FBN-1, a fibrillin-related protein, is required for resistance of the epidermis to mechanical deformation during C. elegans embryogenesis
Figures
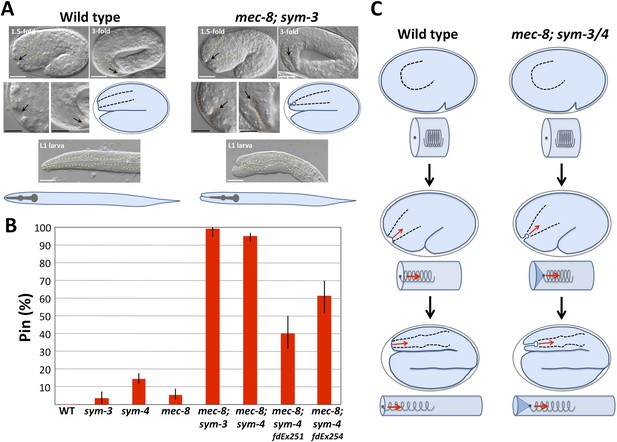
mec-8; sym-3 and mec-8; sym-4 mutants exhibit an abnormal ingression of the anterior epidermis.
(A) Whereas wild-type 1.5-fold embryos display only a shallow ingression of the anterior epidermis (sensory depression) and little or no ingression by the threefold stage, mec-8; sym-3 and mec-8; sym-4 (data not shown) mutants contain a deep keyhole-shaped ingression that increases in depth between the 1.5-fold and 3-fold stages. mec-8; sym-3 and mec-8; sym-4 (data not shown) L1 larvae also contain an ingressed pharynx (Pin) and associated deformities in the head region. Yellow dashed lines indicate lateral pharyngeal borders; orange dashed lines, the sensory depression or keyhole; black arrows, posterior extent of ingression. White scale bars = 10 µm, black bars = 5 µm. (B) Quantification of the Pin phenotype in single and double mutants and in mec-8; sym-4 double mutants containing multi-copy extrachromosomal arrays (fdEx251 and fdEx254) that express the fbn-1e cDNA isoform under the control of the native fbn-1 promoter. Error bars represent 95% CIs. For additional details, see Table 1 and Supplementary file 1. (C) Spring-and-cylinder model in which the pharynx exerts an inward-pulling force at the anterior epidermis throughout the mid-to-late stages of embryonic morphogenesis. In embryo representations, pharyngeal borders are indicated by black dashed lines; in cylindrical representations, the pharynx is represented by a spring that is attached to the anterior epidermis at the dark blue dot. Early comma, 1.5-fold and 3-fold stages of embryogenesis are depicted. Red arrows indicate the inward-pulling force on the epidermis that results from the resistance of the pharynx to stretching.
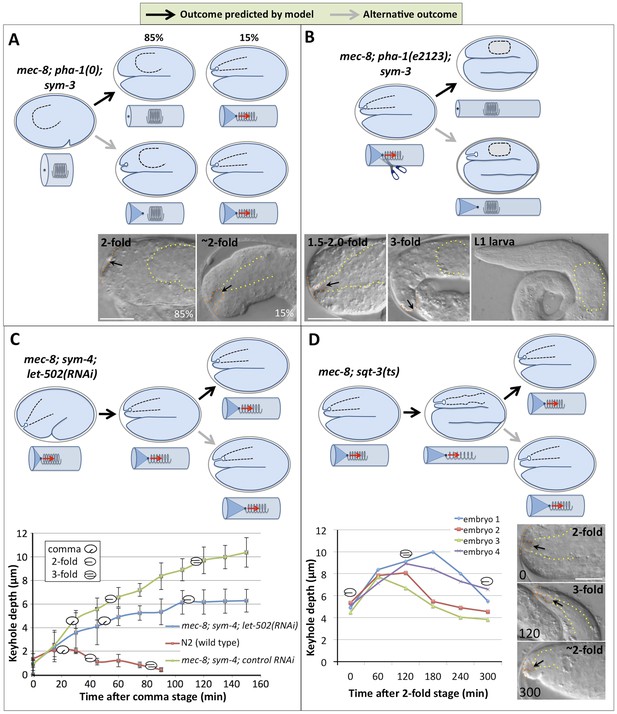
Genetic and phenotypic analyses support an extension spring model for pharyngeal elongation.
(A–D) Predicted models and outcomes for testing the hypothesis that the elongating pharynx exerts an inward-pulling force on the anterior epidermis. Black arrows in models show the predicted (and observed) outcomes; gray arrows, alternative outcomes. For panels with DIC images, yellow dashed lines indicate lateral pharyngeal borders; orange dashed lines, the sensory depression or keyhole; black arrows, posterior extent of ingression. White scale bars = 10 µm. For additional details, see Table 1 and Supplementary file 1. (A) In mec-8; pha-1(tm3671); sym-3 mutants that fail to establish a connection between the pharynx and epidermis (85%), deep ingressions or keyholes are not observed, whereas mutants that form an initial attachment (15%) form a stereotypical keyhole. (B) Detachment of the pharynx from the epidermis after the twofold stage in mec-8; pha-1(e2123); sym-3 mutants leads to loss of the anterior ingression by the threefold embryonic stage and suppression of Pin in L1 larvae. (C) Whereas the depth of the keyhole in mec-8; sym-4 mutants steadily increases between the 2-fold and 3-fold stages of embryogenesis, inhibition of embryonic elongation past the twofold stage by let-502(RNAi) prevents further deepening of the ingression. Error bars indicate 95% CIs, and diagrammed embryos denote the approximate stages of development for each genotype; n = 5 for each genotype at each time point. (D) Reversal of embryonic elongation in mec-8; sqt-3(e2117ts) mutants leads to a decrease in keyhole depth. Each line in the plot represents a different embryo; diagrammed embryos denote the approximate stages of development. For these experiments, rare mec-8; sqt-3(ts) mutants that exhibited a keyhole at the twofold stage (∼5%) were analyzed for reasons of experimental convenience.
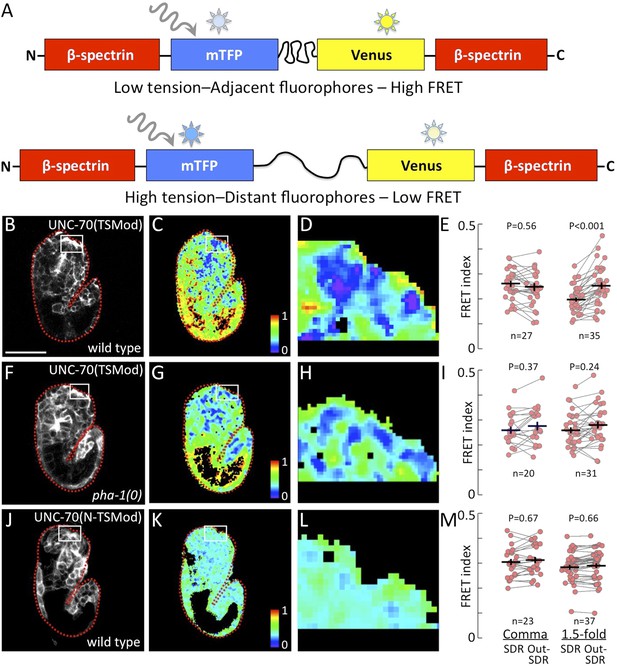
Pharyngeal attachment leads to increased forces at sensory depression.
A FRET-based TSMod inserted into the C. elegans β-spectrin gene (unc-70) was used to assess forces in live embryos. (A) Schematic for how UNC-70(TSMod) detects tension. FRET occurs when the donor fluorophore (mTFP) transfers energy to a nearby acceptor fluorophore (Venus) within the same peptide. When UNC-70(TSMod) experiences mechanical tension, a flexible linker separating mTFP and Venus is lengthened, leading to reduced FRET efficiency. (B–D) Representative images of wild-type and (F-H) pha-1(tm3671) strains that express UNC-70(TSMod). (J–L) Representative images of wild-type embryos expressing the no force control UNC-70(N-TSMod). Panels B, F, J depict 1.5-fold embryos after direct excitation of the Venus acceptor fluorophore. Panels C, D, G, H, K, L show FRET measurements where purple pixels indicate regions of highest tension (low FRET). Small white-framed boxes in panels B, C, F, G, J, K indicate the sensory depression region (SDR), which is enlarged in panels D, H, L. Red dashed lines in panels B, C, E, F, J, K outline the embryos. Scale bar in B = 30 µm. (E, I, M) FRET indices for the SDR and the region outside the sensory depression (Out-SDR). Individual embryos are represented by red circles, which are connected by lines to indicate values acquired from the same embryo. p-values depicted were calculated using a T-test (also see Supplementary file 2). Numbers at the bottom indicate the number of embryos that were analyzed for each condition. Each point is an average of ∼3–5 frames from a z-stack encompassing the embryo (see ‘Materials and methods’ for details).
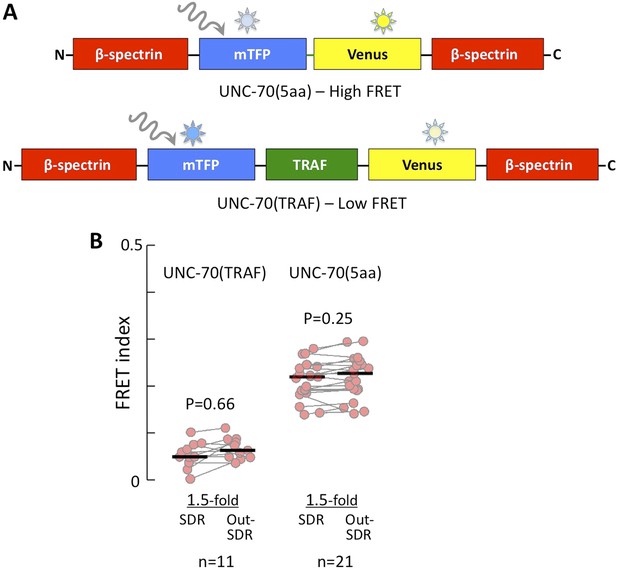
FRET index of low and high FRET controls.
(A) Schematic of UNC-70(5aa) and UNC-70(TRAF) FRET controls. (B) Quantification of low (UNC-70(TRAF)) and high (UNC-70(5aa)) FRET controls in 1.5-fold embryos.
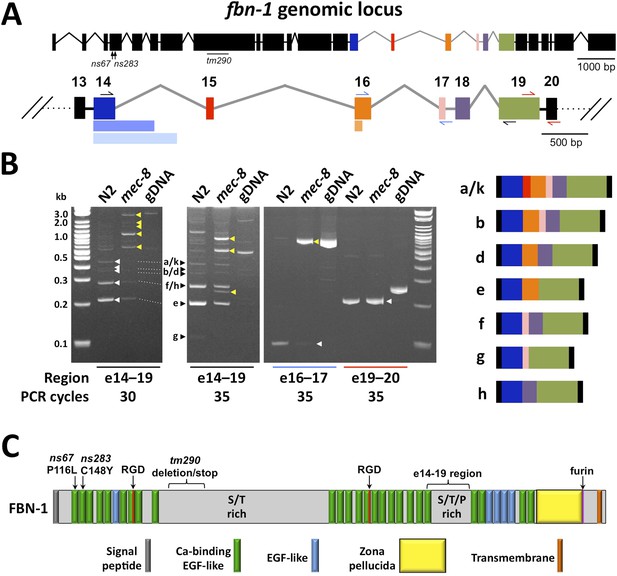
Splicing between a subset of fbn-1 exons is strongly misregulated in mec-8 mutants.
(A) A schematic of the fbn-1 genomic locus is shown with alternatively spliced exons (e14–19) indicated by colored blocks and enlarged below. Single-sided arrows indicate PCR primers used in panel B. Lighter-shaded rectangles below exons 14 and 16 indicate alternative 3′ splice sites for these exons. (B) PCR of the indicated regions of fbn-1 using wild-type (N2) and mec-8 cDNAs derived from embryos and wild-type genomic DNA (gDNA) as templates. White and black arrowheads indicate bands that correspond to known fbn-1 isoforms (depicted on right) based on size estimations for PCR products (in basepairs): a/k = 476, b = 407, d = 341, e = 200, f = 248, g = 107, h = 182. Yellow arrowheads indicate aberrant fbn-1 mRNA products that are present or are strongly enriched only in mec-8 mutants. (C) Schematic of FBN-1 (a isoform) showing the locations of protein domains and the amino acid positions affected by fbn-1 mutant alleles. For an annotated amino acid sequence, see Figure 4—figure supplement 2.
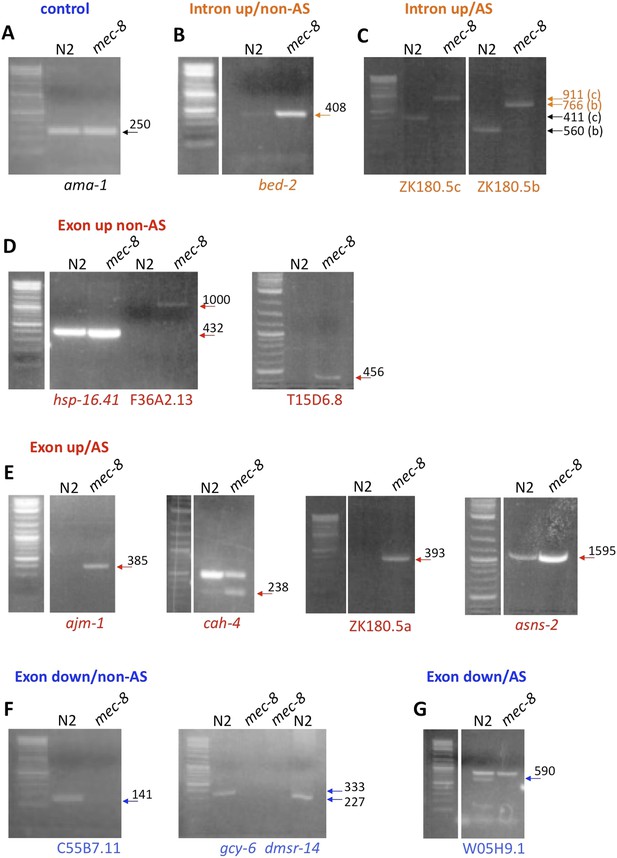
Examples of gene regions differentially expressed in mec-8 mutants and confirmed by RT-PCR.
Total RNA used for tilling arrays was reverse transcribed using oligo-dT primers and amplified by PCR using specific primers for each gene region (see Supplementary file 3). Each gene region was amplified using 30 cycles. Sizes are indicated in base pairs; arrows indicate bands that correspond to the expected sizes in mec-8 mutants based on tiling array results. For size reference, a 2-log DNA ladder (NEB) was used. (A) Control PCR with ama-1. (B) A gene (bed-2) with up-regulated introns (intron up) and that is not alternatively spliced (non-AS). (C) Intron up, AS gene (ZK180.5c and b). (D) Exon up, non-AS genes (hsp-16.41, F36A2.13, T15D6.8). (E) Exon up, AS genes (ajm-1, cah-4, ZK180.5a, asns-2). (F) Exon down, non-AS genes (C55B7.11, gcy-6, dmsr-14). (G) Exon down, AS gene (W05H9.1).
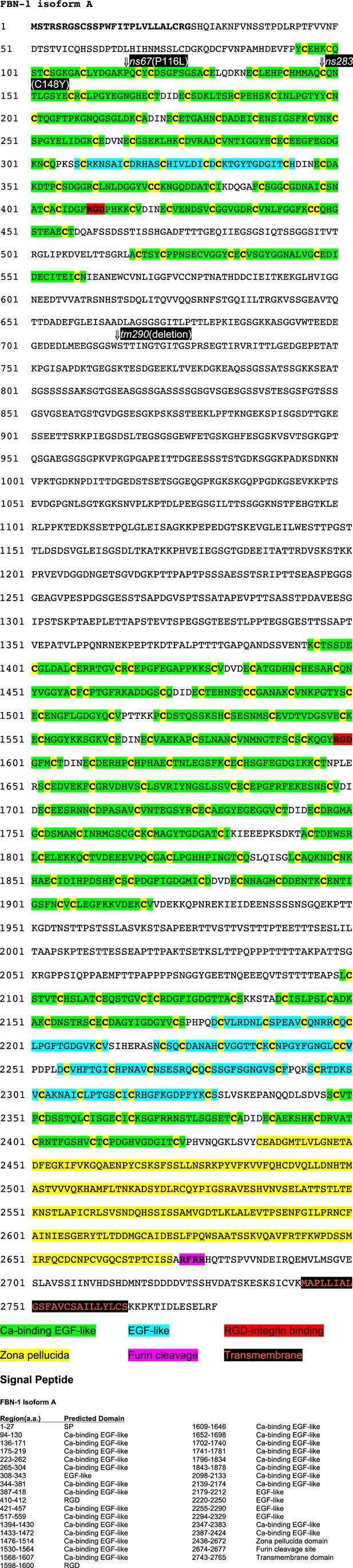
Amino acid sequence of FBN-1.
Annotated peptide sequence for FBN-1 (a isoform). Locations of domains and genetic mutations are indicated.
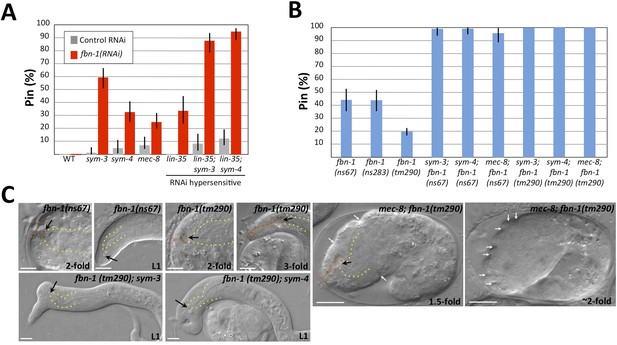
Morphogenesis defects of fbn-1 mutants are strongly enhanced by mutations in sym-3, sym-4 and mec-8.
(A) RNAi feeding of fbn-1 was carried out in the indicated backgrounds including strains hypersensitized to RNAi. Control RNAi strains contained the vector plasmid pPD129.36. (B) The Pin phenotype was scored in fbn-1 mutant alleles and in selected double mutants with fbn-1 and mec-8, sym-3 or sym-4. Error bars in A and B represent 95% CIs. For additional information, see Table 1 and Supplementary file 1. (C) Representative images for select single and compound mutants. Note the presence of strong head malformations in fbn-1(tm290); sym-3 and fbn-1(tm290); sym-4 larvae. Also note that the strong epidermal malformations observed in fbn-1(tm290); mec-8 mutants are suppressed by let-502(RNAi). White arrows indicate ingressions or furrows throughout the epidermis; red arrows, detached anterior cells in fbn-1(tm290); mec-8 mutants. Yellow dashed lines indicate lateral pharyngeal borders; orange dashed lines, the sensory depression or keyhole; black arrows, posterior extent of ingression. White scale bars = 10 µm.
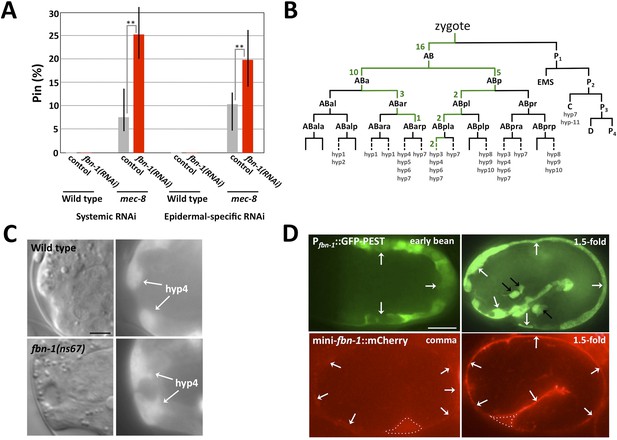
The fbn-1 gene is required in the epidermis and specifies a component of the embryonic sheath.
(A) Systemic and epidermal-specific RNAi of fbn-1 was carried out in wild-type (N2) and strain NR222, respectively, and in both backgrounds containing the mec-8(u74) allele. Note that both systemic and epidermal-specific fbn-1(RNAi) led to an increased percentage of Pin animals in the mec-8 background. Error bars indicate 95% CIs; **p < 0.01. (B) Schematic of the C. elegans embryonic lineage and findings from the fbn-1 mosaic analysis. Strains used for the analysis were WY1059, fbn-1(tm290); sym-4(mn619); fdEx249[fbn-1(+); sur-5::GFP], and WY1068, mec-8(u74); fbn-1(tm290); fdEx249. Green numbers indicate the number of L4 or adult mosaic animals that were not Pin but contained the fbn-1(+) rescuing array within that lineage only. (C) Wild-type and fbn-1(ns67) embryos expressing Pfbn-1::GFP-PEST (a convenient marker for embryonic epidermal cells). hyp4 cells within the focal plane are indicated and show aberrant morphologies in mutant embryos that contain a keyhole. (D). Expression of Pfbn-1::GFP-PEST and mini-fbn-1::mCherry (Δfbn-1–49-2418) reporters. In the Pfbn-1::GFP-PEST panels, epidermal cells are indicated with white arrows. Black arrows indicate several cells positive for Pttx-3::GFP, which was used as an injection marker and is not expressed in epidermal cells. In the mini-fbn-1::mCherry panels, the apical surface of embryonic epidermal cells (sheath) is indicated by white arrows. mini-fbn-1::mCherry is also detected in the extra-embryonic space (white dashed triangles). White scale bar = 10 µm, black bar = 5 µm.
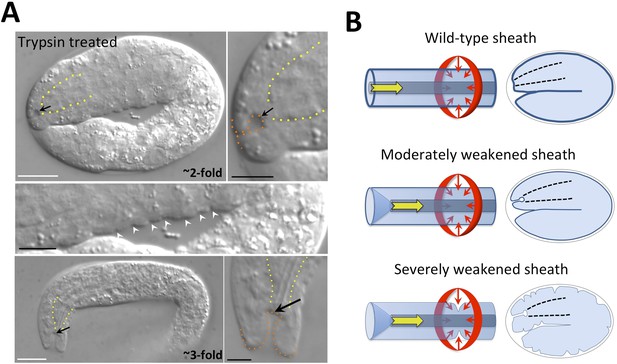
The embryonic sheath is critical for resistance to biomechanical forces.
(A) Wild-type embryos were treated with chitinase to remove part or all of the eggshell and then with trypsin to digest the sheath. Note the presence of a keyhole in both twofold and threefold trypsinized embryos (black arrows) and multiple ingressions or furrows in the epidermis of a twofold embryo (white arrowheads). Yellow dashed lines indicate lateral pharyngeal borders; orange dashed lines, the sensory depression or keyhole. White scale bars = 10 µm, black bars = 5 µm. (B) Model for the circumferential squeezing force (red arrows) and pharyngeal pulling force (yellow arrow) that act on the embryonic sheath. When the sheath is moderately weakened, such as when fbn-1 function is partially impaired, a keyhole phenotype is observed, suggesting that the anterior epidermis is particularly sensitive to a reduction in sheath integrity as a result of the pharyngeal pulling force. In cases where the sheath is more severely compromised, the depth of the keyhole may further increase, and the embryonic epidermis develops ingressions or furrows where circumferential constricting forces are acting.
Tables
Ingression depths of the anterior epidermis
| Ingression depth (μm) ± 95% CI (range; n) | ||
|---|---|---|
| Genotype | 1.5-fold | 3.0-fold |
| N2 | 2.12 ± 0.23 (1.09–3.12; 20) | 0.26 ± 0.096 (0.0–0.67; 20) |
| sym-3(mn618) | 2.39 ± 0.40 (0.81–4.24; 24) | 0.48 ± 0.56 (0.0–6.55; 22) |
| sym-4(mn619) | 2.74 ± 0.64 (0.72–5.67; 21) | 1.28 ± 1.06 (0.0–6.74; 22) |
| mec-8(u74) | 2.33 ± 0.46 (0.72–4.34; 20) | 2.42 ± 1.50 (0.0–10.43; 26) |
| mec-8; sym-3* | 4.25 ± 0.89 (2.77–5.72; 18) | 9.82 ± 0.68 (7.84–12.00; 15) |
| mec-8; sym-4 | 4.27 ± 1.16 (2.09–6.45; 16) | 9.19 ± 0.83 (7.07–10.14; 12) |
| Pha-1(tm3671) | 0.87 ± 0.18 (0.45–1.18; 16) | NA |
| mec-8; pha-1(tm3671); sym-3* | 0.83 ± 0.11 (0.40–1.19; 17) | NA |
| pha-1(e2123) | 2.15 ± 0.27 (1.04–3.34; 19) | 0.10 ± 0.07 (0.0–0.59; 16) |
| mec-8; pha-1(e2123); sym-3* | 5.27 ± 0.53 (3.89–7.46; 14) | 0.60 ± 2.35 (0.0–10.29; 19) |
| fbn-1(ns67) | 3.18 ± 0.85 (0.60–6.02; 13) | 5.34 ± 1.31 (0.0–9.24; 20) |
| fbn-1(ns67); sym-3 | 5.20 ± 0.41 (3.82–6.71; 20) | 11.73 ± 0.85 (8.59–16.34; 19) |
| fbn-1(ns67); sym-4 | 5.98 ± 0.55 (4.25–7.66; 12) | 12.84 ± 0.78 (9.65–16.37; 27) |
| mec-8; fbn-1(ns67) | 5.03 ± 0.47 (3.76–7.19; 19) | 9.84 ± 0.55 (6.94–12.24; 21) |
| fbn-1(tm290) | 6.25 ± 1.81 (0.99–12.17; 16) | 7.63 ± 3.66 (0.0–24.17; 17) |
| fbn-1(tm290); sym-3 | 5.65 ± 0.61 (2.59–7.29; 19) | 15.05 ± 1.56 (9.12–26.06; 24) |
| fbn-1(tm290); sym-4 | 5.54 ± 0.86 (3.52–9.12; 17) | 13.10 ± 1.72 (7.22–19.47; 20) |
| mec-8; fbn-1(tm290) | 9.84 ± 0.55 (5.47–15.18; 31) | NA |
-
*
Because these strains give rise to a high frequency of viable mnEx169(−) progeny in the first generation following loss of the array (F1 escapers), next-generation progeny (F2) from mnEx169(−) F1 parents were scored. NA, Non-Applicable; these genotypes led to embryonic arrest prior to the 3-fold stage.
Additional files
-
Supplementary file 1
Percentage of Pin Animals.
- https://doi.org/10.7554/eLife.06565.014
-
Supplementary file 2
Statistical analysis of FRET data.
- https://doi.org/10.7554/eLife.06565.015
-
Supplementary file 3
mec-8 tiling array data. (A) Gene Regions. Gene regions determined to be differentially expressed in mec-8 animals as compared with wild type. Genes for which all exons were up or down regulated are in bold. Region indicates the specific exons or introns of a transcript that were up or down regulated in mec-8 animals. Note that exon or intron numbering is specific to individual isoforms (as indicated) and may differ between isoforms. For example, the region encompassing intron 14 in fbn-1b, d, e corresponds to introns 14–15 in fbn-1a, k and the region encompassing intron 15 in fbn-1e corresponds to introns 16–18 in fbn-1a, k. Start and End indicate the flanking genomic positions of the differentially expressed exons or introns. Difference is the length in nucleotides of the exons or introns that were differentially expressed. Probes indicate the actual number of 25-nt probes that hybridized the indicated region. D/25 is the theoretical number of 25- nucleotide (nt) probes that cover the length of the genomic fragment. N2 avg and mec- 8 avg are the averaged intensities from three replicates. N2 SD and mec-8 SD are the standard deviation of the three replicates. Ratio is the average intensity of mec-8 divided by the average intensity of N2. SD ratio is the standard deviation of the mec- 8/N2 ratio of intensity. AS (alternatively spliced) indicates whether the gene is alternatively spliced. Gene ratios are color coded according to groups: orange, introns up; green, exons down; red, exons up. fbn-1 and mec-8 gene names are highlighted in red. In some cases different regions from the same gene are separated based on ratio rankings. (B) Transcripts. Transcripts differentially expressed in mec-8 animals compared to wild type. Average ratio is the average of the ratios of each gene region of the transcript. (C) Gene lists. Alphabetical list of all differentially expressed genes including category subdivisions. (D)Alternatively (Alt.) Spliced Genes. Alphabetical list of all differentially expressed genes including category divisions and information regarding alternative splicing. (E) PCR Primers. Sequences of primers (5′–3′) used to amplify candidate gene regions. Expected sizes for candidates that are differentially expressed in mec-8 mutants are indicated. Orange indicates introns upregulated in mec-8; red, exons upregulated in mec-8; blue, exons downregulated in mec-8. For genes that contain intronic regions that are upregulated in mec-8, sizes with and without, respectively, intronic sequences are shown.
- https://doi.org/10.7554/eLife.06565.016





