Recurrent gain of function mutation in calcium channel CACNA1H causes early-onset hypertension with primary aldosteronism
Figures
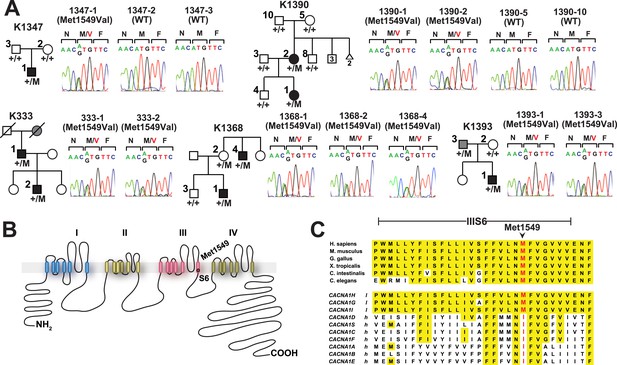
Kindreds with hypertension and primary aldosteronism (PA) with CACNA1HM1549V mutation at conserved position of S6 domain.
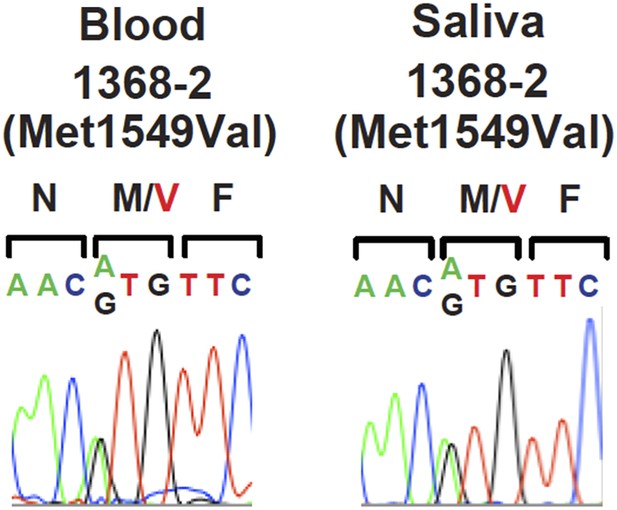
(A) Pedigrees of kindreds with CACNA1HM1549V mutation are shown. Studied subjects with early-onset hypertension are shown as black filled symbols, and subjects with early-onset hypertension by family history (K333) or low renin with normal blood pressure (K1393) are shown as grey filled symbols. Genotypes are indicated below each symbol (+/+ denotes wild type sequence; +/M denotes heterozygosity for CACNA1HM1549V variant). Corresponding Sanger sequencing results for selected subjects are depicted to the right. (B) Transmembrane structure of CaV3.2 (encoded by CACNA1H), the pore-forming subunit of a voltage-gated Ca2+ channel, is shown. These channels have four internal homologous repeats (I–IV), each with six transmembrane segments (S1–S6) and a membrane-associated loop between the pore-forming S5 and S6 segments. The p.Met1549Val mutation is located in S6 of repeat III. (C) Conservation of CACNA1HM1549 in CACNA1H orthologs and paralogs. The amino acid sequences of the S6 segment of domain III of CACNA1H, orthologs and paralogs are shown. The S6 segment, including Met1549, is virtually completely conserved (highlighted in yellow) among orthologs and all paralogs that are activated by small changes in membrane potential (l, low voltage-activated) but not those activated by large changes (h, high voltage-activated). M1549 is part of the Met-Phe-Val sequence that is implicated in rapid channel inactivation (Marksteiner et al., 2001).
-
Figure 1—source data 1
Source data corresponding to Figure 1.
- https://doi.org/10.7554/eLife.06315.004
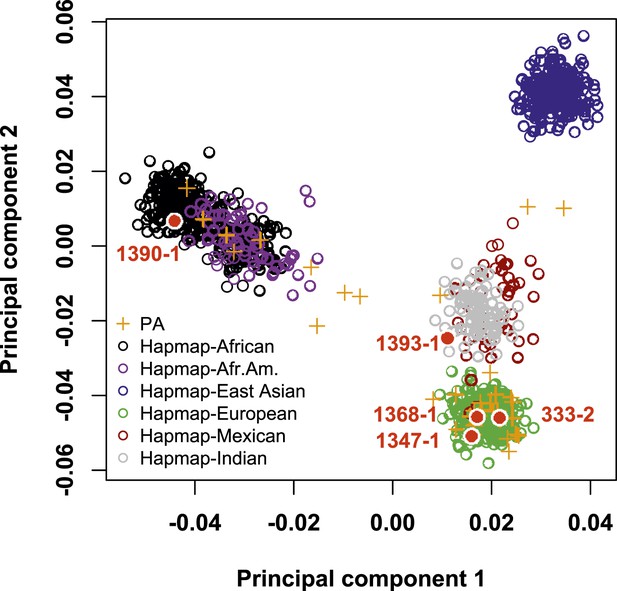
Cohort population structure by principal component analysis (PCA).
PCA of subjects referred for PA. Individuals in the cohort (orange crosses) mostly cluster with HapMap subjects of European and African American subjects. The five individuals with CACNA1H mutation (filled red circles) are of African American (1390-1), Hispanic (1393-1) and European (1368-1, 1347-1 and 333-2) origin, respectively, by history and PCA. Source files are available in Figure 1—source data 1.
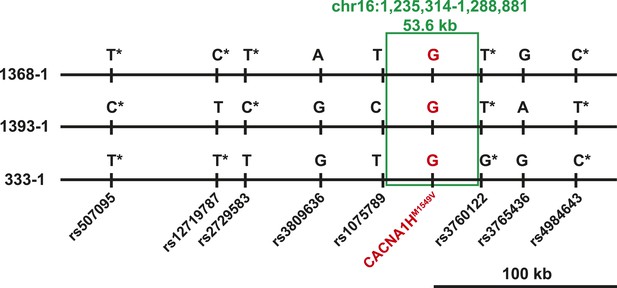
Shared haplotypes in subjects with inherited CACNA1HM1549V variant.
Haplotypes of three affected individuals from kindreds without proven de novo occurrence of CACNA1HM1549V variant were phased using BEAGLE (‘Materials and methods’) (Browning and Browning, 2007). This analysis identified a very small maximum interval shared among all three individuals (∼53.6 kb, green box) flanked by rs1075789 and rs3760122. If only homozygous discordant calls (*) were considered in the absence of phasing, the maximum interval shared by all three subjects would be 127.1 kb and the longest pairwise shared haplotype would be 200.0 kb between 1393-1 and 333-1.
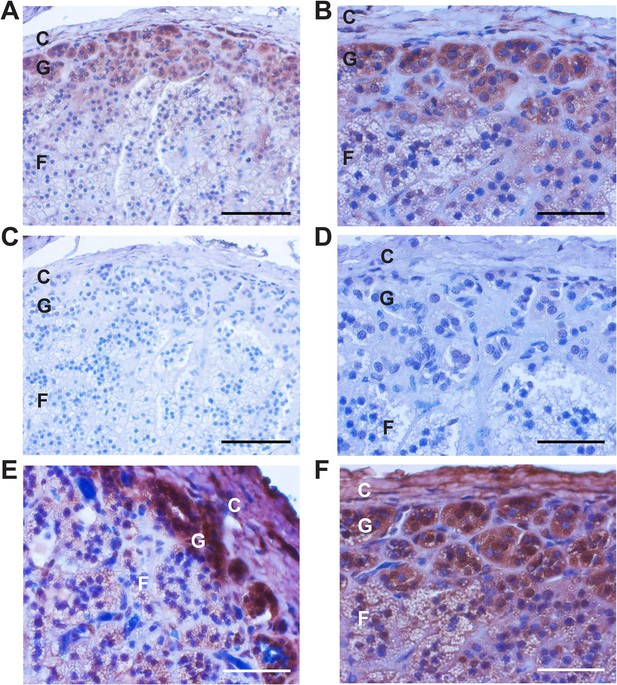
Immunohistochemistry of CaV3.2 in normal human adrenal gland.
Sections of normal human adrenal are shown. C denotes adrenal capsule; G, glomerulosa; F, fasciculata. (A) Normal adrenal gland stained with hematoxylin and an antibody to CaV3.2 (Alomone). (B) Higher power image of adrenal in panel (A). (C, D) Absence of staining after preincubation of the antibody with the antigenic peptide, demonstrating specificity. (E) A second normal human adrenal gland stained for CACNA1H as in (A, B). (F) Gland from (A–D) stained with a second α-CACNA1H antibody (Santa Cruz). Scale bars, 100 μm (A, C); 50 μm (B, D, E, F). The results demonstrate expression of CaV3.2 in the normal zona glomerulosa, which is only several cells in depth.
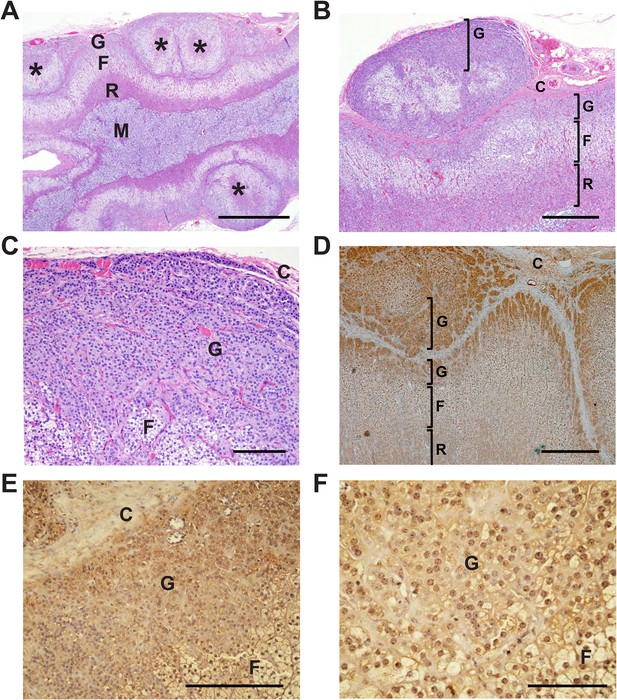
Glomerulosa hyperplasia in adrenal gland of subject 1390-2 with CACNA1HM1549V mutation.
C denotes adrenal capsule; G, glomerulosa; F, fasciculata; R, reticularis; M, medulla. (A) Low power image stained with hematoxylin and eosin. Scale bar 1000 μm. (B, C) Higher power images of adrenal from panel (A), scale bars 400 μm (B) or 100 μm (C). The mutant adrenal shows marked zona glomerulosa hyperplasia, with micronodular invasion of the capsule (denoted by *). (D) Same adrenal gland stained with hematoxylin and antibody to CaV3.2 (Santa Cruz), demonstrating specific staining of zona glomerulosa. Scale bar, 400 μm. (E, F), higher power images stained with second antibody to CaV3.2 (Alomone). Scale bars 250 μm (E) or 100 μm (F). CaV3.2 is expressed in the hyperplastic zona glomerulosa.
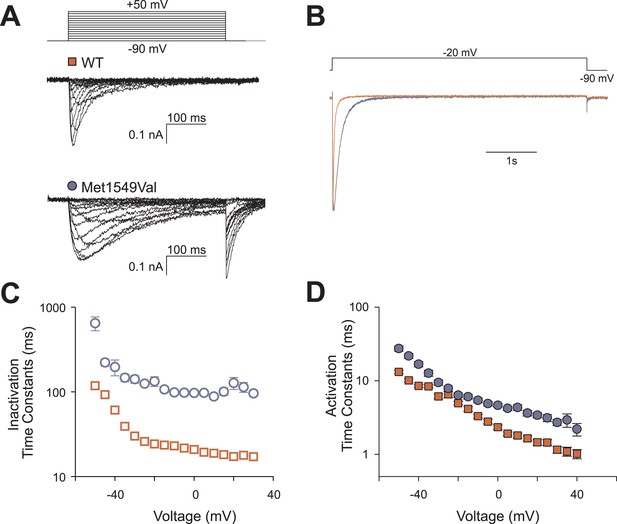
CACNA1HM1549V impairs channel inactivation.
Whole-cell patch clamp recordings were performed in HEK293T cells transfected with CACNA1HWT or CACNA1HM1549V. (A) Cells were held at −90 mV, and voltage steps between −90 and +50 mV were applied to elicit calcium currents, followed by a step to −90 mV to evoke tail currents. Representative recordings show rapid activation and inactivation of CACNA1HWT currents and delayed inactivation of CACNA1HM1549V. Tail currents are exclusively present in CACNA1HM1549V and suggest the presence of non-inactivated mutant channels at the end of the depolarizing pulse. (B) Tail currents are still present after a 5-s pulse to −20 mV. The fraction of non-inactivated channels after 5 s was determined by dividing the peak amplitude at −20 mV before and after 5 s long pulses to voltages between −90 and −20 mV in 5 mV increments (CACNA1HM1549V: 6.7 ± 1%, N = 12; CACNA1HWT: 2.4 ± 0.5%, N = 9; p = 0.004, protocol not shown in figure). (C) Exponential fits of the current decay between −50 and +30 mV provide inactivation time constants. Data from CACNA1HM1549V are shown in blue circles, CACNA1HWT data are shown in red squares. The mutant channel shows almost 10-fold slower inactivation than wild-type (N = 9 for CACNA1HWT, N = 7–14 for CACNA1HM1549V, p < 0.001 across all voltages studied, Mann–Whitney rank sum test). (D) In contrast, activation time constants at different voltages are only slightly slower in CACNA1HM1549V compared to WT (cf. ‘Materials and methods’ for details). Source files are available in Figure 5—source data 1.
-
Figure 5—source data 1
Source data corresponding to Figure 5.
- https://doi.org/10.7554/eLife.06315.012
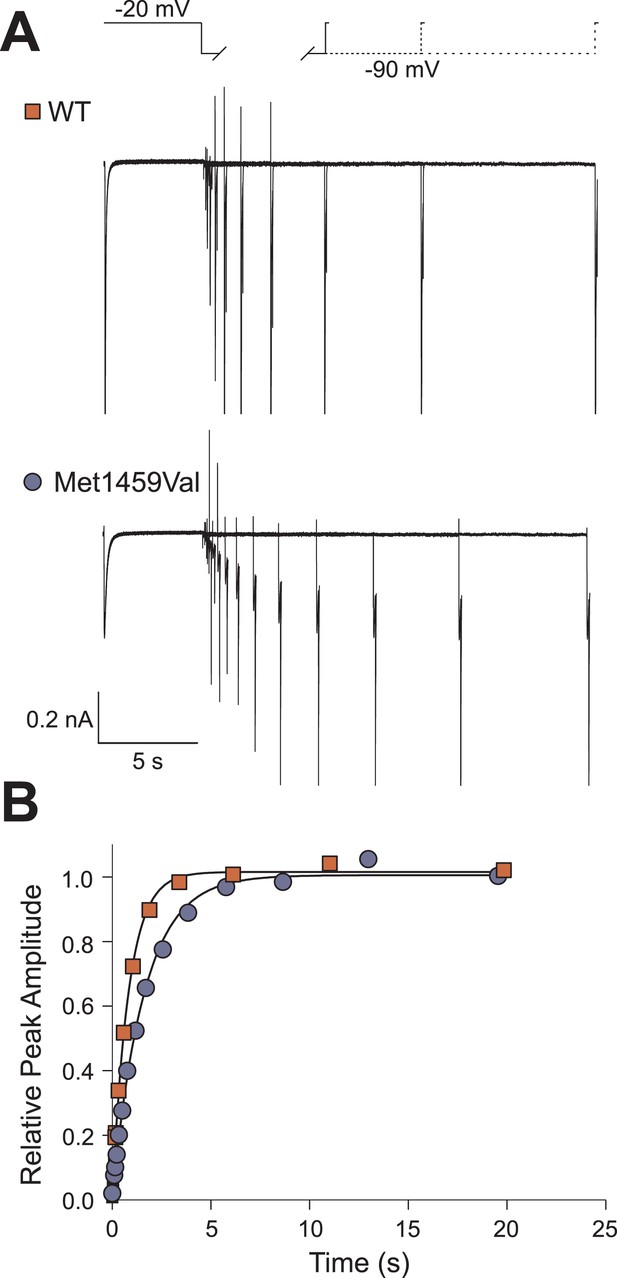
Recovery from inactivation is slightly slower in CACNA1HM1549V.
(A) The recovery from inactivation at −90 mV is decelerated in CACNA1HM1549V. Representative current recordings of CACNA1HWT or CACNA1HM1549V channels. Channels were activated and subsequently inactivated by clamping the membrane potential to −20 mV for 5 s. Afterwards, cells were held at −90 mV for increasing durations followed by short activation at −20 mV. The peak amplitude at the last −20 mV step is dependent on the number of non-inactivated channels that increases upon longer intervals at −90 mV. (B) Monoexponential fits to the plot of the relative peak amplitudes vs the time spent at −90 mV reveal a slight delay in the recovery from inactivation of CACNA1HM1549V channels (time constants for CACNA1HWT: 871.4 ± 52.6 ms, n = 6; CACNA1HM1549V: 1689.0 ± 70.9 ms, n = 10; p = 1 × 10−6). Source files are available in Figure 5—source data 1.
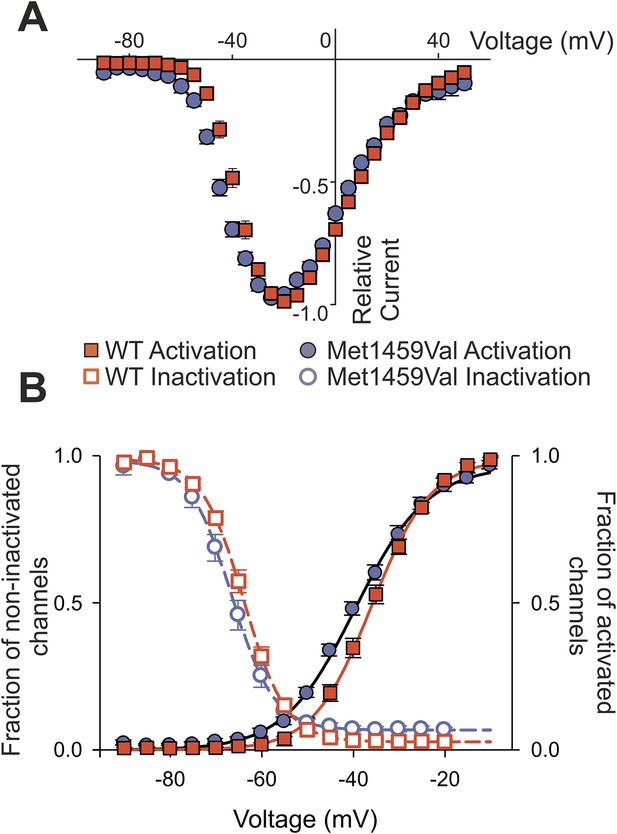
CACNA1HM1549V shifts activation to more hyperpolarized potentials.
(A) Current-voltage plots and (B) activation curves show a shift of V1/2 for activation of the mutant channel to less depolarizing potentials. The voltage dependence of inactivation is shown as open circles or squares. For CACNA1HM1549V, the area under the intersection of activation and inactivation curves (where a fraction of channels show continuous activity) is larger and shifted to more hyperpolarized potentials compared to CACNA1HWT, allowing for increased constitutive Ca2+ influx at potentials close to the resting potential of zona glomerulosa cells (Hu et al., 2012). Source files are available in Figure 6—source data 1.
-
Figure 6—source data 1
Source data corresponding to Figure 6.
- https://doi.org/10.7554/eLife.06315.015
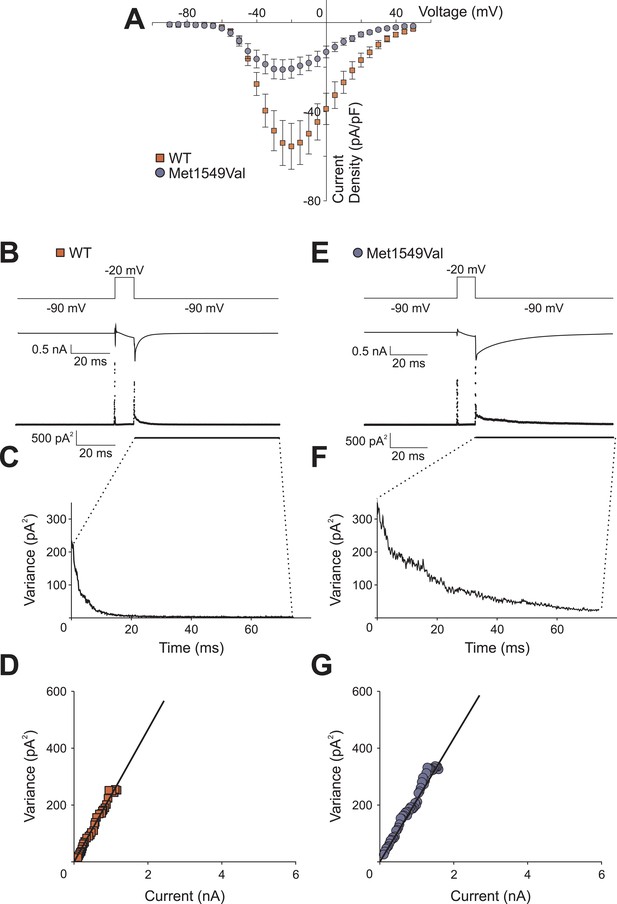
CACNA1HM1549V and CACNA1HWT whole-cell current densities and non-stationary noise analysis.
(A) Whole-cell peak currents were divided by the cell capacitance as determined by the amount of capacitance compensation. CACNA1HM1549V shows a decreased current density compared to CACNA1HWT albeit with large variability. (B, E) Representative mean currents from more than 200 traces recorded from one cell expressing CACNA1HWT (B) or CACNA1HM1549V (E) channels. (C, F) Analysis of the variance at the −90 mV tail pulse reveals a time dependent decrease. (D, G) A plot of the variance vs the current only allows for a small initial part of the expected parabolic distribution to be visible. Linear fits (black line) reveal similar single channel amplitudes (CACNA1HWT: 273.7 ± 3.2 fA, n = 3; CACNA1HM1549V: 285.0 ± 17.3 fA, n = 6; p = 0.67), but cannot be used to determine absolute open probabilities. Source files are available in Figure 6—source data 1.
Tables
Clinical features of index cases with CACNA1HM1549V
| Subject ID | Gender | Age dx | BP (%ile) | Aldo (ng/dl) | PRA (ng/ml/hr) | ARR (ng/dl: ng/ml/hr) |
|---|---|---|---|---|---|---|
| 1347-1 | M | 3 yrs | 160/105 (>99th) | 20 | <0.1 | >200 |
| 1390-1 | F | 7 yrs | 150/90 (>99th) | 66 | 0.2 | 330 |
| 1368-1 | M | 8 yrs | 140/90 (>99th) | 20 | <0.2 | >100 |
| 333-2 | M | 9 yrs | 192/144 (>99th) | 40 | <0.7 | >57 |
| 1393-1 | M | 2 mos | 170/110 (>99th) | 87 | <0.6 | >145 |
-
M, male; F, female; age dx, age at diagnosis of hypertension; yrs, years; mos, months; BP, blood pressure; (%ile), percentile adjusted for age and gender; Aldo, serum aldosterone; PRA, plasma renin activity; ARR, aldosterone:renin ratio, values >20 with aldosterone level >15 are considered indicative of primary aldosteronism (PA).
Laboratory values of carriers and non-carriers of CACNA1HM1549V in kindred 1390
| Subject ID | Gender | Age (yrs) | K+ (mmol/l) | Aldo (ng/dl) | PRA (ng/ml/hr) | ARR (Aldo/PRA) | Direct renin (μIU/ml) | Aldo/direct renin |
|---|---|---|---|---|---|---|---|---|
| Carriers | ||||||||
| 1390-1 | F | 15 | 3.7 | 37 | 0.42 | 88.1 | NA | NA |
| 1390-2 | F | 29 | 3.5 | 22 | NA | NA | 3 | 7.3 |
| Non-Carriers | ||||||||
| 1390-4 | M | 17 | 4.3 | 2 | 1.65 | 1.2 | NA | NA |
| 1390-8 | M | 37 | 3.8 | <1 | 2.25 | <0.4 | NA | NA |
| 1390-5 | F | 62 | 4.1 | 16 | 18.92 | 0.8 | NA | NA |
| 1390-6 | F | 51 | 4.0 | 4 | 1.87 | 2.1 | NA | NA |
| 1390-7 | F | 46 | 4.3 | 3 | 1.67 | 1.8 | NA | NA |
-
M, male; F, female; Age (yrs), Age in years when sample was obtained; K+, serum potassium (reference 3.5–5.5 mmol/l); Aldo, serum aldosterone; PRA; direct renin <5 is indicative of volume-mediated hypertension; ARR, aldosterone:renin ratio, values >20 (using PRA) or values of aldo/direct renin >2.4 with aldosterone level greater than 15 are considered indicative of PA. Blood samples were drawn on the same day, and values were determined in the same laboratory (except for 1390-2, in whom values are pre-adrenalectomy at age 29). See Figure 1 for relationships. 1390-6 and −7 are not included in Figure 1, and are sisters of 1390-5.
Additional files
-
Supplementary file 1
(A) Clinical features of 40 patients with primary aldosteronism. (B) Sequencing statistics of 40 exomes. (C) Previously unreported protein-altering variants that occur in more than one subject. (D) Genes with highest burden of rare (<0.01%) heterozygous variants in cases compared to controls. (E) Genes with highest burden of rare (<0.1%) homozygous, hemizygous or candidate compound heterozygous damaging or conserved variants in cases compared to controls. (F) Illumina sequence reads identifying CACNA1H p.Met1549Val in five unrelated subjects. (G) Demonstration of biological parentage by genotyping of short tandem repeat markers in parent-offspring trios in kindreds 1347 and 1390 confirms that CACNA1H mutations are de novo in these kindreds. (H) Kinship coefficients of affected individuals from kindreds with CACNA1HM1549V variant. (I) Clinical features of family members of index cases with CACNA1HM1549V.
- https://doi.org/10.7554/eLife.06315.017





