RNA-directed remodeling of the HIV-1 protein Rev orchestrates assembly of the Rev–Rev response element complex
Figures
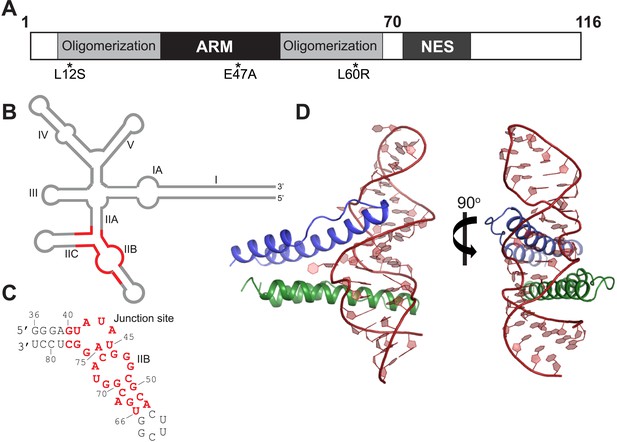
Overall organization and structure of the Rev dimer-RRE complex.
(A) Domain organization of full-length Rev. The protein used to co-crystallize the complex contains residues 1–70 and contains the higher-order oligomerization disrupting mutations, L12S and L60R, and the surface-entropy reducing mutation, E47A. (B) Secondary structure of the RRE (Watts et al., 2009; Bai et al., 2014) with the region used for the co–crystal structure shown in red. (C) Sequence of IIB40 RNA with the region corresponding to the RRE shown in red. The bases are numbered according to older studies (Battiste et al., 1996) for consistency. The RNA contains a terminal UUCG tetra-loop to enhance stability and favor crystallization. See also Figure 1—figure supplement 1 (D) Overall arrangement of the Rev dimer-RNA complex: The RNA (red) is held between the first Rev molecule (blue) bound at IIB and the second Rev molecule (green) bound at the junction site. The N-terminal 11 residues and C-terminal 7 residues of Rev 1–70 are not visible in the structure. See also Figure 1—figure supplement 2, 3.
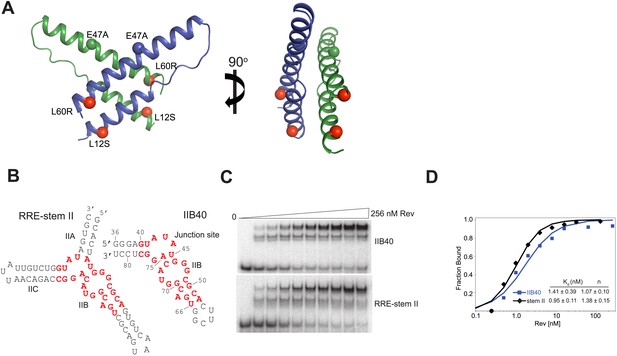
Rev and RRE constructs.
(A) Rev dimer structure (Daugherty et al., 2010b) with the higher-order oligomerization disrupting mutations, L12S and L60R shown in red. Also shown is the surface-entropy reducing mutation, E47A in blue or green. (B) RRE IIB40 is a derivative of the stem II three-helix junction and contains two adjacent Rev-binding sites (red). (C) Gel-shift assays comparing binding of 32P-labeled RRE-stem II or IIB40 to Rev. Free, F, monomer, M and dimer, D complexes are indicated. A doublet/smeared band is observed for the monomer complex with both RNAs and is indicative of conformational heterogeneity. (D) Binding curves calculated from gel-shift assays in (C). Apparent dissociation constants, Kd and Hill coefficient (n) were determined using the equation: Fraction of RNA bound = [Rev]n/(Kdn + [Rev]n) as mean ± s.d. of two replicates.
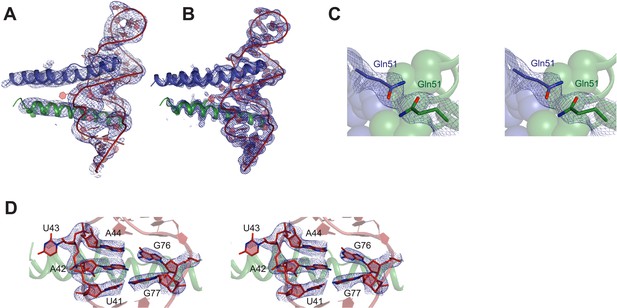
Representative electron density maps.
(A) The final model fit into 5.5 Å density-modified SAD electron density map (contoured at 2.0 σ). (B) 2Fo − Fc electron density map at 3.2 Å after molecular replacement (contoured at 2.0 σ). (C and D) Stereo view of 2Fo − Fc electron density maps located around (C) nucleotides that form the core structure of the junction site (contoured at 2.0 σ) (D) Gln51 which hydrogen bonds across the dimer interface in the RNA-bound form (contoured at 1.0 σ).
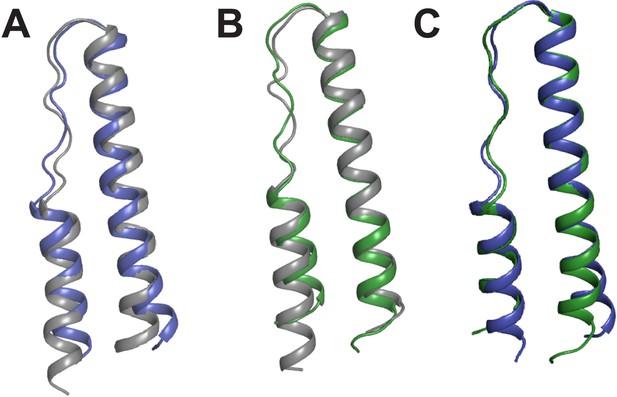
Comparison of Rev monomers.
Superimposition of Rev monomers corresponding to (A) the first Rev molecule in the unbound dimer (grey) and in the dimer-RNA complex (blue) (backbone RMSD = 1.47 Å) (B) the second Rev molecule in the unbound dimer (grey) and in the dimer-RNA complex (green) (backbone RMSD = 1.41 Å) (C) the first (blue) and second Rev (green) molecules from the dimer-RNA structure (backbone RMSD = 0.87 Å).
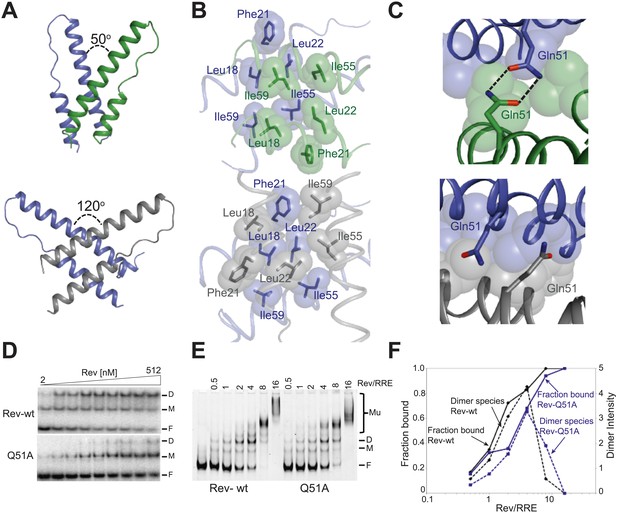
Reorganization of the Rev dimer interface upon RNA binding.
(A) Rev dimer crossing angles differ significantly between RNA-bound (top) and RNA-free (bottom) states (B) Packing of hydrophobic residues at the dimer interface in the RNA-bound (top) and RNA-free (bottom) states. Phe21 is largely excluded from the interface upon RNA binding and, in general, the interface is more loosely packed (Figure 2—figure supplement 1). (C) Gln51 hydrogen bonds across the dimer interface in the RNA-bound conformation (top) but is unable to interact in the free state (bottom). (D) Gel shift assays with 32P-labeled IIB40 RNA and Rev 1–70 dimer show that a monomeric Rev-RNA complex accumulates when Gln51 is mutated to Ala and the dimer affinity is reduced ∼30-fold (Figure 4—figure supplement 1). F corresponds to free RNA, M to the Rev monomer-RNA complex, and D to the Rev dimer-RNA complex. (E) Gel shift assays with the 234-nt RRE and full-length Rev, visualized using SYBR Green II staining, show reduced accumulation of dimer species and a modest loss of binding affinity with the Gln51Ala mutant (Figure 2—figure supplement 1), quantified in panel (F). Mu corresponds to Rev multimer-RNA complexes.
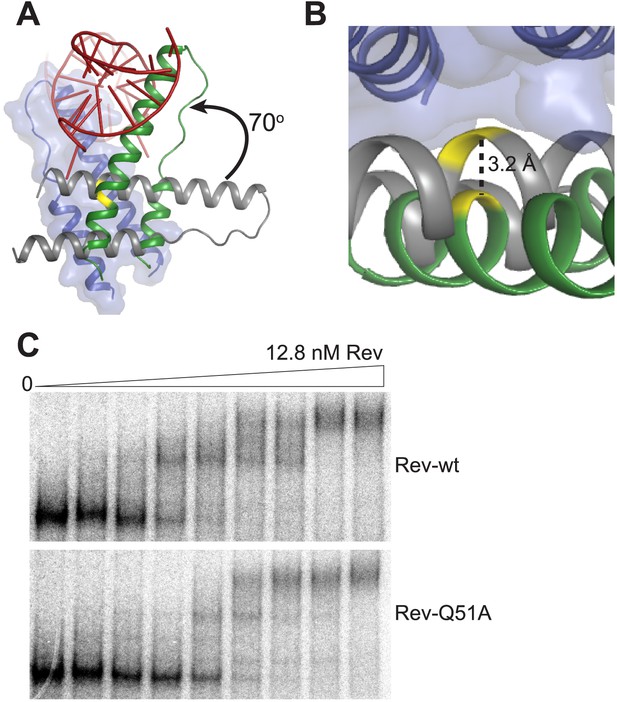
Rev interactions in the free and RNA-bound structures.
(A) Model illustrating the 70° rotation that one subunit of the unbound Rev undergoes in order to contact the RNA at the junction site, pivoting around Ile55 (yellow) (B) View of the dimer interface from the top (RNA-binding side) showing that the second Rev molecule in the RNA-bound dimer is situated ∼3.2 Å further away from the first Rev subunit than in the RNA-free arrangement (C) Gel-shift assay using 32P-labeled full-length RRE (234 nucleotides) and full-length Rev illustrating the modest loss of binding affinity upon mutating Gln51 to Ala. Apparent Kd and n are: wild-type Rev Kd = 145 pM, n = 2.1; Q51A Rev Kd = 280 pM, n = 1.7.
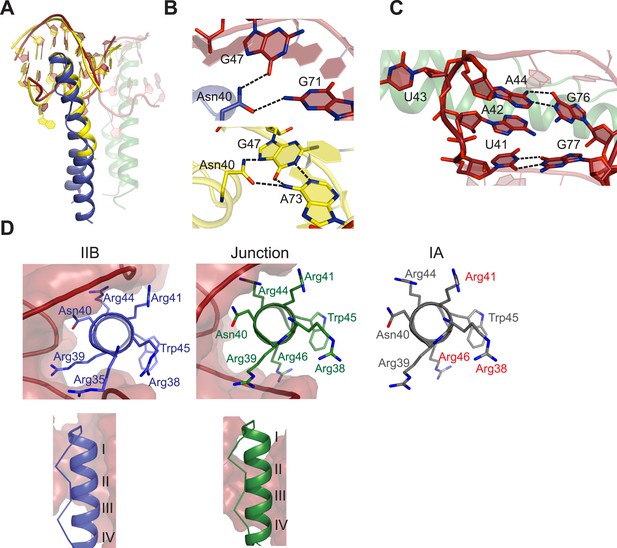
Features of Rev-RNA recognition.
(A) The ARM peptide-IIB NMR structure (yellow) (Battiste et al., 1996) and the Rev dimer-IIB40 structure are nearly superimposable (backbone RMSD = 1.27 Å). (B) Contacts made by Asn40 from the first Rev subunit at the IIB RNA site are different in the Rev dimer-IIB40 (top panel) and ARM peptide-IIB complexes (bottom panel). See also Figure 3—figure supplement 1. (C) RNA structure at the second junction site shows a G-A base-pair and A42-U43 bulge that widens the major groove to accommodate the RNA-binding helix of the second Rev subunit. See also Figure 3—figure supplement 2, 3. (D) Rev- RNA recognition at the three known binding sites in the RRE, shown in the three top panels as views down the helical axis of the Rev ARM at the IIB site, junction site, and stem IA. The binding residues for stem IA are inferred from alanine mutants (in red) (Daugherty et al., 2008). The two bottom panels show side views from the co–crystal structure indicating that turns 1–3 of the ARM helix contact IIB while turns 2–4 contact the junction site.
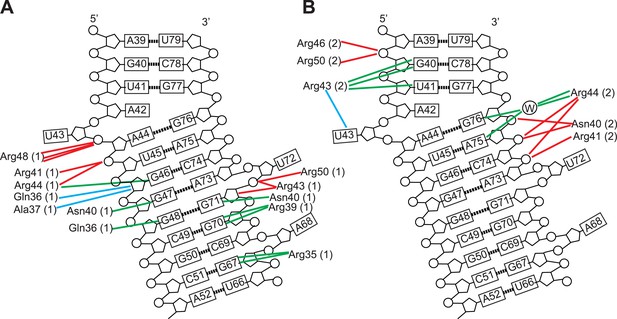
Schematic representation of Rev-RNA contacts at IIB and junction sites.
Schematic representation of Rev-RNA contacts at (A) IIB and (B) the junction site, generated using Nucplot (Luscombe et al., 1997). Red lines indicate hydrogen bonds to the phosphate backbone or sugar, green lines indicate hydrogen bonds to the bases and blue lines indicate Van der Waals contacts.
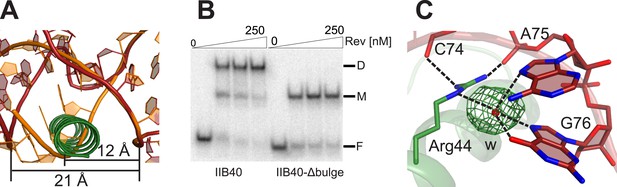
Rev-RNA recognition at the junction site.
(A) Model of IIB40 with a deleted AU bulge from the junction site (orange) superimposed with IIB40 showing that deleting the bulge would result in a steric clash between the RNA in A-form geometry and the second Rev subunit. (B) Gel-shift assay using 32P-labled IIB40 or bulge-deleted RNA and Rev showing an absence of the Rev-dimer-RNA complex with the bulge deleted. (C) Simulated annealing Fo − Fc omit map in green (contoured at 3.0 σ) around the water molecule bridging Arg44 from the second Rev subunit and the RNA junction site.
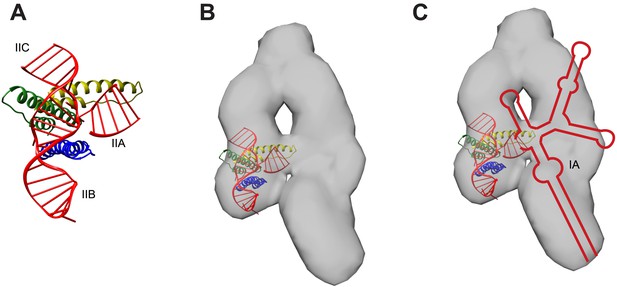
Model for RNA-directed cooperative assembly.
(A) An A-form helix was placed near the junction site to represent the stem IIA helix not present in the crystallized RNA. The second Rev subunit from the crystal structure (green) is oriented to use its higher-order oligomerization surface to recruit a third Rev subunit (yellow) (B) The model generated in (A) was placed within the RRE SAXS envelope (Fang et al., 2013), based on the positioning of sites from the SAXS model. (C) RRE secondary structure from Figure 1B placed within the envelope, indicating how other binding sites might be placed, including the IA site. Figures were generated using UCSF Chimera (Pettersen et al., 2004).
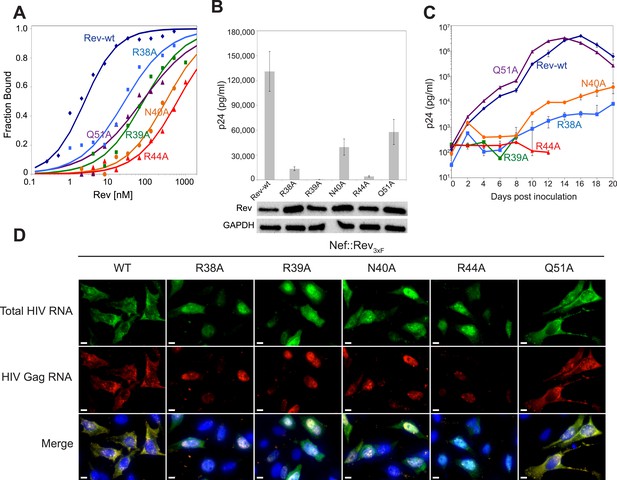
Role of RNA-binding and dimer interface residues in Rev function.
(A) Representative binding curves with Rev 1–70 dimer mutations at positions observed to contact the RNA, calculated from gel shift assays using 32P-labeled IIB40. Apparent dissociation constants, Kd and Hill coefficient (n) are reported in Figure 4—figure supplement 1. (B) p24 production from a Rev-dependent, transiently transfected pCMV GagPol-RRE reporter assay complemented in trans with full length Rev mutants, quantified by ELISA and normalized to both Rev and GAPDH expression levels (representative western blots below show that the mutations do not affect Rev steady state expression levels or stability). Data points are mean ± s.d. of biological triplicates. (C) Viral replication kinetics for HIV-1NL4-3 containing Rev mutants, monitored by p24 release into the culture supernatant. Data points are mean ± s.d. of biological triplicates. See also Figure 4—figure supplement 2 (D) FISH of total HIV RNA or unspliced Gag RNA in the presence of the indicated Rev mutation. Nuclei were stained using DAPI (blue). Nef::Rev3xF denotes infection with virus containing c-terminally 3× flag-tagged Rev in the nef locus. See also Figure 4—figure supplement 3.
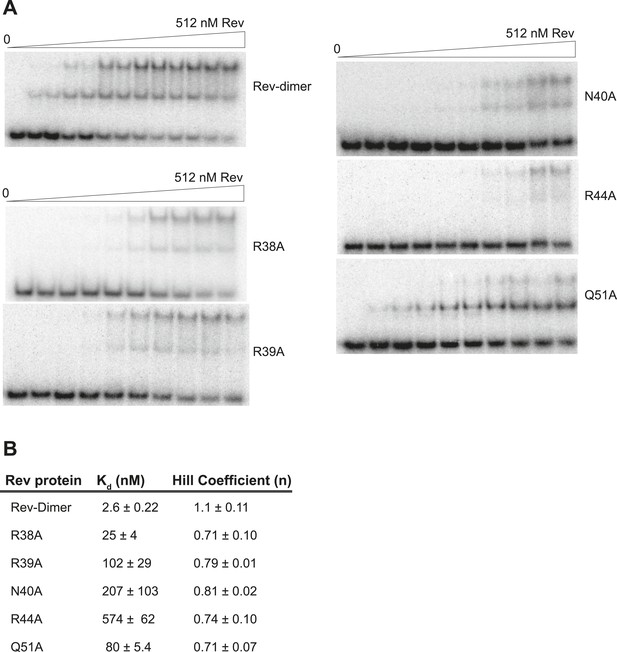
Gel-shift assays with the indicated Rev mutants.
(A) Representative gel-shift assays using 32P-labeled IIB40 and the indicated Rev mutants (B) Apparent binding constants (Kd) and Hill coefficients (n) calculated from fraction of total RNA bound from gel-shift assays using 32P-labeled IIB40 and the indicated Rev mutants. Values reported are mean ± s.d. from two replicates.
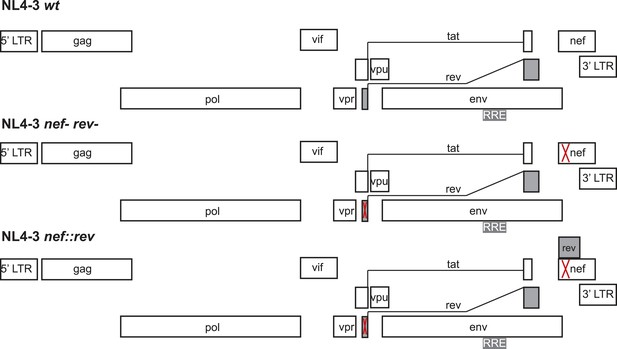
Design of virus constructs.
The endogenous rev locus was deleted by mutating the start codon (ATG) to ACG and introducing a termination codon at residue 23. Both mutations are silent in the overlapping tat open reading frame. The nef locus, dispensable for virus replication ex vivo, was disrupted by introducing SacII and Xba1 restriction endonuclease sites in place of the start ATG codon. Codon-optimized Rev DNA was synthesized (to avoid the introduction of direct nucleotide repeats within the HIV genome that typically results in recombination) with the endogenous Kozak sequence and flanking SacII and XbaI restriction sites to facilitate cloning within the repurposed nef locus.
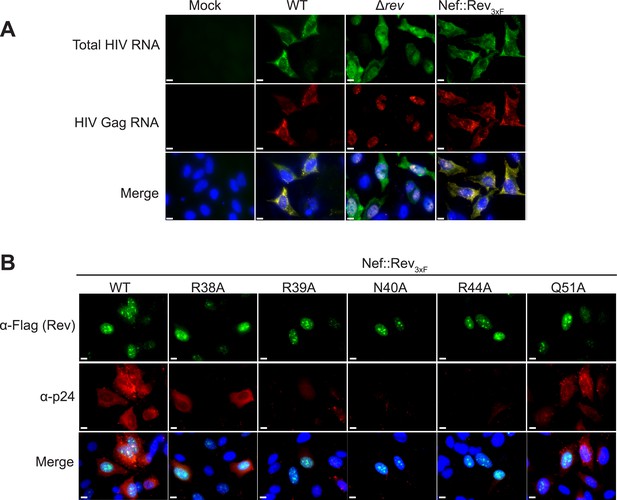
RNA FISH and protein immunofluorescence studies.
(A) FISH of total HIV RNA or unspliced Gag RNA showing that there is no observable phenotypic difference between rev in the endogenous locus and rev engineered into the nef locus. (B) Immunofluorescence staining for 3× flag-tagged Rev and HIV p24 capsid protein, showing that although Rev is expressed and localized in the nuclei, p24 production is attenuated to different degrees in the presence of Rev mutants. Nuclei were stained using DAPI (blue). Nef::Rev3xF denotes infection with virus containing c-terminally 3× flag-tagged Rev in the nef locus.
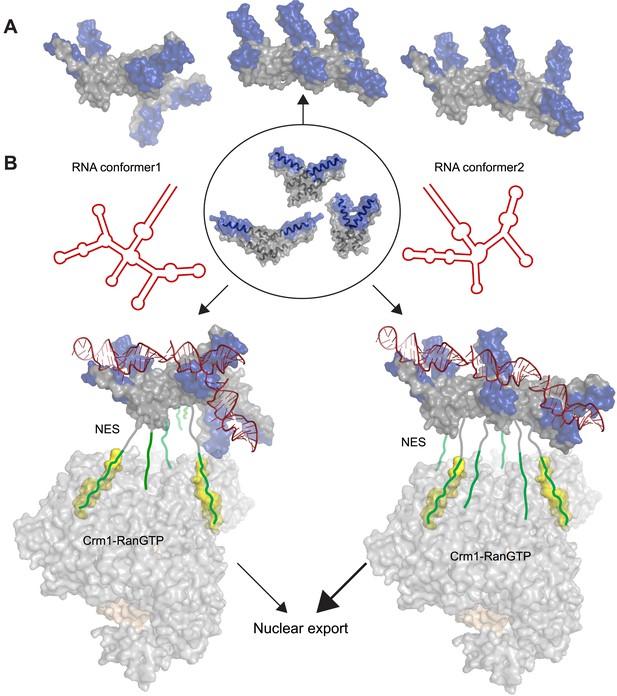
Potential diversity of Rev oligomeric structures and functional implications.
Three types of Rev–Rev interactions observed by crystallography, Rev dimers in the RNA-free (Daugherty et al., 2010b) or RNA-bound states (the current structure) and a Rev dimer using the higher-order oligomer interface (DiMattia et al., 2010) are shown within the circle with Rev in grey and the Rev-ARMs in blue. (A) The higher-order oligomer interface was used to combine Rev dimers in the RNA-free state or RNA-bound state in various arrangements to generate examples of hexamers with different architectures. (B) Models illustrating how changes to the RRE structure (red) can alter the architecture of Rev oligomer. Such changes can alter the Rev-RRE ‘jellyfish’ architecture and spatial distribution of NESs, potentially changing their local effective concentration and avidity for the Crm1 dimer (in grey, with RanGTP in light brown and NES-binding sites in yellow), thereby tuning nuclear export activity. Coordinates of the Crm1-RanGTP dimer are from Booth et al., 2014.
Tables
Diffraction data collection and refinement statistics
| Native | Tungsten derivative | |
|---|---|---|
| Data Collection | ||
| Space group | P 41 3 2 | P 41 21 2 |
| Cell Dimensions | ||
| a, b, c (Å) | 165.3, 165.3, 165.3 | 147.22, 147.22, 199.44 |
| α, β, γ (°) | 90, 90, 90 | 90, 90, 90 |
| Resolution (Å) | 45.85 − 3.2 (3.314 − 3.2)* | 49.34 − 5.55 (5.75 − 5.55) |
| Redundancy | 25.9 (15.9) | 15.2 (13.6) |
| Completeness (%) | 99.72 (98.36) | 99.6 (96.5) |
| I/σI | 21.56 (1.74) | 13 (1.8) |
| R-meas | 0.138 (2) | 0.151 (1.52) |
| Refinement | ||
| Resolution (Å) | 45.85 − 3.2 (3.31 − 3.2) | |
| No. reflections | 342771 (20153) | |
| Rwork/Rfree | 19.3 (27.7)/21.1 (30.7) | |
| No. atoms | ||
| RNA | 849 | |
| Protein | 862 | |
| Ligand/Ion | 5 | |
| Water | 1 | |
| B-factors | ||
| RNA | 113.9 | |
| Protein | 115.3 | |
| Ion | 153.8 | |
| Water | 112.7 | |
| RMS deviations | ||
| Bond lengths (Å) | 0.002 | |
| Bond angles (°) | 0.39 | |
-
*
Statistics for the highest-resolution shell are shown in parentheses.





