A stress assembly that confers cell viability by preserving ERES components during amino-acid starvation
Figures
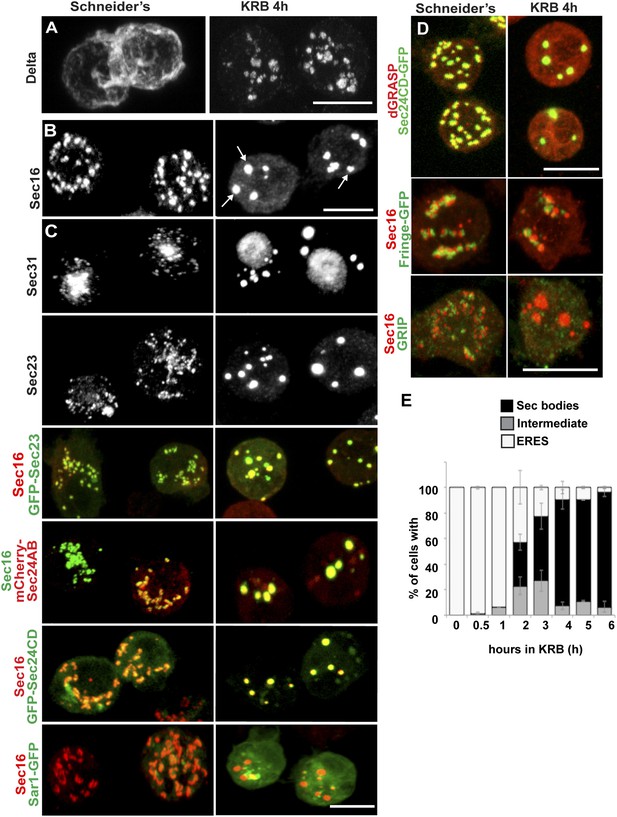
Amino-acid starvation induces the formation of a novel stress assembly in Drosophila S2 cells.
(A) Immunofluorescence (IF) visualization of Delta-myc (using an anti-Delta antibody) in S2 cells in Schneider's (normal growth conditions) or incubated with Krebs Ringer Bicarbonate buffer (KRB) for 4 hr (amino-acid starvation). Note that in Schneider's Delta reaches the plasma membrane whereas it is retained intracellularly in starved cells. (B) IF visualization of endogenous Sec16 in Drosophila S2 cells grown in Schneider's and incubated in KRB for 4 hr. Note the formation of Sec bodies (arrows). (C) IF visualization of Sec31, Sec23 and co-visualization of GFP-Sec23, mCherry-Sec24AB, Sec24CD-GFP, Sar1-GFP with Sec16 in S2 cells in Schneider's and KRB for 4 hr. (D) IF co-visualization of dGRASP/Sec24CD-GFP, GRIP/Sec16, and Fringe-GFP/Sec16 in S2 cells grown in Schneider's and incubated in KRB for 4 hr. (E) Kinetics of Sec body formation in S2 cells incubated in KRB over indicated time (up to 6 hr) expressed as the percentage of cells exhibiting ERES, intermediates (see ‘Materials and methods’), and Sec bodies. Scale bars: 10 μm (A–D).
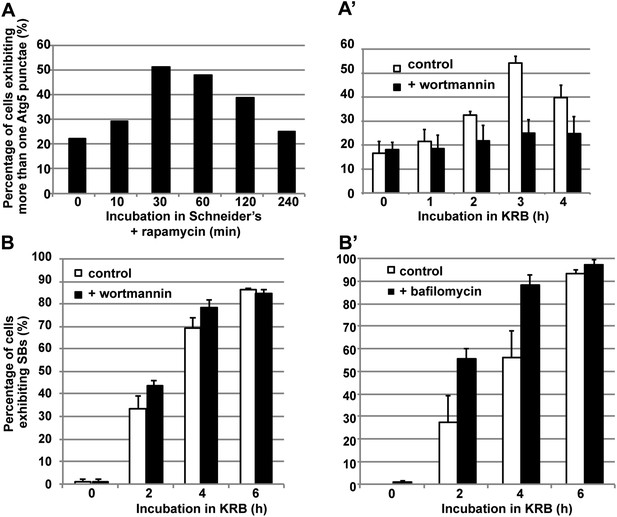
Sec body formation and autophagy.
(A–A′) Quantification of Atg5 punctae formation in S2 cells incubated with rapamycin (A) and incubated with KRB with or without wortmannin (A′) for indicated time points. Note that as expected, autophagy (marked by Atg5) is stimulated by rapamycin and starvation (KRB) and inhibited by wortmannin. (B–B′) Quantification of Sec body formation (marked with Sec16) in cells incubated in KRB with and without wortmannin (B) and with and without bafilomycin (B′).
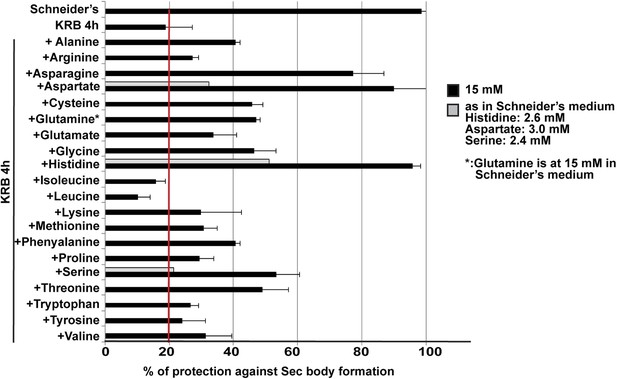
Sec body formation and single amino-acids.
Quantification of the prevention of Sec body formation by specific amino-acids upon incubation in KRB for 4 hr. This is expressed as the percentage of cells exhibiting the normal growth Sec16 localization at ERES. All amino-acids are added at 15 mM except for 3 (gray bars) that were also tested at their concentration in Schneider's medium. Note that at 15 mM, histidine, aspartate, and asparagine significantly decreases Sec body formation.
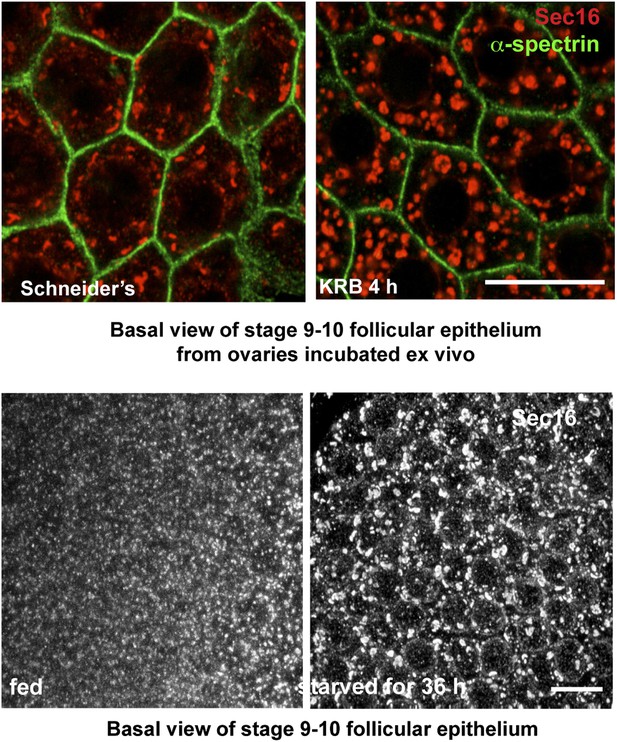
Sec body formation in vivo.
Projection of four equatorial confocal planes of Sec16 and Spectrin (D) in the follicular epithelium covering an egg-chamber from an ovary dissected from a virgin fly fattened for 3 days and incubated ex-vivo in KRB for 4 hr, and from an ovary dissected from a 36-hr starved virgin female. Note that in both cases, Sec16 is found in large punctae reminiscent of Sec bodies. Scale bars: 10 μm.
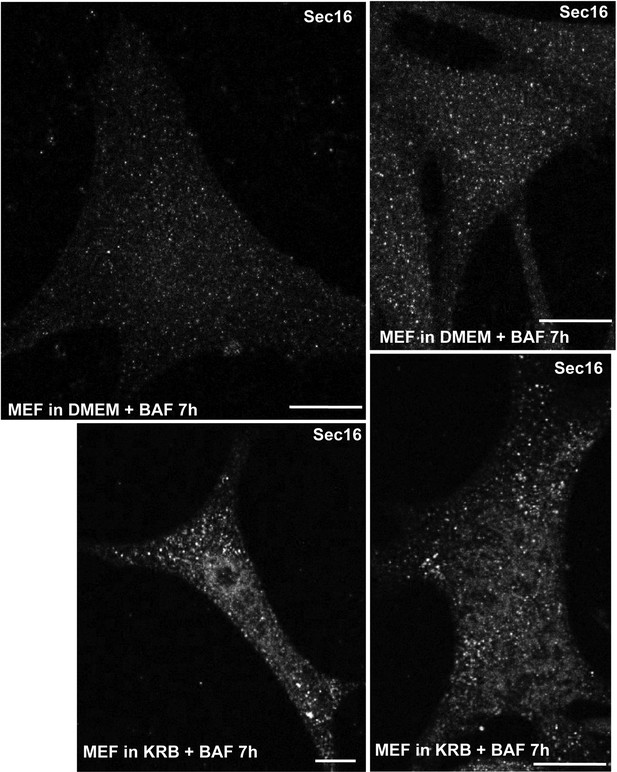
Sec body formation and mammalian cells.
IF visualization of Sec16 in immortalized MEFs incubated in growth medium (DMEM) or KRB plus bafilomycin for 7 hr. Note that Sec16 is remodeled into larger structures. Scale bar: 10 μm.
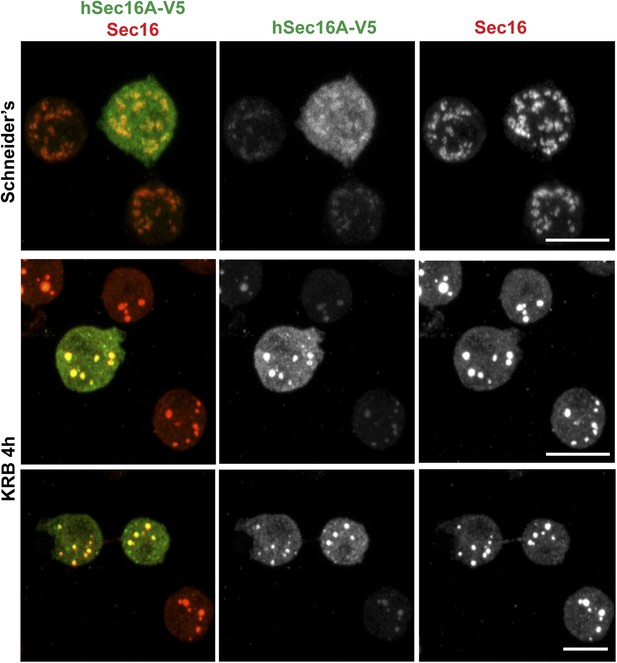
Human Sec16A is incorporated to Sec bodies in starved Drosophila S2 cells.
IF visualization of human Sec16A-V5 transfected in Drosophila S2 cells. Note that it partially localizes to ERES in fed cells but is efficiently incorporated in Sec bodies upon starvation. Scale bar: 10 μm.
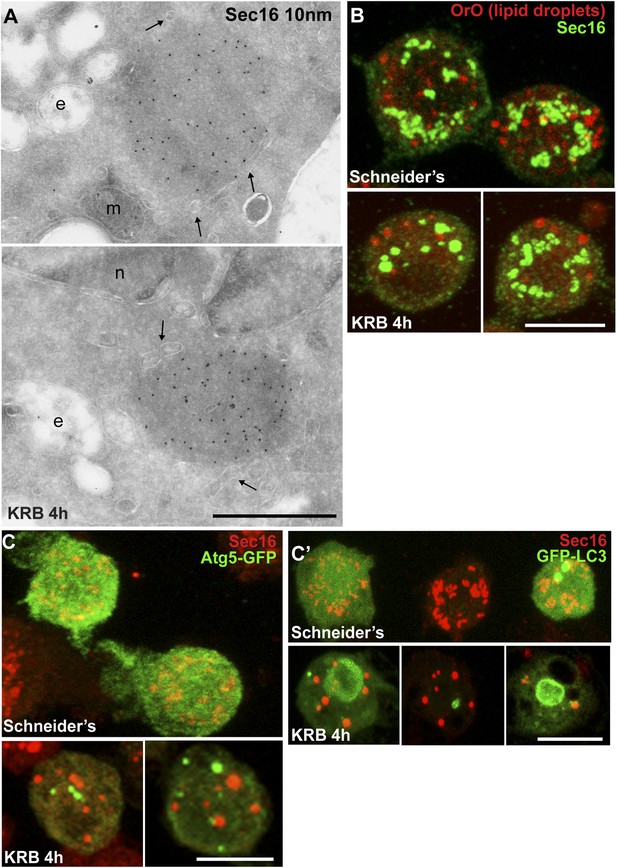
Sec bodies are non-membrane bound structures.
(A) Immuno-electron microscopy (IEM) visualization of Sec16 (10 nm colloidal gold) in Sec bodies in ultrathin sections of S2 cells incubated in KRB for 4 hr. Arrows point to membrane in close proximity of Sec bodies. E, endosomes; n, nucleus; m, mitochondria. (B) Visualization of Sec bodies (Sec16, green) and lipid droplets (marked by oil-red-O, red). Note that 95% of Sec bodies do not co-localize with lipid droplets. (C–C′) Visualization of Sec bodies (Sec16, red) and Atg5-GFP punctae (C) and GFP-Atg8 (C′) after 4 hr starvation. Note that 82% of Sec bodies do not co-localize with Atg5 or Atg8 punctae. Scale bars: 500 nm (A); 10 μm (B, C).
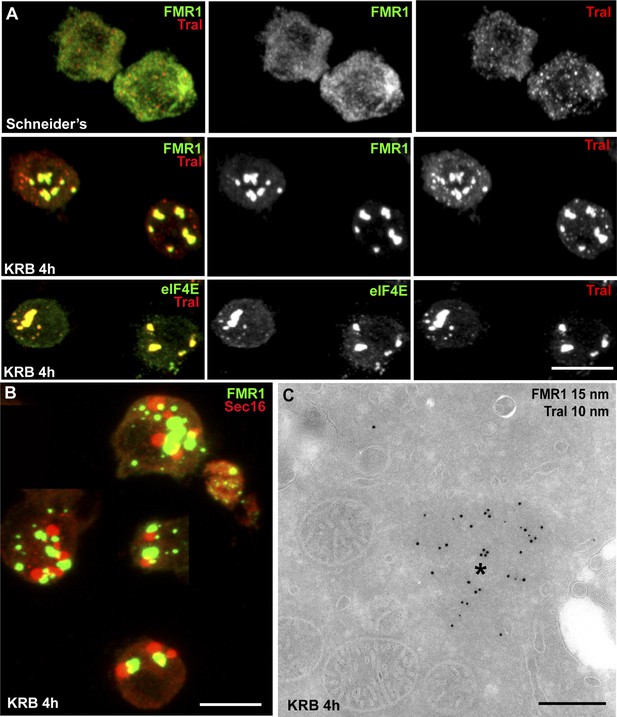
Sec bodies are distinct from Stress Granules and P-bodies.
(A) IF visualization of endogenous FMR1, eIF4E (green), and Tral (red) in cells growing in Schneider's and incubated in KRB for 4 hr. Note that upon starvation, Stress Granules (FMR1, eIF4E) form and P-Bodies (Tral) enlarge to co-localize in SG/PBs. (B) IF visualization of Sec16 (Sec bodies) and FMR1 (SG/PB) in cells incubated in KRB for 4 hr. Sec bodies and SG/PBs are distinct structures but have a spatial relationship. (C) IEM visualization of FRM1 and Tral in ultrathin sections of S2 cell incubated with KRB for 4 hr. Note that the Tral and FMR1 positive SG/PBs (asterisk) are clearly different from Sec bodies (Figure 2A). Scale bars: 10 μm (A, B); 500 nm (C).
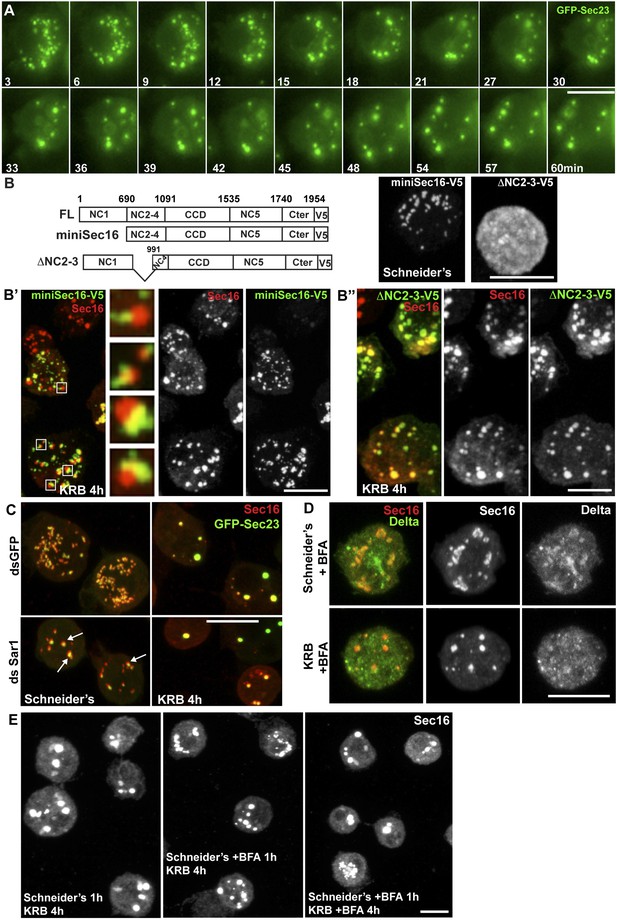
Sec bodies form at ERES but COPII- and COPI-coated vesicle formation is not required.
(A) Stills of a GFP-Sec23 time-lapse video of a cell incubated in KRB (t = 0) for 60 min showing Sec body formation. (B–B″) IF visualization of miniSec16-V5 (B, B′) and ΔNC2-3-Sec16-V5 (B, B″) in S2 cells incubated in Schneider's (B) and KRB for 4 hr. Endogenous Sec16 is in red. Note that the cup-shaped ER forms a cradle for Sec bodies (B′, insert). (C) IF visualization of Sec16 and GFP-Sec23 in mock and Sar1-depleted S2 cells grown in Schneider's and incubated in KRB for 4 hr. Note that the ERES are enlarged in Sar1-depleted cells (arrows) and Sec bodies form in both conditions to the same extent. (D) IF visualizations of Sec16 and Delta-myc in S2 cells incubated with brefeldin A (BFA) in Schneider's and KRB for 3 hr. Note that Delta transport is inhibited in both cases as Delta is retained intracellularly. (E) IF visualization of Sec16 in S2 cells grown in Schneider's and incubated in KRB for 4 hr in the presence or absence of brefeldin A (BFA). Note that pre-incubation with the drug does not affect Sec body formation during starvation. Scale bars: 10 μm.
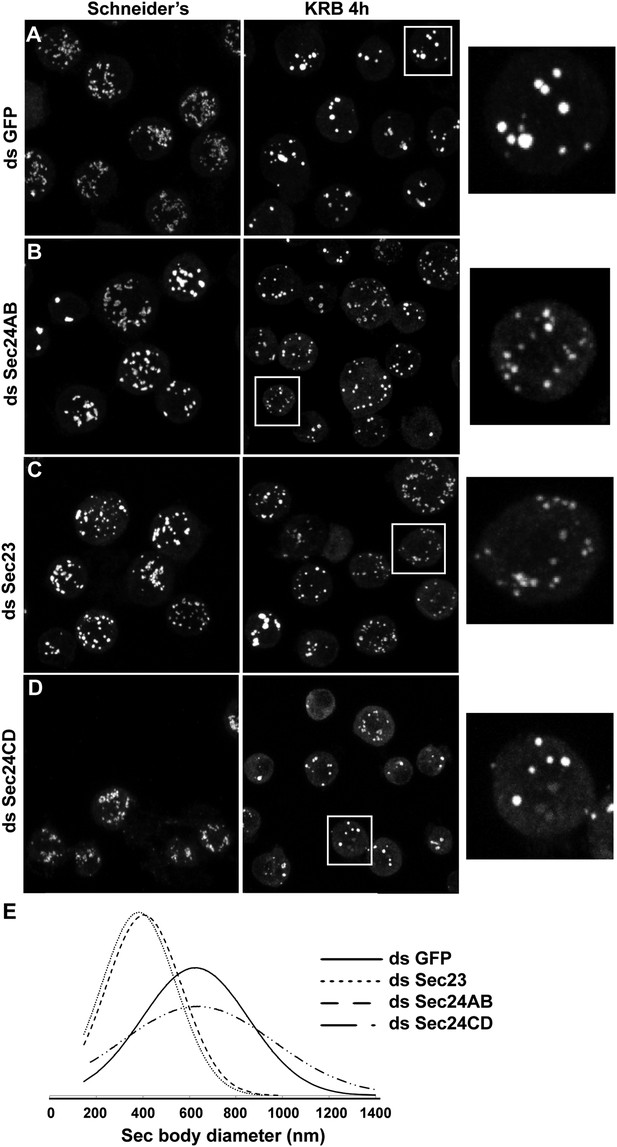
Sec23 and Sec24AB are key factors for Sec body formation.
(A–D) IF visualization of Sec16 in mock (A), Sec24AB (B), Sec23 (C), and Sec24CD (D) depleted S2 cells in Schneider's and incubated in KRB for 4 hr. Note that Sec body formation is inhibited upon Sec24AB and Sec23, but not upon Sec24CD depletion. Boxed areas are shown at higher magnification. (E) Distribution of Sec body size (shown as frequency of observed Sec body diameter) in mock, Sec24AB, Sec24CD, and Sec23-depleted and starved cells (as in A) (dsGFP: 45 cells, 341 Sec bodies; dsSec24AB: 43 cells, 585 Sec bodies; dsSec24CD: 35 cells, 245 Sec bodies; dsSec23: 36 cells, 504 Sec bodies). Note that the Sec body's mean diameter decreases by 1.8-fold upon Sec24AB and Sec23 depletion and that Sec bodies are twice as many. Scale bars: 10 μm.
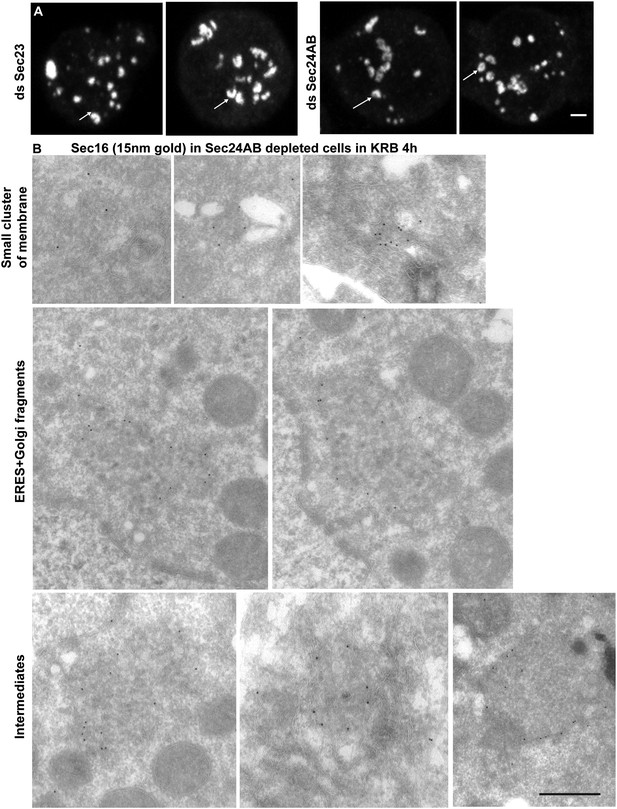
The smaller structures generated in starved Sec24AB-depleted cells are not Sec bodies.
(A) Sec16 positive horseshoe shape structures observed by IF. B: Gallery of Sec16 positive IEM profiles: smaller clusters of membrane, larger ones probably corresponding to ERES mixed with Golgi fragments, intermediate structures (round shape but with membrane in their core and of smaller size than typical Sec bodies). Scale bars: 2 μm (A) and 500 nm (B).
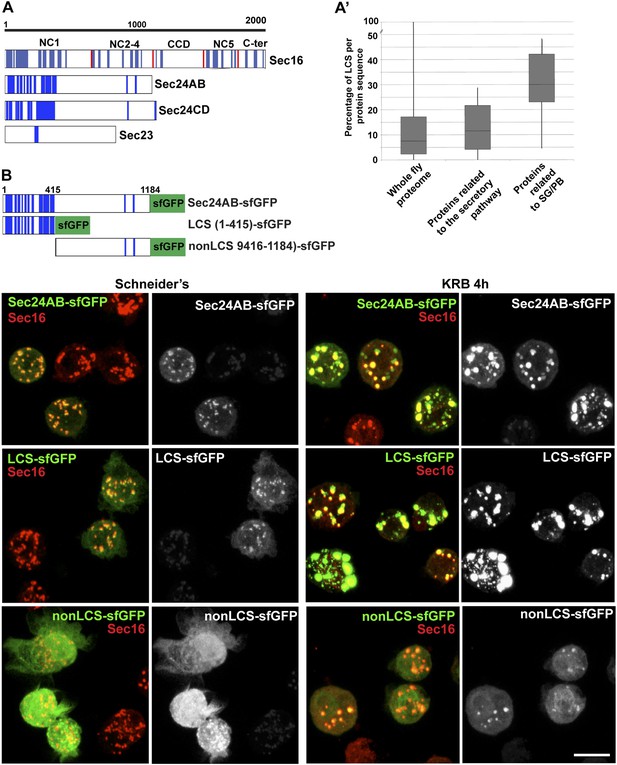
Sec body proteins contain low complexity sequences that are necessary for Sec body recruitment.
(A–A′) Schematic representation of the Low Complexity Sequences (blue bars) in Sec16, Sec24AB, Sec24CD, and Sec23 (A). The red bars mark the boundaries of the Sec16 domains. Genome wide analysis of Low Complexity Sequence (LCS) in the Drosophila proteome, in proteins related to the secretory pathway and proteins related to Stress Granules/P-bodies (A′). B: IF localization of sfGFP-tagged full-length Sec24B, Sec24AB LCS, and Sec24AB nonLCS in S2 cells in Schneider's and KRB for 4 hr, together with endogenous Sec16 (red). Scale bars: 10 μm.
-
Figure 6—Source data 1
Table showing the LCS content of proteins related to Stress Granules and P-bodies as well as the early secretory pathway (related to Figure 6A,A′).
- https://doi.org/10.7554/eLife.04132.016
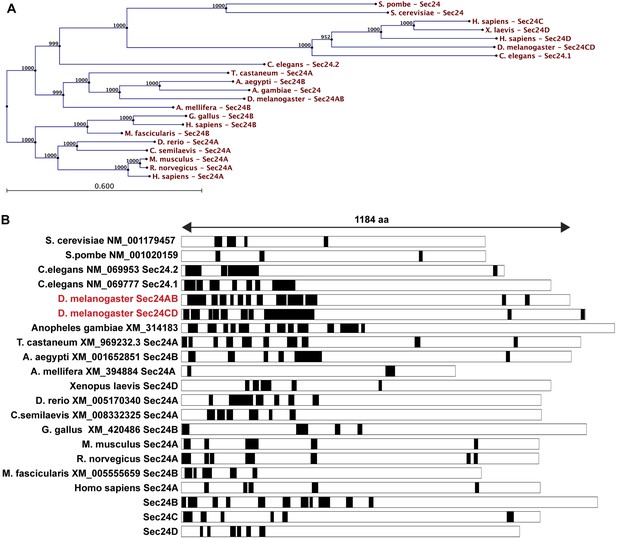
LCS content of Sec24 in different species.
(A) A restricted phylogenic tree of the Sec24 sequences analyzed in (B) Note that S. pombe and S. cerevisiae Sec24 are similar to each other and to H. sapiens Sec24C and D, and D. melanogaster Sec24CD. However, they are distant to D. melanogaster Sec24AB, H. sapiens Sec24A and B. Furthermore, D. melanogaster Sec24AB is distant to D. melanogaster Sec24CD. (B) LCS analysis and schematics in Sec24 sequences of different organisms. Note that most sequences contain a significant percentage of LCS in the N-terminal third of the protein with the exception of S. pombe, A. melifera and G. gallus (related to Figure 6A).
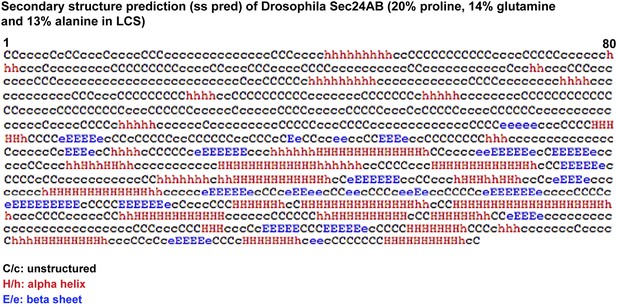

Secondary structure prediction of Drosophila Sec24AB using HH pred.
C/c denotes the unstructured, H/h the alpha helices, and E/e, the beta sheets. Note their absence in the 405 amino-acids of the N-terminus corresponding to LCS.
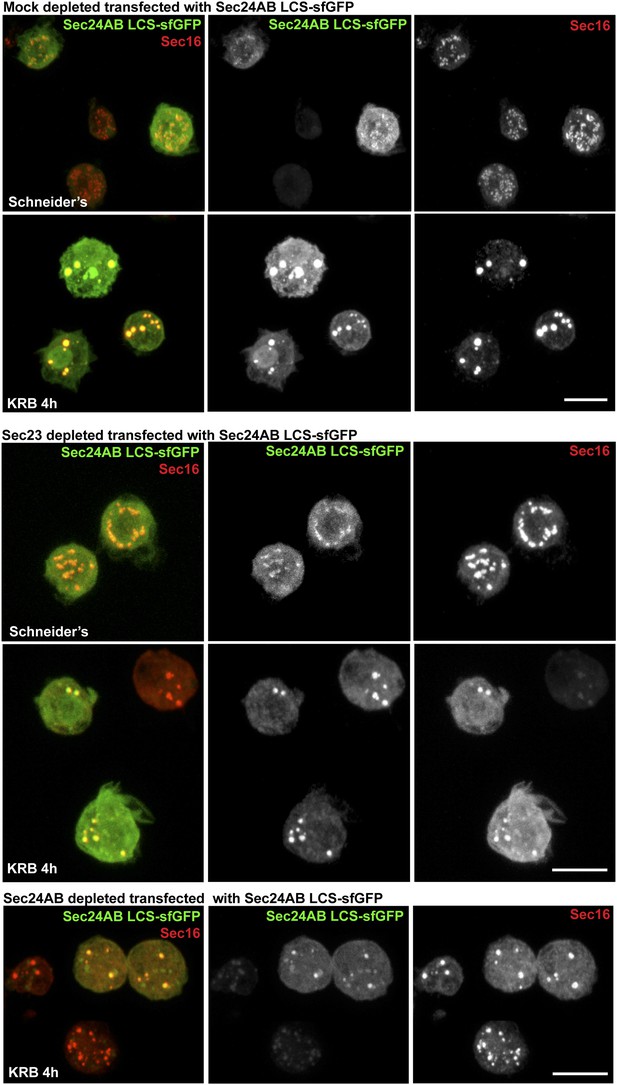
Sec24AB LCS is not sufficient to drive Sec body formation.
S2 cells were depleted of endogenous Sec24AB. When starved (KRB), this resulted in the formation of small structures (as in Figure 5). The transfection of Sec24AB LCS-sfGFP in these depleted cells did not rescue the formation of Sec bodies. Scale bars: 10 μm.
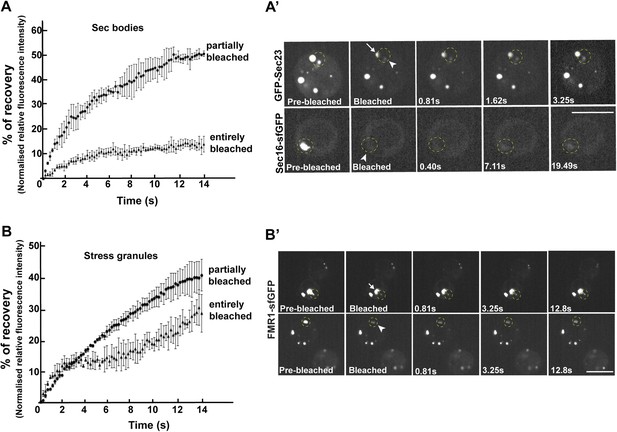
Sec bodies have FRAP properties consistent with liquid droplets.
(A–A′) Percentage of fluorescence recovery after photobleaching (FRAP) over time of individual Sec bodies, marked by GFP-Sec23 (Video 2), and ΔNC1-Sec16-sfGFP (Video 3) in S2 cells incubated in KRB for 4 hr. The triangles in B show the FRAP of Sec bodies that have been entirely bleached (n = 3, arrowheads in B′). The circles show the FRAP of Sec bodies (n = 3, arrows in B′) that have been partially bleached. The dashed circles in B′ indicate the Sec bodies that have been entirely bleached and assessed in stills taken from Video 2 and 3. (B–B′) Percentage of fluorescence recovery after photobleaching (FRAP) over time of individual Stress Granules marked by FMR1-sfGFP (Video 4 and 5) in S2 cells incubated in KRB for 4 hr. The triangles in C show the FRAP of Stress Granules that have been entirely bleached (n = 3, arrowheads in C′). The circles show the FRAP of Stress Granules (n = 3, arrows in B′) that have been partially bleached. The dashed circles in B′ indicate the Stress Granules that have been bleached and assessed in stills taken from Video 4 and 5. Scale bars: 10 μm.
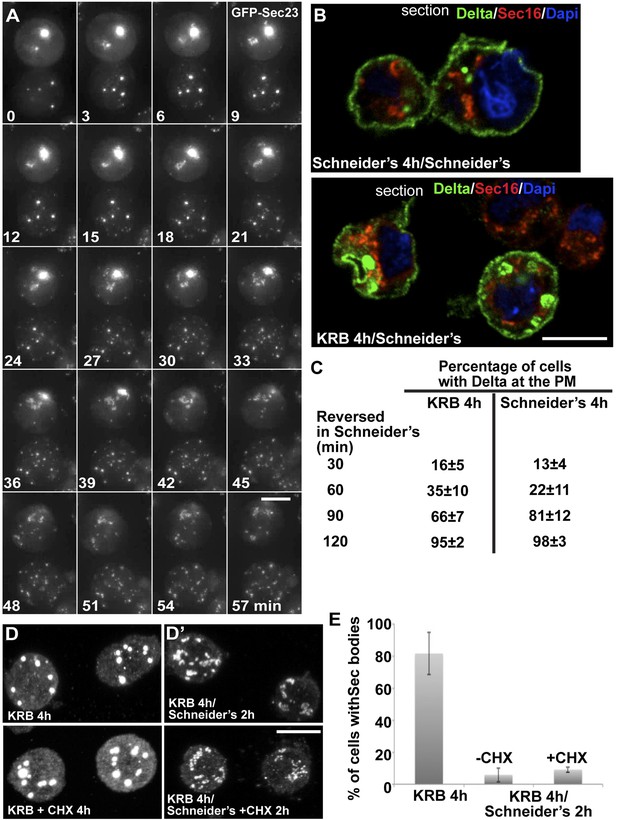
Sec bodies are functionally reversible and act as reservoir for ERES components during starvation.
(A) Stills of a GFP-Sec23 time-lapse video (Video 6) of two cells recovering in Schneider's after 4 hr in KRB (t = 0, up to 60 min). Note that Sec bodies are reversed into ERES. (B) IF localization of Delta in cells that were starved (KRB) or not (Schneider's) followed further incubation in Schneider's for 2 hr. (C) Quantification of the percentage of cells with Delta at the plasma membrane in cells that were either starved (KRB) or not (Schneider's) followed by reversion in Schneider's. Delta was induced for 30, 60, 90, and 120 min while cells were reverted in Schneider's. (D–D′) IF visualization of Sec16 in cells starved in KRB supplemented or not with cycloheximide (CHX, B), and in starved cells further incubated in Schneider's supplemented or not with CHX (C′). Note that neither Sec body formation nor reversal is affected by the presence of CHX. (E) Quantification of the Sec body reversal as described in B′ expressed as the percentage of cells exhibiting Sec bodies.
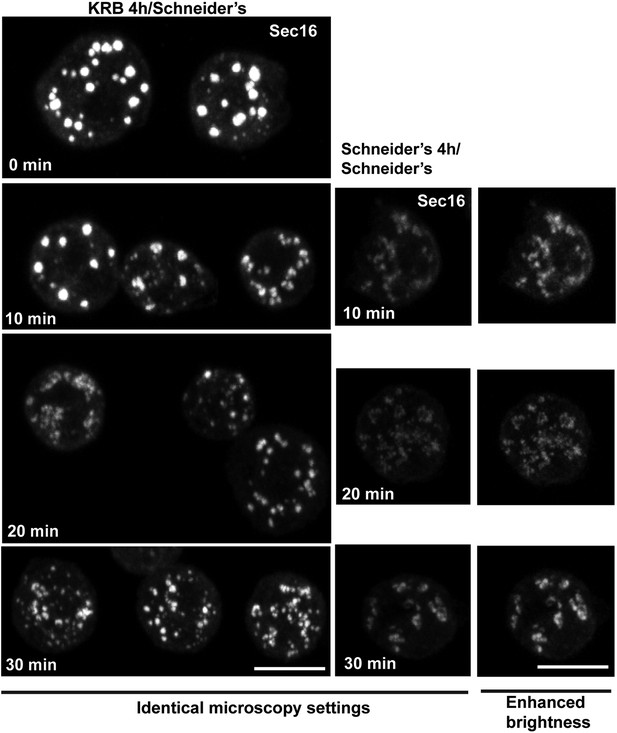
IF localization of Sec16 in cells recovering in Schneider's for 10–30 min after 4 hr in KRB.
At 30 min, 90% of ERES are rebuilt. Note that the intensity of fluorescence of Sec16 is higher than in the cells that were maintained in Schneider's, reflecting the storage of the ERES components in Sec bodies. Scale bars: 10 μm.
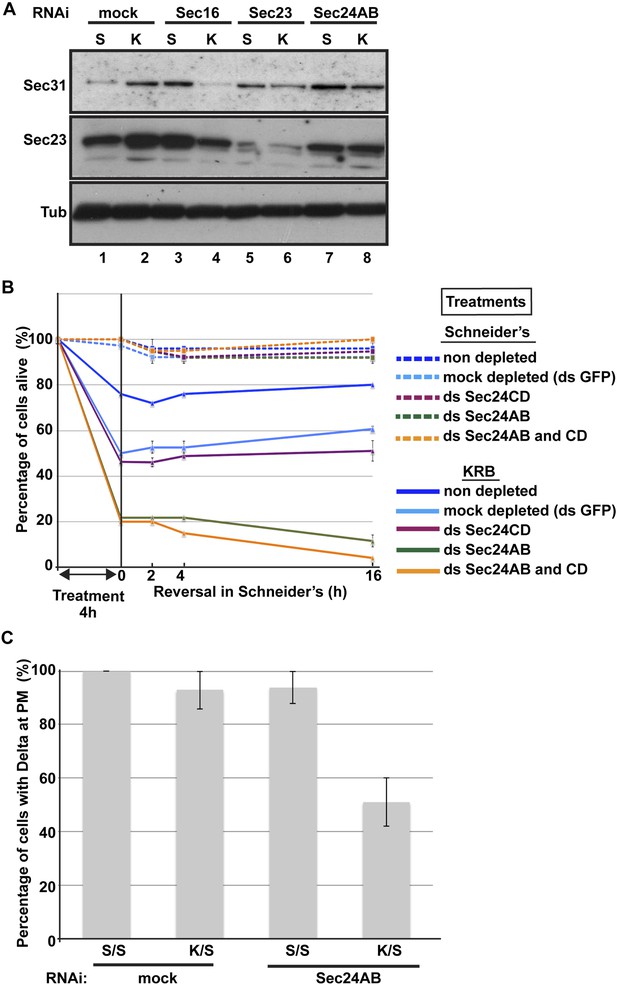
Sec body formation is a pro-survival mechanism.
(A) Western blot of Sec31, Sec23, and tubulin (loading control) of lysates from GFP, Sec16, Sec23, and Sec24AB-depleted cells grown in Schneider's (S) and incubated with KRB for 4 hr (K). (B) Graph of cell viability (expressed as percentage of alive cells). The number of starting cells at t = 0, either non- (control, dark blue lines), mock- (dsGFP, light blue lines), Sec24AB (green lines), Sec24CD (violet lines), and double Sec24 AB and CD (orange lines) depleted, is set at 100%. These cells are incubated in Schneider's (dashed lines) and KRB (solid lines) for 4 hr and further incubated in Schneider's up to 16 hr. Note that the mock depletion (dsGFP, light blue dashed lines) is slightly detrimental to cell survival upon amino-acid starvation when compared to non-depleted (control, dark blue dashed line). (C) Quantification of the percentage of cells with Delta at the plasma membrane in mock-, Sec23, and Sec24AB-depleted cells that were either starved (KRB) or not (Schneider's) followed by reversion in Schneider's. Delta was induced for 90 min while cells were reverted in Schneider's. Error bars in B represent standard error of the mean and in C standard deviation.
Videos
Time-lapse movie of Sec body formation.
GFP-Sec23 time-lapse video of two cells incubated in KRB (t = 0) for 1 hr. One frame was taken every 3 min and the videos are displayed at 7 frame/s (related to Figure 4A).
FRAP of one half-bleached Sec body and one fully bleached.
FRAP video of two GFP-Sec23 positive Sec bodies, one photobleached partially and one entirely, recorded every 10 ms for at least 20 s, and then every minute. The video is displayed at 7 frame/s (related to Figure 7B′).
FRAP of a fully bleached Sec body.
FRAP video of one ΔNC1-Sec16-sfGFP positive Sec body entirely bleached, recorded every 10 ms for at least 20 s, and then every minute. The video is displayed at 7 frames/s (related to Figure 7B′).
FRAP of a fully bleached Stress Granule.
FRAP video of one FMP1-sfGFP positive Stress Granule entirely bleached, recorded every 10 ms for at least 20 s, and then every minute. The video is displayed at 7 frames/s (related to Figure 7B′).
FRAP of a partially bleached Stress Granule.
FRAP video of one FMP1-sfGFP positive Stress Granule partially bleached, recorded every 10 ms for at least 20 s, and then every minute. The video is displayed at 7 frames/s (related to Figure 7B′).
Sec bodies are reversible.
GFP-Sec23 time-lapse video of two cells recovering in Schneider's after 4 hr in KRB (t = 0, up to 66 min). One frame was taken every 3 min and the video is displayed at 7 frame/s (related to Figure 8A).





