Reliable cell cycle commitment in budding yeast is ensured by signal integration
Figures
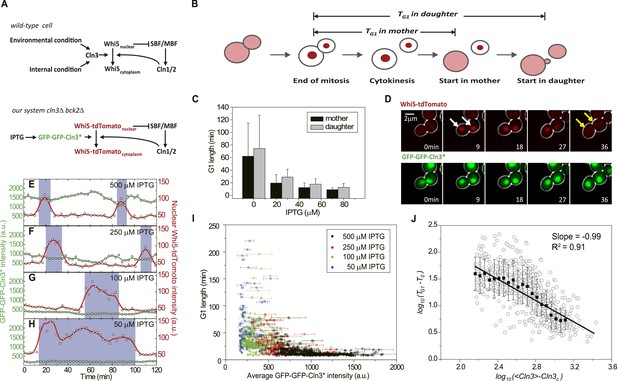
G1 length is inversely proportional to the average Cln3.
(A) Schematic of the Start regulatory network in wild-type (upper panel) and in strains used in this study (lower panel). (B) G1 length is defined as the time interval during which Whi5 resides in the nucleus (Figure 1—figure supplement 1). Whi5 localization is schematically shown in red. (C) Population-averaged G1 length at different IPTG concentrations for cells carrying IPTG-induced CLN3 as the Start signal (Strain YCT2002). Error bars represent standard deviation. (D) Composite bright-field and fluorescence images for cells carrying GFP-GFP-CLN3* and WHI5-tdTomato (Strain YCT2003) under full induction (2 mM IPTG). White arrows indicate the beginning of G1; yellow arrows indicate the timing of Start transition. (E–H) Time courses of the GFP-GFP-Cln3* (green) and nuclear Whi5-tdTomato (red) fluorescent intensities in a representative single cell at each different IPTG concentration. The empty squares and circles denote the raw data of Cln3 and nuclear Whi5 fluorescence, respectively; the green and red lines are the smoothing splines of the raw data; purple boxes show the G1 duration. (I) G1 length vs the average Cln3 fluorescent intensity in G1. Each dot represents a measurement from one cell cycle event; error bars indicate standard deviation. Color groups represent data collected from each of the corresponding IPTG concentrations. (J) G1 length vs the average Cln3 fluorescence in log–log scale (empty circles are calculated from I). <Cln3> represents the average Cln3 fluorescence in G1; T0 and Cln3c are fitted from I as described in ‘Materials and methods’. The solid line is the best linear fit of the binned data (filled squares). Error bars indicate standard deviation.
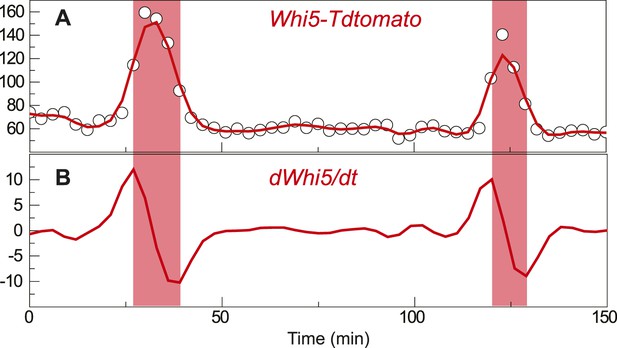
Definition of G1 length.
(A) Nuclear Whi5-tdTomato intensity in a representative single cell; open circle shows the raw data; solid red line shows the spline smoothing. (B) The first derivative of the smoothed data; pink shade area denotes the G1 duration.
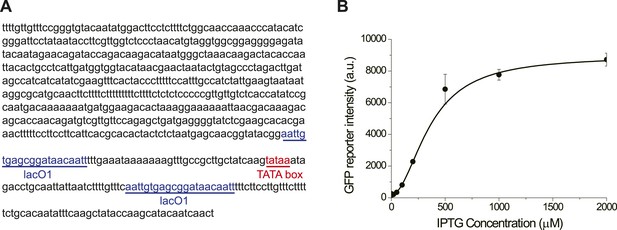
The promoter GlacSpr is repressed by LacI and induced by IPTG.
(A) The sequence of GlacSpr. (B) The dose–response curve of GlacSpr. The transcriptional activity of GlacSpr in the OFF state is a little higher than GAL1pr in glucose, as the result, GlacSpr is not tight enough to completely shut off the cyclin activity of the wild-type Cln3. The cln3Δ bck2Δ cells carrying GlacSpr-CLN3 are viable in the absence of IPTG with prolonged G1 phase.
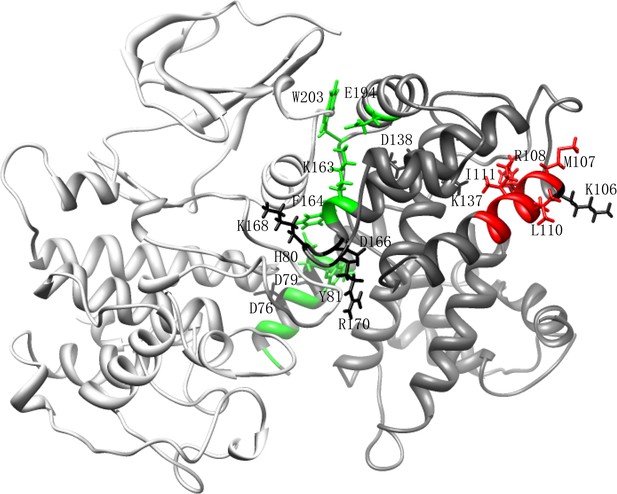
The homologous structure of Cln3-Cdk1 complex.
Cln3 is shown in dim grey and Cdk1 in light grey. The Cln3 in this model only contains amino acid 70-312. The hydrophobic patch is indicated in red, while the interface to Cdk1 is indicated in green. The mutation sites we screened by alanine substitution are shown in sticks. Among those sites, D76, D79, K106, R108, D137, K138, K163, D166, K168, R170, K357, and K359 are in the clustered charge residues; M107, R108, L110, and I111 are in the hydrophobic MRAIL patch; H80, Y81, K163, F164, D166, E194, and W203 are on the interface of Cln3-Cdk1 complex.
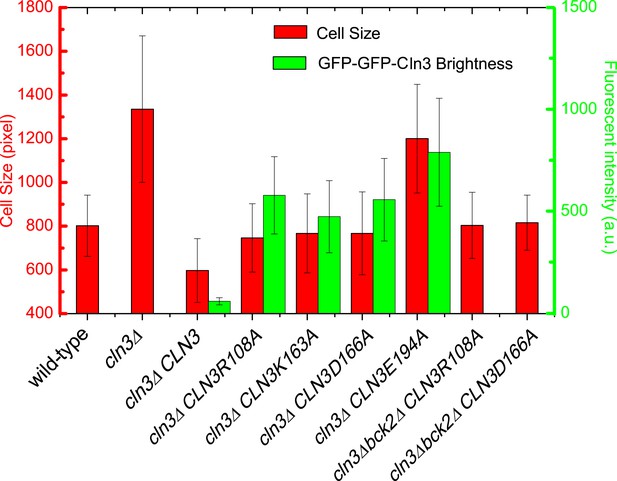
Cell size and GFP brightness of selected Cln3 mutants under full induction.
https://doi.org/10.7554/eLife.03977.007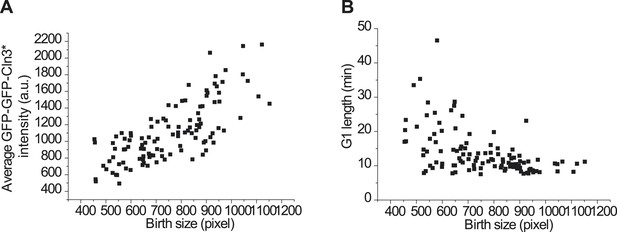
Size control is maintained with the mutant Cln3, Cln3*.
(A) The correlation between birth size and GFP-GFP-Cln3* intensity under full induction, Pearson Coefficient is 0.75. (B) The correlation between birth size and G1 length under full induction, Pearson Coefficient is −0.51 (YCT 2003).
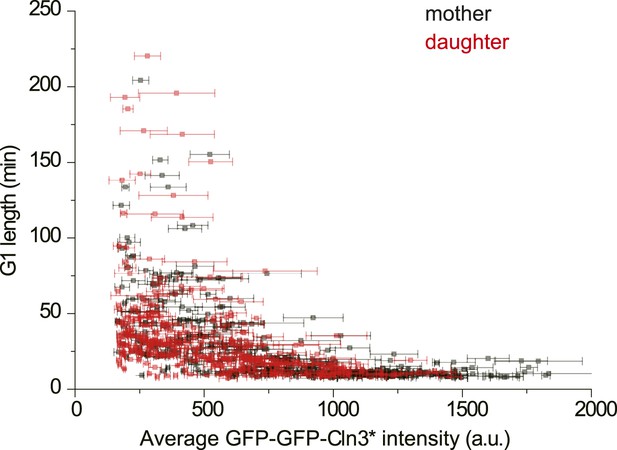
The correlation between G1 length and average Cln3 fluorescence intensity in mother and daughter cells (YCT2003).
https://doi.org/10.7554/eLife.03977.009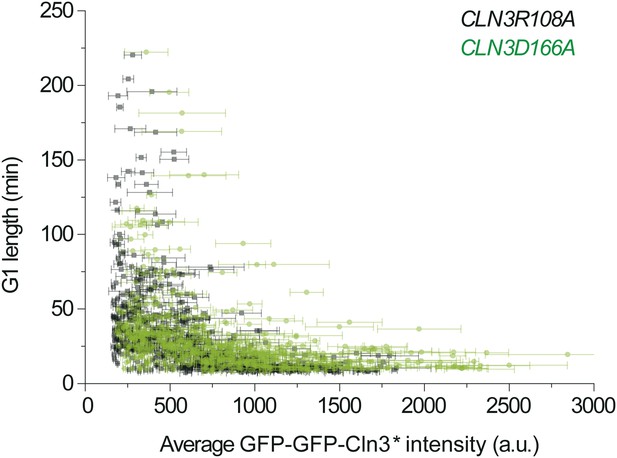
The correlation between G1 length and average Cln3 fluorescence intensity with different Cln3 signals: CLNR108A (YCT2003) and CLN3D166A (YCT2004).
https://doi.org/10.7554/eLife.03977.010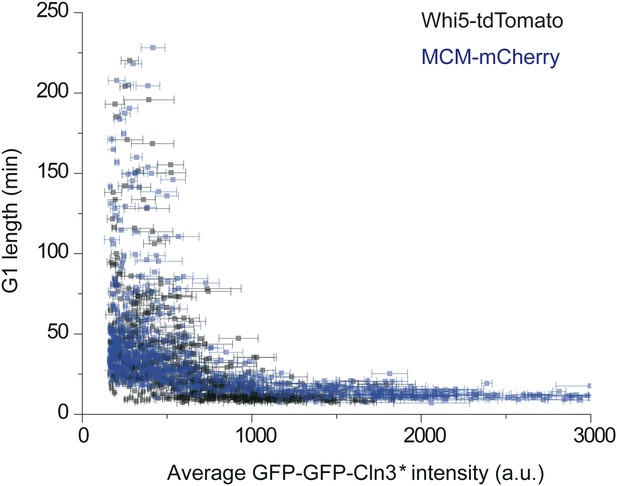
The correlation between G1 length and average Cln3 fluorescence intensity with different G1 length markers: WHI5-tdTomato (YCT2003) and MCM-mCherry (YCT2010).
https://doi.org/10.7554/eLife.03977.011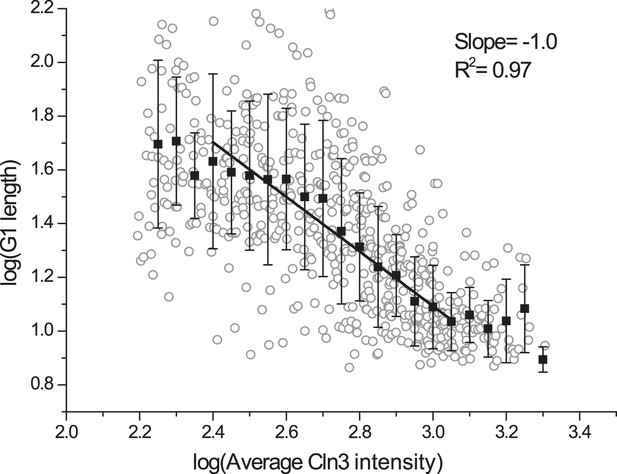
G1 length is inversely proportional to averaged Cln3 intensity without deducting asymptotes.
Open circle denotes raw data in log–log scale; closed square denotes the binned data; solid line is the best linear fit of the middle part; error bars are standard deviation.
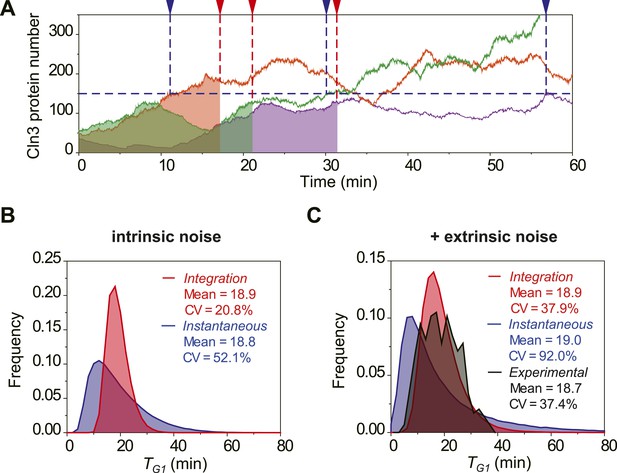
The integration of Cln3 reduces the variability of G1 length.
(A) Representative Cln3 profiles in single cells. Each color represents simulation of a single cell. In the Instantaneous Model, Start is triggered at the time TG1 (vertical blue dash line) when Cln3 profile hits a threshold (here set to 150) for the first time. In the Integration Model, Start is triggered at the time TG1 (vertical red dash line) when the integration of Cln3 (the area under the Cln3 curve as indicated by shadow) reaches a threshold (here set to 1900). The two thresholds are chosen to generate the same average TG1 ≈ 20 min in both models. In generating Cln3 profiles, both intrinsic noise and extrinsic noise are used (Elowitz et al., 2002; Raser and O'Shea, 2004). (B–C) The distributions of TG1 in the Instantaneous (blue) and Integration (red) models, respectively, with intrinsic molecular noise only (B) and with both intrinsic and extrinsic noise (C). The parameters to generate this figure are specified in Figure 2—source data 1. The G1 length distribution from experimental data (Strain YCT2001) is shown in (C).
-
Figure 2—source data 1
Meaning, value and reference of the parameters to generate Figure 2.
- https://doi.org/10.7554/eLife.03977.014
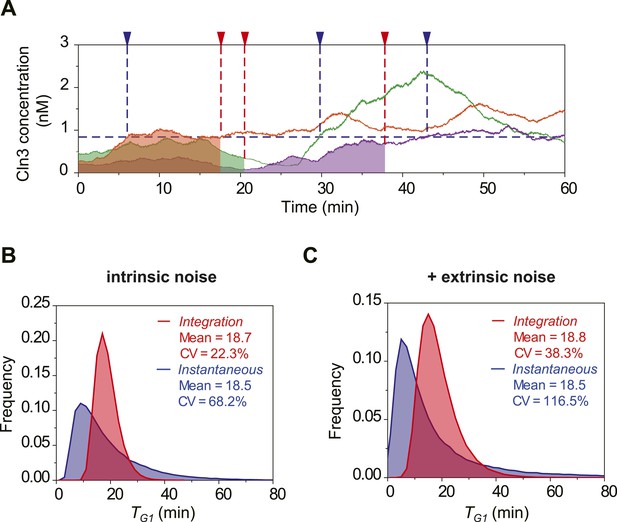
Stochastic simulation of Cln3 profile and Start triggering process with nucleus volume increase.
(A) Representative Cln3 concentration profiles in single cells. Each color represents simulation of a single cell. (B–C) The distributions of TG1 in the Instantaneous (blue) and Integration (red) models, respectively, with intrinsic molecular noise only (B) and with both intrinsic and extrinsic noise (C).
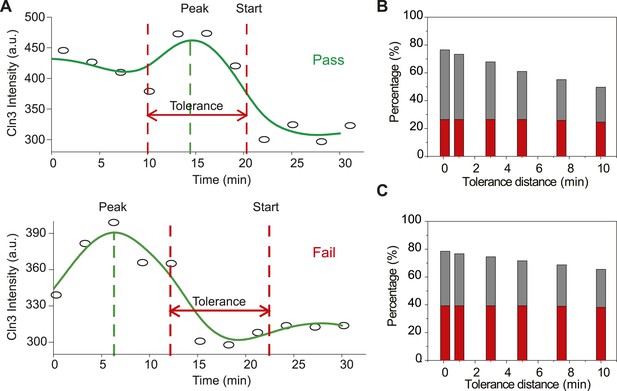
The Instantaneous Model fails with the Cln3 profiles measured in experiment.
(A) Schematic plot of the test. The test is considered a pass if the timing of Start is near the timing of Cln3 peak by the specified tolerance value. The Cln3 profiles are from the real data. Open circles denote the raw data; solid lines are the smoothing splines. (B–C) Test failure percentage. Grey bars indicate the failure percentage and red bars indicate the percentage of cells whose Cln3 peak value is more than 20% larger than Cln3 at Start, for all cells (B) and cells with G1 longer than 30 min (C).
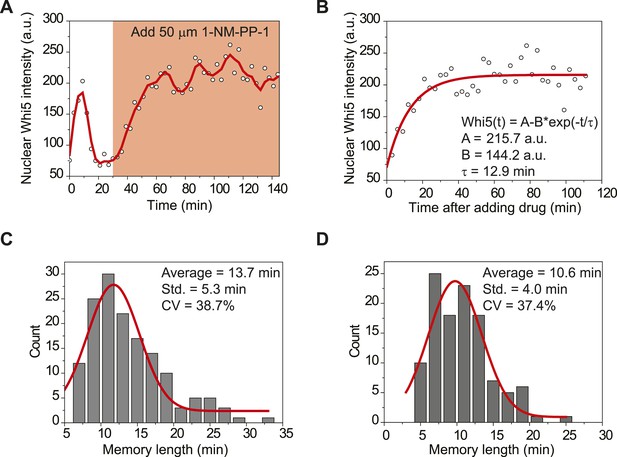
Measuring the memory length through Whi5 nuclear entry when Cdk1 is inactivated.
(A) Dynamics of nuclear Whi5 intensity in a representative single cell (YCT2016). 50 μM 1-NM-PP-1 was added to the medium to inhibit Cdk1 activity at 30 min. Open circles are the raw data; solid line is the smoothing spline. (B) Fitting Whi5 dephosphorylation rate and the memory length from Whi5 nuclear entry when Cdk1 is inactivated. Open circles are from A. Solid line is the least squares best fit of the equation Whi5(t) = A − B × exp(−t/τ), where 1/τ denotes Whi5 dephosphorylation rate and τ is the memory length (Supplementary file 1A). (C–D) The distributions of the memory length in mother (C) and daughter (D) cells, respectively. The red solid lines are Gaussian envelops of the distributions.
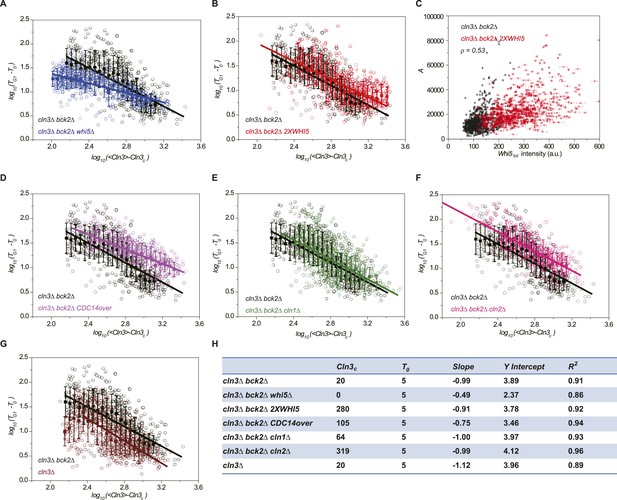
Whi5 acts as the integrator of Cln3-Cdk1 activity.
Correlations between G1 length and the average Cln3 fluorescence in log–log scale in (A) cln3Δ bck2Δ whi5Δ strain (Strain YCT2008) (blue), (B) cln3Δ bck2Δ 2XWHI5 strain (Strain YCT2007) (red), (D) cln3Δ bck2Δ CDC14 overexpressing strain (Strain YCT2009) (purple), (E) cln3Δ bck2Δ cln1Δ strain (Strain YCT2013) (green), (F) cln3Δ bck2Δ cln2Δ strain (Strain YCT2014) (pink) and (G) cln3Δ BCK2+ strain (YCT2015) (wine), in comparison with in cln3Δ bck2Δ strain (Strain YCT2003) (black). Data are also plotted in linear scale in Figure 3—figure supplement 1. <Cln3> represents the average Cln3 fluorescence in G1; T0 and Cln3c are fitted from Figure 3—figure supplement 1 as described in ‘Materials and methods’. The solid lines are linear fits of the binned data (filled squares). Error bars indicate standard deviation. Cln3c, T0, Slope, Y Intercept and R2 of the linear fit for each strain are summarized in (H). (C) The integral A (calculated as TG1 mutiplied by average Cln3 intensity in G1) vs Whi5tot fluorescence intensity in cln3Δ bck2Δ (black) and cln3Δ bck2Δ 2XWHI5 (red) strains. Each dot represents a measurement from one cell cycle event; error bars indicate standard deviation ρ signifies Pearson correlation coefficient.
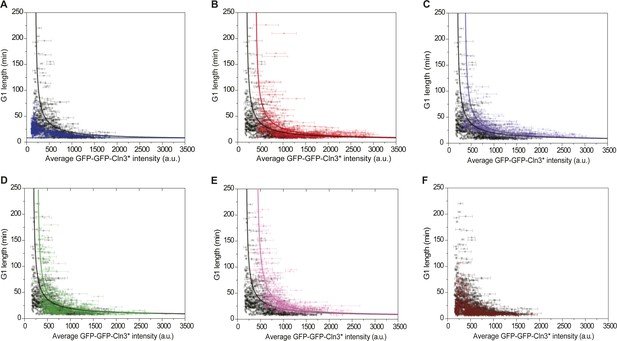
Correlations between G1 length and the average Cln3 fluorescence in (A) cln3Δ bck2Δ whi5Δ strain (Strain YCT2008) (blue), (B) cln3Δ bck2Δ 2XWHI5 strain (Strain YCT2007) (red), (C) cln3Δ bck2Δ CDC14 overexpressing strain (Strain YCT2009) (purple), (D) cln3Δ bck2Δ cln1Δ strain (Strain YCT2013) (green), (E) cln3Δ bck2Δ cln2Δ (Strain YCT2014) (pink) and (F) cln3Δ BCK2+ strain (YCT2015) (wine), in comparison with in cln3Δ bck2Δ strain (Strain YCT2003) (black).
Each dot represents a measurement from one cell cycle event; error bars indicate standard deviation; solid lines are simulation results from the ODE model (Supplementary file 1B). Note we didn't fit the result of cln3Δ BCK2+ strain, because the mechanism of Bck2 related Start triggering is not clear.
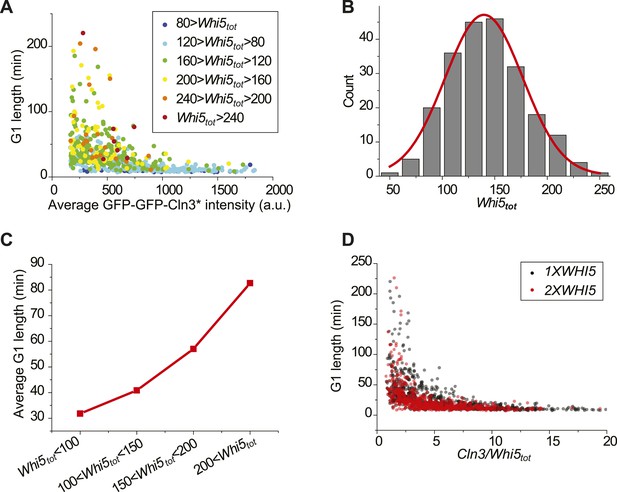
Variation of G1 length in low Cln3 region is due to the variance in Whi5tot.
(A) The Cln3-TG1 correlation indicating Whi5tot intensity of each cell by colors. (B) Whi5tot distribution of cells with Cln3 intensity less than 300. Red line is the Gaussian envelop. (C) Average G1 length binned by Whi5tot intensity of cells with Cln3 intensity less than 300. (D) G1 length verse Cln3 intensity divided by Whi5tot intensity in 1XWHI5 (YCT2003) and 2XWHI5 (YCT2007) strains.
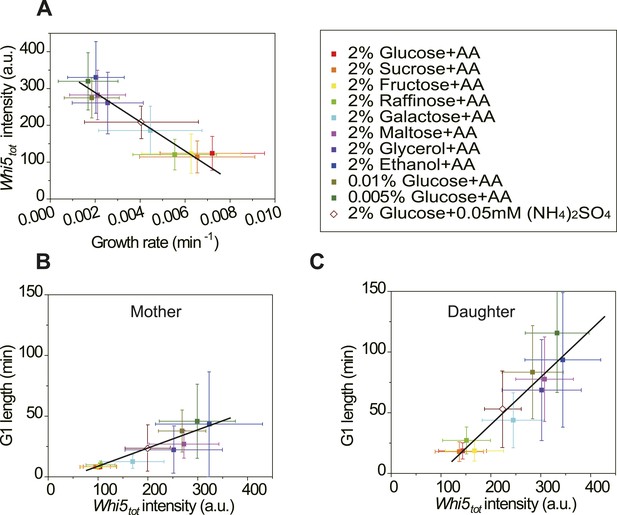
Cells modulate Whi5tot concentration to coordinate cell division with nutrient conditions.
(A) Whi5tot intensity is negatively correlated with growth rate in various nutrient conditions (Strain YCT2001). (B–C) G1 length is positively correlated with Whi5tot intensity for mothers (B) and daughters (C) (Strain YCT2001) in different nutrient conditions. Each color represents one nutrient condition as in A. Each dot in A–C is calculated from all cells in that nutrient condition. (Single-cell data are presented in Figure 4—figure supplement 4,5; statistic results are summarized in Figure 4—source data 1) Black straight lines are guide lines; error bars indicate standard deviation.
-
Figure 4—source data 1
(A)The growth rates and Whi5tot intensities in different nutrient conditions.
(B) The Whi5tot intensities and G1 lengths in different nutrient conditions.
- https://doi.org/10.7554/eLife.03977.022
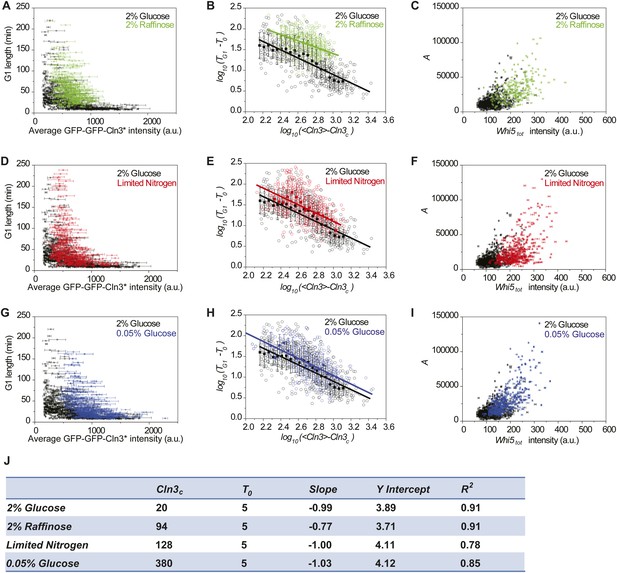
The correlations between Cln3 concentration, Whi5tot intensity and G1 length in suboptimal nutrient conditions.
(A, D and G) The correlations between G1 length and the average Cln3 fluorescence in 2% raffinose (A), limited nitrogen (D) and 0.05% glucose (G), comparing with in 2% glucose (Strain YCT2003). Each dot represents a measurement from one cell cycle event; error bars indicate standard deviation. (B, E and H) The correlations in A, D and H are plotted in log–log scale in (B), (E) and (H), respectively (open circles). The solid lines are linear fits of the binned data (filled squares). Error bars indicate standard deviation. Cln3c, T0, Slope, and Y Intercept are the fitting parameters as described in ‘Materials and methods’ and summarized in (J). Note the linear fit for 2% raffinose is not very meaningful, because its Cln3 only varies in a small range. (C, F and I) The corresponding correlations between the integral A and Whi5tot fluorescence intensity in 2% raffinose (C), limited nitrogen (C) and 0.05% glucose (F), comparing with in 2% glucose. Each dot represents a measurement from one cell cycle event; error bars indicate standard deviation. Each nutrient condition was shown in a separate panel in Figure 4—figure supplement 2.
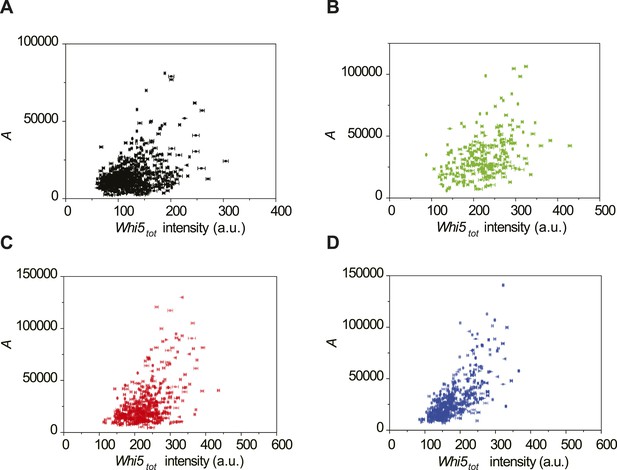
The correlations between the integral A and Whi5tot fluorescence intensity in 2% glucose (A), 2% raffinose (B), limited nitrogen (C) and 0.05% glucose (D) (Strain YCT2003).
https://doi.org/10.7554/eLife.03977.024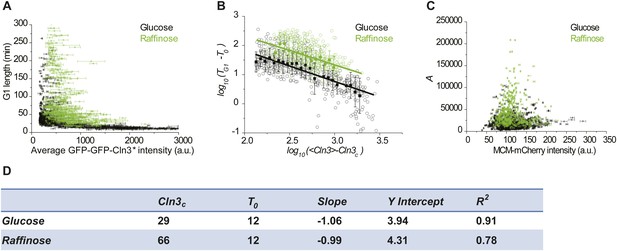
The inverse correlations in raffinose comparing with in glucose with the MCM marker as a measure of G1 length (YCT2010).
(A and B) The Cln3-TG1 correlations with the MCM marker as a measure of G1 length in glucose and raffinose in linear (A) and log–log scale (B). The solid lines in B are linear fits of the binned data (filled squares). Cln3c, T0, Slope, and Y Intercept are the fitting parameters as described in ‘Materials and methods’ and summarized in (D). (C) The correlation between A and MCM-mCherry fluorescence in glucose and raffinose. Error bars indicate standard deviation.
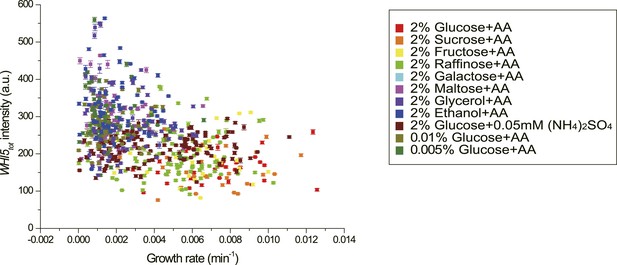
The correlation between growth rate and Whi5tot fluorescence intensity in single cells in different nutrient conditions (YCT2001).
https://doi.org/10.7554/eLife.03977.026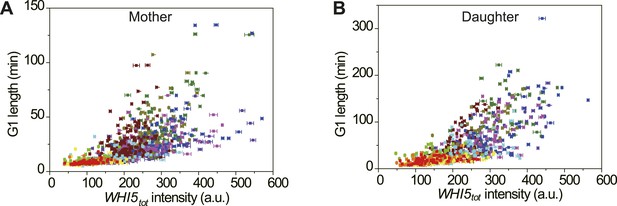
The correlation between Whi5tot intensity and G1 length in single cells in different nutrient conditions in mother (A) and daughter (B) cells (YCT2001).
Legend is the same as in Figure 4—figure supplement 4.
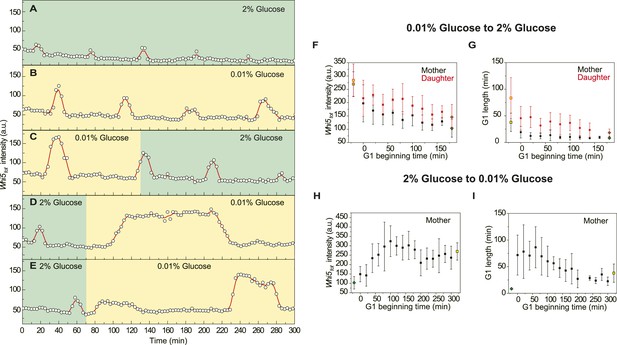
The adaptation of Whi5 and G1 length after nutrient transitions (YCT2001).
(A–E) The raw data of Whi5 dynamics and G1 length from representative single cells in constant 2% (A), constant 0.01% (B), 0.01–2% (C) and 2–0.01% (D–E) glucose media; 0.01% glucose period is marked as yellow and 2% glucose period is marked as green, respectively. (F–I) Whi5tot intensity and G1 length adapt with time after nutrient transitions; yellow squares are steady states in 0.01% glucose whereas green diamonds are steady states in 2% glucose; time 0 is the timing of nutrient transition; G1 beginning time is the time when a G1 phase (not necessarily the first G1 phase) begins after nutrient transition. There is no data for daughter cells in 2–0.01% glucose condition, because only few daughter cells budded in the first 300 min after transition.
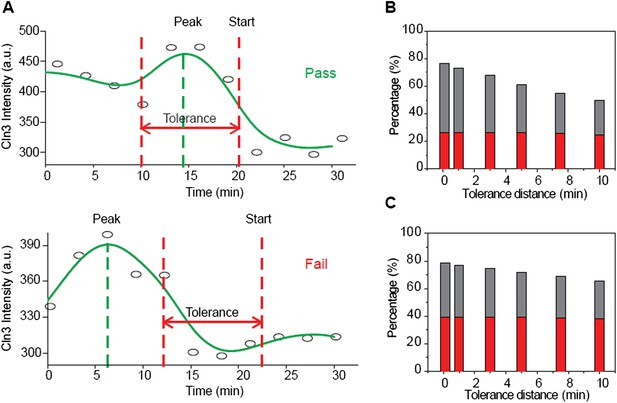
The Instantaneous Model fails with the Cln3 profiles measured in experiment. (A) Schematic plot of the test. The test is considered a pass if the timing of Start is near the timing of Cln3 peak by the specified tolerance value. The Cln3 profiles are from the real data. Open circles denote the raw data; solid lines are the smoothing splines. (B-C) Test failure percentage. Grey bars indicate the failure percentage and red bars indicate the percentage of cells whose Cln3 peak value is more than 20% larger than Cln3 at Start, for all cells (B) and cells with G1 longer than 30 min (C).
Additional files
-
Supplementary file 1
(A) Mathematic model of Whi5 kinetics. (B) ODE model of the Start network.
- https://doi.org/10.7554/eLife.03977.029
-
Supplementary file 2
(A) Yeast Strain list. (B) Plasmid list.
- https://doi.org/10.7554/eLife.03977.030





