Reverse evolution leads to genotypic incompatibility despite functional and active site convergence
Figures
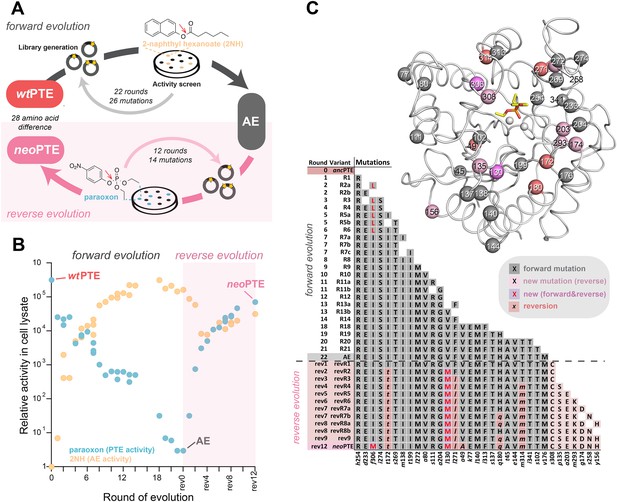
Activity and sequence changes of PTE over the evolution.
(A) Overview of the experimental evolution. Libraries were generated and transformed into Escherichia coli. Proteins were expressed and screened for paraoxon and/or 2NH hydrolysis in bacterial lysates. Several thousand variants were screened per round, theoretically covering most single point mutations in the ∼1000 bp PTE gene. Details are given in Supplementary file 1. (B) Activity changes during the forward (screening for arylesterase hydrolysis) and reverse evolution (screening for re-increase in phosphotriesterase hydrolysis). Steady-state kinetic parameters for all variants are provided in Supplementary file 2A. (C) Type, position, and order of occurrence of the 33 mutations obtained in the evolution. Mutations are shown relative to wtPTE (GenBank accession number KJ680379) with lower case italics denoting the amino acid found in wtPTE. Note that wtPTE was obtained in previous screens for improved expression levels in E. coli and contains six mutations relative to the naturally occurring PTE (I106L, F132L, K185R, D208G, R319S) (Roodveldt and Tawfik, 2005; Tokuriki et al., 2012). The following mutations occurred in individual variants, but were not fixated after DNA shuffling: R7a: a204G, R7c: a102V, R19: a78T, v143A, t311A, revR1: c59Y, s238R, revR5: i176V, revR8a: d264E, revR8b: i296V. All additional variants characterized and sequenced in each round are shown in Supplementary file 1.
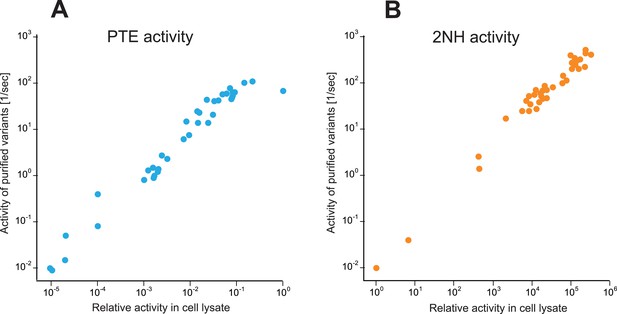
Correlation between activities measured in cell lysate and using purified enzyme for all variants selected over the evolution (Supplementary file 2).
(A) Phosphotriesterase activity. (B) Arylesterase activity. All measurements were carried out at 200 μM substrate. Activities in cell lysate are given relative to wtPTE.
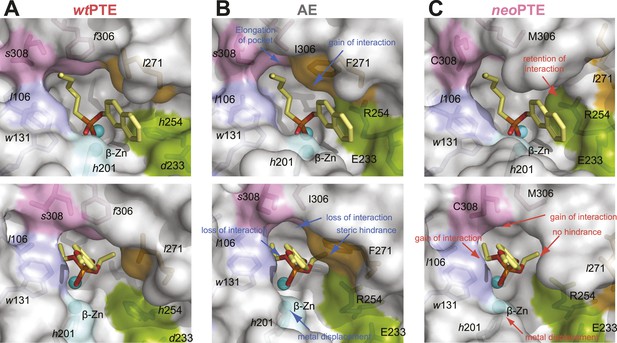
Reshaping of the PTE active site over the evolution.
(A) WtPTE (PDB ID: 4PCP) features an active site which is well adapted for paraoxon hydrolysis, but suboptimal for 2NH. (B) In the forward evolution, selection for arylesterase activity leads to several changes in the binding pocket from wtPTE to AE (PDB ID: 4PCN). (C) The reverse evolution leads to restoration of the ancestral state in neoPTE (PDB ID: 4PBF). The four regions of change are highlighted in different colors. Top row: the 2NH analogue (yellow) was modeled into the three structures by superposition with PTE-R18 in complex with the analogue (PDB ID: 4E3T) (Tokuriki et al., 2012). Bottom row: the paraoxon analogue diethyl 4-methoxyphenyl phosphate (yellow) was modeled into the structures by superposition with Agrobacterium radiobacter PTE in complex with the analogue (PDB ID: 2R1N) (Hong and Raushel, 1996). Amino acids found in wtPTE are shown in lower case italics.
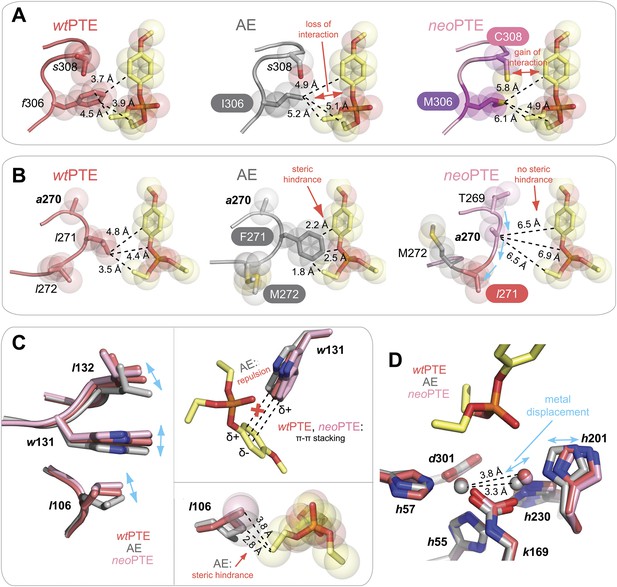
Details of the active site changes.
By overlaying the structures of wtPTE, AE, and neoPTE with the structure of A. radiobacter PTE in complex with the paraoxon analogue diethyl 4-methoxyphenyl phosphate (see Figure 2), regions important for paraoxon binding could be identified and the effect of mutations derived. The loss of phosphotriesterase in AE and its restoration in neoPTE is achieved mainly by changes in shape complementarity between enzyme and substrate, changes in hydrophobicity, and π-π stacking (A–C). It is likely that the movement of the β-metal also influences catalysis, although the exact effects of the metal displacement on catalysis are as yet unclear (D). (A) Interaction between paraoxon and residues 306 and 308. Substitution of the bulky Phe306 by Ile improves 2NH binding in AE, but results in a loss of interaction with paraoxon. In neoPTE, rather than reversion of f306I, s308 is mutated to the more hydrophobic Cys, improving interaction with the para-nitrophenyl group (Figure 2 pink region, Figure 7A). (B) Steric hindrance between Phe271 and paraoxon. Substitution of Leu271 to the larger Phe improves 2NH binding in AE, but causes steric hindrance with paraoxon. In neoPTE, in addition to reversion to the ancestral Leu, repositioning of the loop through a combination of remote mutations results in a ‘downward’ movement of Leu271, further enlarging the pocket (see also Figure 2 orange region, Figure 7A). (C) Shift in position of Leu106/Trp131/Leu132 (Figure 2 purple region, Figure 7B). While wt- and neoPTE feature edge-to-face π-π stacking between Trp131 and the para-nitrophenyl ring, in AE the shift in position brings Trp131 closer to the partially positive edges of the ring, resulting in electrostatic repulsion. Moreover, the shift in Leu106 brings it closer to the ethoxy group of the substrate in AE. (D) Shift in position of the β-metal. The inter-metal distance is reduced from 3.8 Å in wtPTE to 3.3 Å in AE through a movement of His201 and the β-metal. In neoPTE, the original spacing seen in wtPTE is restored (Figure 2 light blue region, Figure 7C), perhaps because the decreased distance in AE destabilized the transition state for paraoxon hydrolysis.
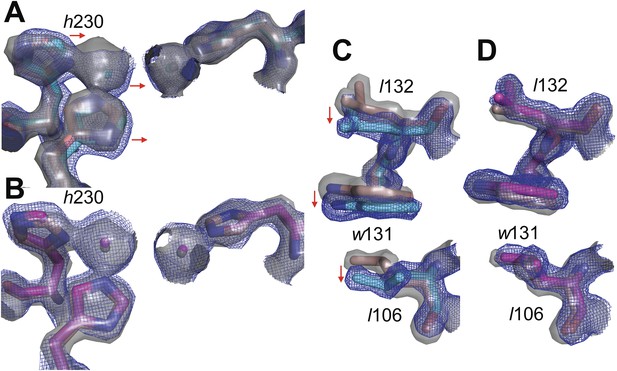
Overlay of electron density maps for the active sites of (A) wtPTE (salmon) and AE (cyan) and (B) wtPTE (salmon) and neoPTE (magenta).
Electron density for wtPTE is shown as grey isosurface (2 s), while electron density of AE and neoPTE is shown as isomesh (2 s), for contrast between the two maps. Comparison between (A) and (B) illustrates the shift of the β-metal ion closer to the α-metal ion in AE, and the shift back to the wtPTE position in neoPTE. Likewise comparison between the positions of Leu106, Trp131 and Leu132 in (C) wtPTE (salmon) and AE (cyan) and (D) wtPTE (salmon) and neoPTE (magenta) illustrates that these sidechains adopt alternative positions in AE, but return to their original conformations in neoPTE.
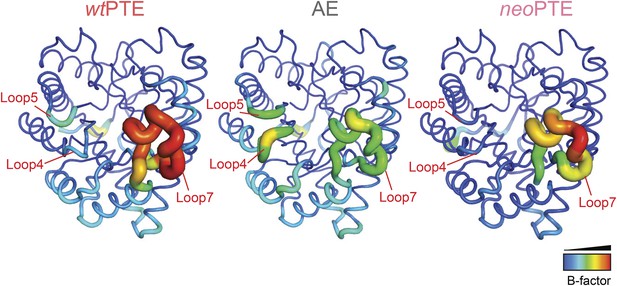
Development of B-factors over the evolution.
In wtPTE, loop 7 shows the maximum B-factor. The forward evolution for 2NH activity resulted in stabilization of loop 7, whereas flexibility of loops 4 and 5 increased. In neoPTE, the original dynamics of the structure were restored as shown by the increased flexibility of loop 7 as well as the reduced B-factor of loops 4 and 5.
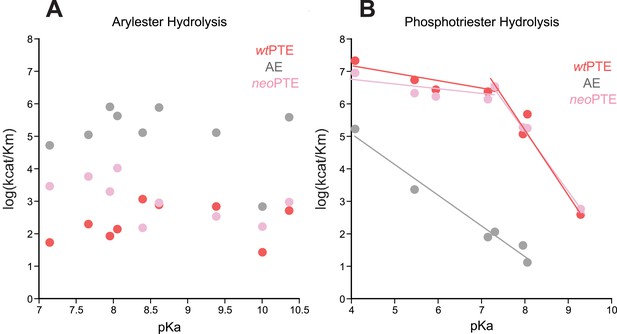
Linear free energy relationships of wtPTE, AE, and neoPTE.
(A) Arylester hydrolysis. The kcat/KM of all three variants is independent of the leaving group pKa. (B) Phosphotriester hydrolysis. In neoPTE, the break in leaving group dependence around pH 7, which is characteristic for wtPTE (Hong and Raushel, 1996; Tokuriki et al., 2012), is restored. Information about each substrate is provided in Supplementary file 2.
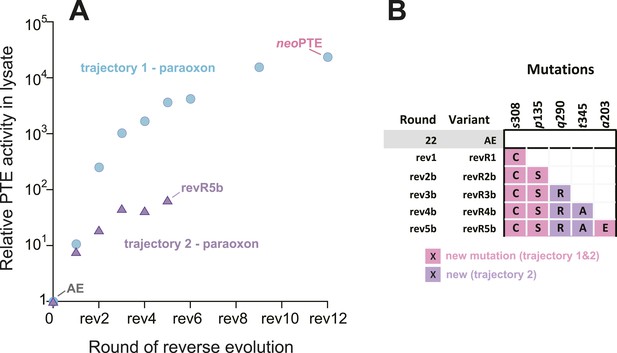
An alternative experimental evolution, where fixation of back-to-wild-type reversions was prohibited, failed to restore the original level of PTE activity.
(A) Activity changes in the alternative trajectory. After five rounds, PTE activity plateaued at a 65-fold improvement (trajectory 2), 340-fold lower than the main trajectory (trajectory 1). (B) Mutations accumulated in the alternative trajectory. All clones containing reversions, which occurred frequently, were removed after sequencing and thereby prohibited from fixating. Three of the five new mutations also fixated in the main trajectory. The mutations c59Y and s238R occurred in variants revR1 and revR2b, but were not fixated after DNA shuffling. Amino acids found in wtPTE are shown in lower case italics. All additional variants characterized and sequenced in each round are shown in Supplementary file 1.
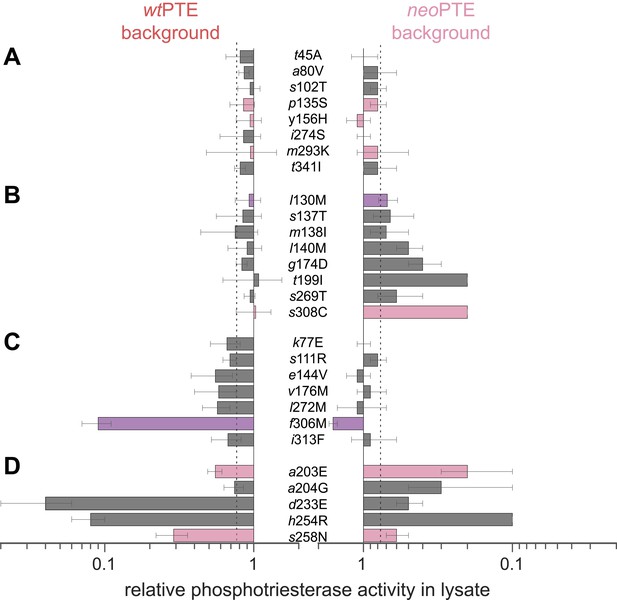
Genotypic incompatibility between wtPTE and neoPTE.
(A–D) The effect of the 28 amino acid exchanges separating the two enzymes was tested in the background of wtPTE and neoPTE, respectively. Activities are given relative to the parent mutational background, wtPTE or neoPTE. Amino acids found in wtPTE are shown in lower case italics. Color code as in Figure 1. (A) Compatible exchanges, neutral in both backgrounds. (B, C) Partially incompatible exchanges, neutral in one background but deleterious for another. (D) Mutually incompatible exchanges. Mutations causing a >1.3-fold change compared to the respective parent mutant (dotted line) are considered non-neutral. p-values compared to each parent (Supplementary file 2B) and p-values for the effect of each mutation Figure 4—source data 1. in the two backgrounds were calculated. Note that the effect of i313F, which causes a significant decrease in wtPTE, is statistically not significant between wtPTE and neoPTE.
-
Figure 4—source data 1
Comparison of the effect of mutations in wt- and neoPTE.
Fold-changes between the two backgrounds as well as p-values calculated according to the t-test are given. [a] Only mutations with an average >1.3-fold difference between backgrounds AND a p-value <0.05 are considered significant. Non-significant values are underlined. These belong to the ‘PANEL A’ series, which consists of mutations that are neutral in both cases. The only exception is i313F from PANEL C, which is significantly different from wtPTE, but not significantly different between wt- and neoPTE.
- https://doi.org/10.7554/eLife.06492.012
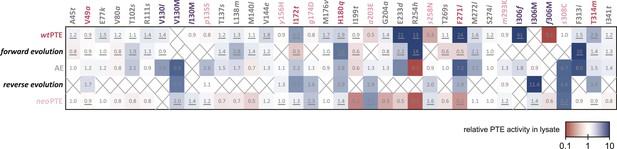
Changes in phosphotriesterase activity upon mutations in five different backgrounds: wtPTE in the forward evolution, AE in the reverse evolution, and neoPTE.
Thirty-three positions were mutated in the entire evolution, two of which (130 and 306) were mutated to two different amino acids. Amino acids found in wtPTE are shown in lower case italics. Numbers indicate the fold change in activity caused by a mutation in a certain background (Supplementary file 2B–F). Mutations causing a >1.3-fold change compared to the respective parent mutant are considered non-neutral. p-values compared to each parent were calculated (Supplementary file 2B,D,F). The mutations T341i in AE, l140M and t199I in the forward evolution, and V49a and s258N in the reverse evolution are not significant (p-values >0.05). Therefore, out of 144 mutations, only five show a >1.3-fold effect, but are statistically not significant. Boxes that are crossed out indicate that a mutation did not occur in this background. For direct comparison, the activity changes resulting from a mutation are adjusted to the same direction—from the amino acid found in AE to the respective other amino acid (label at the top). To illustrate, the effect of R254h was measured as follows: AE and neoPTE contain Arg254, and thus the effect of R254h is directly calculated based on the comparison between AE and AE-R254h (Fold changeR254h = ActicityAE-R254h/ActivityAE) and between neoPTE and neoPTE-R254h (Fold changeR254h = ActicityneoPTE-R254h/ActivityneoPTE). However, because wtPTE and the forward evolution background already contain His254, the effect of introducing this amino acid has to be calculated ‘in reverse’ by first assuming to remove this mutation and then adding it back in, that is, based on the comparison between wtPTE-h254R and wtPTE (Fold changeR254h = ActicitywtPTE/ActivitywtPTE-h254R). All mutational effects that were calculated in this ‘reverse’ way are underlined. Note that wtPTE-h254R is identical to the round 1 variant and therefore the effect in the forward evolution is the same as in the wtPTE background. Because R254h did not occur in the reverse evolution, no effect could be calculated in this background and the respective box is crossed out.
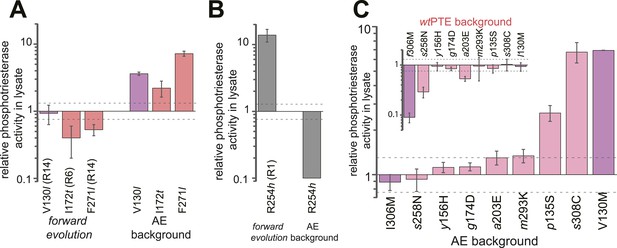
Epistasis between mutations in the forward evolution restricts some reversions while permitting others as well as new mutations.
(A) Several reversions change their effect from unfavorable upon their initial occurrence in the forward evolution to favorable in AE. (B) Other reversions change their effect from favorable to unfavorable. Note that, in the forward evolution, mutations occurred in the opposite direction as shown (l130V, t172I, l271F, and h254R), but are given in the same direction as AE for direct comparison. Phosphotriesterase activity was too low to be determined in AE + R254h, but at least 10-fold reduced. (C) The effect of new mutations changes from wtPTE (small panel) to AE (large panel). Relative activities were calculated by comparing a variant containing a certain mutation with one lacking only this mutation. Mutations causing a >1.3-fold change compared to the respective parent mutant (dotted line) are considered non-neutral. p-values compared to each parent (Supplementary file 2B) and p-values for the effect of each mutation in the two respective backgrounds shown in each panel were calculated (Figure 6—source data 1, 2). Note that the mutation m293K, which causes a significant increase in AE, does not have a significantly different effect in the two backgrounds. Amino acids found in wtPTE are shown in lower case italics. Color code as in Figure 1. All other mutational effects in the different backgrounds are given in Figure 5 and Supplementary file 2.
-
Figure 6—source data 1
Comparison of the effect of mutations in the forward evolution and in AE (panels A, B).
Fold-changes between the two backgrounds as well as p-values calculated according to the t-test are given. [a] Only mutations with an average >1.3-fold difference between backgrounds and a p-value <0.05 are considered significant. [b] Phosphotriesterase activity was too low to be determined in AE + R254h, but at least 10-fold reduced.
- https://doi.org/10.7554/eLife.06492.015
-
Figure 6—source data 2
Comparison of the effect of mutations in wtPTE and AE (panel C).
Fold-changes between the two backgrounds as well as p-values calculated according to the t-test are given. [a] Only mutations with an average >1.3-fold difference between backgrounds and a p-value <0.05 are considered significant. Non-significant values are underlined. Note that the effect of m293K, which causes a significant increase in AE, is statistically not significant between wtPTE and AE.
- https://doi.org/10.7554/eLife.06492.016
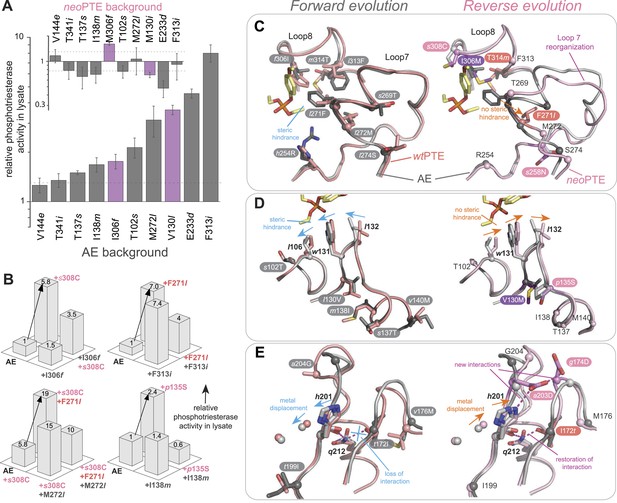
Convergence to the original active site configuration in the reverse evolution through rewiring of the molecular interaction network leads to genetic incompatibility.
(A, B) Epistasis during the reverse evolution causes irreversibility and incompatibility. (A) The activity change of mutations that were favorable in the initial stage of reverse evolution, but not reverted. neoPTE background: small panel; AE background: large panel. Color code as in Figures 1, 2. Mutations causing a >1.3-fold change compared to the respective parent mutant (dotted line) are considered non-neutral. p-values compared to each parent (Supplementary file 2B) and p-values for the effect of each mutation in the two respective backgrounds shown in each panel were calculated (Figure 7—source data 1). (B) Combinations of mutations that constrained the evolutionary trajectory due to sign epistasis. Phosphotriesterase activity is shown on a linear scale. p-values are given in Supplementary file 2G. Note that the two mutants AE + F271l + s308C and AE + M272l + s308C have non-significant p-values compared to the ‘double mutant’ in this series, AE + F271l + M272l + s308C. However, determination of kcat/KM values confirms sign epistasis in this series (see also Supplementary file 2G). (C–E) Amino acid changes in the forward (left panel) and reverse evolution (right panel). (C) Reorganization of loops 7 and 8. A new mutation, s258N, caused the reorganization (see also Figure 2—figure supplement 1A,B). (D) Different combinations of remote mutations in loop 3 resulted in identical positioning of Leu106, Trp131, and Leu132 in wtPTE and neoPTE (see also Figure 2—figure supplement 1C). (E) Rewiring the interaction network in neoPTE by remote mutations in loops 4 and 5 led to β-metal displacement (see also Figure 2—figure supplement 1D). Amino acids found in wtPTE are shown in lower case italics.
-
Figure 7—source data 1
Comparison of the effect of mutations in wtPTE and AE (panel A).
Fold-changes between the two backgrounds as well as p-values calculated according to the t-test are given. [a] Only mutations with an average >1.3-fold difference between backgrounds and a p-value <0.05 are considered significant. Non-significant values are underlined.
- https://doi.org/10.7554/eLife.06492.018
Additional files
-
Supplementary file 1
Description of the directed evolution rounds.
- https://doi.org/10.7554/eLife.06492.019
-
Supplementary file 2
Kinetic parameters of all variants.
- https://doi.org/10.7554/eLife.06492.020
-
Supplementary file 3
Crystallographic information.
- https://doi.org/10.7554/eLife.06492.021





