A shift in anterior–posterior positional information underlies the fin-to-limb evolution
Figures
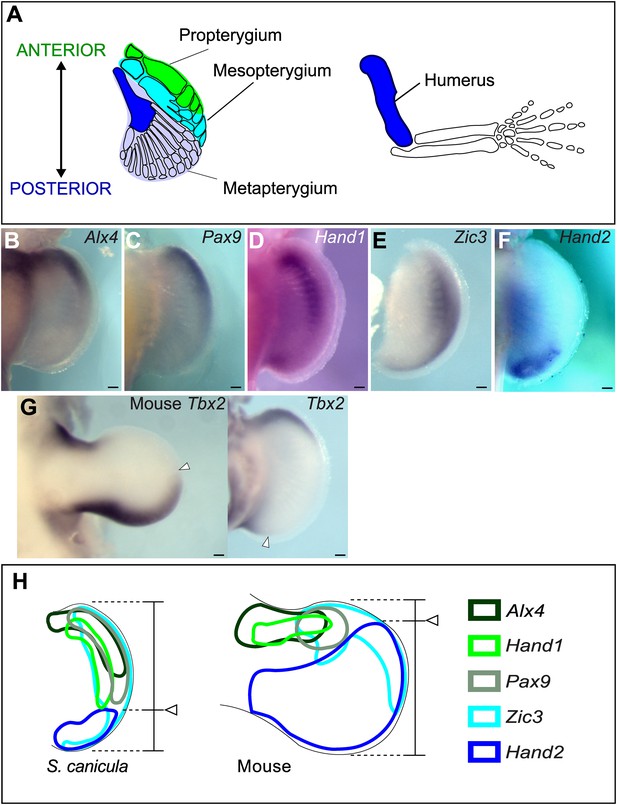
Anterior–posterior patterning in Scyliorhinus canicula pectoral fin buds.
(A) Skeletal patterns of S. canicula pectoral fin and mouse limb. Blue colours, homologous elements. (B–G) In situ hybridisation for Alx4 (B), Pax9 (C), Hand1 (D), Zic3 (E), Hand2 (F), and Tbx2 (G) in S. canicula pectoral fin buds at stage 30 and mouse limb bud at E11.5 (left panel in G). Arrowheads in G, anterior boundary of posterior Tbx2 expression. Dorsal view; anterior is to the top. Scale bars, 100 μm. (H) Schematic of the gene expression patterns. Arrowheads, Hand2 expression boundary. Expressions of mouse limb buds at E11.5 are after EMBRYS database (Yokoyama et al., 2009; Fernandez-Teran et al., 2003).
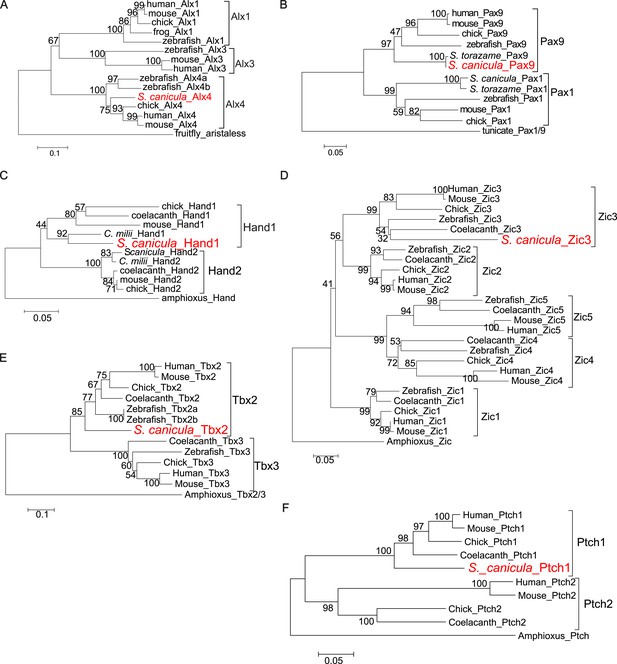
Molecular phylogenetic trees of relevant S. canicula genes.
(A–E), Trees for Alx4 (A), Pax9 (B), Hand1 (C), Zic3 (D), Tbx2 (E), and Ptch1 (F) were generated from amino acid sequences of the homeodomain and neighbouring sequences (A), the paired box and C-terminal sequences (B), Helix-loop-helix domain and C-terminal sequences (C), N-terminal sequences (D), and C-terminal sequences (E and F). The neighbour-joining method was used for constructing the trees. The numbers at nodes indicate bootstrap probabilities with 1000 replicates.
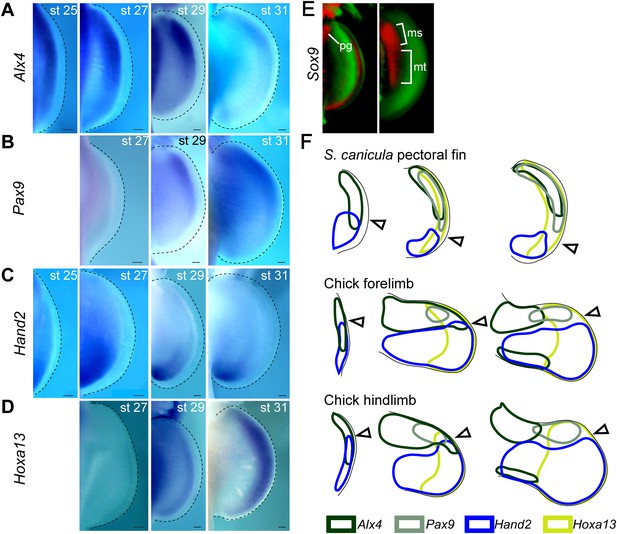
Temporal expression analysis of Alx4, Pax9, Hand2, and Hoxa13 in S. canicula pectoral fins.
(A–D) Expression of Alx4 (A), Pax9 (B), Hand2 (C), and Hoxa13 (D) at the indicated stages. Dorsal view of right pectoral fin buds of S. canicula embryos (anterior to the top). (A) Alx4 was initially expressed throughout the fin buds at stage 25, but the posterior expression of Alx4 was reduced by stage 27. Alx4 expression was then restricted to the anterior two-thirds of the fin bud at stage 29 and to the anterior half of the fin bud at stage 31. (B) Pax9 expression was not detected at stage 27 but was present in anterior distal fin buds at stage 29. This expression persisted until at least stage 31. (C) Hand2 was initially expressed throughout the fin bud, although slightly more-robust expression was observed at the posterior side. Subsequently, Hand2 expression became posteriorly restricted by stage 27. (D) Hoxa13, which is a marker for a late developmental stage in tetrapod limb buds, was detected since stage 29 in a distal domain. (E) OPT scans of Sox9 expressions (red) stained with propidium iodide (green) at stage 28 (left) and late stage 29 (right). pg, pectoral girdle; ms, mesopterygium; mt, metapterygium. (F) Comparison of Alx4 (black), Pax9 (grey), Hand2 (blue), and Hoxa13 (light green) expression patterns between S. canicula fin buds and chick forelimb and hindlimb buds (see Figure 1—figure supplement 3 for detailed gene expression in chick limb buds). st, stage. Scale bars, 100 μm.
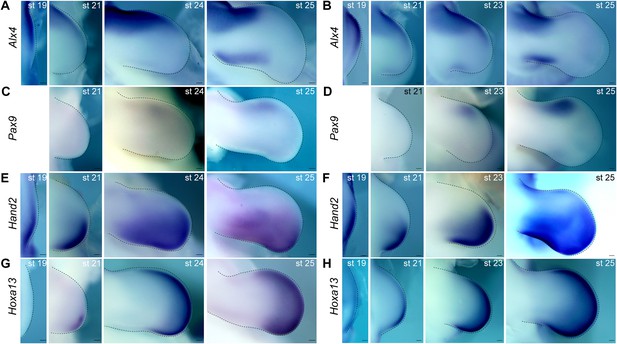

Temporal expression analysis of Alx4, Pax9, Hand2, and Hoxa13 in chick limb buds.
(A–H) Expression of Alx4 (A, B), Pax9 (C, D), Hand2 (E, F), and Hoxa13 (G, H) at the indicated stages. Dorsal views of chick forelimb (A, C, E, G) and hindlimb (B, D, F, H) buds (anterior to the top; distal to the right). (A, B) At stage 19, Alx4 expression was broad but was slightly weaker in the posterior side of forelimb (A) and hindlimb (B) buds. At later stages, its expression became restricted to the anterior one-third of the limb buds, with subsequent further restriction to the anterior proximal region both in forelimb (A) and hindlimb (B) buds. Alx4 expression also appeared in the posterior distinct region of chick forelimb (A) and hindlimb (B) buds at stage 25. (C, D) Pax9 expression was first present in the anterior region at stage 24 in forelimb buds (C) and stage 23 in hindlimb buds (D) but was more distally restricted than Alx4 expression. At stage 25, expression of Pax9 was robust, but the extent of its overlap with Alx4 expression was limited both in forelimb (C) and hindlimb (D) buds. (E, F) Hand2 expression was initially complementary with Alx4 and subsequently with Pax9 expression in forelimb (E) and hindlimb (F) buds. (G, H) At stage 21, Hoxa13 expression fully overlapped with the distal Hand2 expression domain in forelimb (G) and hindlimb (H) buds. At later stages, co-expression of Pax9 and Hoxa13 was observed in the anterior distal portion of the limb buds, but this overlap was not as extensive as that between the Hand2 and Hoxa13 expression domains. st, stage. Scale bars, 100 μm.
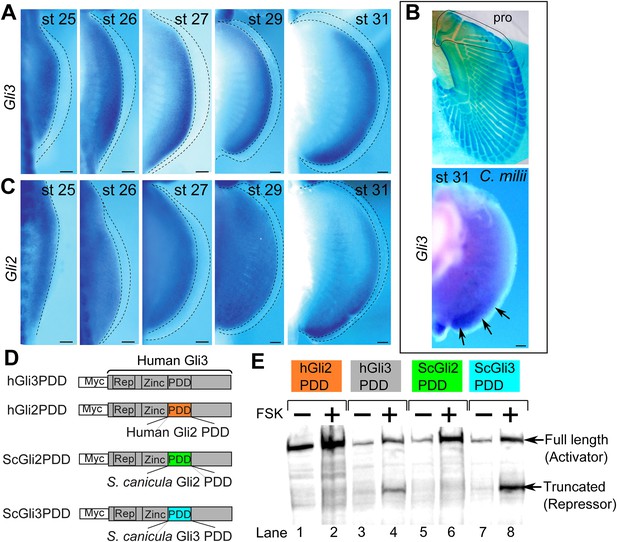
Expression and processing of Gli3 and Gli2 in S. canicula embryos.
(A) Expression of Gli3 in S. canicula pectoral fins. (B) Alcian blue staining of C. milii pectoral fin at stage 35 (top, the ventral view of a right fin flipped horizontally) and Gli3 expression at stage 31 (bottom, a left pectoral fin flipped horizontally). pro, propterygium. (C) Expression of Gli2 in S. canicula pectoral fin buds. Scale bars, 100 μm. (D) The Gli3 chimera constructs. hGli3 PDD, full-length human Gli3 (grey box) with Myc tags. hGli2, ScGli2 and ScGli3 PDD, chimeric Gli3 genes recombined at the processing determinant domain (PDD) with human Gli2, S. canicula Gli2 and Gli3, respectively. (E) Protein processing of the chimeric constructs in cell cultures treated with either FSK (+) or DMSO (−). Truncated Gli3 is detected only in hGli3 PDD (lane 4) and ScGli3 PDD (lane 8).
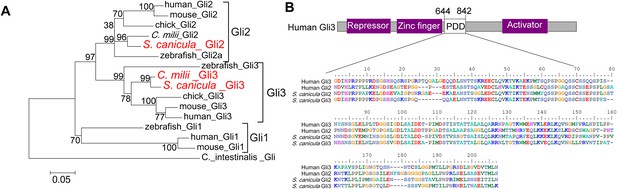
Phylogenetic tree of Gli2 and Gli3, and PDD amino acid sequences.
(A) Trees for Gli2 and Gli3 were generated from amino acid sequences of the zinc finger domain and PDD. The neighbour-joining method was used for constructing the trees. The numbers at nodes indicate bootstrap probabilities with 1000 replicates. (B) Human Gli3 (the upper diagram) is composed of an N-terminal repressor domain (Repressor), a DNA-binding domain (Zinc finger), and a C-terminal activator domain (Activator). The alignment shows the homologous PDD domains from both human and S. canicula Gli2 and Gli3.
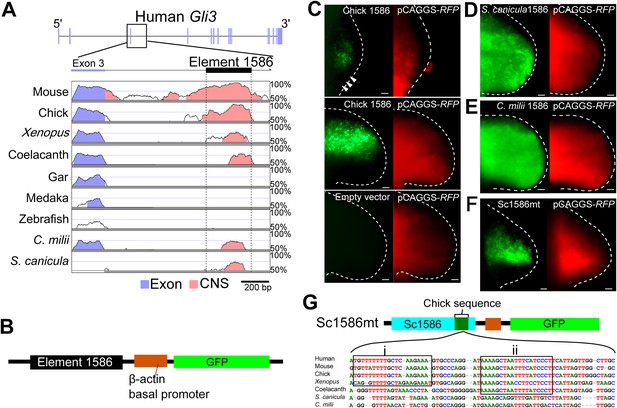
The Gli3 limb-specific enhancer of S. canicula and C. milii.
(A) VISTA plots of Gli3 intron 3 from indicated animals. Blue vertical bars, exons of human Gli3; black rectangle, element 1586. Regions with >70% identity are indicated: blue, exon; pink, non-coding sequences. (B) The enhancer construct. (C) GFP expression in chick forelimb buds driven by chick element 1586 at stage 19 (top, n = 3/3), stage 23 (middle, n = 14/14), and empty vector (bottom, n = 0/7). pCAGGS-RFP (right). (D–F) GFP expression driven by element 1586 of S. canicula (D, n = 11/11), C. milii (E, n = 10/10) and Sc1586mt (F, n = 4/4). Scale bars, 100 μm. (G) Scheme of Sc1586mt, S. canicula enhancer (blue) partially replaced by chick sequence (green) and alignment. Boxes indicate tetrapod (i) and sarcopterygian (ii) specific sequences.
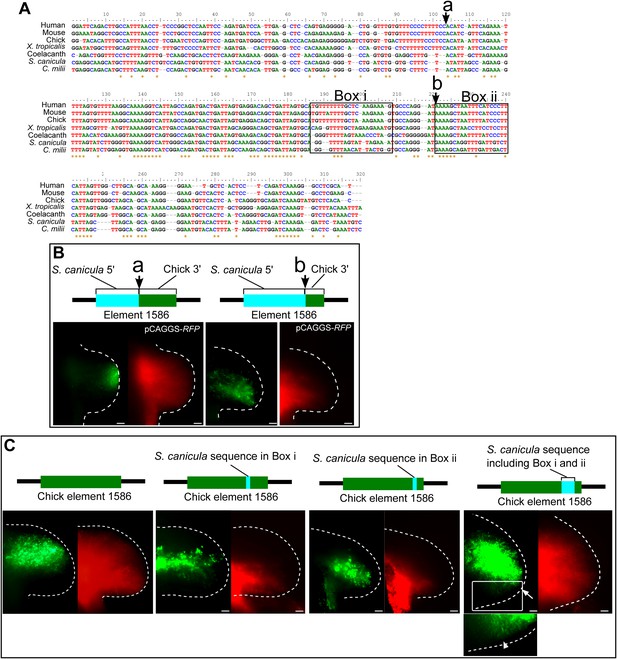
Detailed functional analyses of element 1586.
(A) Alignment of element 1586 sequences. Points where chick and S. cacnicula enhancers are recombined in (B) are indicated by arrows a and b. Box i and Box ii are sequences that are replaced with S. canicula sequence in (C), activities of element 1586 recombined at the indicated points. Note that the posterior repressive activity is not altered at arrow a, but partially destroyed at arrow b, suggesting that the repressive sequences are located between arrow a and b and after arrow b. (C) Activities of chick element 1586 that are partially recombined with S. canicula sequences at Box i and Box ii in (A). The sequences were chosen because of specific conservation among tetrapods (Box i) and sarcopterygians (tetrapod and coelacanth, Box ii). Each of replacement can slightly alter the posterior repression of GFP (n = 3/3). When the chick enhancer is replaced by both sequence (the most right panel), the distal GFP expression is significantly shifted to the posterior limb buds (n = 4/4, white arrow). And weak expressions are also detected in the posterior proximal limb buds (n = 2/4. arrow head in the bottom panel, magnified view of the above white box with enhanced contrast), suggesting that the sequences at Box i and Box ii are partially responsible for the posterior repression. Scale bars, 100 μm.
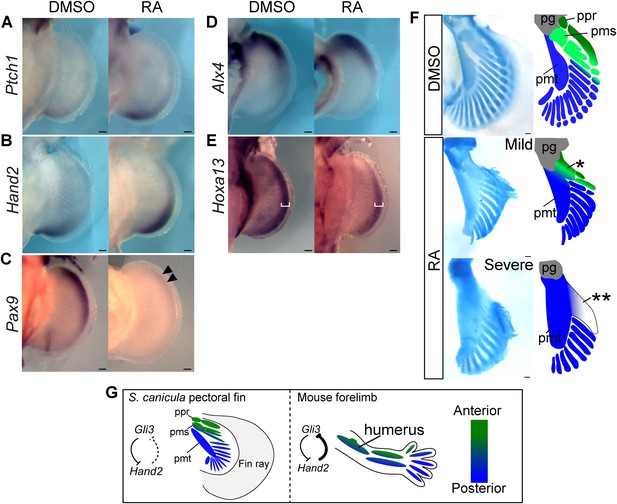
RA treatment causes ectopic activation of Shh signalling and loss of anterior skeletal elements.
(A–E) In situ hybridisation of S. canicula pectoral fin buds for Ptch1 (A; left fins flipped horizontally), Hand2 (B), Pax9 (C), Alx4 (D), and Hoxa13 (E) treated with 1% DMSO or 1 μg/ml retinoic acid (RA) (n = 2/2 for each for each except n = 4/4 for Hand2). Arrowheads in C, a weak expression of Pax9. White brackets in E, width of Hoxa13 expression domain along the proximal-distal (PD) axis. (F) Pectoral fin skeletal patterns of 1% DMSO control (n = 4) and 1–2 μg/ml RA (n = 4). Right panels, schematics of interpretive skeletal patterns. *, an anterior proximal radial; **, a fused radial attached to the metapterygium; pg, pectoral girdle; ppr, pms and pmt, proximal propterygium, mesopterygium and metapterygium. Scale bars, 100 μm. (G) Comparison between S. canicula fin and mouse limb. Green and blue colours represent anterior–posterior (AP) positional information. ppr, pms, and pmt denote proximal propterygium, mesopterygium, and metapterygium, respectively.





