Transcription dynamically patterns the meiotic chromosome-axis interface
Figures
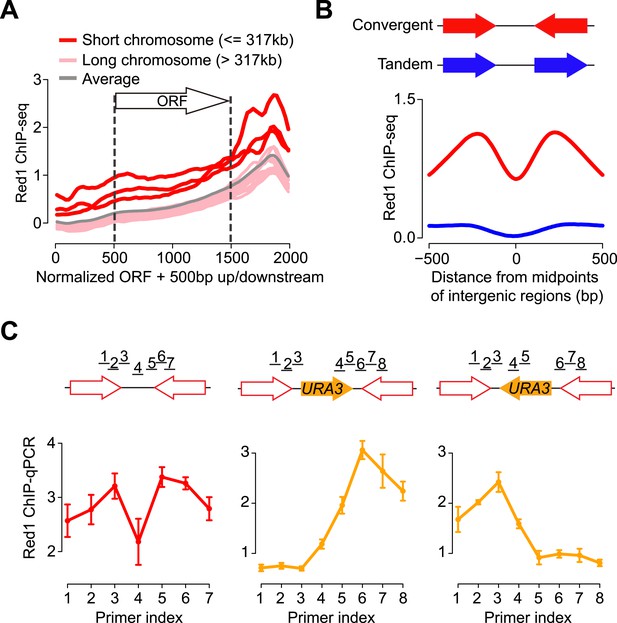
Red1 preferentially localizes to the 3′ ends of convergent genes.
(A) Red1 distribution was plotted as an average across all genes on each of the 16 chromosomes as well as for the whole genome. The coding regions of genes were normalized to lengths of 1 kb (500 bp–1500 bp on x-axis). 500 bp upstream of the start codon and downstream of the stop codon were included in the plot. (B) Comparison of Red1 binding between convergent and tandem gene pairs. Average signals of Red1 binding were plotted within a 1 kb region from the midpoints of the intergenic regions. Signals were averaged at each bp among all the convergent (red) and tandem (blue) gene pairs across the genome. (C) qPCR analysis of Red1 binding at the YKL077W/YKL075C convergent gene pair before and after insertion of a URA3 transcription unit. Schematic shows the insertion of URA3 in two orientations between YKL077W and YKL075C. ChIP signals were normalized against input and an internal control; the control primer was chosen at the promoter region of YKL077W. Failure to detect a peak doublet upon URA3 insertion may be due to primer positions or because the URA3 transcript does not lead to overlapping transcription (see Figure 2).
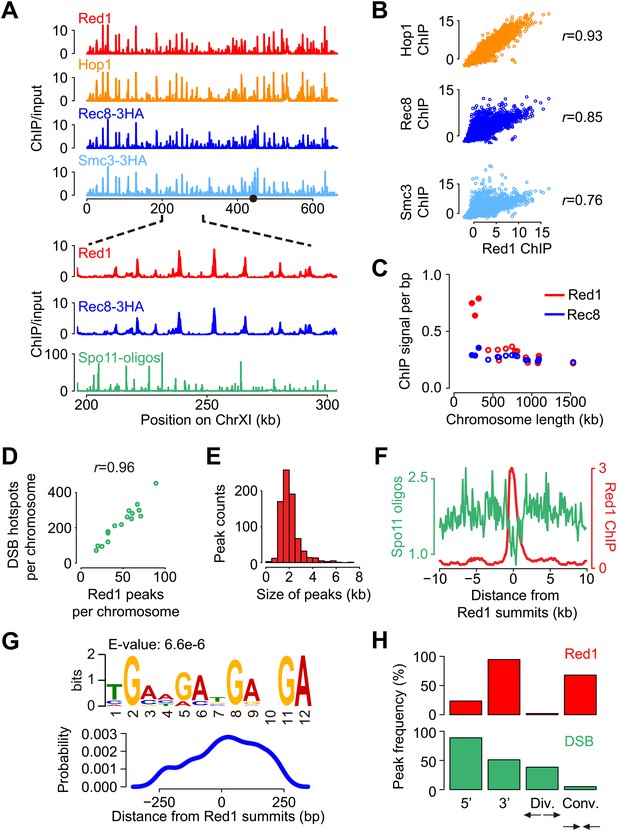
Genome-wide localization of meiotic axis proteins and cohesin.
(A) Distribution of axis proteins and cohesins as determined by ChIP-seq. Chromosome XI is shown as an example to show co-enrichment with the exception of the centromere (indicated by black circle). Bottom panels are a zoom-in to show that axis proteins in general do not localize to DSB hotspots as measured by Thacker et al. (2014). (B) Pairwise correlation between Red1 ChIP and cohesin ChIP signals. (C) Red1 and Rec8 ChIP signal per bp as a function of chromosome length. 25 kb to either side of the centromeres were excluded from this analysis to avoid biases caused by the strong centromere-proximal enrichment of Rec8 (see A). The three shortest chromosomes are displayed as solid dots. (D) Numbers of Red1 peaks (see ‘Materials and methods’ for peak calling) were correlated (Pearson's r) with the numbers of DSB hotspots on each chromosome. (E) Distribution of widths for 774 Red1 peaks. (F) Spo11 oligos (green) were strongly depleted at the axis protein associated regions (red). Average Spo11 oligo density is plotted as a function of distance from Red1 peak summits. Red1 signals were averaged among all the peaks. (G) Percentage of Red1 peaks (top) and DSB hotspots (bottom) in different positions relative to genes. Peaks were defined as a 500 bp-surrounding region of summits. 5′ regions of genes were defined as 500 bp upstream of the start codons, and 3′ regions of genes were defined as 250 bp on either side of the stop codons. Divergent (Div.) or convergent (Conv.) regions refer to the intergenic areas between two adjacent start or stop codons, respectively. (H) Putative motif at axis sites. Top: motif derived from Red1 ChIP-seq using MEME-ChIP (Machanick and Bailey, 2011). Bottom: probability of motif occurrence around Red1 peak summits.
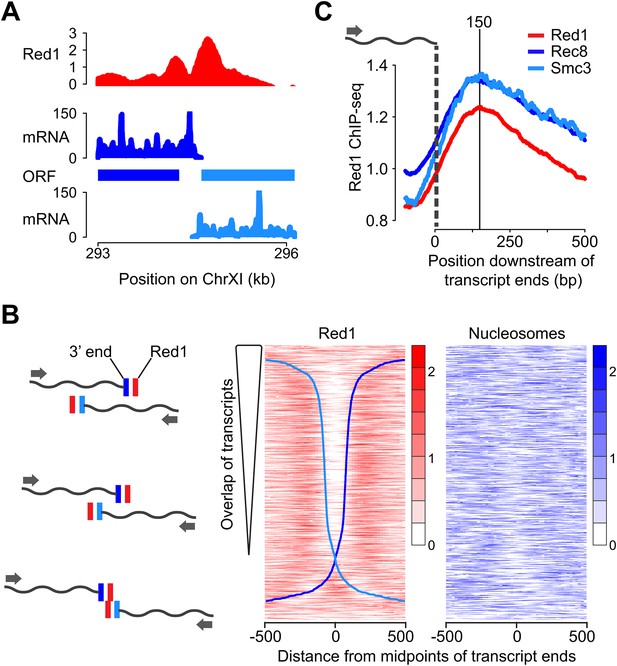
Red1 accumulates next to transcript ends of convergent gene pairs.
(A) Example of overlapping transcripts from a convergent gene pair. ORFs and transcripts of YKL077W and YKL075C were plotted with respect to their positions on chromosome XI. (B) Two-peak pattern of Red1 binding at convergent gene pairs correlates with the amount of transcript overlap. Convergent gene pairs were ranked by the extent of transcript overlap (Brar et al., 2012a). Red1 signals (red) were plotted within 1-kb regions centered at the midpoints between of transcript ends. Nucleosome occupancy signals (blue) (Pan et al., 2011a) were also plotted as described above. The two blue curves represent the positions of transcript ends of each gene (dark blue = forward direction, light blue = reverse direction; see schematic to the left). (C) Enrichment of cohesin subunits and axis proteins was averaged for all convergent transcript ends and plotted as a function of distance from the transcript ends.
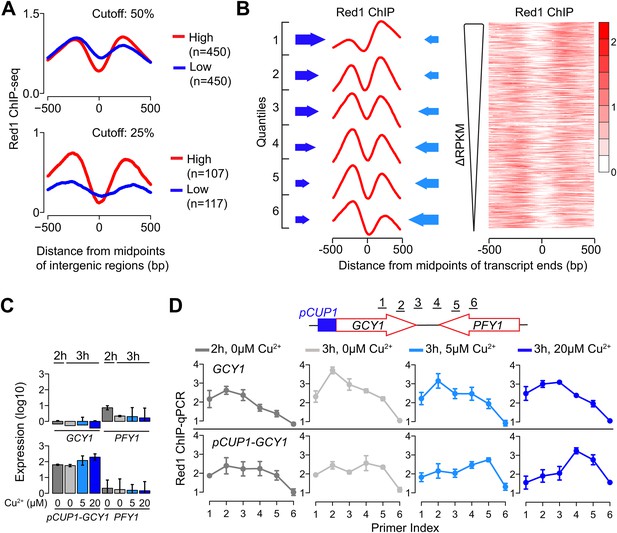
Transcriptional activity dictates Red1 binding patterns.
(A) Convergent gene pairs were identified, in which both genes were strongly transcribed (red) or both genes were weakly transcribed (blue) and average Red1 signal was determined around the midpoints of the intergenic regions. The upper panel represents a cutoff of 50% (both genes were among the top 50% highly transcribed or among bottom 50% lowly transcribed genes); the lower panel represents a cutoff of 25%. (B) The Red1 binding signals are biased toward the more weakly transcribed gene at convergent gene pairs. Convergent gene pairs were ranked according to their differences in RPKM values (ΔRPKM = RPKMforward − RPKMreverse), and Red1 binding signals were plotted (right panel). The left panel shows the average Red1 binding signal in six quantiles of ranked ΔRPKM. Arrow sizes schematically represent the relative transcriptional activities of convergent gene pairs. (C) Gene expression of the GCY1/PFY1 convergent gene pair in response to different concentrations of copper was measured by RT-qPCR at the indicated time points. Copper was added to the sporulation medium at 2 hr, and samples were collected at 2 hr and 3 hr. Transcription levels of both convergent genes were examined in a wild-type strain and a strain harboring a pCUP1 promoter insertion upstream of GCY1. Error bars: S.D. of three independent replicates. (D) ChIP-qPCR analysis of Red1 binding from the same experiment as in (C). Schematic depicts relative position of primer pairs. Error bars: S.D.
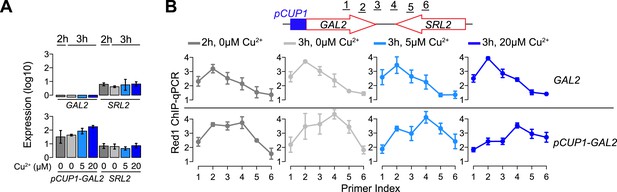
Red1 position changes in response to transcription.
(A) Gene expression of the GAL2/SRL2 convergent gene pair in response to different concentrations of copper was measured by RT-qPCR at the indicated time points. Copper was added to the sporulation medium at 2 hr, and samples were collected at 2 hr and 3 hr. Transcription levels of both convergent genes were examined in a wild-type strain and a strain harboring a pCUP1 promoter insertion upstream of GAL2. Error bars: S.D. of three independent replicates. (B) ChIP-qPCR analysis of Red1 binding from the same experiment as in (A). Schematic depicts relative positions of primer pairs. Error bars: S.D.
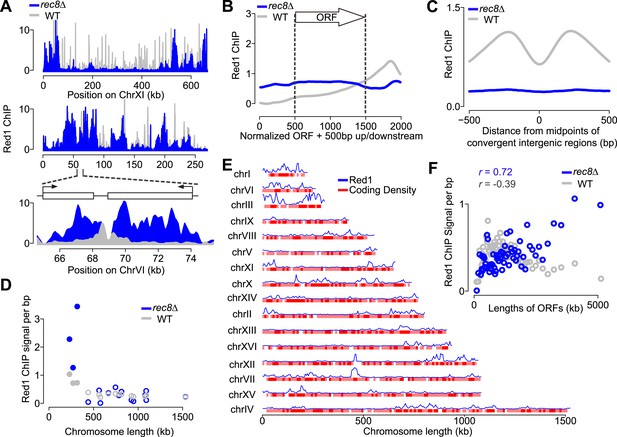
Transcriptional focusing depends on cohesin.
(A) Chromosomal localization of Red1 in rec8Δ mutants (blue) and WT (gray) on chromosomes XI and VI. Bottom panel shows zoom-in for a convergent gene pair on chromosome VI. (B) Red1 enrichment along genes in rec8Δ mutants and WT. (C) Average Red1 accumulation in convergent gene regions in rec8Δ mutants and WT. (D) Average Red1 ChIP signal per bp in rec8Δ mutants and WT as a function of chromosome length. (E) Alignment of chromosomal Red1 binding in rec8Δ mutants (blue) with regions exhibiting high coding density (red). Red1 signals were plotted using a 5-kb smoothing window. Coding density was calculated using 0 (intergenic) or 1 (coding) at each nucleotide position and then smoothed using a 10-kb window and plotted as a heatmap. (F) Red1 signal per bp in rec8Δ mutants and WT as a function of gene lengths (ORFs). Genes were clustered into 64 groups of 100 genes according to similar gene lengths.
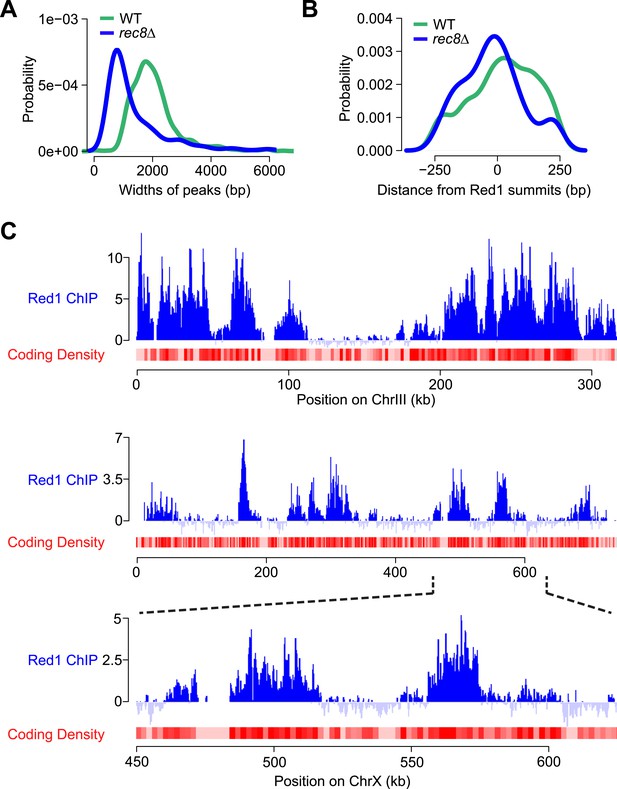
Red1 binding in rec8Δ mutants.
(A) Distribution of widths of Red1 peaks in WT (green) and rec8Δ mutants (blue). (B) Probability of GAN motif occurrence around Red1 peak summits in WT (green) and rec8Δ mutants (blue). (C) Examples of correlation of Red1 binding and coding density along chromosomes III and X. Red1 signals were plotted unsmoothed. Coding density was calculated as in Figure 4E but using a smoothing window of 2 kb.
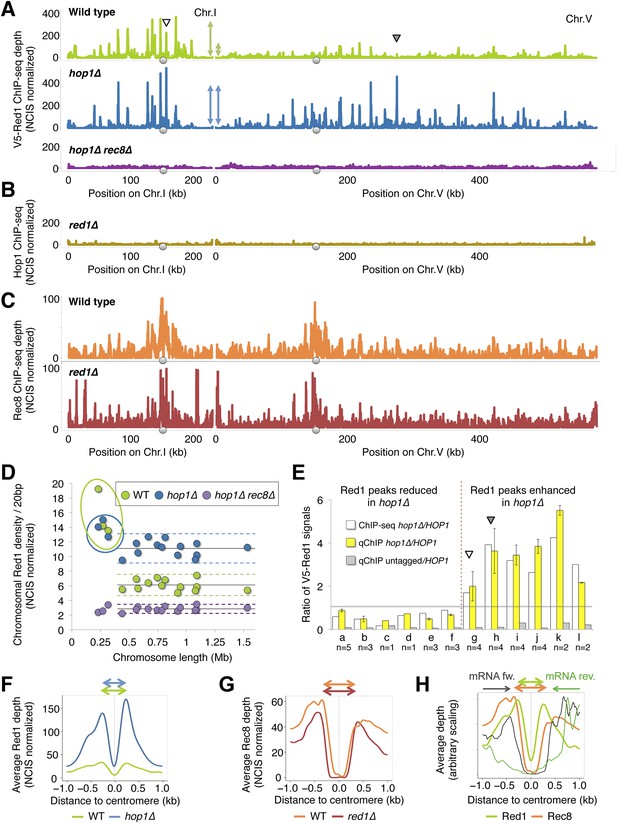
Chromosomal localization of Red1 requires Rec8 and Hop1 redundantly.
(A) Red1 binding profiles in WT (green), hop1Δ mutants (blue), and rec8Δ hop1Δ mutants (purple) from ChIP-seq experiments (NCIS normalized). A small (Chromosome I) and a medium sized chromosome (Chromosome V) are shown as examples to illustrate the reduced Red1 density on chromosome V in WT but not in hop1∆ mutants (indicated schematically by the double-headed arrows). Arrowheads indicate increased Red1 binding confirmed by qPCR in (E). (B) Genome-wide Hop1 ChIP-seq profiles in red1∆ mutants (gold) (NCIS normalized). Chromosomes I and V are shown. (C) Genome-wide Rec8 ChIP-seq profiles in WT (orange) and red1∆ mutants (red) (NCIS normalized). Chromosomes I and V are shown. (D) Red1 density/20 bp (sum of Red1 signal after NCIS normalization divided by chromosomal length) plotted against chromosomal length. The three smallest chromosomes exhibit highly increased binding in WT (green dots), but this difference is largely abolished in hop1∆ mutants (blue dots). Red1 density is reduced to noise level and no longer shows biased distribution in hop1∆ rec8∆ mutants (purple dots). Horizontal lines indicate the mean of 16 chromosomes (continuous lines), plus minus twofold standard deviation (dashed lines). (E) White bars: ratio of V5-red1 ChIP-Seq signal (hop1∆/HOP1) at six cohesin peaks that decrease (a-f: chr3 219k, chr1 171k, chr1 195k, chr6 216k, chr1 95k, chr1 134k) and six cohesin peaks that increase (g-l: chr1 156k, chr5 274k, chr4 435k, chr4 712k, chr7 506k, chr16 576k) upon HOP1 deletion (data from the experiment shown in (A)). Yellow bars: ratio of V5-Red1 signals (hop1∆/HOP1) obtained from ChIP-qPCR primer pairs located at the indicated positions. Gray bars: ratio of V5 signals (untagged/V5-Red1, HOP1) obtained at the corresponding positions. The gray line separates reduction (below 1) from increase (above 1). Arrowheads indicate positions on chr1 and chr5 shown in (A). DNA from each biological repeat was evaluated both at increasing and decreasing sites to exclude systematic bias in DNA preparation. Each single repeat confirmed the direction of change at each location. n indicates the number of repeats. (F) Average densities of V5-Red1 ChIP-seq signals over all 16 yeast centromeres, aligned at their midpoints. The averaged peaks are fourfold higher in hop1∆ mutants (blue) than in HOP1 cells (green). The fold increase at centromeres exceeds the ∼twofold increase observed for the entire chromosomes (see D). (G) Rec8-HA (orange) and Rec8-HA red1∆ (red) flank the centromere at a slightly wider distance (around 345 bp) than Red1 (270 bp). (H) V5-Red1 (light green) and Rec8-HA (orange) are shown as in (G). The average transcription levels from 4 hr after meiotic induction (Brar et al., 2012a) around centromeres (forward—black, reverse—dark green) reveal a transcription free pocket, in which Red1 resides. Meiotic cohesin peaks almost coincide with the ends of transcripts. Rec8-HA and V5-Red1 are shown on two different scales.
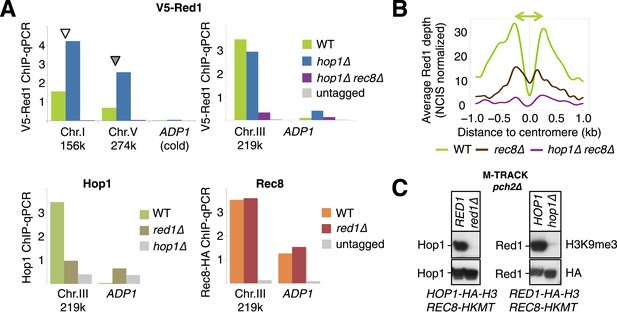
Analysis of axis protein binding.
(A) ChIP-qPCR analysis of axis proteins at selected positions in the indicated mutant backgrounds was performed with the precipitated DNA of the respective ChIP-seq experiment, before library amplification, for independent assessment of relative scaling of different profiles as shown in Figure 5. Gray bars indicate signal independent of the respective tag (V5-Red1 or Rec8-HA) or antibody (hop1∆). X-axis labels indicate genomic positions of primer pairs. (B) Averaged densities of V5-Red1 ChIP-seq signals over all 16 yeast centromeres, aligned at their midpoints. Two rather low V5-Red1 peaks (green) flank the centromere midpoint at a distance of around 220 bp. rec8∆ mutants (brown) or rec8∆ hop1∆ double mutants (purple) show reduced peaks. (C) The meiotic axis remodeler Pch2 is not responsible for the loss of interaction of Hop1 (or Red1) with Rec8 in red1∆ (or hop1∆) mutants, respectively. Upper panels: detection of interaction at 4 hr in SPM using anti-H3K9me3 antibody. Lower panels: total Hop1-HA-H3 (Red1-HA-H3) protein as determined by Western against the HA tag.
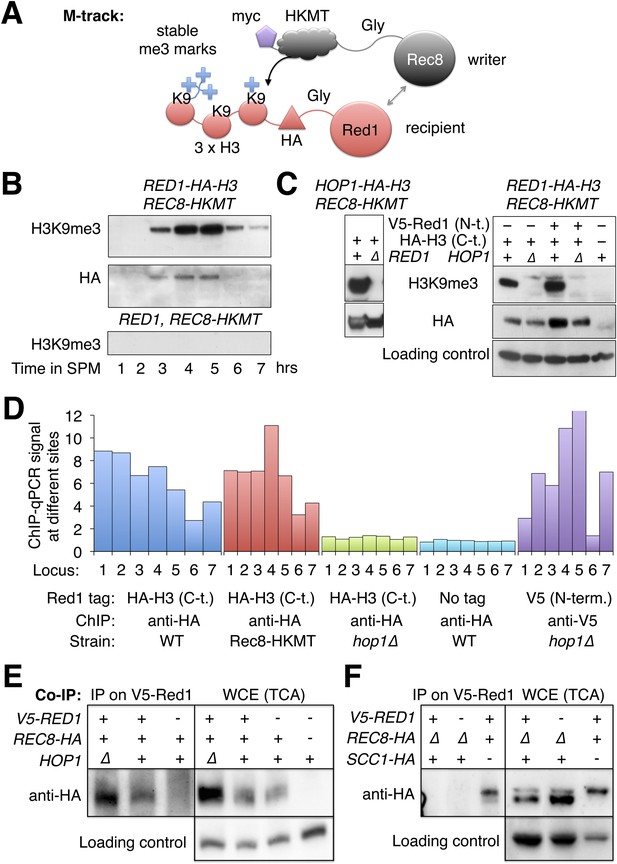
Red1 and Hop1 interact with Rec8.
(A) Illustration of the physical proximity assay M-track. The writer (Rec8) is tagged at on its C-terminus with a human derived histone lysine methyltransferase (HKMT), which will transfer up to three methyl groups to lysine 9 of a small histone H3 fragment presented by the recipient (Red1) depending on the lifespan of the interaction. Using an H3-K9me3 specific antibody, the interaction between writer and recipient can be visualized (Zuzuarregui et al., 2012). (B) Stable proximity between Red1 and Rec8. Upper panel: detection of interaction between Rec8 and Red1 in a meiotic time course using anti-H3-K9me3 antibody. Middle panel: total Red1-HA-H3 protein using anti-HA antibody. Lower panel: K9me3 antibody signal depends on the H3 tag. (C) Left panels: proximity between C-terminally tagged Hop1 and Rec8 depends on Red1. Upper panel: detection of proximity between Rec8 and Hop1 at 4 hr in SPM using anti-H3-K9me3 antibody. Lower panel: total Hop1 protein as determined using anti-HA antibody. Right panels: proximity between C-terminally tagged Red1 and Rec8 depends on Hop1. Upper panel: detection of proximity between Rec8 and Red1 at 4 hr in SPM using anti-H3-K9-me3 antibody. Middle panel: total Red1-HA-H3 protein as determined by Western against the HA tag. Lower panel: loading control. (D) When the C-terminus of Red1 is tagged, Hop1 becomes essential for the chromosomal recruitment of Red1. qChIP (using anti-HA antibody) of Red1-HA2-H3 (blue), Red1-HA2-H3 in the presence of Rec8-HKMT (red), Red1-HA2-H3 in hop1Δ mutants (green) and untagged Red1 (turquoise) at 7 peak sites (see Supplementary file 1B). The values for V5-Red1 in hop1Δ mutants (purple) were taken from the profiles shown in Figure 5A and put in proportion to Red1 WT (see also qPCR for V5-Red1 in Figure 5—figure supplement 1A). (E) Left panel: at 4 hr in SPM Red1 was immuno-precipitated with anti-V5 antibody and co-precipitating Rec8 was detected with anti-HA antibody. Lanes from left to right are V5-RED1 hop1∆, V5-RED1 HOP1, RED1 HOP1. Right panel: Western blot (TCA) of the cultures at the time of the IP. Lanes from left to right are V5-RED1 hop1∆, V5-RED1 HOP1, RED1 HOP1, RED1 HOP1. Rec8 was tagged with HA at the C-terminus except in the rightmost lane, where it was untagged. Swi6 was stained on the stripped blot as a loading control. (F) Same type of experiment as in (E), but in two strains Scc1-HA was expressed from the REC8 promoter, while REC8 was deleted. Lanes from left to right are V5-RED1 pREC8-SCC1-HA rec8∆, RED1 pREC8-SCC1-HA rec8∆, V5-RED1 SCC1 REC8-HA. Right panel: Western blot (TCA) of the same cultures at the time of the IP. Swi6 served as loading control.
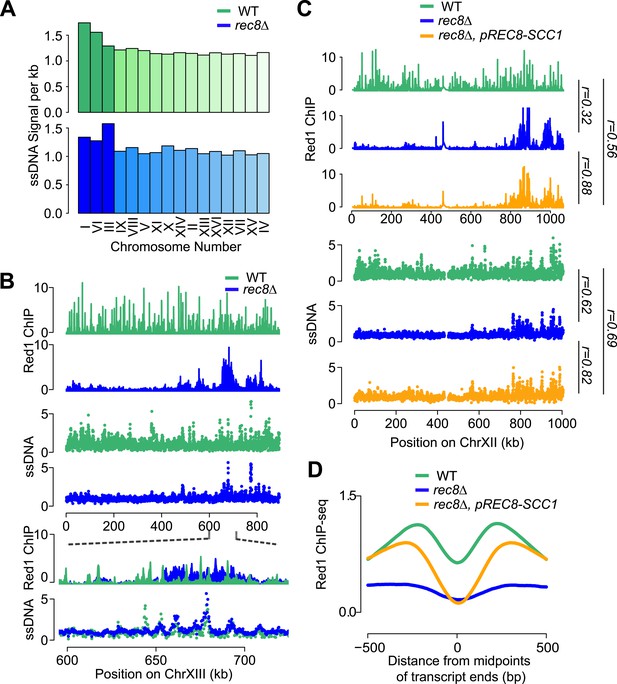
Mitotic cohesin rescues transcriptional focusing but not genome-wide defects of rec8Δ mutants.
(A) Relative DSB hotspot intensity of each chromosome in WT (green) and rec8Δ mutants (blue) as determined using ssDNA arrays. Chromosomes are sorted by increasing size. (B) DSB activity spatially correlates with Red1 enrichment along chromosomes. Distribution of Red1 enrichment (top panels) and DSB hotspots (middle panels) along chromosome XIII in WT (green) and rec8Δ mutants (blue). Zoom-in in bottom panels shows that DSB hotspots location is unaltered at finer scales in rec8Δ mutants despite severely altered Red1 distribution. (C) Ectopic expression of mitotic cohesin does not rescue the defects in Red1 localization or DSB formation of rec8Δ mutants. Distribution of Red1 enrichment (top panels) and DSB hotspots (bottom panels) along chromosome XII in WT (green) and rec8Δ (blue) and rec8Δ pREC8-SCC1 (orange). (D) Ectopic expression of mitotic cohesin partially rescues transcriptional focusing of Red1 in rec8Δ mutants. Average Red1 accumulation in convergent gene regions in WT (green), rec8Δ (blue), and rec8Δ pREC8-SCC1 (orange).
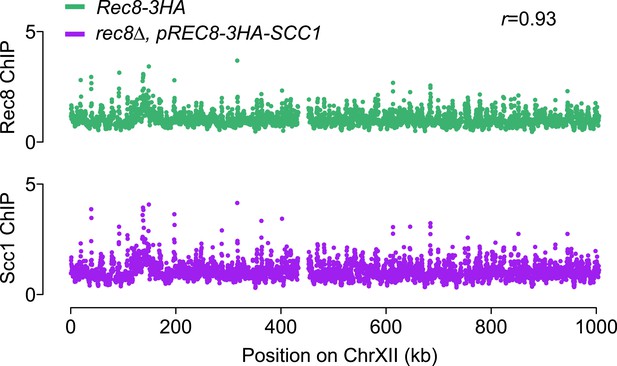
Ectopically expressed Scc1 localizes to the same sites as Rec8 in meiosis.
Top panel: ChIP–chip analysis of Rec8-3HA in otherwise WT cells (green). Bottom panel: ChIP–chip analysis of 3HA-Scc1 in rec8Δ, pREC8-3HA-SCC1 mutant cells (purple). Signal along chromosome XII is depicted to allow comparison with the profiles shown in Figure 7C.
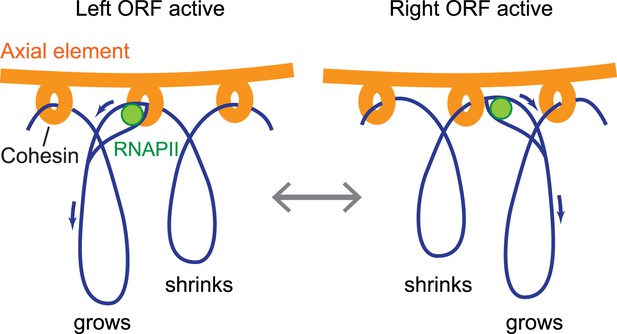
Model of the interface between axial elements and DNA.
Using the topological protein–DNA interaction of cohesin, a robust linkage between a protein rod (the axial element) and a transcriptionally active chromosome can be established. The protein axis forms on top of cohesin, while the elongating RNAPII, likely unable to pass through the cohesin molecule, may pull the DNA through the complex to avoid a premature transcription block. Blue arrows indicate the direction of DNA flow, going in the opposite direction of transcription. Based on the range of transcriptional overlap, loop movements will in most cases not exceed 500 bp and are not drawn to scale.
Additional files
-
Supplementary file 1
Strains and primers. (A) Strains used in the work. (B) Primers used for qPCR and RT-qPCR.
- https://doi.org/10.7554/eLife.07424.016
-
Supplementary file 2
Python code for sequence extraction for motif analysis.
- https://doi.org/10.7554/eLife.07424.017





