Mechanistic insight into the conserved allosteric regulation of periplasmic proteolysis by the signaling molecule cyclic-di-GMP
Figures
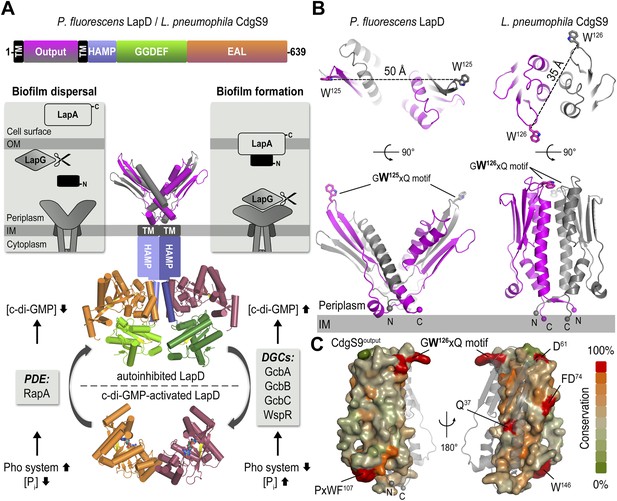
The LapD signaling system and structures of the LapD periplasmic domain.
(A) Overview of LapD-mediated signaling. The primary structure and domain organization of the LapD receptor is shown (top panel). The cartoon (bottom panel) summarizes the current model of LapD-mediated regulation of biofilm formation in P. fluorescens by the differential recruitment of the periplasmic protease LapG in response to exogenous inorganic phosphate (Pi) availability (reviewed in Boyd and O'Toole, 2012). Low bioavailability of Pi induces the expression of phosphodiesterases (PDEs) via the Pho system, which results in low intra-cellular concentrations of c-di-GMP and LapD adopting an autoinhibited state. Auto-inhibited LapD cannot sequester the periplasmic protease LapG, allowing the LapG enzyme to cleave the cell-surface adhesin LapA, releasing it from the cell surface. Conversely, when environmental Pi is present in sufficient concentration, a subset of diguanylate cyclases (DGCs) produce intra-cellular c-di-GMP to levels sufficient for binding to the EAL domain of LapD, thus activating this receptor. Activated LapD binds to and sequesters LapG in the periplasm, which promotes biofilm formation by preventing cleavage of LapA. (B) Crystal structures of the periplasmic output domain from P. fluorescens LapD and L. pneumophila CdgS9. Two orthogonal views of each output domain structure are shown in ribbon representation, with the two protomers colored pink and gray, respectively (P. fluorescens LapDOutput, PDB 3PJV; Navarro et al., 2011). Distances between the tryptophan residues in the conserved GWxQ motif of each protomer are indicated. The relative position of the inner cell membrane (gray bar) and connection to the flanking transmembrane (TM) helices are indicated. (C) Surface conservation of the output domain. The sequences of 18 orthologs of LapD were aligned and the sequence conservation was mapped on to the solvent-accessible surface of CdgS9Output. The surface is colored according to the degree of conservation.
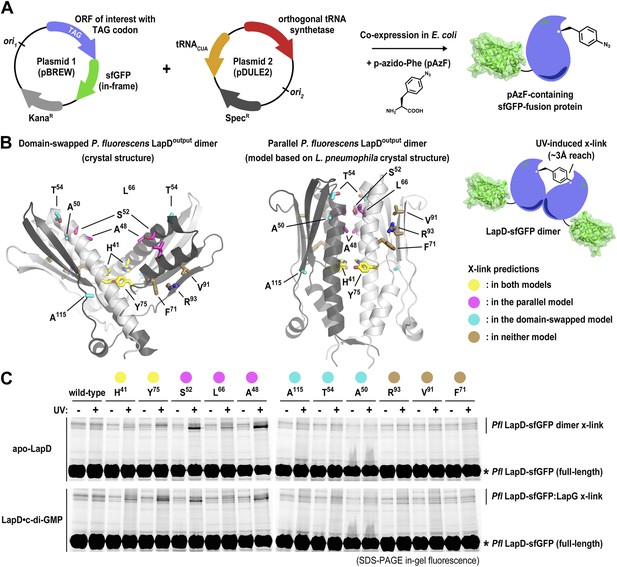
Distinguishing between the ‘open’ domain-swapped and ‘closed’ parallel conformations of the output domain of LapD.
(A) Non-natural amino acid incorporation into protein. The UV-photo-activatable non-natural amino acid para-azido-phenylalanine (pAzF) was site-specifically incorporated into the output domain of full-length P. fluorescens LapD, which was fused C-terminally to superfolder GFP (sfGFP), using a two-plasmid amber-suppression expression system. (B) Cross-linking strategy based on structural models. Two different models of the output domain of LapD are shown with the eleven sites of pAzF incorporation shown as sticks and color-coded according to structure-based cross-linking predictions. The left panel shows the domain-swapped, open conformation determined previously (PDB 3PJV; Navarro et al., 2011), while the right panel shows a homology model of the ‘closed’, parallel conformation based on the crystal structure of CdgS9Output. (C) Crosslinking patterns support a ‘closed’, parallel model. Cells expressing full-length LapD fused to sfGFP with pAzF incorporated at the indicated positions were harvested by centrifugation and lysed by sonication. Membranes were isolated by ultracentriguation and resuspended in buffer lacking detergent supplemented with and without c-di-GMP. After equilibration, half of each sample was exposed to short-wave UV light. These membrane suspensions were immediately solubilized in 2% SDS and subjected to SDS-PAGE analysis. Gels were imaged by fluorescence. As indicated, cross-links appear as band-shifts with a higher molecular weight compared to LapD-sfGFP.
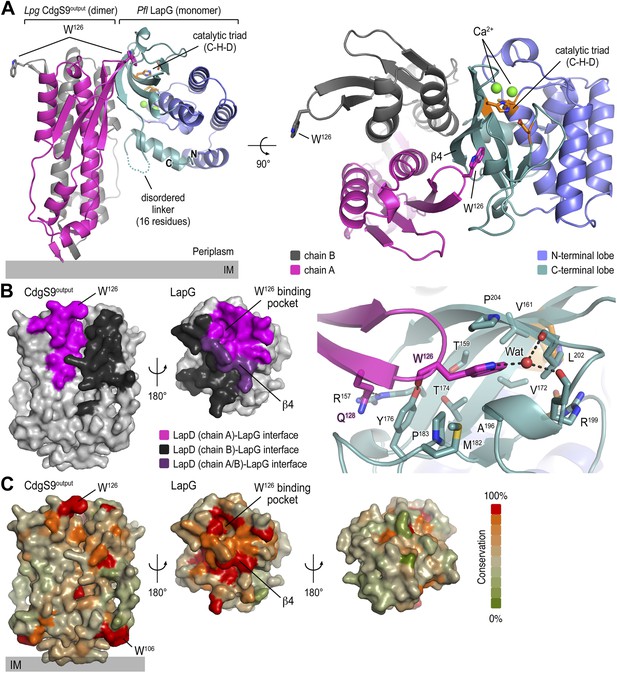
Crystal structure of a P. fluorescens LapG and L. pneumophila CdgS9Output complex.
(A) Overview of the complex structure. The LapG-CdgS9Output complex is shown as a ribbon representation, with the two protomer chains of CdgS9Output colored in pink and gray, and the N- and C-terminal lobes of LapG shown in slate and cyan, respectively. The conserved catalytic triad (cysteine-histidine-aspartate; C-H-D) is shown as sticks with the carbon atoms in orange. The highly conserved tryptophan residue (W126) in each protomer of CdgS9Output is presented as sticks, calcium ions of LapG are shown as green spheres. The relative position of the inner cell membrane (gray bar) is indicated. Two orthogonal views are shown. (B) Interaction interface between CdgS9Output and LapG. The interface footprints on LapG and the CdgS9 half-sides (pink, purple and black) were highlighted on the accessible surface area of the individual proteins (left panel). The binding pocket for CdgS9 W126 in LapG is shown as a close-up view (right panel). A 180° rotation was applied to LapG from the complex structure to view the interface. (C) Surface conservation of CsgS9Output and LapG interaction interface. Based on the alignment of 18 sequences of LapD and LapG orthologs, the sequence conservation was mapped onto the accessible surface of each protein. The surface is colored according to the degree of sequence conservation. For LapG, two views, separated by a 180° rotation, are shown.
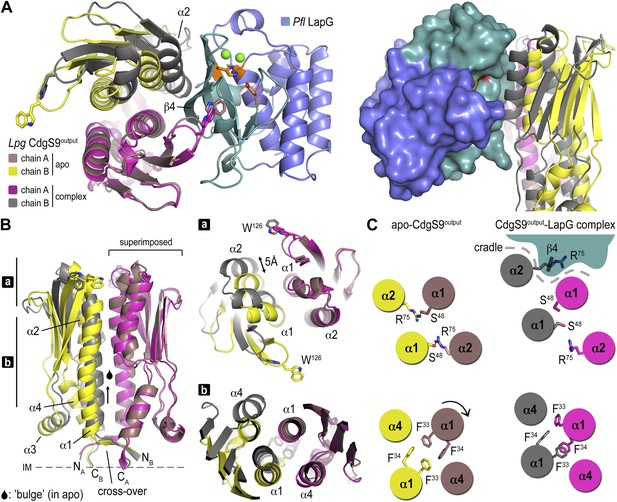
Conformational changes in CdgS9Output between the apo-state and in complex with LapG.
(A) Overview of the superposition of apo-CdgS9Output and LapG-bound CdgS9Output. Chain A of the CdgS9Output dimer was used as the reference for the superposition. Chains A and B of apo-CdgS9Output are colored tan and yellow, while chains A and B of LapG-bound CdgS9Output are colored pink and gray, respectively. The N- and C-terminal lobes of LapG are colored in slate and cyan, respectively. Both cartoon (left panel) and surface (right panel) representations of LapG are shown. (B) Components of the conformational change. The superposition in (A) is shown as a side-view, depicting the helical motions of α1, α2, and α4. Two perpendicular views are shown, with the right panel showing slices of the top view. (C) Cartoon model. The structural transition from an apo-state (yellow/tan) to a LapG binding-competent state (pink/gray) of CdgS9Output based on the analysis shown in panels (A) and (B) is illustrated. The top pair depicts changes at the distal tip of the output domain; the bottom pair illustrates changes in the membrane proximal region of the output domain.
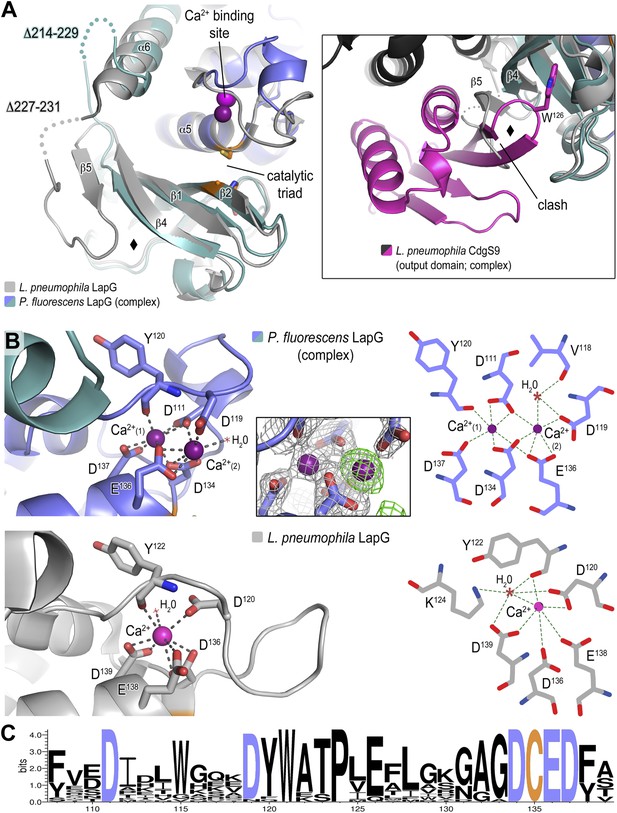
Structure of P. fluorescens LapG in complex with CdgS9Output.
(A) The structure of L. pneumophila LapG shown in gray (PDB: 4FGO, Chatterjee et al., 2012) was superimposed on that of P. fluorescens LapG, colored in slate and cyan, when in complex with CdgS9Output. A close-up view of the differences observed in the C-terminal lobe of the two LapG structures is shown (right inset). Disordered regions not resolved in the crystal structures are shown as dots. Calcium ions are represented as magenta spheres. The catalytic triad residues (C-H-D) are shown as sticks with the carbon atoms colored in orange. (B) The Ca2+ binding sites of LapG. P. fluorescens (top panel, blue cartoon) and L. pneumophila LapG (bottom panel, gray cartoon) coordinate two or one calcium ion, respectively. In P. fluorescens LapG, an Fo–Fc omit map contoured at 3σ (green mesh, inset of top panel) shows strong evidence for a second calcium ion not seen in the L. pneumophila structure, which is made possible by local conformational differences in the so-called calcium binding loop that bring D111 of P. fluorescens into position to coordinate the additional calcium ion. A LIGPLOT (Wallace et al., 1995) (right panels) depicts the coordination of calcium ions. (C) Conservation of the di-calcium binding site. A WebLOGO plot (Crooks et al., 2004) generated from an alignment of 18 LapG ortholog sequences demonstrates the conservation of coordinating residues (blue font) as well as the nucleophilic cysteine (yellow font).
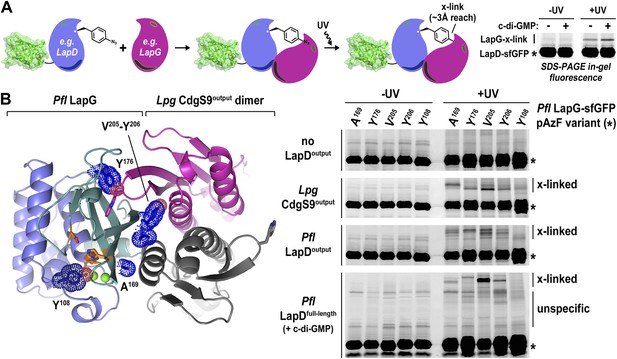
Validation of the crystallographic CdgS9Output-LapG complex interface.
(A) Schematic overview of the Lapd-LapG cross-linking assay. The right panels show a representative experiment in which detergent-solubilized LapD-sfGFP is incubated with non-fluorescent LapG-containing pAzF incorporated at a site predicted to be at the LapD–LapG binding interface in the presence and absence of c-di-GMP. Upon UV irradiation, a fluorescent band corresponding to LapD-sfGFP covalently linked to LapG is observed, which is more intense in the presence of c-di-GMP indicative of more LapD–LapG binding. Other experiments were carried out with a LapG-sfGFP/LapD pairs. (B) Structure-guided cross-linking of LapD variants and LapG. The complex structure (left panel) highlights four sites on LapG (A169, Y176, V205, and Y206) that span the width of the binding interface plus an additional fifth site (Y108) that is not at the interface in which pAzF was incorporated. Each of these five LapG-sfGFP pAzF-containing variants were purified and incubated with purified non-fluorescent CdgS9Output, LapDOutput and c-di-GMP activated, detergent-solubilized full-length P. fluorescens LapD prior to irradiation with UV light. Samples were analyzed by SDS-PAGE and gels imaged by fluorescence as described in Figure 2.
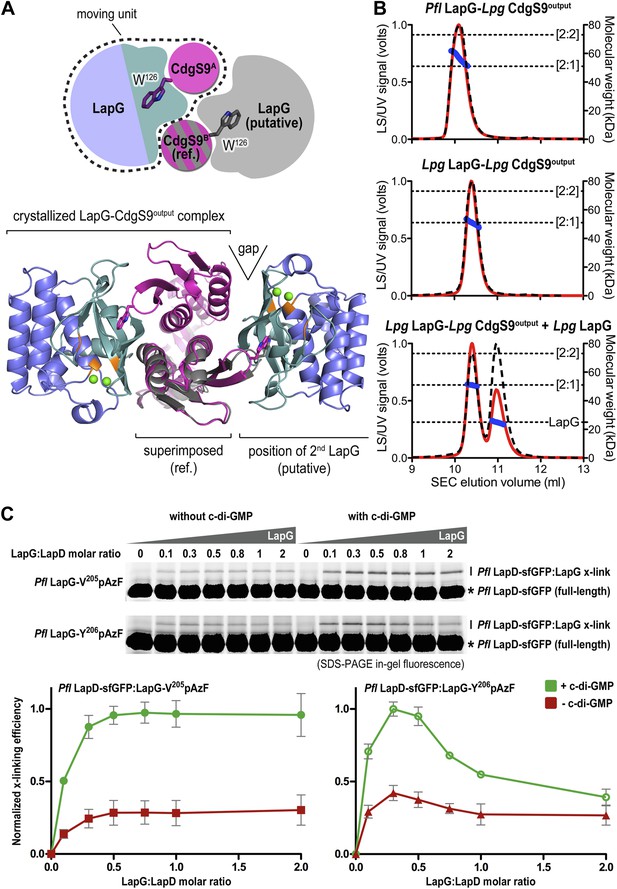
Stoichiometry of the LapD–LapG complex.
(A) Structural model of a 2:2 CdgS9Output–LapG complex. The CdgS9Output domain that engages LapG via its GW126xQ loop was superimposed onto the output domain of the complex structure, which is not bound to LapG. The open arrow indicates poor shape complementarity at the putative second LapG binding site. (B) Stoichiometry of CdgS9Output-LapG complex in solution. SEC-MALS was used to determine the absolute molecular weight of the CdgS9Output-P. fluorescens LapG complex used for crystallization (top panel), a co-expressed complex of L. pneumophila LapG bound to CdgS9Output (middle panel), and the latter complex incubated with an excess L. pneumophila LapG (bottom panel). Red traces and dashed lines show the signal of the light scattering and UV detector, respectively. Dotted, vertical lines indicate the theoretical molecular weight of the 2:1 and 2:2 output domain-protease complexes based on their primary sequence. Blue points across the elution peaks refer to the calculated molecular weight as the proteins elute from the gel filtration column. (C) Assessment of LapG binding to full-length LapD by covalent cross-linking. Purified, non-fluorescent LapG possessing pAzF at positions V205 (top gel) or Y206 (bottom gel) was titrated into a fixed quantity of detergent solubilized LapD-sfGFP in the absence (left side of gel) or presence (right side of gel) of c-di-GMP (20 μM) and allowed to incubate for 20 min. Samples were irradiated with UV light for 5 min and analyzed by SDS-PAGE. Band intensities of LapD-sfGFP cross-linked to LapG from three independent experiments were measured, normalized to non-crosslinked LapD-sfGFP, averaged, and plotted as function of the LapG:LapD molar ratio (±SD), which showed maximum crosslinking at ∼0.3–0.5 LapG molecules to 1 LapD molecule (bottom panels).
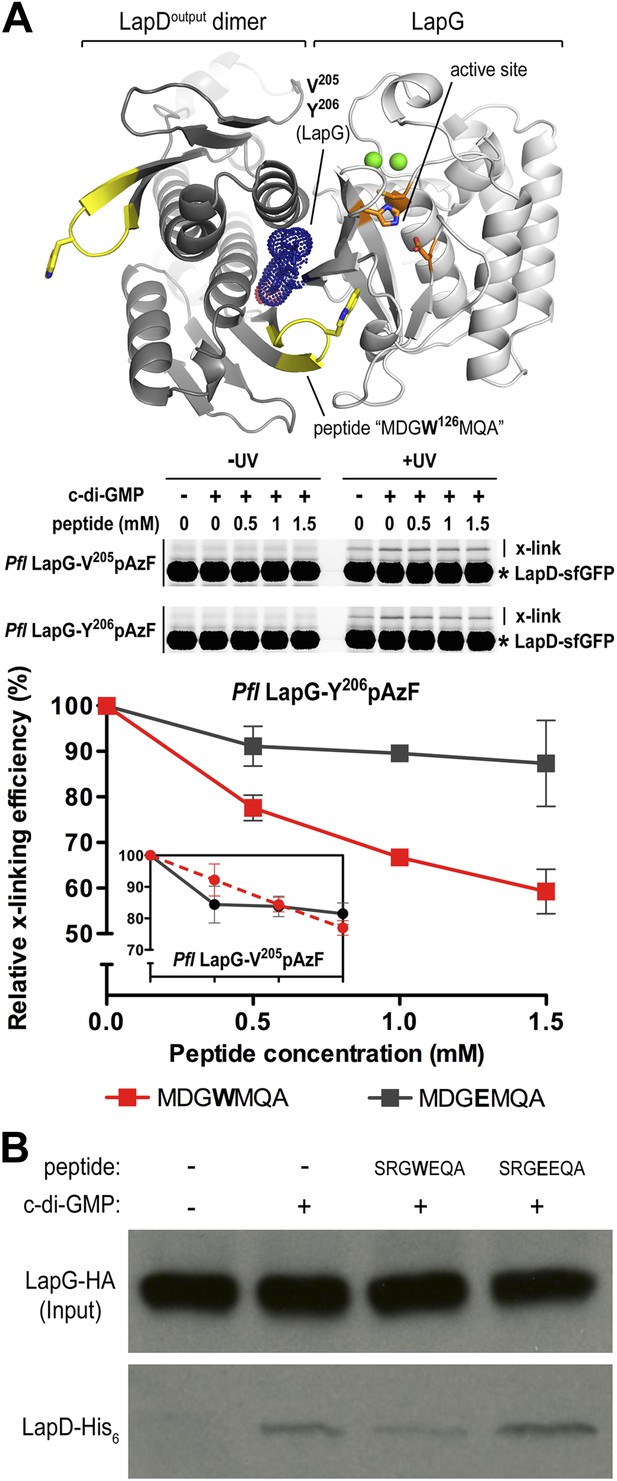
Attenuating LapD–LapG interactions with a peptide containing the GWxQ binding motif.
(A) Cross-linking assay using purified proteins. The CdgS9Output-LapG crystal structure depicts the GWxQ β-hairpin motif of CdgS9Output (yellow cartoon) as well as the two sites of pAzF incorporation (blue mesh) on LapG used to assess LapD–LapG interactions (top panel). To assess the effects of the MDGW126MQA peptide on LapD–LapG interactions, detergent-solubilized, full-length LapD-sfGFP was incubated with purified non-fluorescent LapG containing pAzF at V205 (inset in bottom panel) or Y206 (main graph in bottom panel), and increasing concentrations of peptide. Half of the sample was subjected to UV irradiation for 5 min prior to analysis by SDS-PAGE. The average intensity of the fluorescent band corresponding to LapD-sfGFP crosslinked to LapG from three independent measurements was normalized to non-crosslinked LapD-sfGFP, averaged, and plotted as a function of peptide concentration (±SD) (red line; bottom panel). The peptide MDGE126MQA was used as a negative control (gray line; bottom panel). (B) Pull-downs of LapG-HA from lysates of P. fluorescens. LapD-His6 and LapG-HA were co-expressed in a P. fluorescens strain with deleted lapD and lapG genes. Western blots using HA (top panel) and His6 (bottom panel) specific primary antibodies were used to assess the levels of LapD-His6 that co-purify with LapG-HA in the presence or absence of peptides and c-di-GMP.
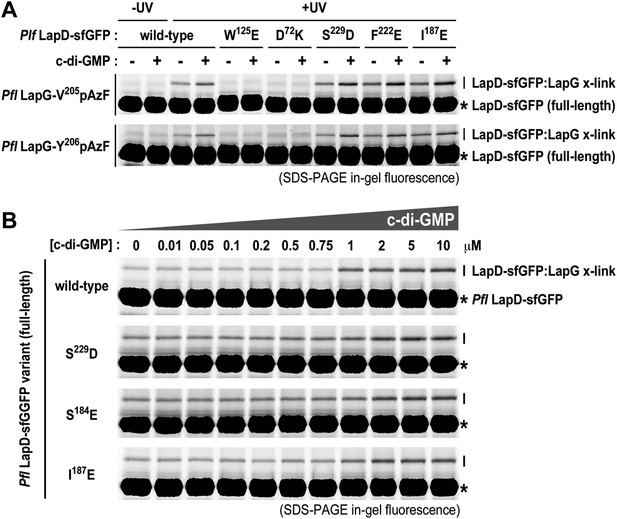
Mutations in the HAMP domain and S helix sensitize LapD toward c-di-GMP.
(A) Effect of structure-guided point mutations on LapD function. Detergent solubilized LapD-sfGFP variants (2 μM) were incubated with non-fluorescent LapG (1 μM) containing pAzF at positions V205 (upper gel) and Y206 (lower gel) in the presence and absence of c-di-GMP prior to exposure to UV light and SDS-PAGE analysis. Fluorescently imaged gels indicated LapD mutations W125E and D72K abolish LapD–LapG interactions while HAMP domain and S helix mutations S229D, F222E, and I187E have higher levels of interaction with LapG in the absence and presence of c-di-GMP compared to wild-type. (B) C-di-GMP sensitivity of LapD. C-di-GMP was titrated into a fixed amount of detergent solubilized LapD-sfGFP (2 μM) variants and non-fluorescent LapG (1 μM) containing pAzF at position V205, after which crosslinking was initiated by UV irradiation and analyzed by SDS-PAGE. A sharp transition in crosslinking is observed between 0.75 μM and 1 μM c-di-GMP for wild-type LapD-sfGFP but not for the other activating mutations, which have higher basal levels of crosslinking and a less pronounced transition.
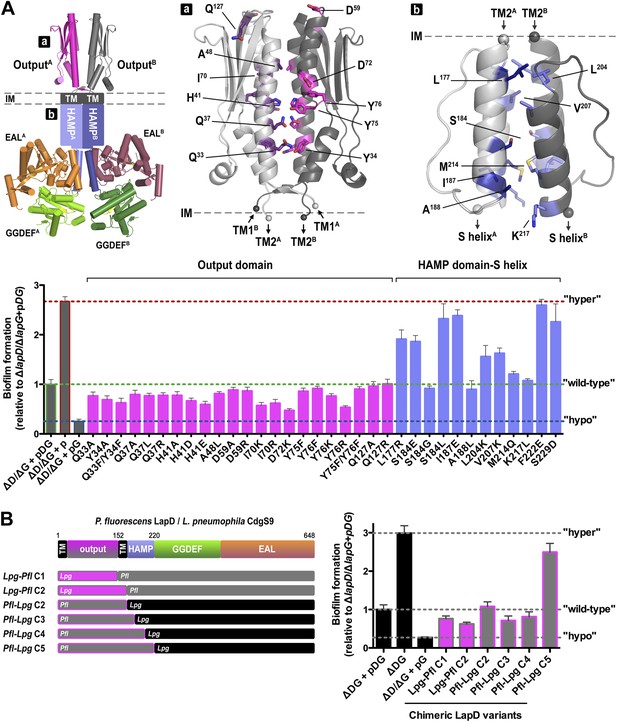
Mutational analysis of LapD regulation.
(A) Biofilm phenotypes of LapD point mutants. A structural model of full-length LapD is shown, which includes the closed, parallel output domain that is linked to the cytoplasmic S helix-GGDEF-EAL domain module via transmembrane and HAMP domains (left panel). Homology models of output domain (middle panel) and a HAMP domain (right panel) of P. fluorescens LapD were used to design mutations predicted to alter the activity of LapD. Biofilm phenotypes of these LapD variants were assessed using previously established in vivo assays (lower panel). Representative data from 8–12 wells for at least three biological replicates are shown as means ±SD. (B) Biofilm phenotypes of chimeric constructs. LapD constructs were created that contained either the periplasmic or cytoplasmic domains of L. pneumophila CdgS9 replacing the respective native P. fluorescens LapD domains (left panel). ΔD/ΔG: P. fluorescens strain lacking lapD and lapG genes, pDG: dual expression plasmid for LapD and LapG, p: empty plasmid, pG: expression plasmid for LapG.
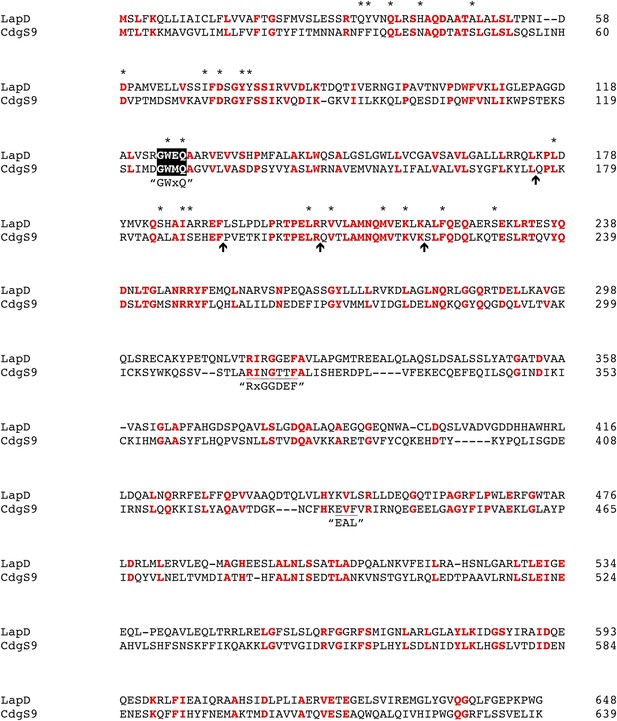
Pairwise alignment of LapD and CdgS9.
Strictly conserved residues are highlighted in red. The positions of point mutations used in this study are marked by asterisks. Arrows indicate positions of the chimeric swap points. Other functional motifs are highlighted and labeled.
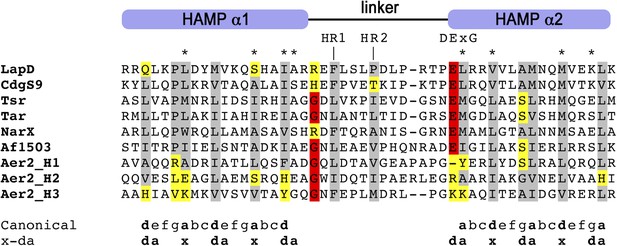
Multiple-sequence alignment of HAMP domains.
The alignment highlights conserved hydrophobic residues (gray), other residues implicated in HAMP domain switching (red), and divergent substitutions (yellow). The hydrophobic positions in helices 1 and 2 are central elements for the conformational transitions from a complementary x-da to a knobs-into-holes supercoil packing through a 26° helical rotation within the HAMP domain's four-helix bundle, according to the gear-box model (Hulko et al., 2006). Only a few positions show non-conservative changes in LapD and CdgS9, while the overall alignment indicates that these HAMP domain adopt a canonical fold established for Af1503 (PDB 2L7H) (Ferris et al., 2011) and Aer2 (PDB 3LNR) (Airola et al., 2010). Residues that were targeted for site-directed mutagenesis of LapD are highlighted by asterisks (*).
Videos
Composite structural overview.
The video shows several interpolations between the apo-state and the LapG-bound output domain dimer of CdgS9, based on the crystallographic data presented here. It depicts the conformational changes between the two states of the periplasmic output domain and how they lead to LapG recruitment.
Tables
Data collection and refinement statistics
| CdgS9Output (SeMet) | LapG/CdgS9Output (SeMet) | LapG/CdgS9Output (native) | |
|---|---|---|---|
| Data collectiona | |||
| Space group | P6122 | P21 | P21 |
| Unit cell axes | |||
| a, b, c (Å) | 61.58, 61.58, 147.53 | 59.08, 74.69, 62.88 | 75.85, 73.67, 88.22 |
| α, β, γ (ο) | 90.0, 90.0, 120.0 | 90.0, 101.8, 90.0 | 90.0, 92.9, 90.0 |
| Resolution Limits (Å) | 30–2.14 (2.22–2.14) | 30–0.29 (2.37–2.29) | 44–2.10 (2.17–2.10) |
| Unique Observations | 9611 (924) | 22,884 (2231) | 55,783 (4959) |
| Completeness | 98.6 (97.6) | 93.8 (92.2) | 97.7 (87.3) |
| Multiplicity | 21.9 (18.6) | 7.2 (5.4) | 4.3 (3.2) |
| Average I/σ | 35.4 (10.5) | 20.1 (2.3) | 12.4 (2.8) |
| Rsym (%) | 7.7 (30.9) | 8.9 (67.1) | 8.2 (38.7) |
| Refinement | |||
| Rcryst/Rfree (%) | 21.9/24.3 | 17.0/21.1 | |
| No. protein molecules | 1 | 6 | |
| No. protein residues | 131 | 886 | |
| No. water molecules | 90 | 677 | |
| Total number atoms | 1158 | 7826 | |
| rmsd bond angles (o) | 1.02 | 1.05 | |
| rmsd bond lengths (Å) | 0.004 | 0.004 | |
| <B> protein (Å2) | 41.2 | 34.0 | |
| <B> water (Å2) | 59.1 | 38.8 | |
| <B> calcium (Å2) | N/A | 35.3 | |
| Ramachandran Plot (%) | |||
| Favored | 98 | 98 | |
| Outliers | 0 | 0 | |
| PDB code | 4U64 | 4U65 | |
-
a
Numbers in parentheses correspond to values in the highest resolution.





