Expression levels of MHC class I molecules are inversely correlated with promiscuity of peptide binding
Figures
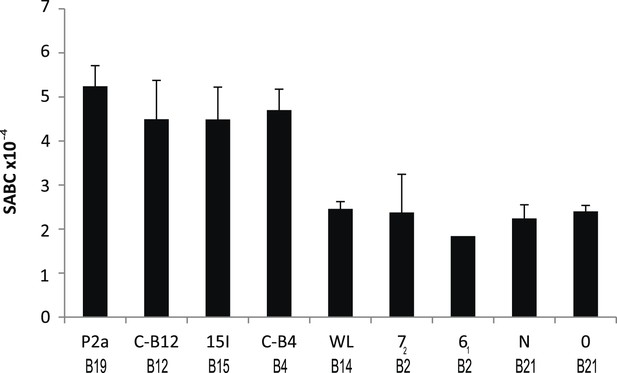
Cell surface expression levels of class I molecules vary markedly between chicken haplotypes, as determined by a quantitative flow cytometric assay.
Spleen cells from various inbred experimental chicken lines (with MHC haplotypes indicated) were stained with the monoclonal antibody F21-2 against chicken major histocompatibility complex (MHC) class I heavy chain and the specific antigen binding capacity (SABC, which reflects number of epitopes per cell calculated in reference to specific antibody-binding calibration beads). Results are means of triplicate stains, with error bars indicating standard deviation.
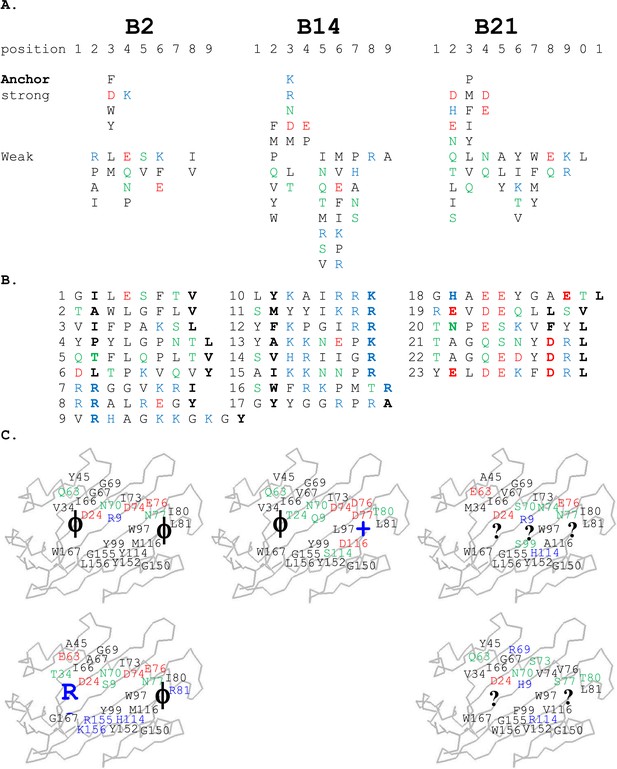
Peptides isolated from class I molecules of B2, B14, and B21 chickens show promiscuity of peptide binding.
For all panels, amino acids are in single letter code, with basic residues shown in blue, acidic in red, polar in green, hydrophobic in black. (A) Sequences of peptides bound to class I molecules isolated from three chicken strains determined from peptide pools showing apparent anchor, strong and weak signals. (B) Sequences of individual peptides, with confirmed anchor residues in bold. (C) Peptide anchor residues in large letters (or question marks for unknown) superimposed on a model of class I α1 and α2 domains with those residues of the major (above) and minor (below) class I sequences that are both polymorphic and potentially peptide contacts indicated as smaller letters; numbering based on human class I (HLA-A2) sequence.
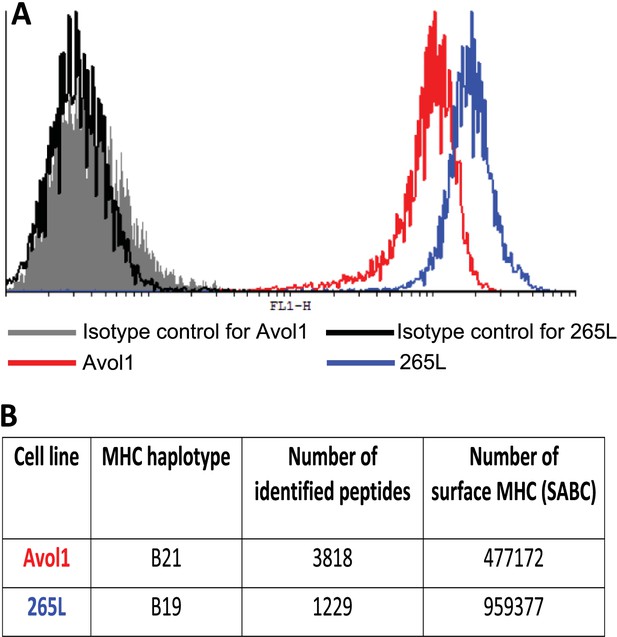
There is an inverse correlation between the cell surface expression levels of class I molecules and the variety of peptides isolated from class I molecules.
(A) The B19 cell line 265L and the B21 cell line AVOL-1 were analysed by flow cytometry by staining with the mAb F21-2 to chicken class I molecules. AVOL-1 had slightly more autofluorescence, so the settings on the FACScan were adjusted so that the mean fluorescence intensity of the isotype control sample was the same as for 265L. The histogram shows the fluorescence intensity in the FL1 channel on the x-axis and the number of events on the y-axis. (B) In the same flow cytometry experiment, the calibration beads from the QIFIKIT were stained separately with the secondary antibody for calibration curves to calculate the SABC, which reflects the absolute numbers of epitopes on the cell surface. As a separate experiment, the class I molecules were isolated from each cell line by affinity chromatography with F21-2 and analysed by LC-MS/MS. Table shows the SABC and the number of different peptides found for each cell line.
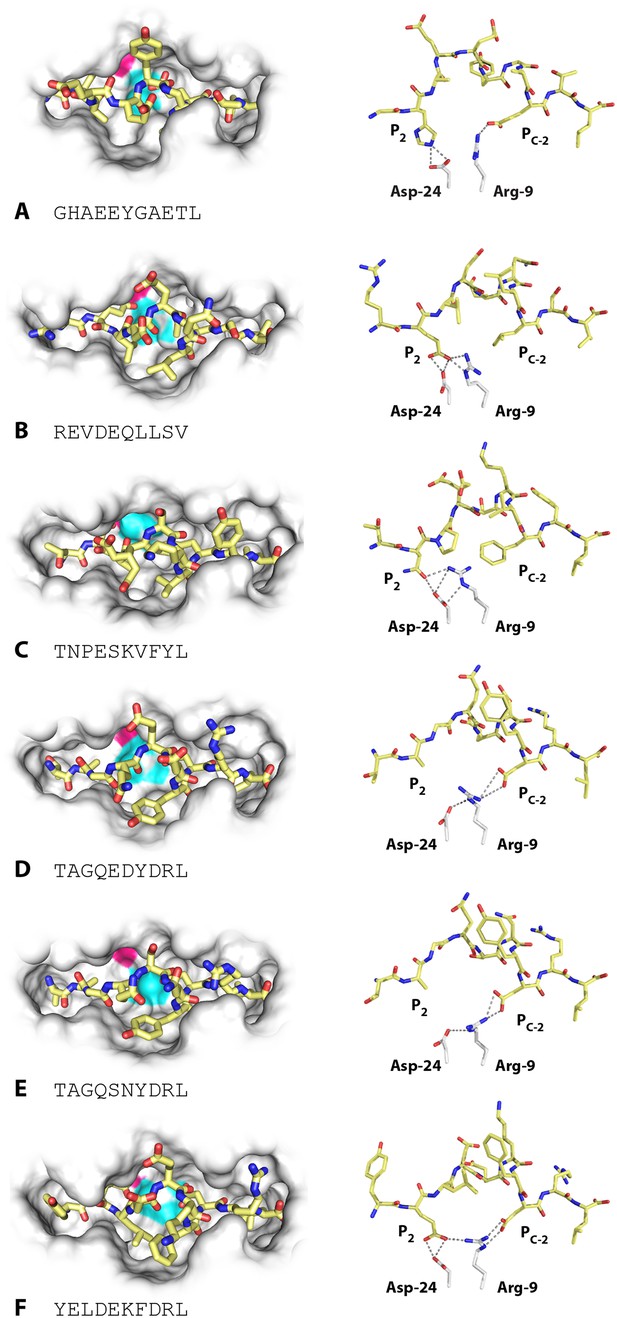
Structures of BF2*2101 with different peptides show several modes of promiscuous binding through remodelling of the binding site.
Left panels, top down view with peptide as sticks (N-terminus to the left; carbon atoms, yellow; nitrogen atoms, blue; oxygen atoms, red) and class I molecule as solid surface (grey except for positions of Asp24 side chain oxygen atoms in pink and Arg9 side chain nitrogen atoms as cyan). Right panels, side view from α2 domain side with peptide, Asp24 and Arg9 as sticks (hydrogen bonds, dotted lines; carbon atoms of Asp24 and Arg9, white; all else as in left panels). (A) GHAEEYGAETL (peptide P316; PDB 3BEV); (B) REVDEQLLSV (P330; 3BEW); (C) TNPESKVFYL (P458; 2YEZ); (D) TAGQEDYDRL (P394; 4D0B); (E) TAGQSNYDRL (P399; 4D0C); (F) YELDEKFDRL (P400; 4CVZ).
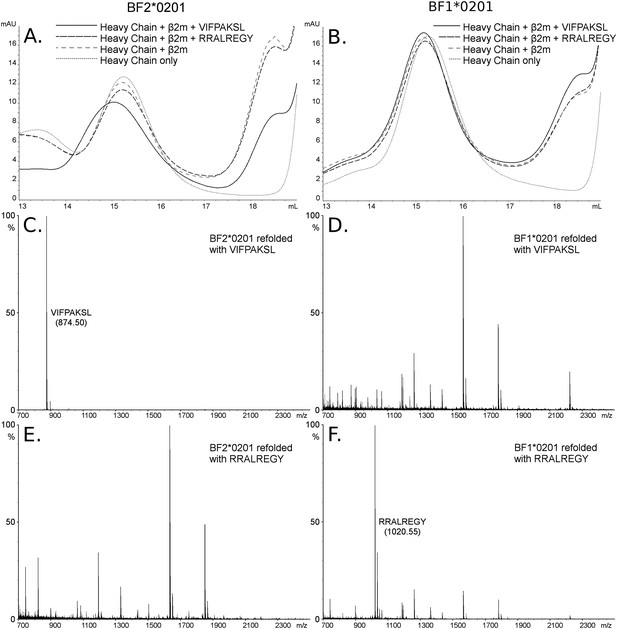
The dominantly expressed class I molecule BF2*0201 binds VIFPAKSL but not RRALREGY, while the minor class I molecule BF1*0201 binds RRALREGY but not VIFPAKSL.
(A and B) Size exclusion chromatography (SEC) traces for BF2*0201 or BF1*0201 heavy chains expressed in bacteria refolded with or without β2-microglobulin (β2m) and peptide. The heavy chain BF2*0201 refolded with β2m and the appropriate peptide migrates as a native monomer, whereas refolded with the inappropriate or no peptide migrates in the same position as heavy chain alone. In contrast, all these conditions for the heavy chain BF1*0201 result in molecules that migrate roughly the same mobility. (C through F) Mass spectrometry (MALDI-TOF) analysis of the monomer peaks of heavy chain refolded with β2m and peptide shows that VIFPAKSL but not RRALREGY can be recovered from BF2*0201, while RRALREGY but not VIFPAKSL can be recovered from BF1*0201. Note the many peaks for BF1*0201 with VIFPAKSL and for BF2*0201 with RRALREGY, representing background contaminants detected as sensitivity was increased in the search for the synthetic peptide. Comparable results were obtained with YPYLGPNTL, RRALREGY, RRGGVKRI, and the B15 (BF2*1501) peptide KRLIGKRY.
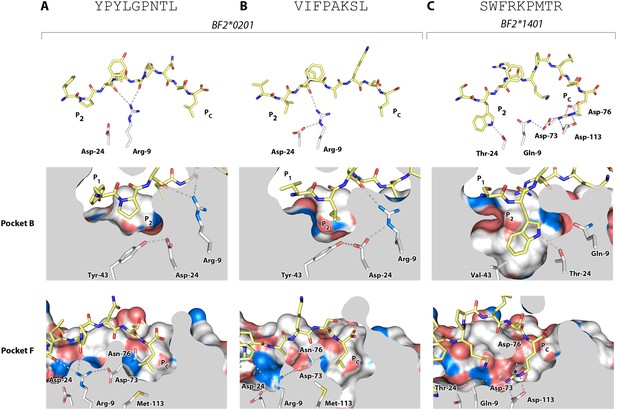
Structures of BF2*0201 and BF2*1401 show promiscuous binding via hydrophobic binding pockets for the anchor residues at peptide position P2 and Pc, with the class I residues at positions 24 and 9 playing supporting roles, and with residues lining the pockets explaining the relative size of anchor residues.
Upper panels, side view from α2 domain side with peptide as sticks (N-terminus of peptide to the left; carbon atoms of peptide, yellow; carbon atoms of class I molecule, white; nitrogen atoms, blue; oxygen atoms, red; hydrogen bonds, dotted lines; carbon atoms of Asp24 and Arg9, white). Middle (pocket B) and bottom (pocket F) panels, side view cut-away from α2 domain side with peptide and selected class I residues as sticks (numbering based on chicken class I sequence) and with rest of MHC molecule as solid surface. (A) YPYLGPNTL bound to BF2*0201 (peptide P377; PDB 4CVX); (B) VIFPAKSL bound to BF2*0201 (P473; 4D0D); (C) SWFRKPMTR bound to BF2*1401 (P479; 4CW1).
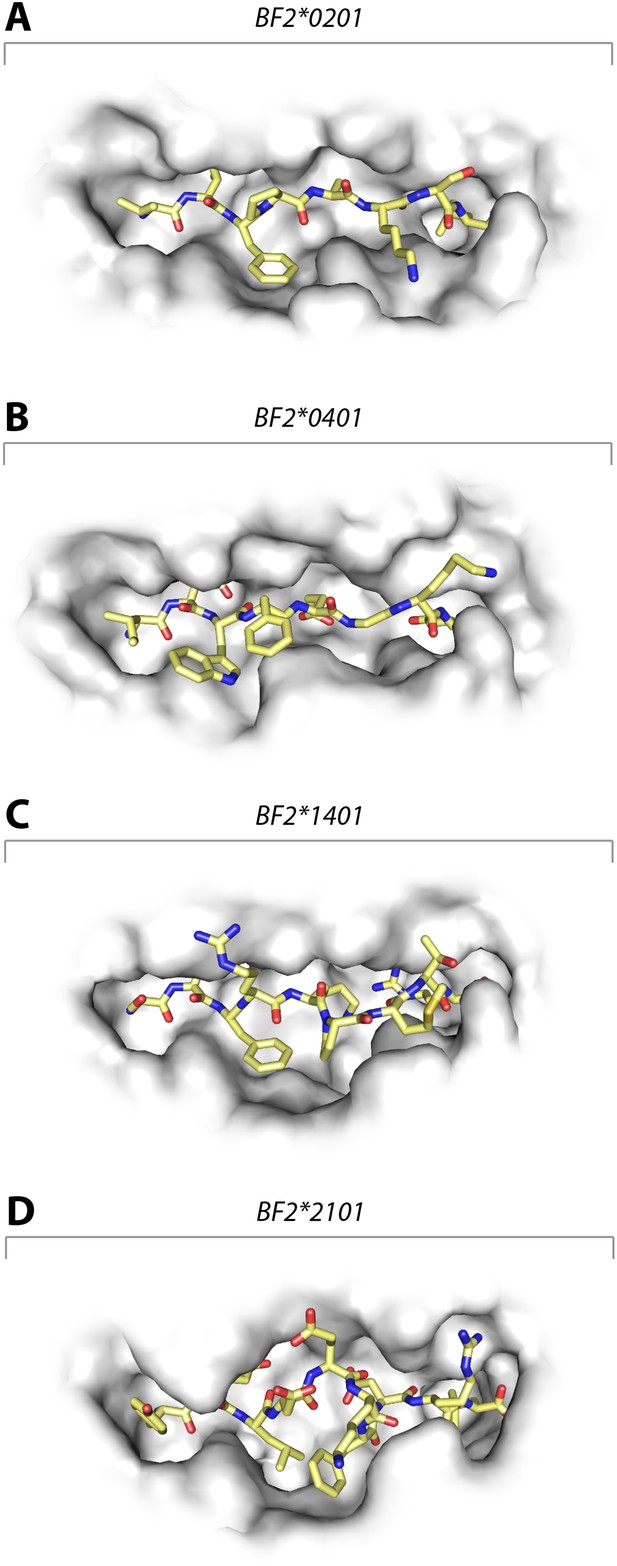
Structures of chicken class I molecules show differences in the width of the peptide-binding groove, with the fastidious BF2*0401 having the narrowest groove and the promiscuous BF2*2101 being the widest in the centre of the groove.
Top down view with peptide as sticks (N-terminus to the left; carbon atoms, yellow; nitrogen atoms, blue; oxygen atoms, red) and class I molecule as grey solid surface. (A) VIFPAKSL bound to BF2*0201 (P473; 4D0D); (B) IDWFEGKE bound to BF2*0401 (IE8; 4G43); (C) SWFRKPMTR bound to BF2*1401 (P479; 4CW1); (D) YELDEKFDRL bound to BF2*2101 (P400; 4CVZ).
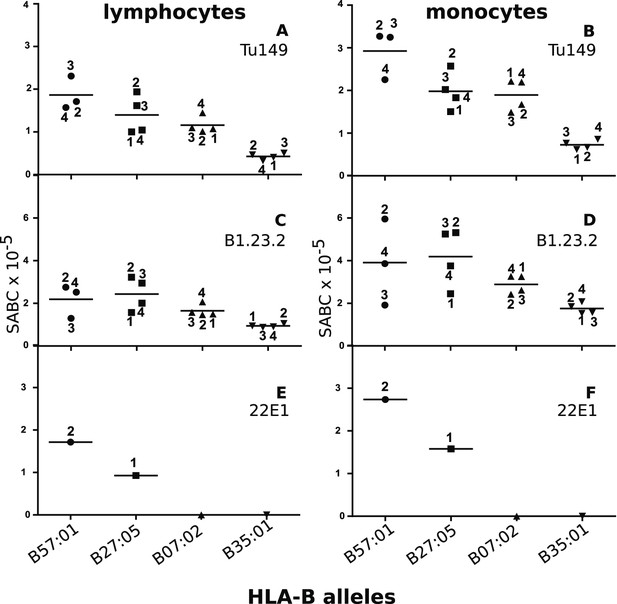
Human class I molecules show an inverse correlation between cell surface expression level and peptide binding promiscuity.
Levels of specific antibody binding capacity (SABC) for different mAb: (A and B) Tu149, (C and D) B1.2.23, (E and F) 22E-1 for (A, C and E) ex vivo lymphoctyes and (B, D and F) ex vivo monocytes. Each point represents the sample from a particular donor (identified with anonymous labels that correlate with haplotypes in Table 2; donor B57:01/01 failed to donate); bars indicate the mean for each HLA-B allele.
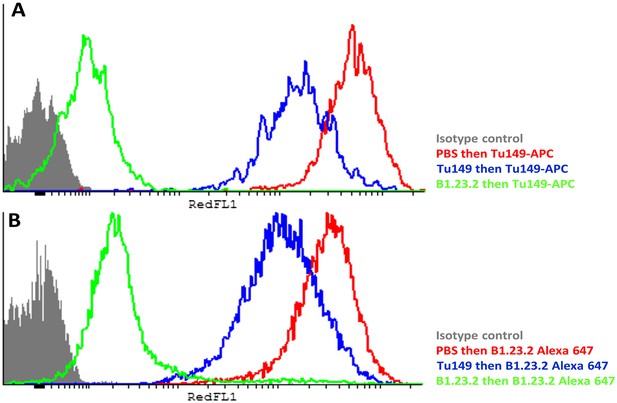
The epitopes on HLA-B57:01 for mAb B1.23.2 and Tu149 are overlapping, with B1.23.2 having a much higher affinity.
Win cells were treated with saturating amounts of unlabelled Tu149 (blue), saturating amounts of unlabelled B1.23.2 (green) or PBS (red), and then stained with either A Tu149 conjugated to APC or B B1.23.2 conjugated to Alexa Fluor 647. Results were compared to staining with only isotype control (grey).
Tables
Data collection and refinement statistics
| PDBID | 4d0b | 4d0c | 4cvz | 2yez | 4cvx | 4d0d | 4cw1 |
|---|---|---|---|---|---|---|---|
| Data collection | |||||||
| Space group | P212121 | P212121 | P212121 | P212121 | P65 | P212121 | P212121 |
| Cell dimensions | |||||||
| a, b, c (Å) | 60.9 | 60.5 | 60.5 | 60.6 | 173.9 | 88.1 | 62.2 |
| 69.2 | 68.9 | 69.0 | 69.0 | 173.9 | 92.5 | 90.58 | |
| 95.4 | 94.8 | 93.7 | 94.8 | 87.5 | 223.6 | 144.8 | |
| α, β, γ (°) | 90 | 90 | 90 | 90 | 90 | 90 | 90 |
| 90 | 90 | 90 | 90 | 90 | 90 | 90 | |
| 90 | 90 | 90 | 90 | 120 | 90 | 90 | |
| Resolution (Å) | 56.03–2.80 (2.95–2.80) | 51.00–2.82 (2.89–2.82) | 50.84–2.39 (2.56–2.39) | 41.03–2.45 (2.58–2.45) | 75.66–3.30 (3.56–3.30) | 88.08–3.13 (3.21–3.13) | 72.38–2.58 (2.65–2.58) |
| Rsym or Rmerge | 0.118 (0.30) | 0.12 (0.30) | 0.15 (0.91) | 0.07 (0.22) | 0.17 (0.56) | 0.18 (0.69) | 0.14 (0.59) |
| I/σI | 8.1 (2.8) | 8.2 (1.9) | 11.9 (2.1) | 9.5 (1.5) | 11.1 (3.8) | 8.8 (2.1) | 8.8 (2.3) |
| Completeness (%) | 91.2 (89.8) | 90.8 (76.0) | 99.9 (100) | 85.9 (53.0) | 100 (99.9) | 99.6 (98.8) | 99.3 (98.4) |
| Redundancy | 4.0 (3.6) | 3.8 (1.9) | 6.4 (6.5) | 2.8 (1.6) | 7.3 (7.5) | 4.6 (4.4) | 5.5 (4.2) |
| Refinement | |||||||
| Resolution (Å) | 56.03–2.80 | 50.92–2.81 | 50.84–2.39 | 41.03–2.45 | 75.66–3.30 | 88.08–3.13 | 72.38–2.58 |
| No. reflections | 9361 | 8967 | 16,086 | 8913 | 21,707 | 32,916 | 26,228 |
| Rwork/Rfree | 27.5/29.6% | 27.5/28.2% | 25.9/26.9% | 25.2/27.3% | 23.8/26.2% | 29.2/29.9% | 28.6/29.1% |
| Number of atoms | |||||||
| Protein | 3061 | 3057 | 3079 | 3091 | 6052 | 12,102 | 6044 |
| Ligand/ion | 0 | 4 | 24 | 0 | 0 | 0 | 0 |
| Water | 5 | 31 | 42 | 30 | 8 | 9 | 77 |
| B-factors | |||||||
| Protein | 30.30 | 34.24 | 30.23 | 32.22 | 77.86 | 47.02 | 40.02 |
| Ligand/ion | – | 26.32 | 46.25 | – | – | – | – |
| Water | 14.33 | 12.88 | 24.38 | 8.61 | 34.87 | 34.31 | 29.50 |
| Root mean square (r.m.s.) deviations | |||||||
| Bond lengths (Å) | 0.008 | 0.007 | 0.007 | 0.007 | 0.004 | 0.007 | 0.007 |
| Bond angles (°) | 0.89 | 0.86 | 0.85 | 0.83 | 0.759 | 0.82 | 0.84 |
-
Highest resolution shell is shown in parenthesis.
Anonymized donors with class I alleles and inferred HLA-C expression
| Donor | HLA-A alleles | HLA-B alleles | HLA-C alleles | HLA-C expression |
|---|---|---|---|---|
| 5701/02 | 02:01, 02:01 | 57:01, 57:01 | 06:02, 15:02 | high, unknown |
| 5701/03 | 01:01, 01:01 | 57:01, 57:01 | 06:02, 06:02 | high, high |
| 5701/04 | 01:01, 03:01 | 57:01, 57:01 | 06:02, 07:01 | high, low |
| 2705/1 | 03:01, 68:01 | 27:05, 27:05 | 01:02, 02:02 | high, high |
| 2705/2 | 02:06, 11:01 | 27:05, 27:05 | 01:02, 03:03 | high, low |
| 2705/3 | 02:01, 02:01 | 27:05, 27:05 | 01:02, 12:03 | high, high |
| 2705/4 | 02:01, 02:12 | 27:05, 27:05 | unknown | |
| 0702/1 | 02:01, 03:01 | 07:02, 07:02 | 07:02, 07:02 | low, low |
| 0702/2 | 03:01, 03:01 | 07:02, 07:02 | 07:02, 07:02 | low, low |
| 0702/3 | 02:01, 03:01 | 07:02, 07:02 | 07:02, 07:02 | low, low |
| 0702/4 | 03:01, 11:01 | 07:02, 07:02 | 07:02, 07:02 | low, low |
| 3501/1 | 11:01, 11:01 | 35:01, 35:01 | 04:01, 04:01 | high, high |
| 3501/2 | 11:01, 11:01 | 35:01, 35:01 | 04:01, 04:01 | high, high |
| 3501/3 | 03:01, 03:01 | 35:01, 35:01 | 04:01, 04:01 | high, high |
| 3501/4 | 11:01, 11:01 | 35:01, 35:01 | 04:01, 04:01 | high, high |
Crystallization conditions, data collection, and refinement information
| Structure (PDB ID) | Crystallization conditions | Cryo-protectant | Beamline | Data processing | Refinement program |
|---|---|---|---|---|---|
| 4 d0b | 0.1M MMT buffer, pH 5.0, 25% PEG 1500 | 15% ethylene glycol | I02 (Diamond Light Source) | AUTOPROC SUITE with XDS & SCALA | AUTOBUSTER |
| 4d0c | 0.05M KH2PO4, 20% PEG 8000 | 20% ethylene glycol | I02 (Diamond Light Source) | XIA2 with XDS & AIMLESS | AUTOBUSTER |
| 2yez | 0.1M MMT buffer, pH 4.0, 25% PEG 1500 | 15% ethylene glycol | I03 (Diamond Light Source) | AUTOPROC SUITE with XDS & SCALA | AUTOBUSTER |
| 4cvz | 0.1M sodium acetate, pH 5.0, 1.5 M ammonium sulphate | 8M sodium formate | I04-1 (Diamond Light Source) | XIA2 with XDS & AIMLESS | AUTOBUSTER |
| 4cvx | 0.1M MMT buffer, pH 7.0, 25% PEG 1500 | 15% ethylene glycol | I04 (Diamond Light Source) | XIA2 with XDS & AIMLESS | REFMAC |
| 4 d0d | 0.1M MMT buffer, pH 7.0, 25% PEG 1500 | 15% ethylene glycol | I04 (Diamond Light Source) | XIA2 with XDS & AIMLESS | AUTOBUSTER |
| 4cw1 | 0.1M MIB buffer, pH 5.0, 25% PEG 1500 | 15% ethylene glycol | ID29 (ESRF) | XIA2 with XDS & AIMLESS | AUTOBUSTER |
Ramachandran statistics
| Structure (PDB ID) | Ramachandran outliers number (%) | Ramachandran favoured number (%) |
|---|---|---|
| 4cvz | 0 (0%) | 371 (98.15%) |
| 4 d0b | 0 (0%) | 356 (94.43%) |
| 4d0c | 0 (0%) | 361 (95.76%) |
| 2yez | 1 (0.26%) | 364 (96.04%) |
| 4cvx | 0 (0%) | 710 (95.69%) |
| 4 d0d | 0 (0%) | 1430 (96.30%) |
| 4cw1 | 0 (0%) | 730 (98.25%) |





