COMP-1 promotes competitive advantage of nematode sperm
Figures
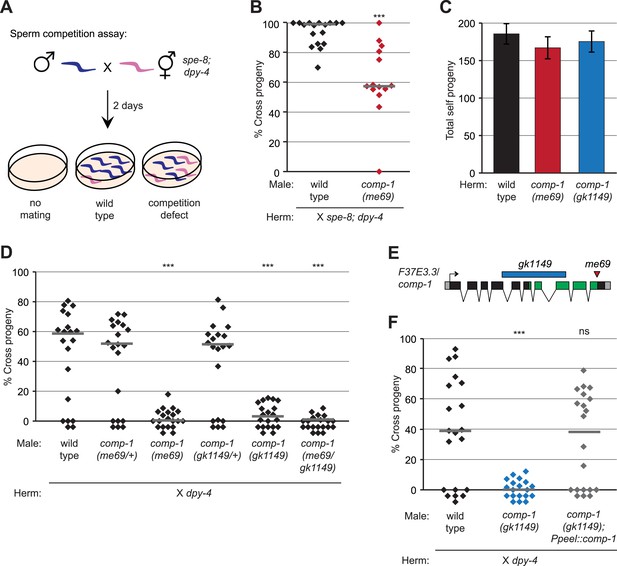
Isolation of the male precedence mutant me69 in a genetic screen.
(A) Screening assay for mutants with reduced male precedence, showing outcomes for mating failure, mating by wild-type males, and mating by males with less-competitive sperm. (B) me69 males have decreased precedence in the screen assay. Males were mated to spe-8(hc53); dpy-4 hermaphrodites, and offspring were scored as Dumpy (self) or non-Dumpy (cross) progeny. (C) me69 and gk1149 mutant hermaphrodites have normal self fertility. Total self progeny of unmated hermaphrodites were counted. Error bars, 95% confidence intervals; p > 0.05 (Student's t test). (D) Mutants for comp-1 have defects in male precedence. Males were mated to dpy-4 hermaphrodites, and offspring were scored as Dumpy (self) or non-Dumpy (cross). (E) Schematic of the F37E3.3 gene showing the kinase-like domain (green) predicted by the Conserved Domains Database (Marchler-Bauer et al., 2011) and the locations of the me69 and gk1149 alleles. Boxes, exons; black, coding regions; gray, 5′ and 3′UTRs. (F) Expression of the F37E3.3 gene in sperm rescues the comp-1 male precedence defect. Male precedence was assayed for comp-1(gk1149); jnSi168[Ppeel::comp-1] and control strains as in Figure 1D. (B, D, F) Each point represents the result of an individual cross; lines indicate medians. ***, p < 0.001; **, p < 0.01; ns, not significant (Kolmogorov–Smirnov test; all comparisons are to wild-type). In addition to the genotypes shown, all males were homozygous (B, C, F) or heterozygous (D) for him-5(ok1896), and control strains in (F) harbored oxSi221.
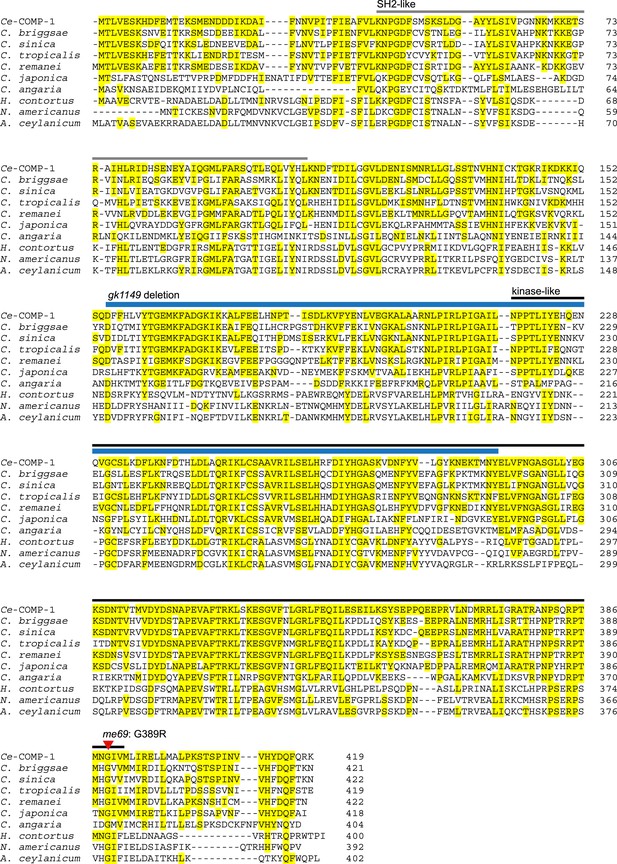
COMP-1 is highly conserved within the Caenorhabditis genus and present in related parasitic species.
Alignment of C. elegans COMP-1 with its orthologs from other nematode species. Yellow highlighting represents amino acids conserved with the C. elegans protein. Bars above the sequence indicate the positions of the divergent kinase-like (black) and SH2-like (gray) domains, as predicted in the Conserved Domains Database (Marchler-Bauer et al., 2011); an alternate prediction for the kinase-like domain includes amino acids 123–394 (Manning, 2005). Positions of the me69 and gk1149 alleles are shown in red and blue as in Figure 1E. For C. japonica, C. sinica, and N. americanus, the predicted COMP-1 proteins present in current databases were incomplete or had regions with poor similarity. In each case, examination of sequence from the comp-1 region allowed us to generate new predicted proteins showing similarity across the full-length Ce-COMP-1 and the other orthologs. For C. sinica, we obtained sequence from the comp-1 region, which revealed a minor assembly error in Csp5.Scaffold_00675.g14294.tt (PRJNA194557). For C. japonica, previous annotation (WormBase) of the comp-1 region predicted two overlapping gene models encoding proteins similar to the N and C terminus of COMP-1; our new predicted Cja-COMP-1 fuses JA64544 and CJA40432. For N. americanus, we were able to identify additional exons upstream of the comp-1 gene model predicted from NECAME_15795. Other accession numbers: Acey_s0303.g1899.t2 (PRJNA231479), Cang_2012_03_13_00293.g8644.t2 (PRJNA51225), CBP27128, RP29840, Csp5.Scaffold_00675.g14294.tt (PRJNA194557), Csp11.Scaffold630.g17815.ti (PRJNA53597), HCOI01934100.t1 (PRJEB506).
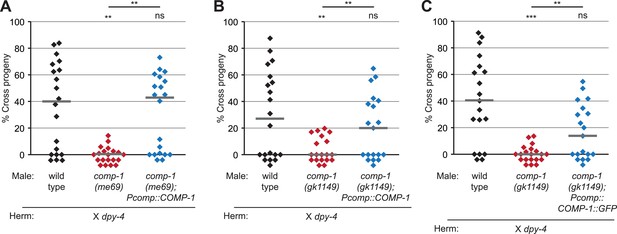
COMP-1 transgenes rescue the male precedence defects of comp-1 mutants.
(A, B) The jnSi109[Pcomp-1::COMP-1] transgene, which contains a 3.9 kb region surrounding F37E3.3, rescues the precedence defect of (A) comp-1(me69) and (B) comp-1(gk1149) males in crosses to dpy-4 hermaphrodites. (C) Expression of COMP-1::GFP rescues the precedence defect. comp-1(gk1149); jnSi171[Pcomp-1::COMP-1::GFP] males have a wild-type precedence pattern in crosses to dpy-4 hermaphrodites. Precedence assays were performed as in Figure 1D. ***, p < 0.001; **, p < 0.01; ns, not significant (Kolmogorov–Smirnov test). Lines indicate medians. In addition to the indicated genotypes, control strains contained the transgene oxSi221[Peft-3 ::GFP].
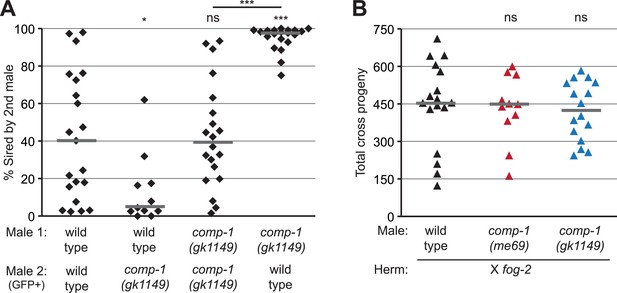
The comp-1 mutant has defects in male–male sperm competition.
(A) comp-1 male sperm are outcompeted by wild-type male sperm. Wild-type and/or comp-1(gk1149) males were mated sequentially to fog-2 hermaphrodites; second-mated males harbored the transgene mIs11(GFP+). Offspring were scored for GFP, and the percentage of GFP-positive progeny produced 0–16 hr after second-male mating is shown. (B) comp-1 mutant males have wild-type levels of fertility in the absence of competition. Males were crossed to fog-2 hermaphrodites and total progeny were counted. (A, B) Individual data points are shown; lines indicate medians. *, p < 0.05; ***, p < 0.001; ns, not significant (Kolmogorov–Smirnov test; comparisons are to wild-type unless indicated by a line linking the two data sets).
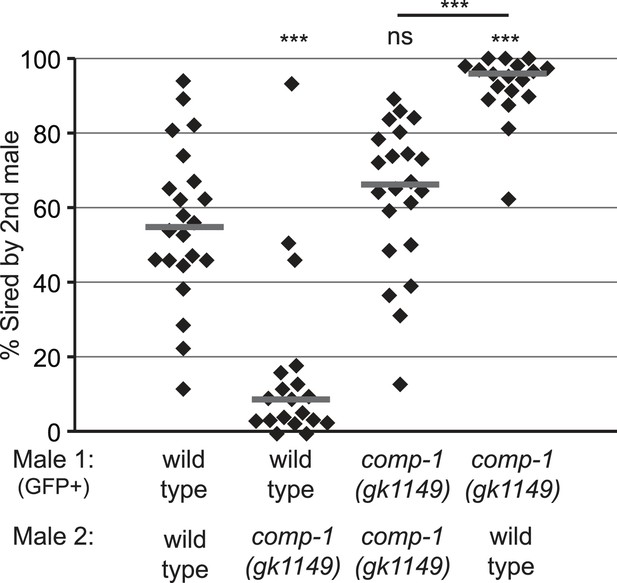
The comp-1 mutant has defects in male–male sperm competition.
comp-1 male sperm are outcompeted by wild-type male sperm. Wild-type and/or comp-1(gk1149) males were mated sequentially to fog-2 hermaphrodites; first-mated males harbored the transgene mIs11(GFP+). Offspring were scored for GFP, and the percentage of GFP-positive progeny produced 0–16 hr after second-male mating is shown. GFP-marked males show an apparent slight disadvantage, which is observed consistently but is not statistically significant. Lines indicate medians. ***, p < 0.001; ns, not significant (Kolmogorov–Smirnov test).
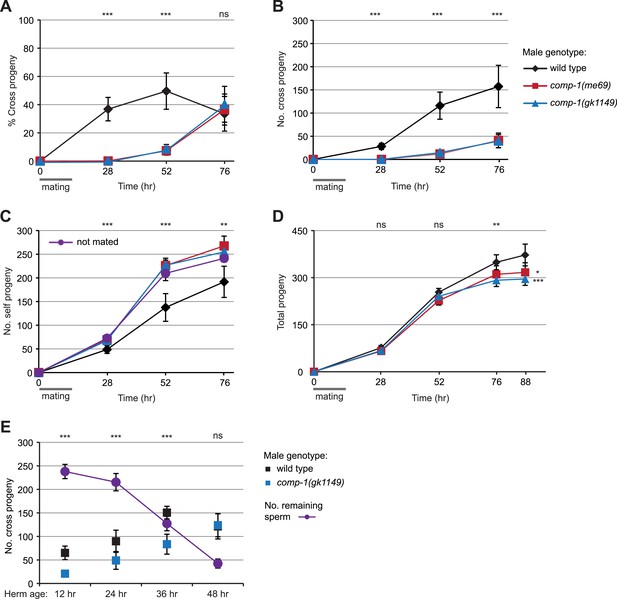
comp-1 male sperm have long-term precedence defects.
(A) Crosses with comp-1 males result in a low percentage of cross progeny. (B) The number of cross progeny sired by comp-1 increases at late time points. (C) Crosses with comp-1 males do not suppress production of self progeny. Purple line indicates self progeny of unmated hermaphrodites. (D) Crosses with comp-1 males result in decreased progeny numbers as compared to those with wild-type males. (A–D) Males were crossed to dpy-4 hermaphrodites for 16 hr (gray line); progeny were collected throughout the recipients' reproductive lifespans and scored as self or cross progeny. All graphs are from a single data set that is representative of three repeats. For B–D, cumulative progeny numbers are shown. (E) comp-1 male sperm are used at wild-type levels in crosses to sperm-depleted hermaphrodites. Males were crossed to staged dpy-4 recipients for 24 hr and progeny generated during the mating period were scored as self or cross progeny. ‘No. remaining sperm’ indicates the number of self sperm present within recipients at each stage, inferred from brood counts of unmated dpy-4 hermaphrodites performed in parallel. Data points indicate averages; error bars, 95% confidence intervals. **, p < 0.01; ***, p < 0.001; ns, not significant (Kolmogorov–Smirnov test, comparing wild-type to each comp-1 mutant).
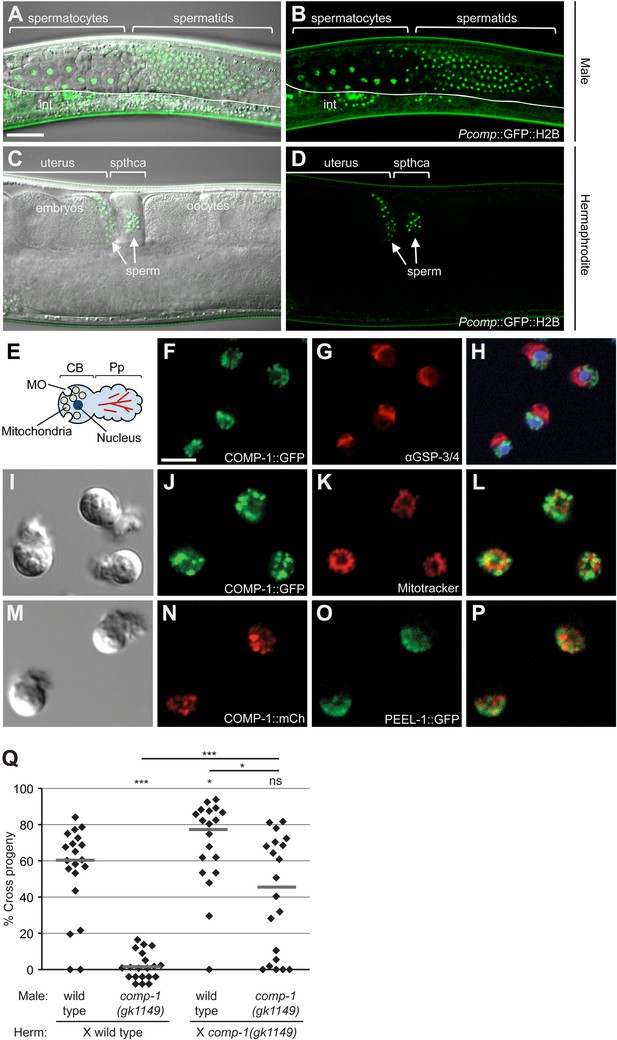
COMP-1 is expressed and functions in sperm of both males and hermaphrodites.
(A–D) Images of jnSi118[Pcomp-1::GFP::H2B] adult males (A, B) and hermaphrodites (C, D), which express the comp-1 reporter in developing sperm. int, intestinal autofluorescence. Scale bar (A–D), 30 μm. (E) Schematic of structural organization of spermatozoa. (F–H) COMP-1 does not colocalize with GSP-3/4, which is in the pseudopod. Images of jnSi171[COMP-1::GFP] male spermatozoa fixed and stained with α-GSP-3/4 antibody (red) and DAPI (blue). Scale bar (F–P), 5 μm. (I–L) COMP-1 does not colocalize with mitochondria. Images of jnSi171[COMP-1::GFP] male spermatozoa stained with Mitotracker. (M–P) COMP-1 does not colocalize with PEEL-1::GFP, which is at the membranous organelles. Images of jnSi143[COMP-1::mCherry]; jnSi177[PEEL-1::GFP] male spermatozoa. (Q) comp-1 functions in both male and hermaphrodite sperm. Wild-type and comp-1(gk1149) males were tested against wild-type and comp-1(gk1149) hermaphrodites in the short-term precedence assay. Lines indicate medians. *, p < 0.05; ***, p < 0.001; ns, not significant, Kolmogorov–Smirnov test.
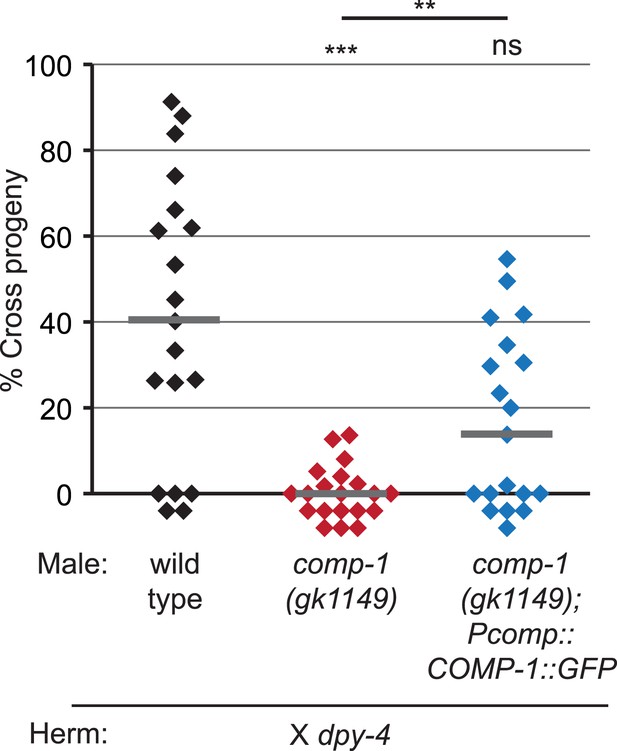
A COMP-1::GFP transgene rescues the male precedence defects of comp-1 mutants.
Expression of COMP-1::GFP rescues the precedence defect. comp-1(gk1149); jnSi171[Pcomp-1::COMP-1::GFP] males have a wild-type precedence pattern in crosses to dpy-4 hermaphrodites. Precedence assays were performed as in Figure 1D. ***, p < 0.001; **, p < 0.01; ns, not significant (Kolmogorov–Smirnov test). Lines indicate medians. In addition to the indicated genotypes, control strains contained the transgene oxSi221[Peft-3 ::GFP].
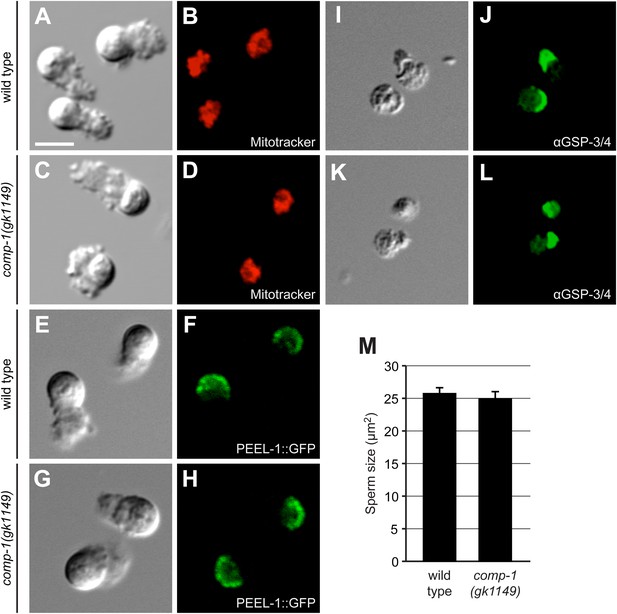
comp-1 sperm have normal organization and size.
(A–D) Wild-type (A, B) and comp-1(gk1149) (C, D) spermatozoa stained for mitochondria using Mitotracker. (E–H) Wild-type (E, F) and comp-1 (G, H) spermatozoa expressing PEEL-1::GFP (membranous organelles). (I–L) Wild-type (I, J) and comp-1(gk1149) (K, L) spermatozoa fixed and stained with α-GSP-3/4 antibody (green). (A–L) Scale bar, 5 μm. (M) comp-1 male spermatid size is not significantly different from wild-type. Cross sectional areas through the center of spermatids were measured. Error bars, 95% confidence interval; p = 0.41, Student's t test.
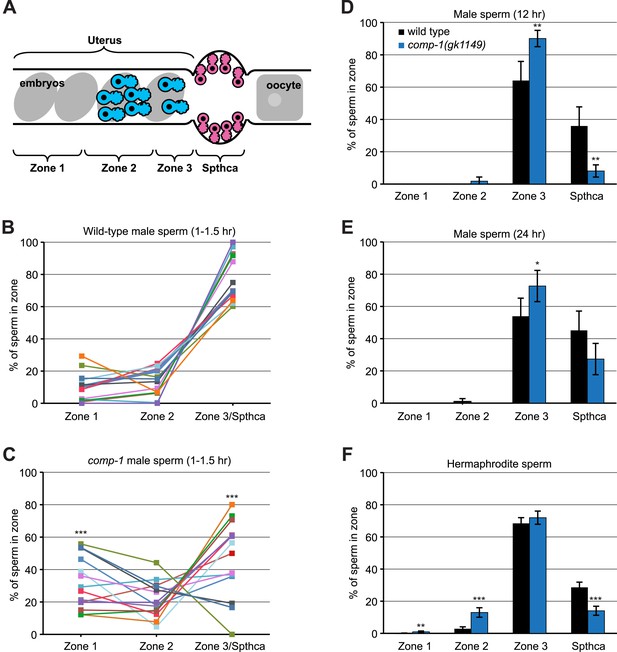
comp-1 sperm have defects in migration and spermathecal accumulation.
(A) Schematic of hermaphrodite gonad arm showing zones used to quantify sperm position. (B, C) Localization of wild-type (B) and comp-1(gk1149) (C) Mitotracker-labeled male sperm 1–1.5 hr after transfer to hermaphrodites. Percentage of total male sperm is shown. (D, E) Localization of jnSi118[GFP::H2B] male sperm 12 hr (D) and 24 hr (E) after transfer to hermaphrodites. Percentage of male sperm in the focal plane with maximum sperm in the spermatheca is shown. (F) Localization of hermaphrodite self sperm in 24 hr post-L4 unmated hermaphrodites. Animals were stained with DAPI to facilitate counting of sperm cells. Percentage of total hermaphrodite sperm is shown. Error bars, 95% confidence intervals. *, p < 0.05; **, p < 0.01; ***, p < 0.001; Student's t test.
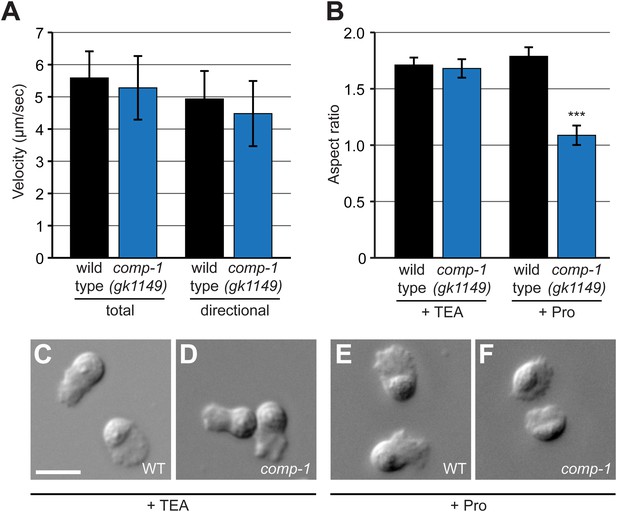
comp-1 male sperm can migrate normally but have context-dependent defects in cell morphology.
(A) comp-1(gk1149) sperm can migrate in vivo at speeds equivalent to wild-type sperm. Mitotracker-labeled males were crossed to N2 hermaphrodites and time-lapse images of sperm migrating through zone 2 were collected. Velocity and directional velocity toward the spermatheca were measured using ImageJ. (B–F) comp-1(gk1149) spermatids show reduced pseudopodial extension after activation by Pronase. (B) Quantification of aspect ratios of wild-type and comp-1(gk1149) sperm treated with either TEA or Pronase. (C–F) Representative images of wild-type (C, E) and comp-1(gk1149) (D, F) sperm treated with TEA (C,D) or Pronase (E,F). Error bars, 95% confidence interval; ***, p < 0.001, Kolmogorov–Smirnov test. Scale bar, 5 μm.
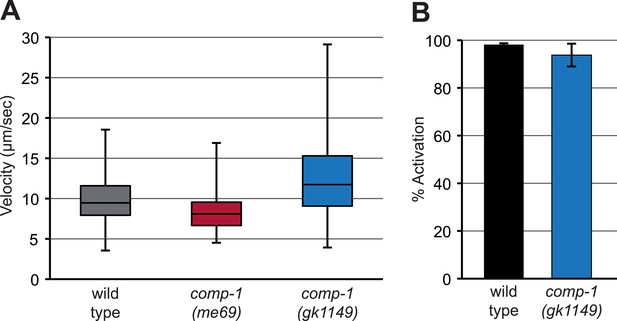
comp-1 sperm can crawl and be activated in vitro.
(A) comp-1 sperm can crawl at wild-type velocities in vitro. Wild-type and comp-1(gk1149) spermatids were treated with TEA for 30 min and velocity was obtained from time-lapse images collected every 30 s. As we observed a high level of variability among different samples for each genotype, the range of observed values is shown using a box-and-whiskers plot. For each genotype, n = 5–6 samples, 65–130 cells. (B) comp-1 sperm activate in Pronase. Wild-type and comp-1(gk1149) spermatids were treated with Pronase for 30 min and scored for activation based on the presence or absence of a pseudopod. Error bars, 95% confidence intervals.
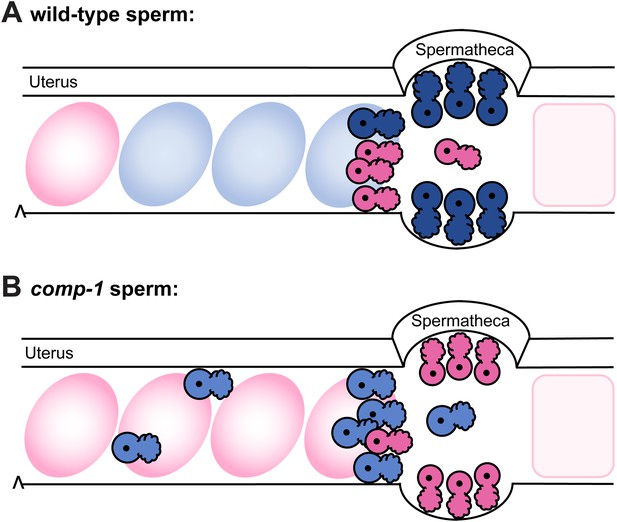
Model: comp-1 sperm have localization defects that result in failure to compete with wild-type sperm.
(A) Wild-type male sperm (blue) migrate to the region of the spermathecae, where they displace hermaphrodite self sperm (pink) and preferentially fertilize oocytes. Oocytes fertilized by male sperm are shown in blue; oocytes fertilized by self sperm are shown in pink. (B) comp-1 mutant male sperm (light blue) migrate to the spermathecae, but remain outside while wild-type sperm (pink) are present, and are thus excluded from opportunities to fertilize oocytes. They also show delayed migration to the spermathecal region and increased localization in the periphery of the female reproductive tract.
Tables
me69 is linked to chromosome I
| Marker* | Genetic position* | Genomic position† | Haw/+ frequency‡ |
|---|---|---|---|
| WBVar00240399 | I:0.91 | I:6350803 | 1/16 |
| WBVar00172772 | II:0.12 | II:6789208 | 8/16 |
| WBVar00067953 | III:−0.31 | III:8318640 | 10/16 |
| WBVar00188750 | IV:1 | IV:4625317 | 3/16 |
| WBVar00240687 | V:0.88 | V:8177520 | 9/16 |
-
*
-
†
WormBase WS243 (accessed 30 August 2014).
-
‡
me69; him-5 males were crossed to CB4856 Hawaiian hermaphrodites, F1 males were crossed back to me69; him-5 hermaphrodites, and F2 males were assayed for precedence defects in crosses to spe-8; dpy-4 hermaphrodites. Animals scoring as mutant (me69 homozygotes) were scored by PCR and restriction digest for centrally-located SNPs on each chromosome. Animals lacking Hawaiian alleles at all loci tested were considered self progeny and excluded from analysis.
Mapping of me69 on chromosome I
| No. F2s* | WBVar 00240 394† | WBVar 00240 397 | WBVar 00240 399 | WBVar 00155 231 | WBVar 00240 416 | WBVar 00240 407 | WBVar 00159 097 | WBVar 00240 414 | WBVar 00161 629 |
|---|---|---|---|---|---|---|---|---|---|
| 825026 | 5482531 | 6351803 | 8646304 | 10614690 | 11472093 | 12433167 | 13066381 | 14154889 | |
| 16 | B/B | B/B | B/B | ||||||
| 6 | H/B | B/B | B/B | ||||||
| 2 | H/B | B/B | B/B | B/B | B/B | B/B | B/B | H/B | H/B |
| 3 | B/B | B/B | B/B | B/B | B/B | B/B | B/B | H/B | H/B |
| 1 | H/B | H/B | H/B | B/B | B/B | B/B | B/B | B/B | B/B |
-
F2 males from the cross described in Table 1 were scored for SNPs across chromosome I. Animals were either homozygous Bristol (B/B) or heterozygous for the Hawaiian allele (H/B) at each SNP.
-
*
Number of F2 males showing each pattern.
-
†
SNP designation and genomic position on chromosome I. Wicks et al. (2001); WormBase.
Construction of entry plasmids used to generate targeting constructs
| Fragment description | Fragment length | Forward primer | Reverse primer | Vector | Plasmid name |
|---|---|---|---|---|---|
| comp-1 promoter | 712 | GGGACAACTTTGTATAGAAAAGTTGCCAGTTCCTCGCCTAGCTTTC | GGGACTGCTTTTTTGTACAAACTTGATGCTTTTGATTCGATAGATGATCC | pDONR P4-P1r | pJMH1 |
| comp-1 coding region | 1921 | GGGGACAAGTTTGTACAAAAAAGCAGGCTCAATGACGTTGGTCGAATCGAAAC | GGGACCACTTTGTACAAGAAAGCTGGGTCTTATTTGCGCTGGAATTGATC | pDONR 221 | pJMH2 |
| comp-1 coding region without stop codon | 1918 | GGGGACAAGTTTGTACAAAAAAGCAGGCTCAATGACGTTGTCGAATCGAAAC | GGGACCACTTTGTACAAGAAAGCTGGGTATTTGCGCTGGAATTGATC | pDONR 221 | pJMH3 |
| comp-1 3’ region | 561 | GGGGACAGCTTTCTTGTACAAAGTGGAAGAACTTACGGAAGAATATG | GGGGACAACTTTGTATAATAAAGTTGATGCGTTCTCATCAGGCTTC | pDONR P2r-P3 | pJMH4 |
| peel-1 coding region* without stop codon | 3279 | GGGGACAAGTTTGTACAAAAAAGCAGGCTGCTTAATGCGCTTTGGTAAG | GGGGACCACTTTGTACAAGAAAGCTGGGTCTGGATTTTCAACACTTGGATC | pDONR 221 | pJMH20 |
-
*
Description of targeting constructs used to generate transgenic worm strains
| Construct | Position 1 pDONR P4-P1r | Position 2 pDONR 221 | Position 3 pDONR P2r-P3 | Destination vector | Locus |
|---|---|---|---|---|---|
| Pcomp-1::comp-1::comp-1 3’ region | pJMH1 | pJMH2 | pJMH4 | pCFJ150* | ttTi5605 |
| Pcomp-1::H2B::GFP::comp-1 3’ region | pJMH1 | pCM1.35† | pJMH4 | pCFJ150 | ttTi5605 |
| Ppeel-1::comp-1::tbb-2 3’ region | Ppeel-1 [4-1]‡ | pJMH2 | pCM1.36† | pCFJ150 | ttTi5605 |
| Ppeel-1::comp-1::GFP::unc-54 3’ region | Ppeel-1 [4-1] | pJMH3 | pGH50§ | pCFJ150 | ttTi5605 |
| Ppeel-1::comp-1::mCherry::unc-54 3’ region | Ppeel-1 [4-1] | pJMH3 | mCherry::unc-54 3’ region§ | pCFJ150 | ttTi5605 |
| Ppeel-1::peel-1::GFP::unc-54 3’ region | Ppeel-1 [4-1] | pJMH20 | pGH50 | pCFJ212* | cxTi10816 |
-
*
-
†
-
‡
-
§





