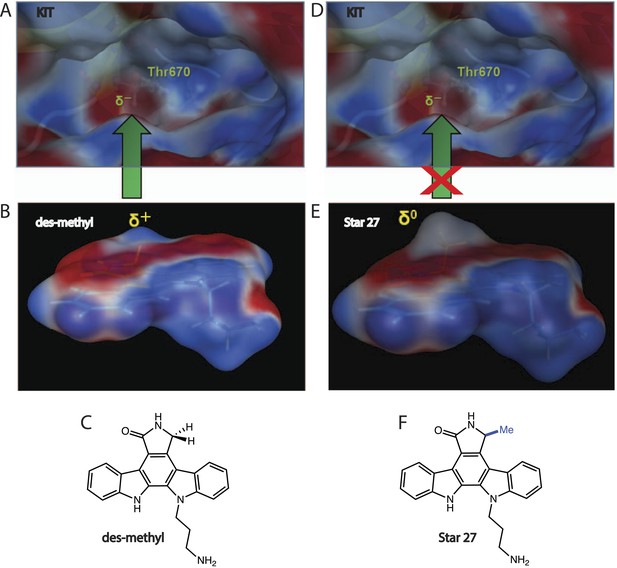
Electronic model of selectivity.
Star 27-des methyl (PKC412-relevant) and Star 27 are depicted in 3D with electron potential mapping (EPM) to highlight the difference of a single methyl group. (A–C) Star 27-des methyl maintains a partial positive charge at the C7 position which matches the partial negative charge on the Thr 670 gatekeeper of KIT, potentially explaining PKC412's relative affinity for KIT. (D–F) Star 27 maintains a neutral charged surface at C7 due to the presence of the methyl group, potentially explaining its lack of binding to KIT.
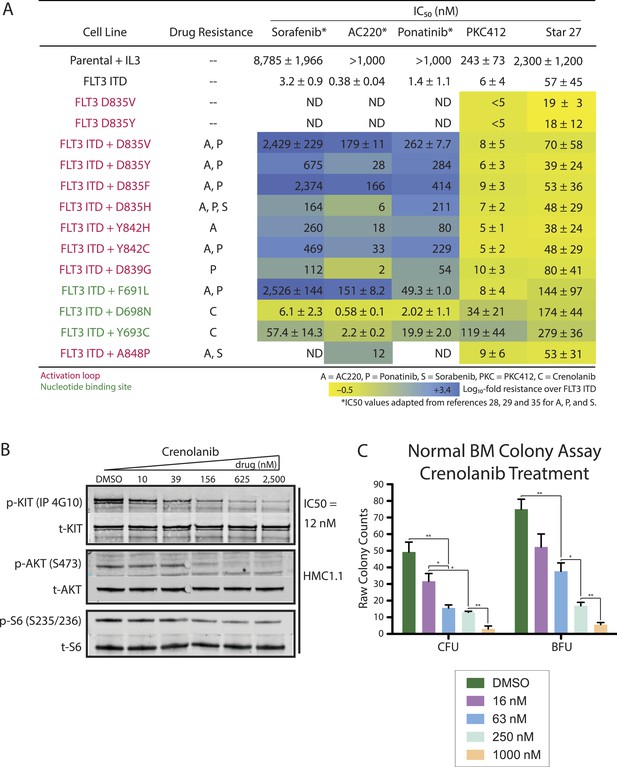
Heatmap of cellular EC50 values for leading clinical therapies (S = Sorafenib, A = AC220, P = Ponatinib, PKC412) and Star 27 against a panel of AML-relevant human- and mouse-derived drug-resistant cell lines.
(A) (adapted from Guo et al., 2007; Kampa-Shittenhelm et al., 2013; Smith et al., 2013). Log10-scale fold resistance between Ba/F3 FLT3 ITD and drug-induced mutations (ITD + KD double mutant) shown via heat map. Point mutations corresponding to each drug-relevant resistance shown. Star 27 maintains potency between FLT3 ITD and resistance mutants comparable to PKC412. Mutations derived independently from both saturating mutagenesis and patient-derived samples [Smith et al., 2012]. Replicates shown are the result of at least two or three attempts, each in triplicate, and error ranges represent the standard error of the mean. (B and C) Crenolanib's effect on p-KIT inhibition and of colony growth in normal BM. (B) Crenolanib inhibits p-KIT in HMC1.1 cells at 12 nM IC50. This level of inhibition translates to downstream kinases, with inhibition of p-AKT (S473) and p-S6 (S235/S236). (C) Crenolanib potently inhibits normal BM colony formation at <63 nM and ca. 63 nM IC50's of CFU and BFU colonies, respectively.





