Low cost, high performance processing of single particle cryo-electron microscopy data in the cloud
Figures
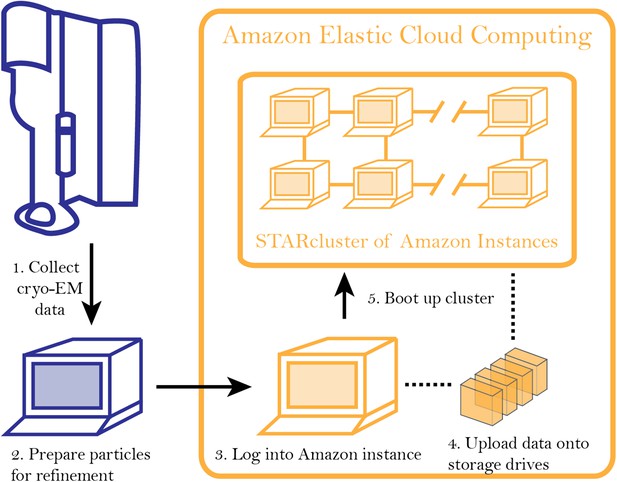
Workflow for analyzing cryo-EM data on Amazon's cloud computing infrastructure.
After collecting cryo-EM data (Step 1), particles are extracted from the micrographs and prepared for further analysis (Step 2). After logging into an ‘instance’ (Step 3), data are uploaded to a storage server (elastic block storage) (Step 4). At this point, STARcluster can be configured to launch a cluster of 2–30 instances that is mounted with the data from the storage volume (Step 5). A detailed protocol can be found at an accompanying Google site: http://goo.gl/AIwZJz.
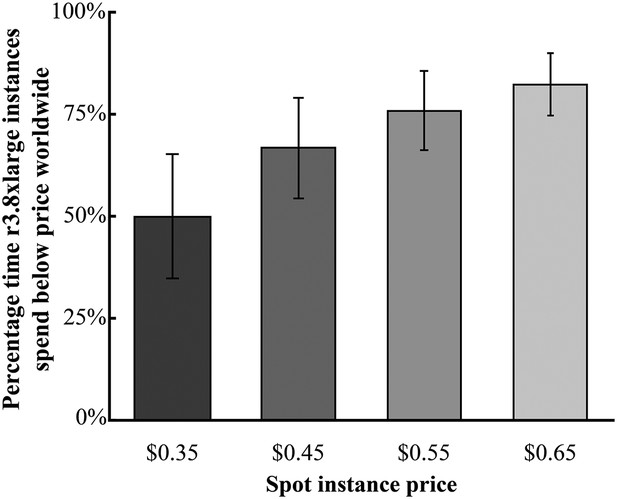
Global availability of Amazon r3.8xlarge spot instances.
Shown is the average percentage time spent by the r3.8xlarge type of instance when the current spot instance price was less than the queried price. The data are averaged over all Amazon's regions worldwide (except for SA-East-1, which does not offer r3.8xlarge instances). Spot instance prices were calculated over a 90-day period from 1 January 2015—1 April 2015, where the average is shown ± the s.e. Source data: Figure 2—source data 1.
-
Figure 2—source data 1
Global spot instance price data from 1 January 2015 to 1 April 2015.
- https://doi.org/10.7554/eLife.06664.005
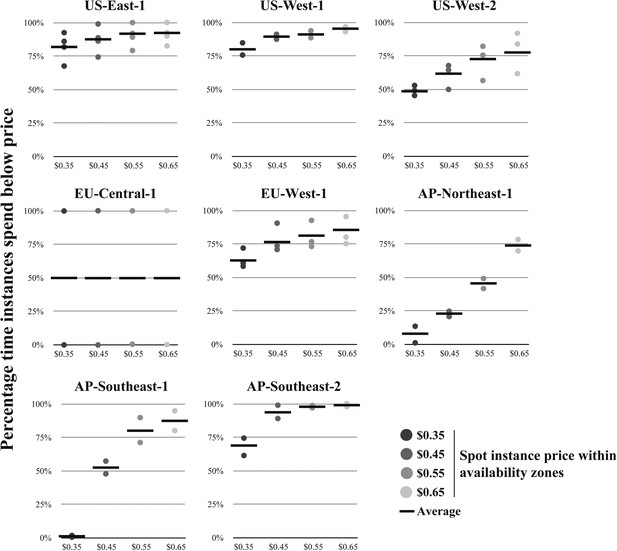
Availability of virtual machines within regions at specified spot instance prices.
For each Amazon region (excluding SA-East-1, which does not offer r3.8xlarge instances), r3.8xlarge spot instance prices were retrieved for each availability zone, where separate availability zones are shown as separate data points for a given spot instance price. (Note: each region can have different number of availability zones). From the spot instance prices, the percentage time of the spot instances that were spent below the specified spot instance price were calculated. The average value is shown as a solid black line. Source data: Figure 2—source data 1.
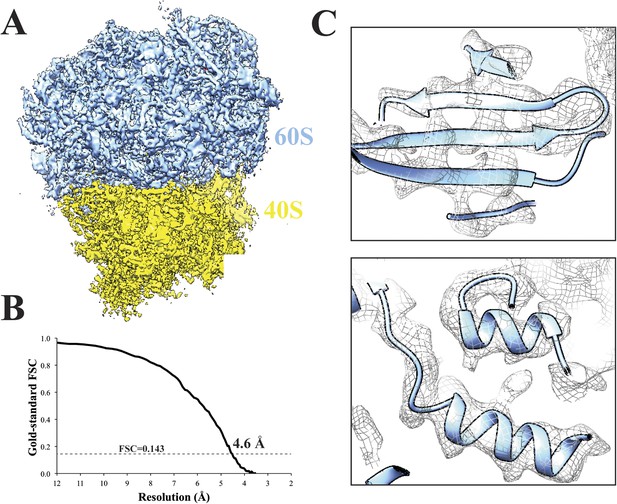
Cryo-EM structure of 80S ribosome at an overall resolution of 4.6 Å.
(A) Overall view of 80S reconstruction filtered to 4.6 Å while applying a negative B-factor of −116 Å2. (B) Gold standard FSC curve. (C) Selected regions from the 60S subunit. Cryo-EM maps were visualized with UCSF Chimera (Pettersen et al., 2004). Source data: Dryad Digital Repository dataset (http://datadryad.org/review?doi=doi:10.5061/dryad.9mb54) (Cianfrocco and Leschziner).
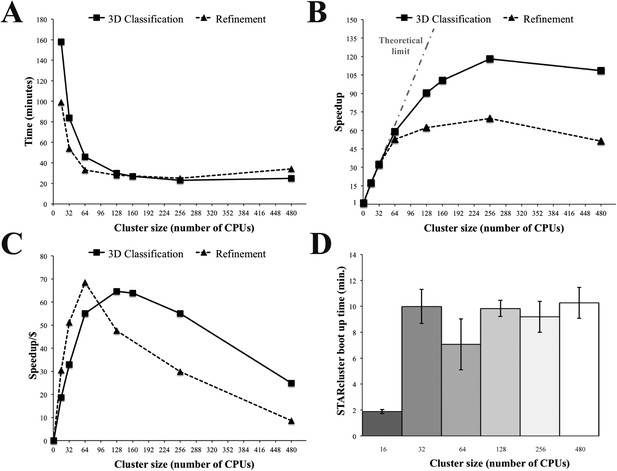
Relion performance on STARcluster configurations of Amazon instances.
(A) Processing times (minutes) for Relion to perform 3D Classification or 3D refinement on 80S ribosome dataset. (B) Speedup for each cluster size relative to a single CPU (black line) shown alongside performance estimate for a perfectly parallel cluster using Amdahl's Law (curve labeled ‘Theoretical limit’). For cluster sizes ≤ 64 CPUs, Relion exhibits near-perfect performance on STARcluster configurations, while cluster sizes > 64 show that Relion's performance reaches a maximum at 256 CPUs for both 3D classification and 3D refinement. (C) Speedup/Cost is plotted against cluster size, where Speedup/Cost is defined as the speedup observed divided by the cost associated with Amazon's pricing at $0.35/hr/16 CPUs. (D) Average STARcluster boot up time (± s.d.) was measured for clusters of increasing size (n = 5). Source data: Figure 4—source data 1.
-
Figure 4—source data 1
Performance analysis statistics for Relion 3D classification and 3D refinement on STARcluster configurations.
- https://doi.org/10.7554/eLife.06664.009
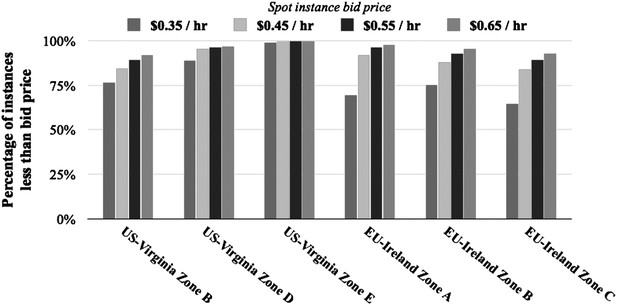
Percentage of instances below bid price over last 90 days. Shown are the percentages of r3.8xlarge instances that are below the spot instance price across different regions and zones.
Tables
Cost of 80S ribosome 3D classification and refinement on a 128 CPU STARcluster configuration at increasing spot instance price.
Spot instance bid price | $0.35 | $0.45 | $0.55 | $0.65 |
80S ribosome 3D classification and refinement cost | $28.89 | $37.14 | $45.39 | $53.64 |
Additional files
-
Supplementary file 1
Step-by-step tutorial describing how to use Amazon's EC2 environment to analyze cryo-EM data.
- https://doi.org/10.7554/eLife.06664.010
-
Supplementary file 2
Comparison of estimated processing times and costs for recent near-atomic cryo-EM structures on Amazon's EC2. Source data: Supplementary File 3.
- https://doi.org/10.7554/eLife.06664.011
-
Supplementary file 3
Source data for tables in Supplementary file 2.
- https://doi.org/10.7554/eLife.06664.012





