Siglec receptors impact mammalian lifespan by modulating oxidative stress
Figures
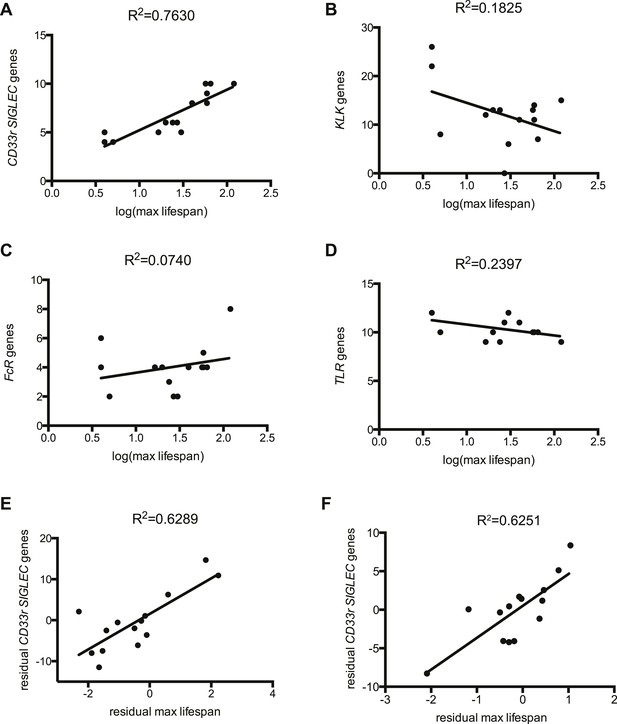
Correlation between gene numbers in gene families and maximum lifespan in mammals.
Numbers of CD33rSiglecs (A), KLK (B), IgG Fc receptors (C), and TLRs genes (D) and maximum lifespan in 14 mammalian species listed in Figure 1—figure supplement 1. (E and F) Correlation of CD33rSIGLECs and maximum lifespan after correction for average adult body weight and phylogeny. PGLS: λ = 1, phylogenetic tree I (E) or tree II (F) were used. The Pearson's correlation coefficient (R2) for each plot is indicated.
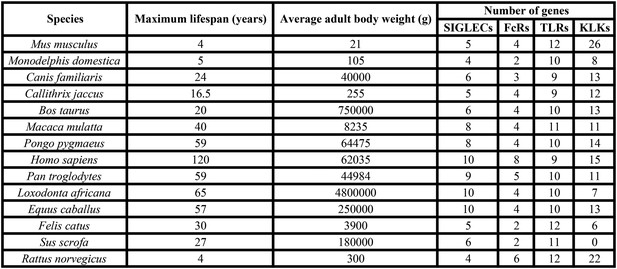
Data of 14 mammalian species used for analysis of correlation.
Maximum lifespan and average adult body weight were retrieved from the AnAge database. The number of genes of each family was either found in the literature or searched following the methodology described in the ‘Materials and methods’.
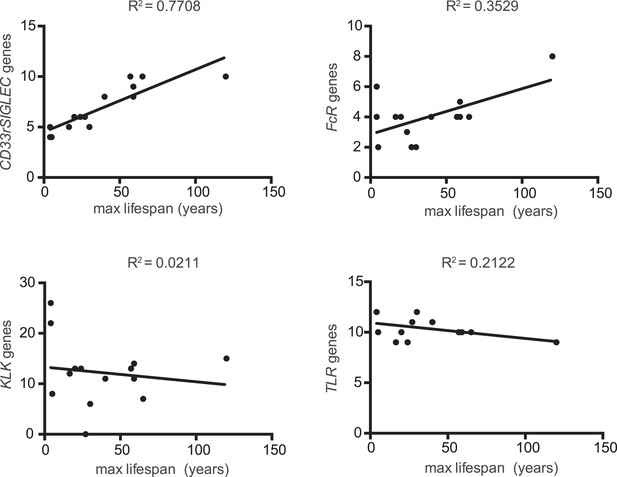
Correlation between number of genes of each family and maximum lifespan.
Data are shown in a linear scale, whereas are presented in a logarithmic scale in Figure 1 for statistical reasons. The correlation coefficient (R2) is indicated for each plot.
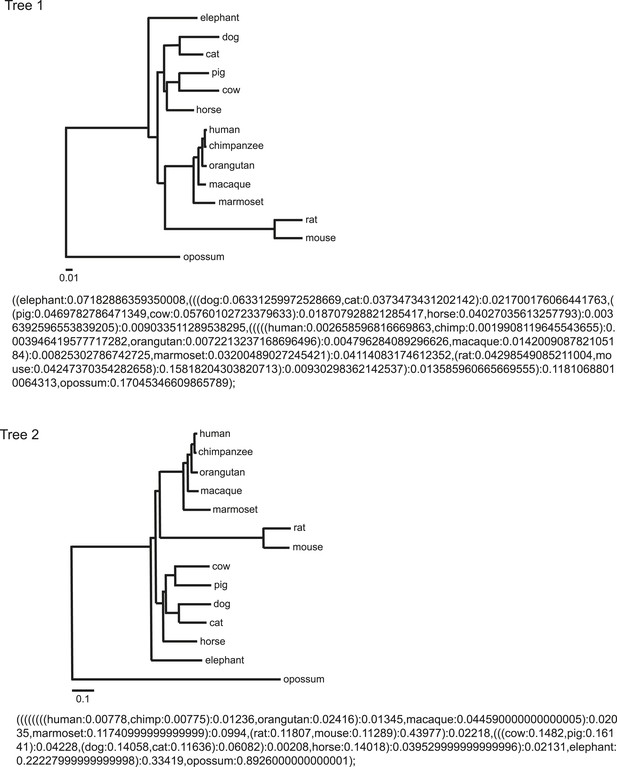
Phylogeny and tree branch information.
Phylogeny and tree branch information used in this study were obtained from Prasad et al. (2008) and trimmed by Archaeopteryx 0.957 (Zmasek and Eddy, 2001; Han and Zmasek, 2009) down to the same 14 species. The topology is also indicated in the Newick tree format.
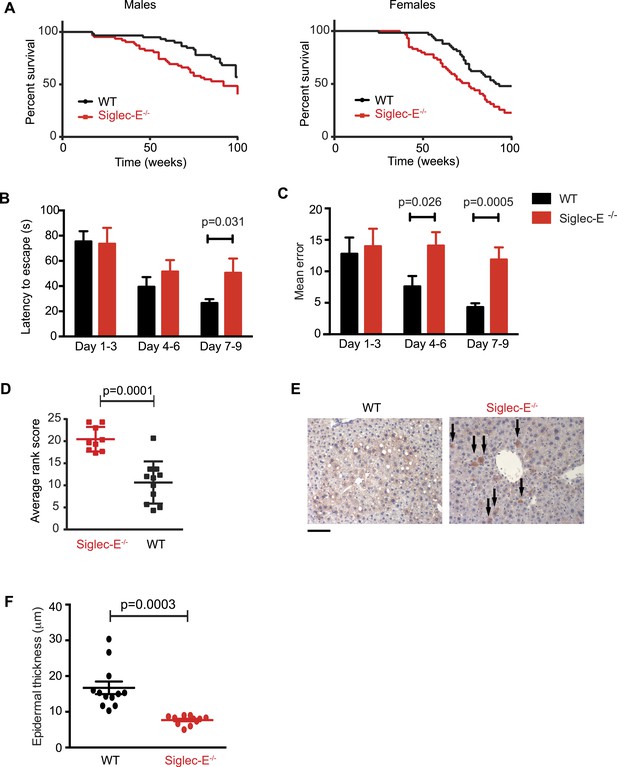
Absence of immunomodulatory Siglec-E aggravates aging phenotypes and reduces lifespan in mice.
(A) Survival curves of WT and Siglec-E−/− male (n = 59–62) and female (n = 58–59) littermates. Data are from two independent cohorts (cohort 1 included 31 WT and 38 Siglec-E−/− males, 31 WT and 33 Siglec-E−/− females. Cohort 2 included 28 WT and 23 Siglec-E−/− males, 27 WT and 26 Siglec-E−/− females). Log-rank test analysis showed significant differences in the survival curves both in males and females (males: χ2 = 5.833, d.f. = 1 and p = 0.0157; females: χ2 = 8.821, d.f. = 1 and p = 0.0030). (B and C) Mice at 80 weeks were assessed for spatial learning and memory via Barnes maze. Latency to escape (B) and number of errors before finding the escape box (C) are indicated in 3-day interval. Error bars reflect mean ± s.e.m. (n = 11). p was calculated with a Student's t test. (D) Hair graying of males was evaluated by three independent observers in a blind test. Average rank scores for each mouse are indicated. Error bars reflect mean ± s.e.m. (n = 9–11). p was calculated with a Student's t test. (E) Surviving mice were sacrificed at 100 weeks of age. Representative field of β-galactosidase staining of liver. Arrows indicate cells with increased localized staining. Scale bar is 100 μm. (F) Skin epidermal thickness was measured for WT and Siglec-E−/−. Mean and s.e.m. are indicated, n = 12, p was calculated with a Student's t test.
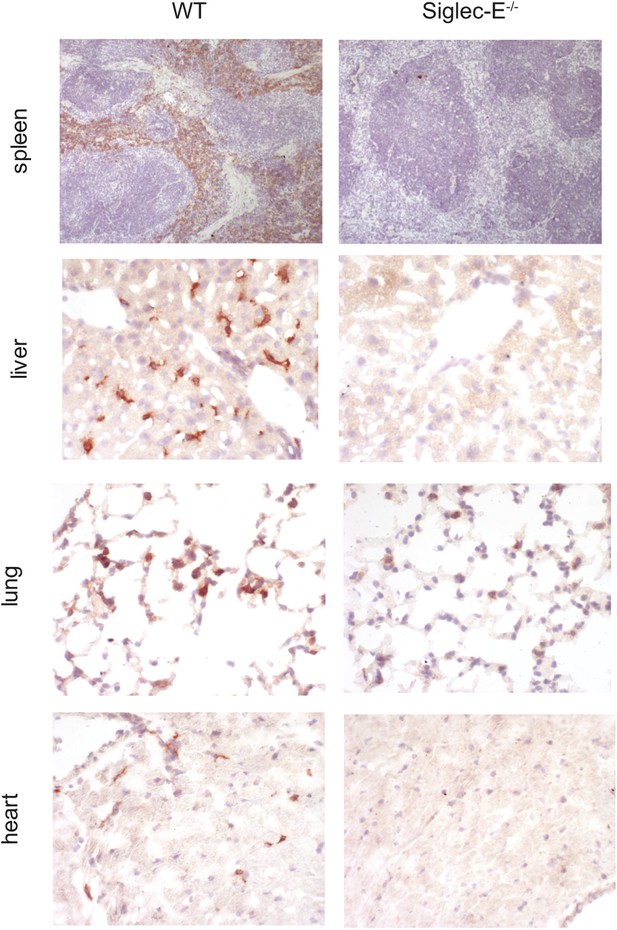
Expression of Siglec-E in mouse tissues.
Sections of frozen organs from WT and Siglec-E−/− animals were stained with anti-Siglec-E antibodies. Siglec-E is expressed in splenocytes and in tissue macrophages in liver, heart, and kidney.
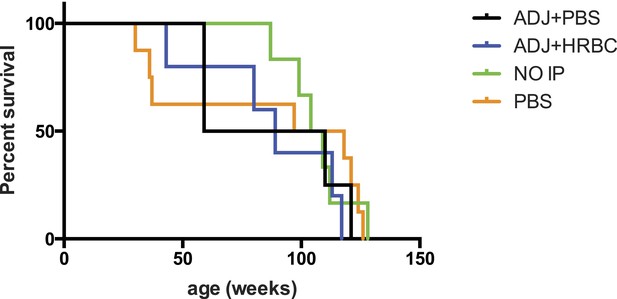
Exposure to human red blood cell membranes does not impact survival of Siglec-E−/− mice.
Survival curves of mice intraperitoneally (IP) injected with human red blood cell membranes and Freund's adjuvant (ADJ + HRBC). Controls mice were injected with adjuvant and PBS (ADJ + PBS) or PBS only. There are no significant differences between groups. Log-rank (Mantel–Cox) test: Chi square = 1.310, df = 3, p = 0.7268. Gehan-Breslow-Wilcoxon test: Chi square = 0.6019, df = 3, p = 0.8960.
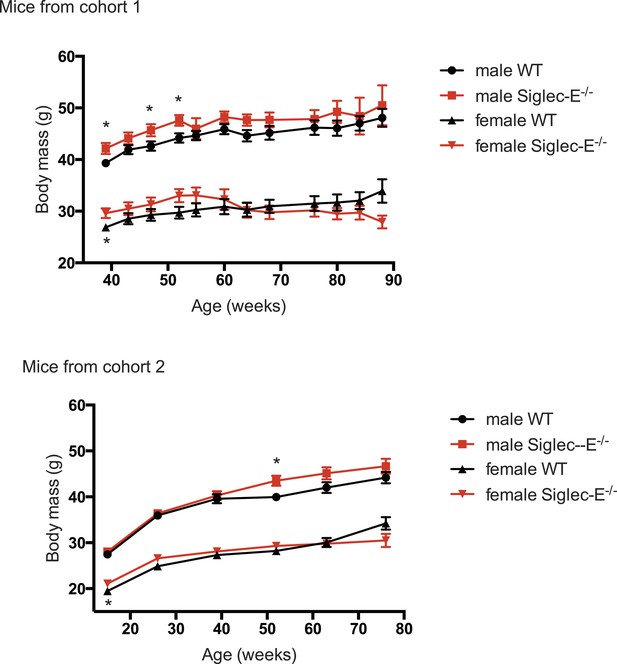
Deletion of Siglec-E does not alter body weight increase.
Weights of mice were recorded for mice from the two cohorts characterized in this study. Indicated are mean ± sem, * indicates p < 0.05.
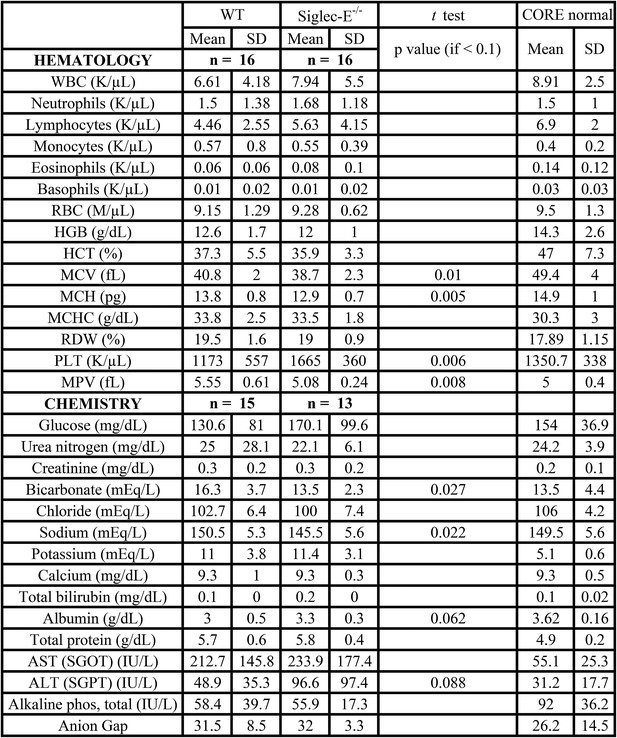
Hematology and serum chemistry values of male mice at the termination of the study.
https://doi.org/10.7554/eLife.06184.013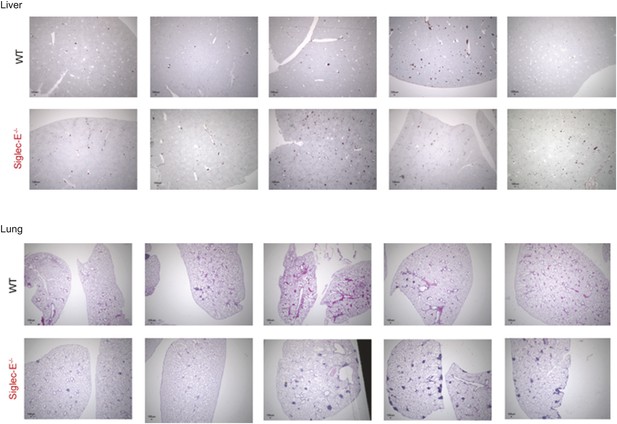
Absence of Siglec-E is associated with overall increased inflammation in liver and lung.
Paraffin sections of liver or lungs from WT and Siglec-E−/− animals were analyzed for inflammation using anti-CD45 immunohistochemistry and demonstrate many more areas with accumulation of inflammatory cells in Siglec-E−/− age mice.
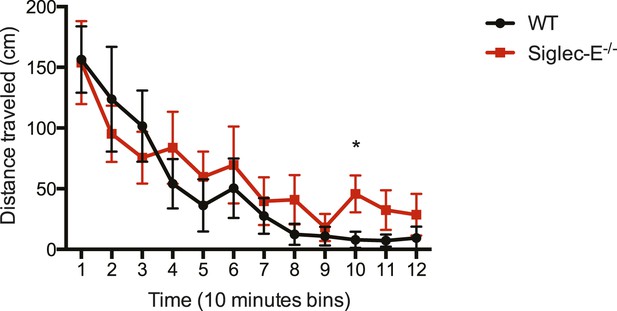
Deletion of Siglec-E does not affect locomotor activity.
WT and Siglec-E−/− mice show habituation of activity (a significant decrease) across the 2 hr test in the activity chamber. There are no differences between the groups. Indicated are mean ± sem, * indicates p < 0.05, n = 7–11.
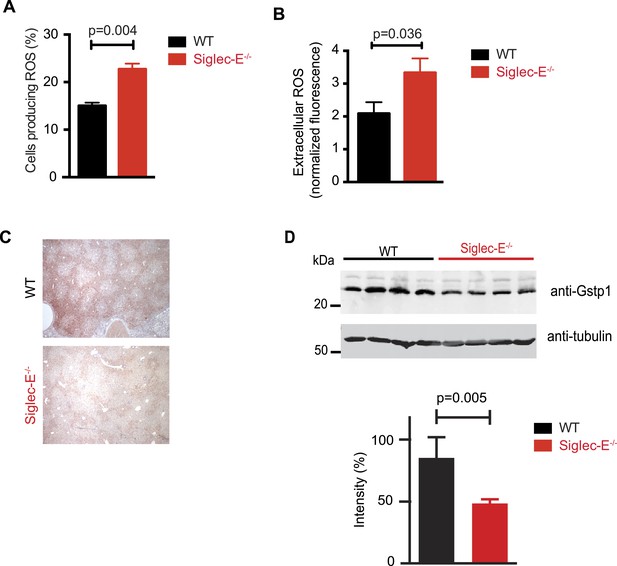
Altered ROS homeostasis in mice lacking Siglec-E.
(A) Neutrophils purified from bone marrow were incubated with immunocomplexes. Cells producing vacuolar ROS were measured by flow cytometry after 60 min. Representative of three experiments, for each n = 3. (B) Neutrophils secrete ROS upon stimulation with PMA for 60 min. Extracellular ROS were detected with a probe that does not cross the plasma membrane (n = 11–12). (C) Representative Gstp1 immunohistochemistry in liver from WT or Siglec-E−/− male mice at 100 weeks. Expression pattern is altered in the knockout mice. (D) Immunoblot analysis and quantification of Gstp1 expression in liver of 100-week-old mice. The level of Gstp1 protein is reduced of about 40%. p was calculated with a Student's t test, n = 4.
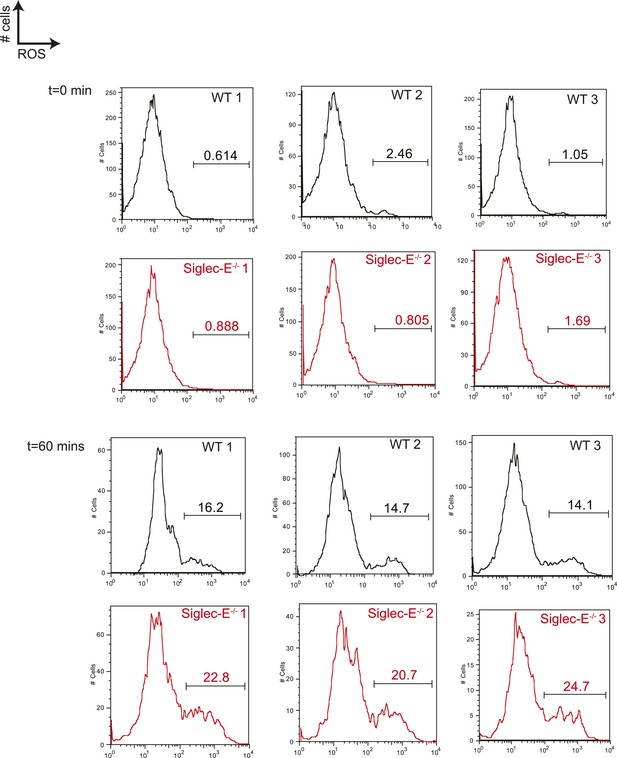
Neutrophils lacking Siglec-E are more prone to oxidative burst.
Neutrophils isolated from bone marrow were incubated with immunocomplexes conjugated with a probe sensitive to ROS. Flow cytometry profiles at 60 min indicate that cells lacking Siglec-E are more prone to produce ROS, whereas the levels of ROS are comparably low before stimulation. Gates indicate cells producing ROS.
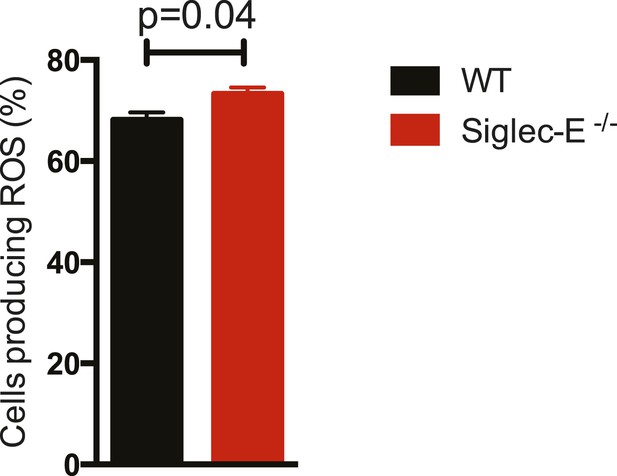
Thioglycollate-elicited neutrophils from Siglec-E−/− produce higher ROS than WT controls.
Neutrophils isolated from peritoneum were incubated with immunocomplexes conjugated with a probe sensitive to ROS. The percentage of cells producing ROS after 60 min of incubation is higher when Siglec-E is absent, n = 3.
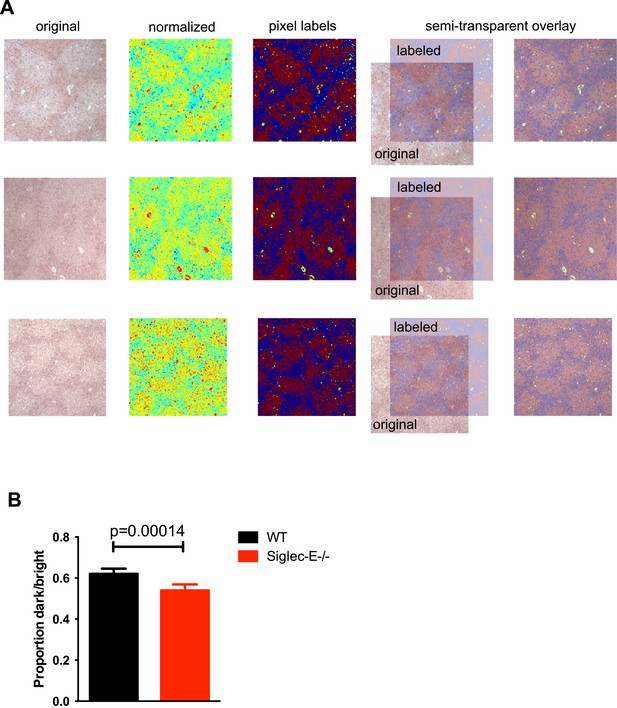
Gstp1 is found in lower levels in the liver of Siglec-E−/− mice.
(A) Representative images generated during analysis of Gstp1 staining of liver tissues. Original images (left) were converted to grayscale to discard color information and only indicate brightness. Normalized images are shown in the second column (with jet color map). Images with pixel labels are in the third column (red: bright pixels, blue: dark pixels, green: discarded from analysis). In the last column, the labeled images are overlayed on the original image to compare pixel labels to the original (30% transparency). Three images per liver sample were acquired and processed in this way. (B) Quantification of the Gstp1 staining, as measured by the proportion of dark/bright areas. Mean ± sem, n = 8.
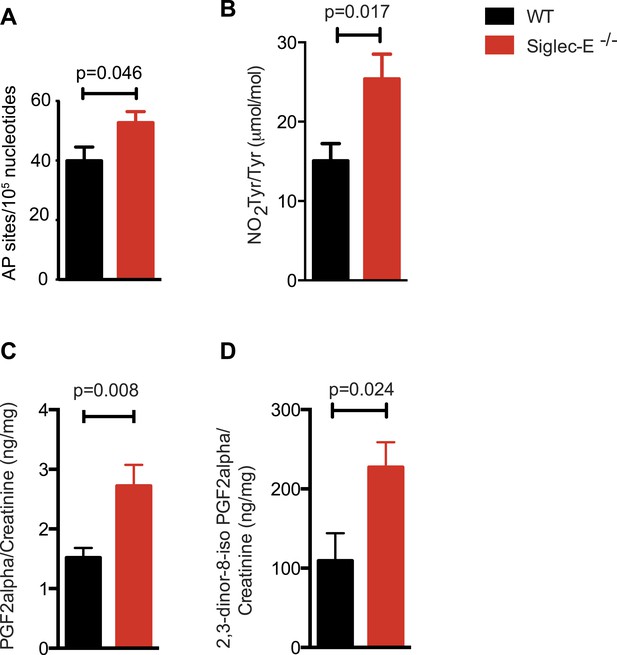
Increased oxidative damage in mice lacking Siglec-E.
(A) Oxidative damage of DNA (AP sites, apurinic/apyrimidinic sites) was measured in the liver from WT and Siglec-E−/− mice at 100 weeks, n = 10. (B) Nitrotyrosine (NO2Tyr) accumulation in plasma proteins of Siglec-E−/− mice, n = 8. (C and D) Concentrations of F2-isoprostanes in urine derived from free radical-induced oxidation of arachidonic acid. Values were normalized by creatinine levels to account for dilution in urine. F2-Isoprostane levels are significantly higher in Siglec-E−/− mice. Data are mean ± s.e.m., Student's t test.
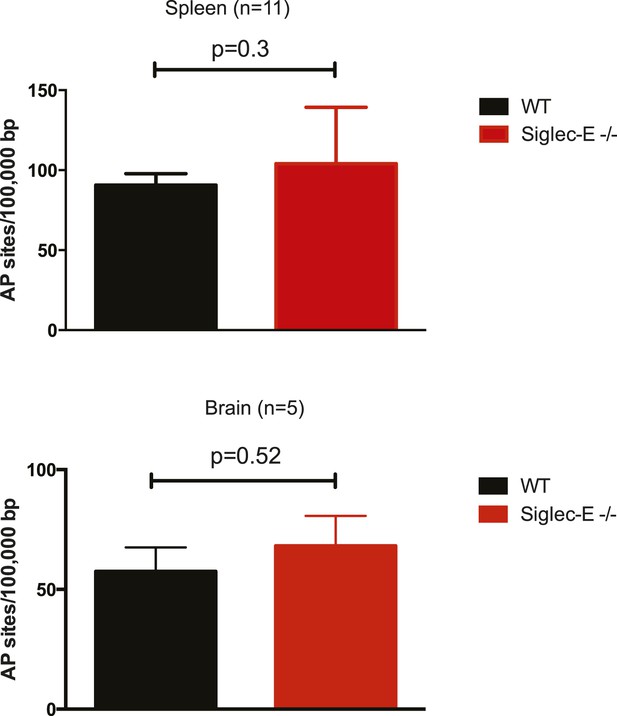
Spleen and brain of aged WT and mutant mice have equivalent levels of DNA damage.
Quantification of apurinic/apyrimidinic (AP) sites in DNA from spleen or brain. Mean ± sem is indicated.
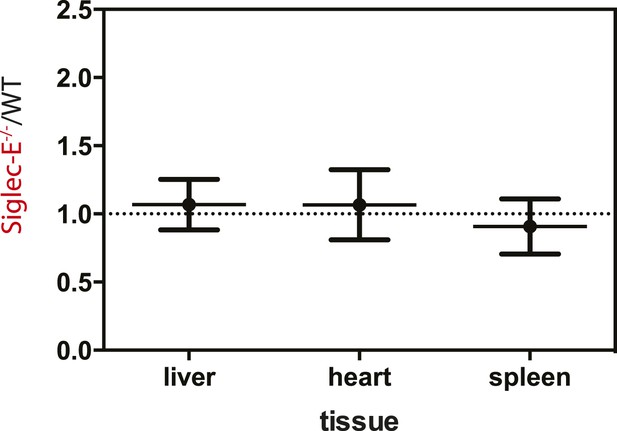
Young Siglec-E−/− mice exhibit levels of DNA damage comparable to WT.
Quantification of AP sites in DNA from liver, heart, or spleen of 10-week-old mice. DNA damage is expressed as ratio between DNA damage in Siglec-E−/− and WT animals, n = 3.
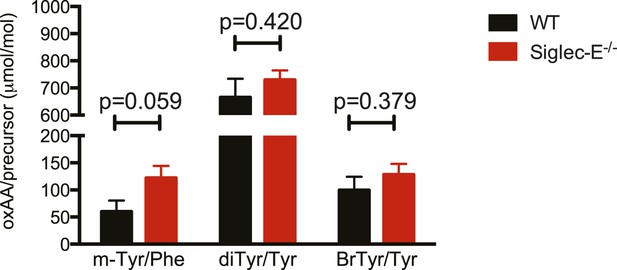
Oxidation of hepatic proteins.
Protein-bound oxidized amino acid content (normalized to the precursor amino acid) in proteins from liver homogenates was determined by stable isotope dilution LC/MS/MS analysis. Methyl-tyrosine (m-Tyr) is a stable adduct of phenylalanine (Phe). Brominated tyrosine (BrTyr) is formed by brominating oxidants. Di-tyrosine (Tyr) is an oxidative crosslink. Indicated is mean ± sem, n = 8.
Tables
Statistical analysis of the correlation between number of genes and maximum lifespan, corrected for phylogeny
| Gene family | PGLS | FIC |
|---|---|---|
| Tree I | ||
| CD33rSIGLECs | 0.00016 | 0.00012 |
| KLKs | 0.49 | 0.17 |
| TLRs | 0.38 | 0.10 |
| IgG Fc receptors | 0.35 | 0.0016 |
| Tree II | ||
| CD33rSIGLECs | 0.00017 | 0.00011 |
| KLKs | 0.65 | 0.23 |
| TLRs | 0.32 | 0.14 |
| IgG Fc receptors | 0.39 | 0.0019 |
-
Phylogenetic comparative analysis was conducted in COMPARE 4.6b using Phylogeny Generalized Least-Squares (PGLS) or Felsenstein's Independent Contrast (FIC) approaches. Student's t-values were computed based on the regression slopes and the standard errors. Two-tailed probability (p) value of a Student's t-test was estimated using a degree of freedom of 11. The phylogenetic relationship of 14 mammalian species represented by Tree I and Tree II are indicated in Figure 1—figure supplement 3. Note that although FIC analysis obtained a significant p value for IgG Fc receptor gene family, the p value increased to 0.56 when human data was excluded. Thus, this correlation is driven by one outlier data point.
Statistical analysis of the correlation between number of genes and maximum lifespan, corrected for body weight
https://doi.org/10.7554/eLife.06184.008| Gene family | t value | p value |
|---|---|---|
| Tree I | ||
| CD33rSIGLECs | 3.521435 | 0.004786 |
| KLKs | 0.360601 | 0.725226 |
| TLRs | −0.701822 | 0.497370 |
| IgG Fc receptors | 1.656519 | 0.125834 |
| Tree II | ||
| CD33rSIGLECs | 3.660917 | 0.003748 |
| KLKs | 0.180227 | 0.860251 |
| TLRs | −0.726465 | 0.482726 |
| IgG Fc receptors | 1.589586 | 0.140235 |
-
Phylogenetic comparative analysis conducted in CAIC package. Average adult body weight and maximum lifespan of 14 mammalian species were log-transformed and phylogenetic regressions were run using pglmEstLambda in the CAIC package in R. This function uses the PGLS method and estimates λ with the average adult body weight controlled for. Student's t-values and two-tailed probability (p) values are shown. The phylogenetic relationship of 14 mammalian species represented by Tree I and Tree II are indicated in Figure 1—figure supplement 3.
Additional files
-
Supplementary file 1
Pathway analysis, calculated with Ingenuity Pathway Analysis software. The p-value is a measure of the likelihood genes in the process appear by chance and it was calculated by right-tailed fisher exact test. The z-score is calculated by IPA software algorithm, and predicts the change in the biological function direction. A z-score ≥ 2 or ≤ −2 is considered significant.
- https://doi.org/10.7554/eLife.06184.024





