Conservation of transcription factor binding specificities across 600 million years of bilateria evolution
Figures
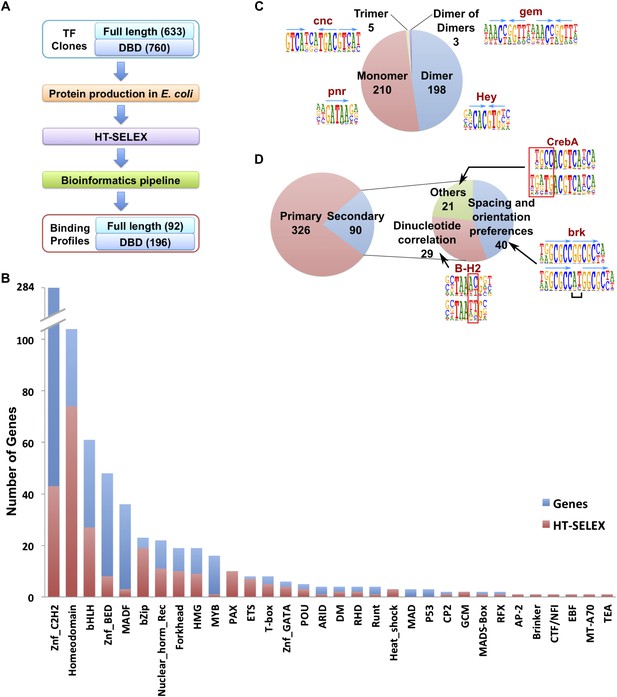
Drosophila HT-SELEX.
(A) Drosophila HT-SELEX pipeline. (B) Coverage of TFs. Number of genes (blue), and number of genes for which we obtained a HT-SELEX model (red) are shown for each TF structural family. Genes that encode more than one domain are counted as members of multiple structural categories. (C) Classification of all binding models into non-repetitive sites (monomer), and sites with two, three or four similar subsequences (dimer, trimer and dimer of dimers, respectively). Logos, for an example, for each type of model are shown, arrows indicate half-sites to highlight multimeric sites. (D) Classification of primary and secondary models (left). Type of secondary model is indicated on the right. Red boxes and black bracket indicate differences between the primary (top) and secondary (bottom) models. Seeds for the generation of the models were identified using the Autoseed algorithm (see ‘Material and methods’ and Figure 1—figure supplements 1–14 for details).
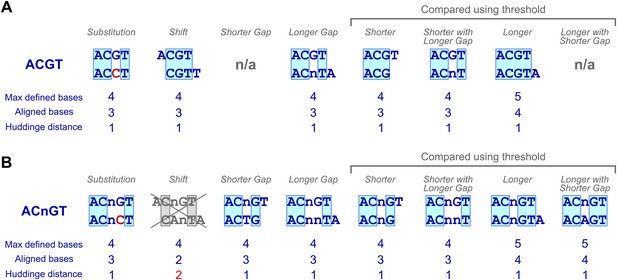
Examples of subsequences that have a Huddinge distance of one.
(A, B) A four base ungapped subsequence ‘ACGT’ and types of subsequences that are at Huddinge distance of one from it (A) and a gapped subsequence with four defined bases ‘ACnGT’ and types of subsequences that are at Huddinge distance of one from it (B). Note that shift of gapped subsequences results in a minimum Huddinge distance of 2 (gray cross). Comparison of subsequences with different number of defined bases (shorter, longer and shorter with gap) requires a threshold to compensate for the fact that shorter subsequences that match perfectly to longer ones are always present at equal or higher numbers than the longer subsequence.
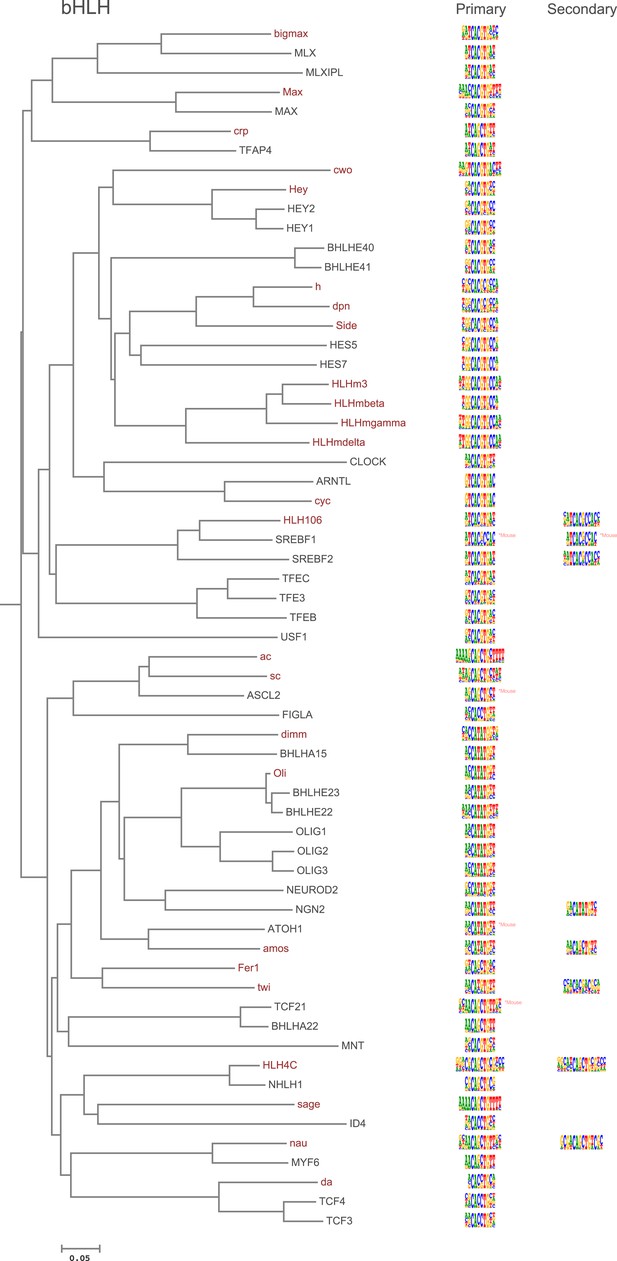
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with bHLH motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif, respectively.
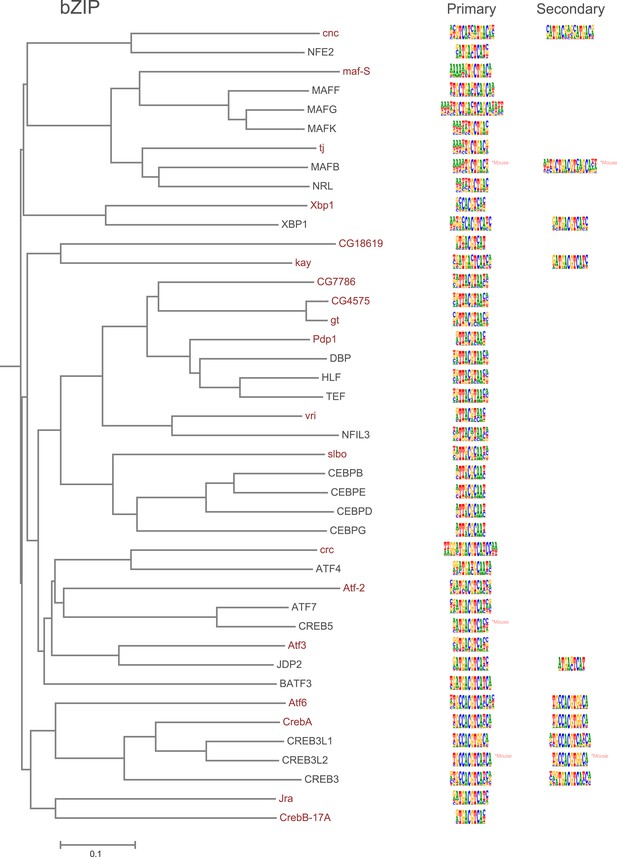
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with bZIP motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
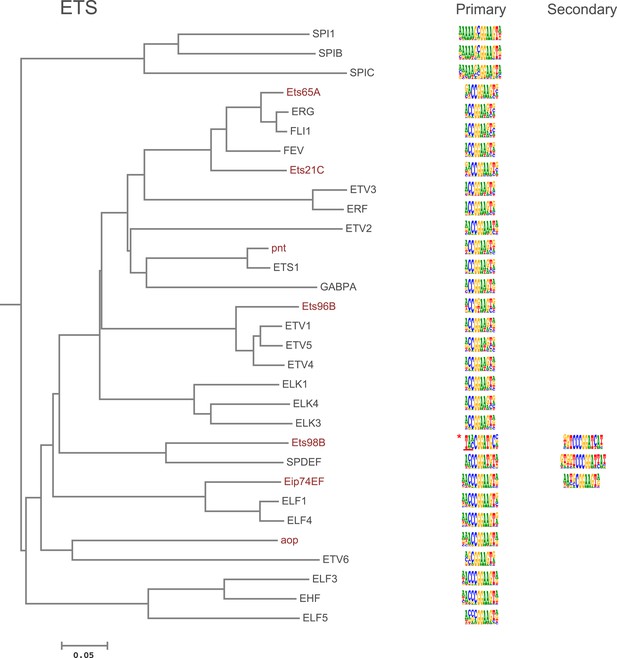
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Ets motifs obtained using HT-SELEX.
Asterisk indicates Ets98B; underlining indicates region where different 5′ sequences were enriched in different experiments. Indicated motif represents sequence obtained from both DBD and full-length protein in one set of experiments. Core and 3′ flank shown were identified in all experiments. Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
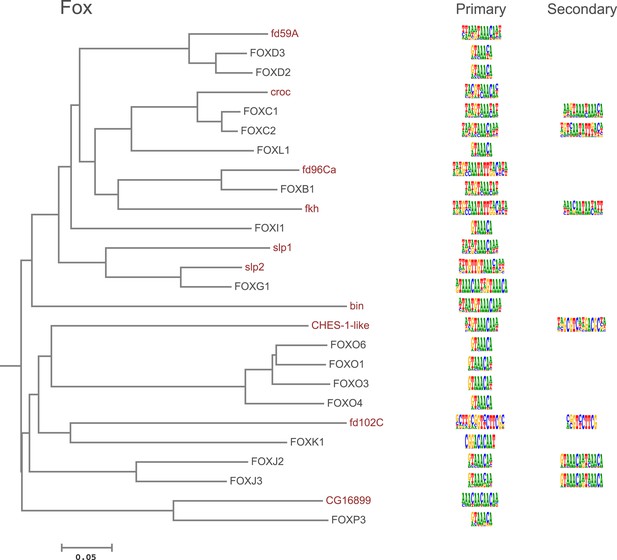
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Fox motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
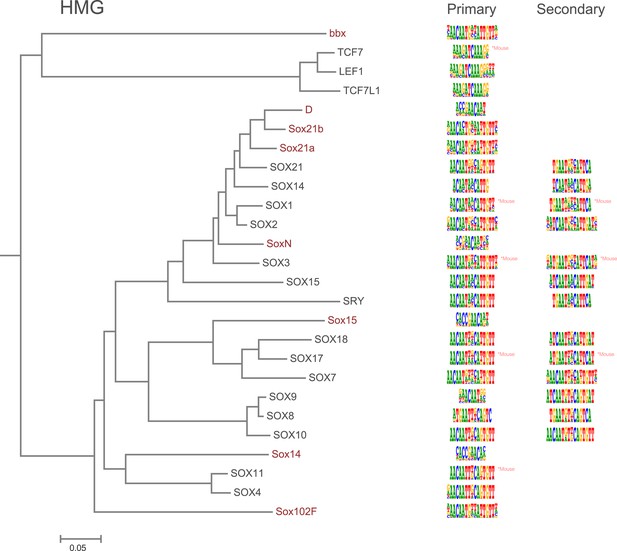
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with HMG motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
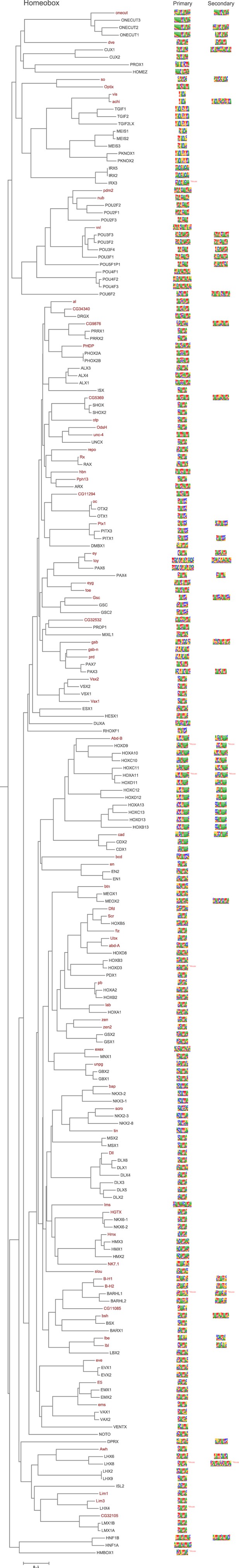

Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Homeobox motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
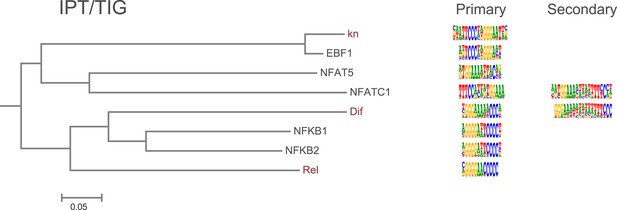
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with IPT/TIG motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
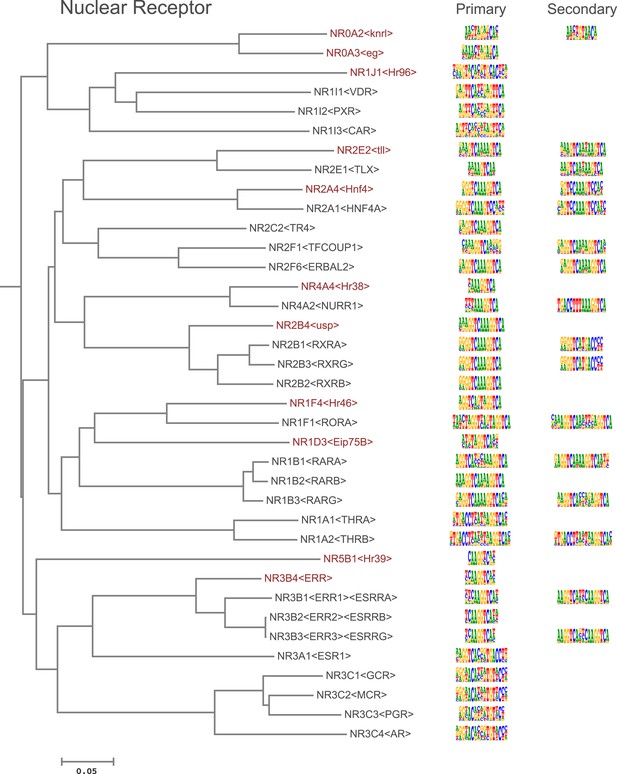
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Nuclear receptor motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
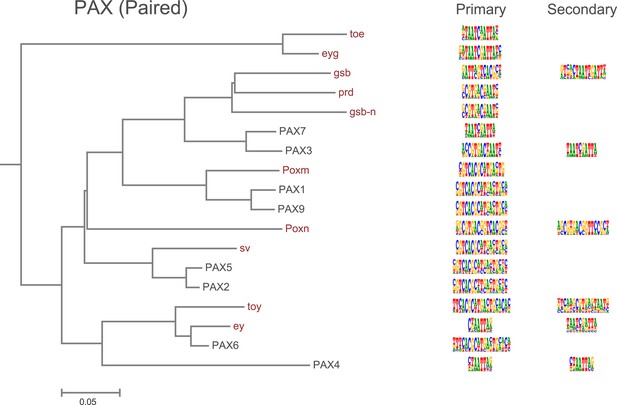
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Pax (based on paired box) motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
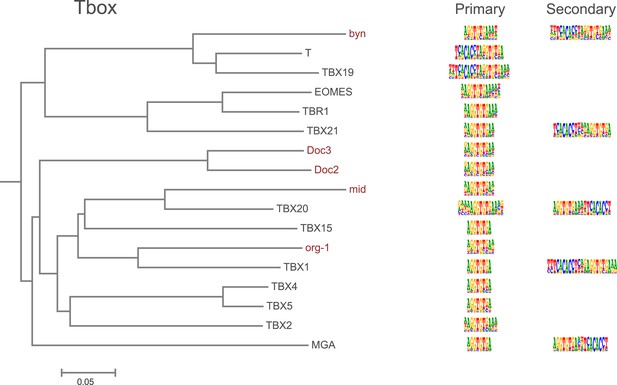
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Tbox motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
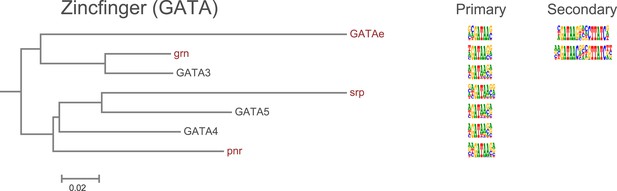
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Zf-GATA motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
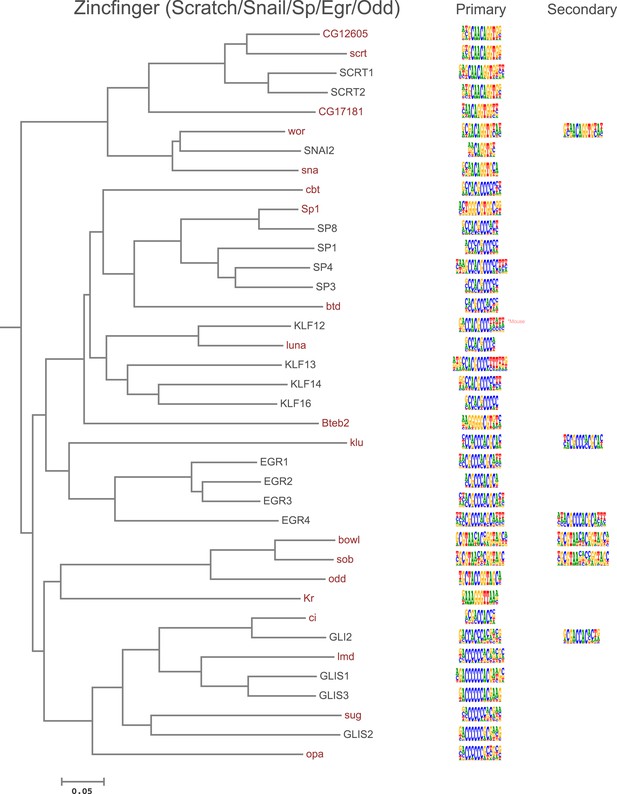
Amino-acid sequence similarity dendrograms for major TF families (human, mouse and Drosophila) annotated with Zf-C2H2 motifs obtained using HT-SELEX.
Drosophila TFs are in red typeface. Left and right columns indicate primary and secondary motif respectively.
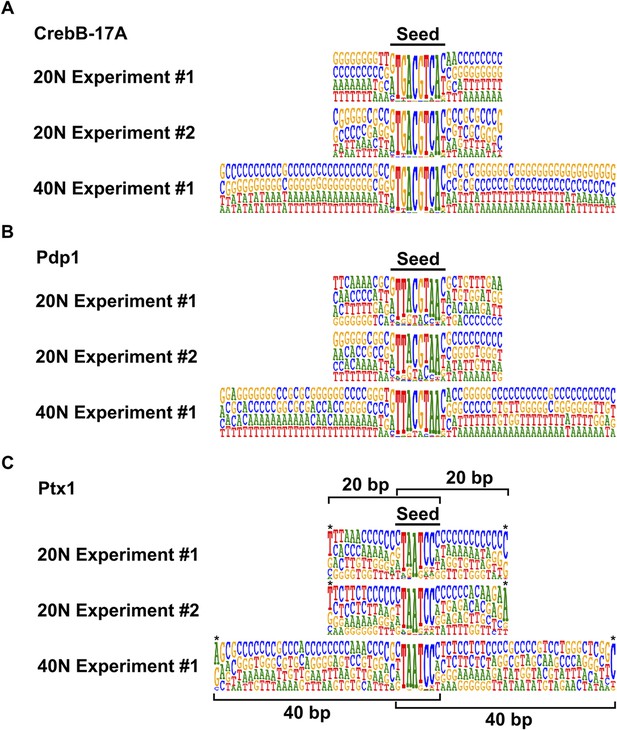
Reproducibility of the HT-SELEX pipeline.
(A) CrebB-17A, (B) Pdp1, and (C) Ptx1. Bar represents the position of the seed. Note that all experiments enrich very similar motifs irrespective of the length of the randomized region, and that flanking sequences adjacent to the seed display no preference. Apparent specificity at the very end of the flanks (asterisks) is an artefact caused by the small number of reads where the seed sequence matches to the very end of the randomized region (position of randomized region in such cases is indicated by brackets).
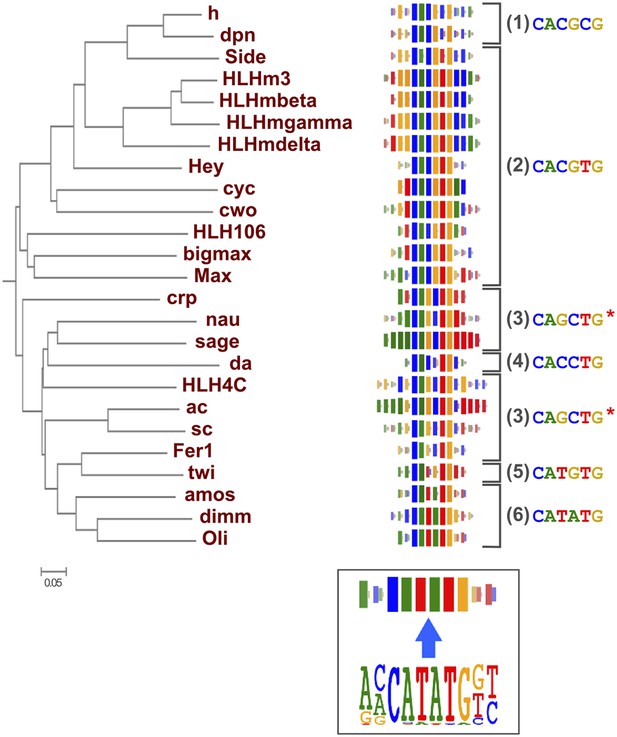
Relationship between similarities of TF DBD amino-acid sequence and binding specificity.
Barcode logos (middle) for bHLH DBDs arranged according to the amino-acid sequence similarity indicate that sequence conservation is predictive of binding specificity. Inset shows an example of a conversion of a sequence logo into a Barcode logo. For each position, the frequency of each base is indicated by the width of the corresponding colored bar; the intensity of color and the height of the bars at each position are determined by the base with the highest frequency. The core recognition sequences are also indicated (right). Note that structurally close TFs recognize the same core sequences and similar flanking sequences (e.g., HLHm3, HLHmgamma, and HLHmdelta). Note also that CAGCTG motif is recognized by two distinct clades (Asterisks). For analysis of other TF families, see Figure 2—figure supplement 1.
-
Figure 2—source data 1
Raw Data for 2D heatmap.
- https://doi.org/10.7554/eLife.04837.019
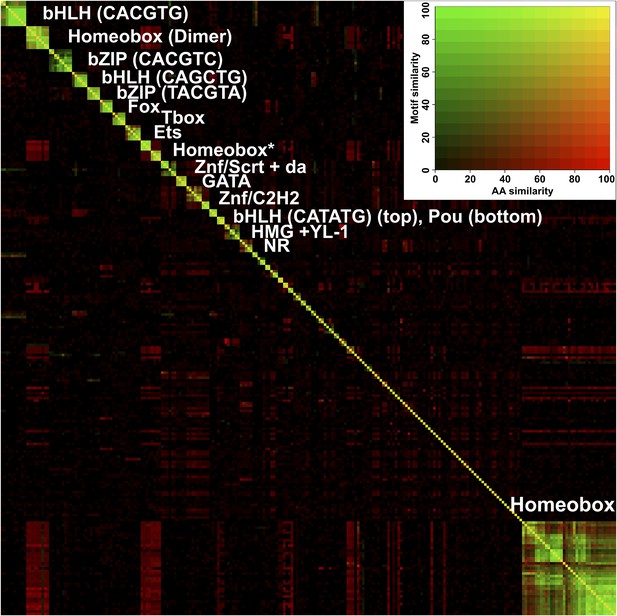
Heatmap showing similarity of binding profiles and amino-acid sequence similarity score (blastp) between all TFs studied.
Rows and columns are ordered by motif similarity measured using all possible 6 bp sequences with variable length gaps in the middle. Subfamilies are indicated by the respective core binding motifs. Homeodomain models containing the canonical (T/C)AATTA site are on bottom right, and models containing TAATCC or TAAA(T/C)G are indicated by an asterisk. Dimeric homeodomain sites (Dimer) are also indicated. Inset shows the two-dimensional color scale, yellow indicates that both motifs and amino-acid sequences are similar. Note that in general, the motif similarity is determined by TF structural family. Note also the red rectangles off the diagonal, indicating that although the homeodomain proteins binding to different motifs have high sequence similarity, the motifs themselves are divergent. See Figure 2—source data 1 for details.
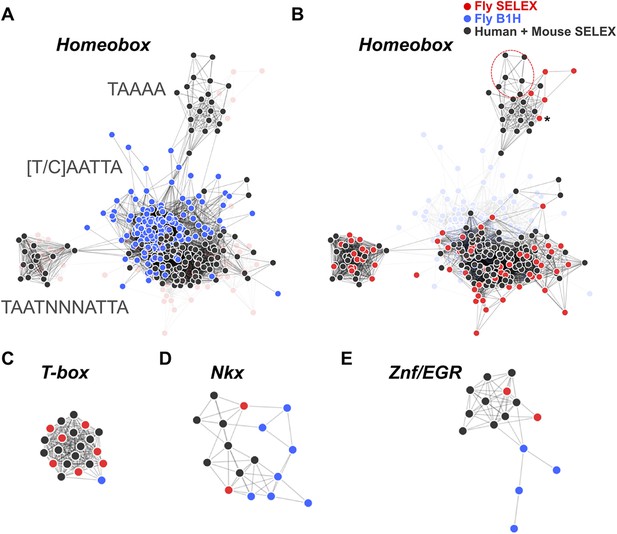
HT-SELEX reveals similarity of binding profiles between fruit flies and humans.
(A) Network graph of previously determined mammalian and fruit fly homeodomain protein DNA binding specificities. Mammalian and fruit fly TF models are represented by black and blue nodes, respectively, and an edge is drawn between similar models. Note that based on existing data, it appears that fruit fly homeodomains (blue) recognize only a subset of the mammalian homeodomain specificities. Partial consensus sequences for the node clusters are also indicated. (B) Increased precision obtained by using HT-SELEX for both fruit fly and mammalian TFs reveals that the range of mammalian homeodomain specificities is covered by fruit fly TFs. Note also that Drosophila has only one posterior homeodomain protein (Abd-B) that recognizes a motif (asterisk) that is similar to motifs bound by human HOX9-12. Drosophila thus lacks a protein whose motif preference is similar to that of human HOX13 proteins (six models inside red oval; see also Jolma et al., 2013). (C, D, E) Analysis of previous models for T-box, Nkx homeodomain and EGR proteins. Note that previous data for mammals (black) and fruit flies (blue) suggest divergence of the fruit fly specificity, whereas the fruit fly HT-SELEX data (red) reveal the similarity of the mammalian and fruit fly binding profiles. See Figure 3—source data 1 for details. See also Figure 3—figure supplements 1–3.
-
Figure 3—source data 1
Raw Data and a script for the Motif network construction.
- https://doi.org/10.7554/eLife.04837.022
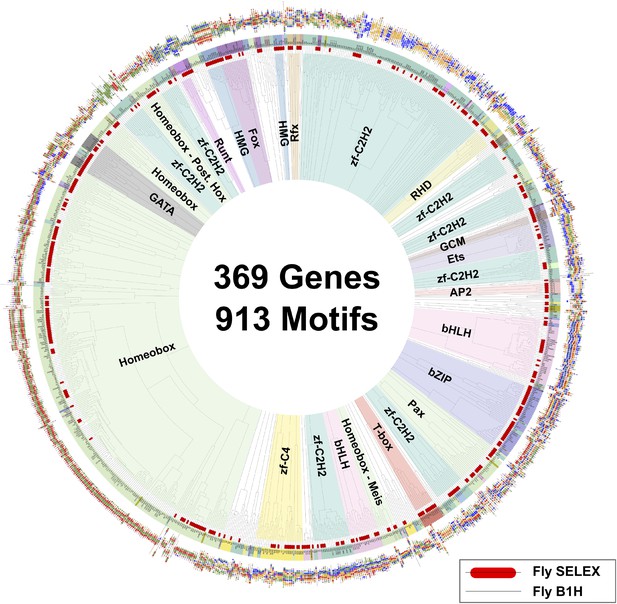
Similarity of binding profiles generated using HT-SELEX and B1H.
Dendrogram shows motif similarities between the fruit fly motif collection in this study and the bacterial one hybrid collection (Noyes et al., 2008; Zhu et al., 2011; Enuameh et al., 2013). Barcode logos of all models are also shown. HT-SELEX models are highlighted by red bars. Note that the B1H motifs and HT-SELEX motifs are similar but that B1H defines a shorter motif than HT-SELEX (see Figure 3—figure supplement 3).
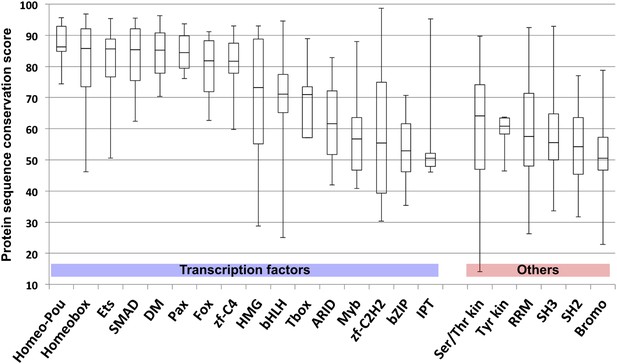
Comparison of amino-acid sequence similarity score (from blastp) of indicated protein domains of Drosophila and human ortholog pairs.
Note that many TF domains are highly conserved, but that others display levels of conservation that are similar to those observed for domains involved in protein–protein, protein-RNA, and enzyme–substrate interactions (kinase domains, RNA recognition module (RRM), SH2, SH3, and Bromodomains). Note also the broad distributions of conservation scores within TF families.
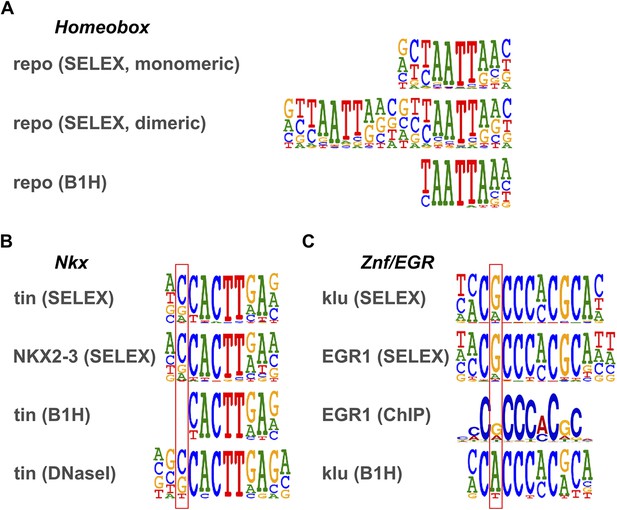
Comparison of motifs defined by HT-SELEX and B1H.
(A) Homeobox family gene repo. HT-SELEX reveals weak flanking preference that is not identified by B1H. HT-SELEX also identifies a homodimeric motif (see Jolma et al., 2013). (B) Comparison between HT-SELEX and B1H motifs for Drosophila Nkx gene tin and human NKX2-3. Main difference is indicated by box. Note that site identified using DNase I (Bergman et al., 2005) supports the flanking specificity revealed by HT-SELEX. (C) Comparison between HT-SELEX and B1H motifs for C2H2 zinc finger protein klu/EGR. Main difference is indicated by box. Note that site identified using MEME analysis of ChIP-seq data from Yan et al. (2013) supports the flanking specificity revealed by HT-SELEX.
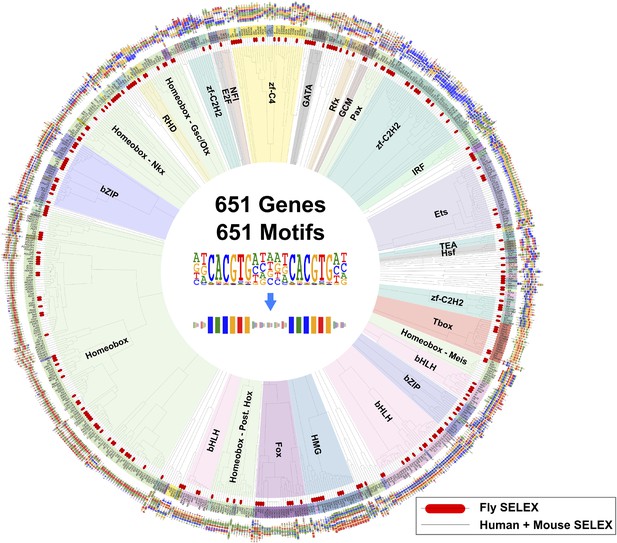
Similarity of primary binding profiles between fruit fly and human + mouse.
Dendrogram shows motif similarities between the fruit fly motif collection in this study and the human and mouse HT-SELEX collection of (Jolma et al., 2010; Jolma et al., 2013). Where both human and mouse motifs exist, human motif is shown. Drosophila models are indicated by red bars. Barcode logos for each factor are also shown. An example of conversion of a sequence logo into a Barcode logo is shown in center. See Figure 4—source data 1 for details.
-
Figure 4—source data 1
Raw Data and a script for the Dendrogram construction.
- https://doi.org/10.7554/eLife.04837.027
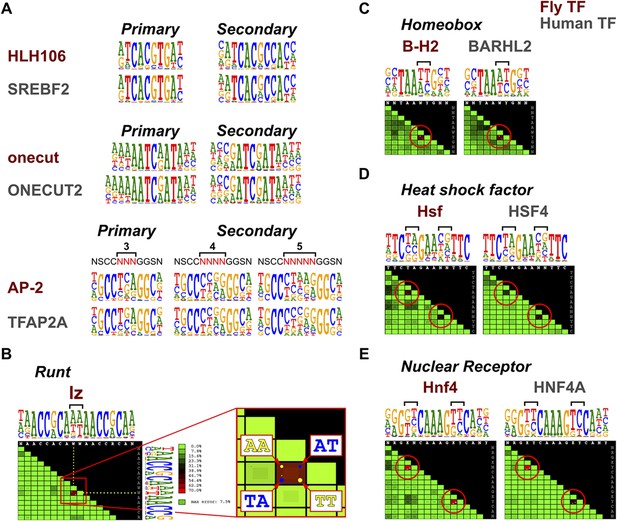
Conservation of TF secondary binding modes and dinucleotide preferences.
(A) Conservation of secondary binding modes. Sequence logos showing primary and secondary binding specificities for the indicated Drosophila (red typeface) and human (gray typeface) transcription factors are shown. Note that similar secondary modes exist for all factors. (B) Heatmap showing interdependency between bases in the binding model of the Runt family TF lz. Color of each tile indicates the deviation of the observed base distribution from a prediction using a mononucleotide (PWM) model that assumes independence of the indicated bases (color scale on the right; red indicates high deviation). Bracket indicates the two bases that show the largest deviation. Inset shows magnification of the tile; dots inside the each tile indicate pairs of bases that are over- (yellow) or under- (blue) represented relative to mononucleotide model prediction. (C, D, E) Conservation of dinucleotide preferences of the indicated orthologous proteins from fruit fly and human. Base positions deviating from mononucleotide model are indicated by red circles on the heatmap and brackets above the sequence logos.
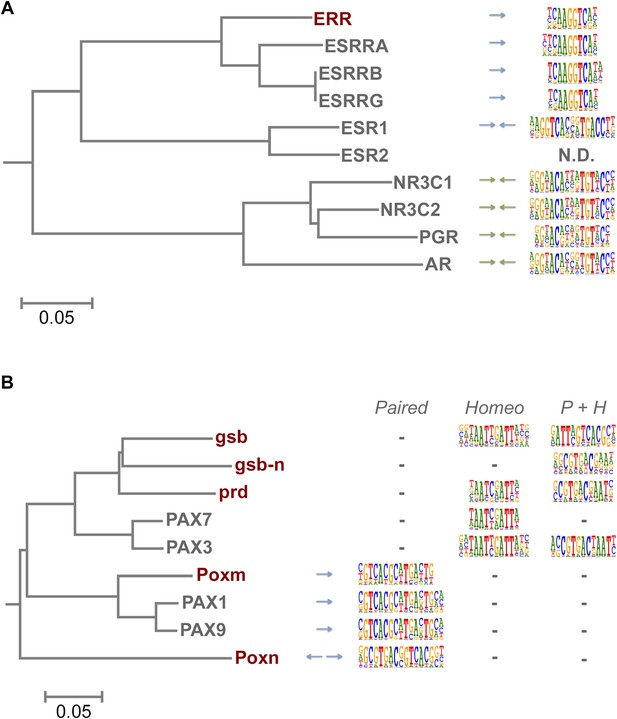
Evolution of TF binding specificity by gene duplication and divergence.
(A) Duplication and divergence has generated three distinct specificities in related nuclear receptors. Dendrogram is drawn based on amino-acid sequence similarity between the DBDs. Binding motifs are shown on the right column. Only one of the specificities (top clade), a monomeric AAGGTCA motif (blue arrow) exists in Drosophila. A human-specific site (middle clade) recognized by ESR1 is a dimer with the same half-site, but in a head-to-head configuration. Another human-specific site (bottom clade) recognized by androgen receptor has a different half-site (G/A)G(A/T)ACA (green arrow). (B) Specificity of the Pax proteins in relation to the sequence similarity of their paired DBDs. Binding motifs (right) are categorized into paired, homeodomain (homeo), and paired and homeodomain (P + H). Note that fruit fly Poxn binds to a site that is not recognized by any of the human PAX proteins. Arrows indicate an orientation of the paired motifs. N.D. = not determined. Complete data for all major families are shown in Figure 1—figure supplement 2–13.
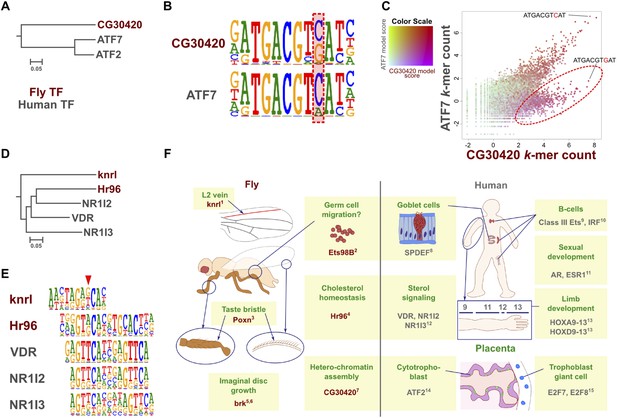
Evolution of TF binding specificity by duplication and divergence.
(A, B, C) The Drosophila bZIP protein CG30420 recognizes a site that is different from those recognized by its human orthologs ATF2 and ATF7. Box in B indicates the position whose specificity is diverged between fruit fly and human. Note that CG30420 recognizes a 10-mer ATGACGTGAT that is not bound by ATF7. Enrichment of 10-mers in CG30420 and ATF7 experiments is shown in panel C, oval indicates 10-mers preferentially recognized by CG30420. Two-dimensional color scale indicates the score of each k-mer against CG30420 and ATF7 PWM models (red indicates strong match to both, violet and green strong matches to only CG30420 and ATF7, respectively). For replicates and structural analysis, see Figure 7—figure supplement 1. (D, E) The diverged binding specificities between Drosophila and human in nuclear receptor subfamily. Whereas, all human NR1I subfamily genes (VDR, NR1I2, NR1I3) recognize a motif containing direct repeat of a GTTCA motif, Hr96 (NR1J subfamily) recognizes tail-to-tail dimer of a different half-site, GT(G/T)CA. Drosophila knrl (NR0 subfamily), which has no human ortholog, recognizes a G(A/G)(G/T)CA motif, which is not recognized by any human nuclear receptor. The diverged position in NR1 subfamily is indicated by red triangle. Dendrograms in A and D show amino-acid sequence similarity of the DNA-binding domains. (F) Summary of biological roles of TFs with divergent specificities. Cell types and biological functions are indicated in green in Drosophila (left) and human (right). For some TFs with multiple functions (e.g., bZIP and HOX proteins, nuclear receptors), only one divergent role is shown for clarity. In addition to their divergent roles, Hr96 and its orthologs have also shared functionality (xenobiotic responses; (King-Jones et al., 2006; Reschly and Krasowski, 2006)). Note that TFs with novel specificities are often associated with cell types that do not exist in the other organism. References: 1(Lunde et al., 1998); 2(Hsouna et al., 2004); 3(Boll and Noll, 2002); 4(Horner et al., 2009); 5(Schwank et al., 2008); 6(Doumpas et al., 2013); 7(Seong et al., 2011); 8(Gregorieff et al., 2009); 9(Bartel et al., 2000); 10(Lu et al., 2003); 11(Oliveira et al., 2004); 12(Pardee et al., 2011); 13(Zakany and Duboule, 2007); 14(Maekawa et al., 1999); 15(Chen et al., 2012). Raw data and scripts are provided at Figure 7—source data 1.
-
Figure 7—source data 1
Raw Data for kmer plot.
- https://doi.org/10.7554/eLife.04837.031
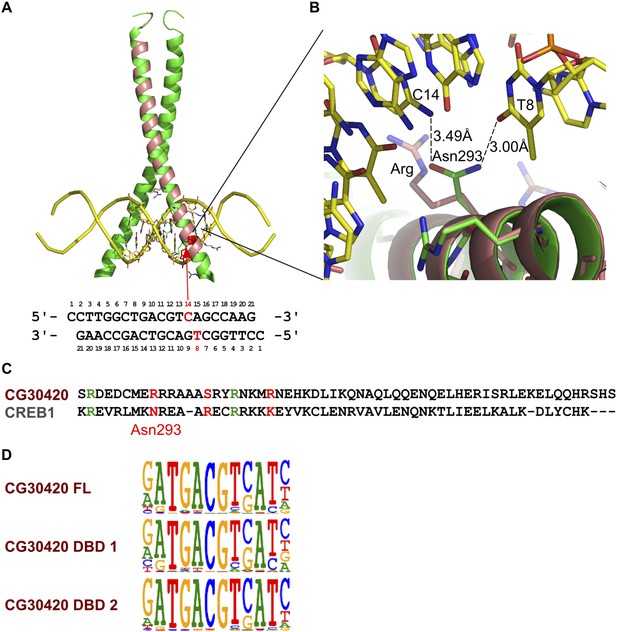
Validation of the difference in specificity between CG30420 and CRE family bZIP proteins.
(A) Ribbon diagram of the CREB bZIP/DNA complex (protein in green, DNA in yellow) and model of DNA recognition helix of CG30420 (in pink). Arrow indicates position of an amino-acid that recognizes bottom strand thymidine (T8), and the top strand cytosine (C14) that located at the position where the specificity difference between CG30420 and bZIP proteins of the CREB family are observed. The DNA sequence co-crystallized with CREB is shown below the structure (PDB entry 1DH3). (B) Close view of the Asn293 that recognizes bottom strand T8 and top strand C14 bases of the CRE site. This residue is replaced in CG30420 by an arginine, a bulky side chain that does not fit to the site, and is most likely directed to a backbone phosphate. This amino-acid substitution most likely explains the observed difference in specificity between CG30420 and CREB family proteins at this position. (C) Sequence alignment of CREB1 and CG30420 (CLUSTAL/omega). Residues involved in DNA interactions are colored (red = divergent, green = similar). (D) Reproducibility of the HT-SELEX for CG30420. Three independent experiments with three different constructs for CG30420 show very similar results.
Additional files
-
Supplementary file 1
(A) DBD and full-length clones. (B) Selex oligos. (C) Summary of HT-SELEX results and comparison to B1H. (D) Experimental details, including seed sequence, multinomial level and SELEX cycles used for signal and background. (E) Drosophila and human PWMs in one table. (F) Count of obtained motifs.
- https://doi.org/10.7554/eLife.04837.033
-
Supplementary file 2
Drosophila and human PWMs as separate files.
- https://doi.org/10.7554/eLife.04837.034
-
Supplementary file 3
Scripts for the total Autoseed, the barcode logo, and the dinucleotide heatmap.
- https://doi.org/10.7554/eLife.04837.035





