The human ARF tumor suppressor senses blastema activity and suppresses epimorphic tissue regeneration
Figures
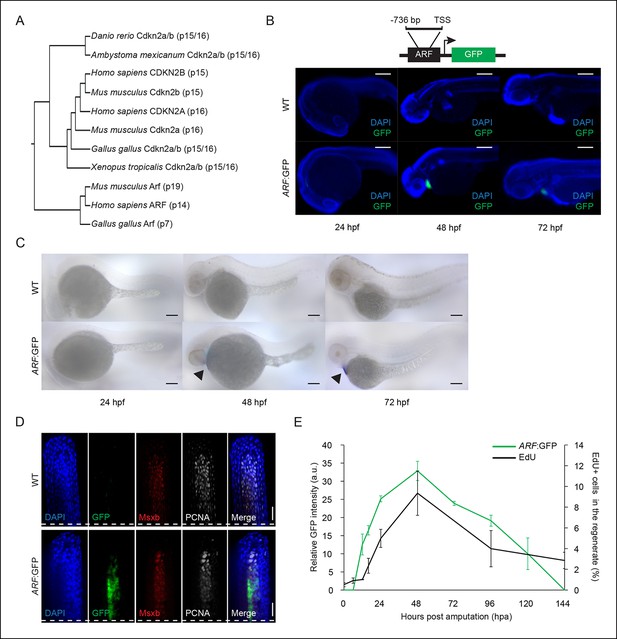
ARF, not normally present in highly regenerative vertebrates, is specifically activated in blastemas of ARF transgenic zebrafish.
(A) Comparison of amino acid sequences of proteins produced by the Cdkn2a/b loci of zebrafish (Danio rerio), amphibians including the axolotl (Ambystoma mexicanum) and western clawed frog (Xenopus tropicalis), chickens (Gallus gallus), and mammals including the mouse (Mus musculus) and human (Homo sapiens). While Cdkn2a and Cdkn2b are conserved and encode Ink4 orthologs, Arf evolved recently and orthologs do not exist in highly regenerative vertebrates including teleost fish and urodele amphibians. (B) Schematic of transgene expressing cytoplasmic Green fluorescent protein (GFP) under the control of the human ARF promoter (top). The promoter consists of human regulatory sequences 736 bp upstream of the transcriptional start site (TSS) of ARF. Immunostaining (wide-field images) for GFP at 24 hpf, 48 hpf, and 72 hpf in wild type (WT) and ARF:GFP embryos (bottom). Scale bars: 200 μm. GFP expression is visible in the hearts of transgenic fish due to presence of a separate transgene used for selection (cmlc2:GFP). (C) Whole-mount in situ hybridization for GFP at 24 hpf, 48 hpf, and 72 hpf in WT and ARF:GFP embryos. Scale bars: 100 μm. Alkaline phosphatase staining is detected in the hearts of transgenic fish (arrow heads) because of the selection transgene as in (B). (D) Confocal images of coronal vibratome sections immunostained for GFP, Msxb, and Proliferating cell nuclear antigen (PCNA) at 2 dpa in WT and ARF:GFP fins. Scale bars: 50 μm. GFP expression is induced in the proliferative blastema of the regenerate, but it is not expressed in the surrounding epithelium. White dashed lines represent amputation planes. (E) GFP intensity (green line) in the regenerates of ARF:GFP transgenic fish relative to WT fish after amputation. The black line represents the percentage of EdU + cells in the regenerates of WT fish after amputation (N=3; secondary axis). Figure supplement 1 shows in vitro ARF promoter assays. Figure supplement 2 shows additional images for panels B, D, and E. Figure supplement 3 shows wound healing in WT and ARF:GFP fins. Results are shown as mean ± standard deviation. hpa: Hours post amputation.
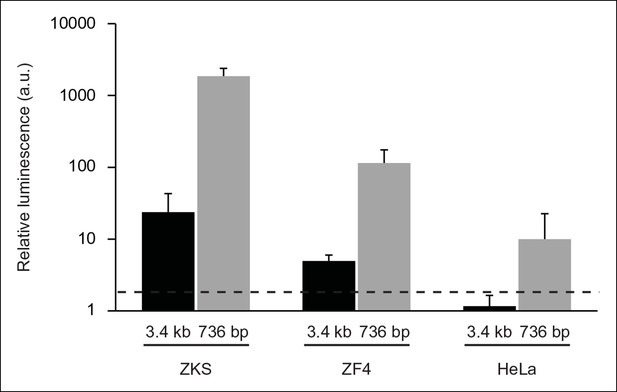
In vitro analysis of ARF promoter constructs in zebrafish and human cells.
Representative luciferase reporter data of three replicates: relative luminescence generated by ARF promoter-firefly luciferase reporter constructs transfected into zebrafish (ZKS, ZF4) and human (HeLa) cells. Two ARF promoter-reporter constructs were tested; one contained sequences up to 3.4 kb upstream of the TSS of ARF (3.4 kb), while the other contained sequences up to 736 bp upstream of the TSS of ARF (736 bp). Relative luminescence was measured by normalizing firefly luciferase values to those detected from a Renilla luciferase construct used as a transfection efficiency control. The relative luminescence values were then normalized to those of the negative control construct, pcDNA. Any values above 2 (black dashed line) are significant (N = 3; pcDNA = 1; p<0.05). Results are shown as mean ± standard deviation. ZKS: Zebrafish kidney stromal.
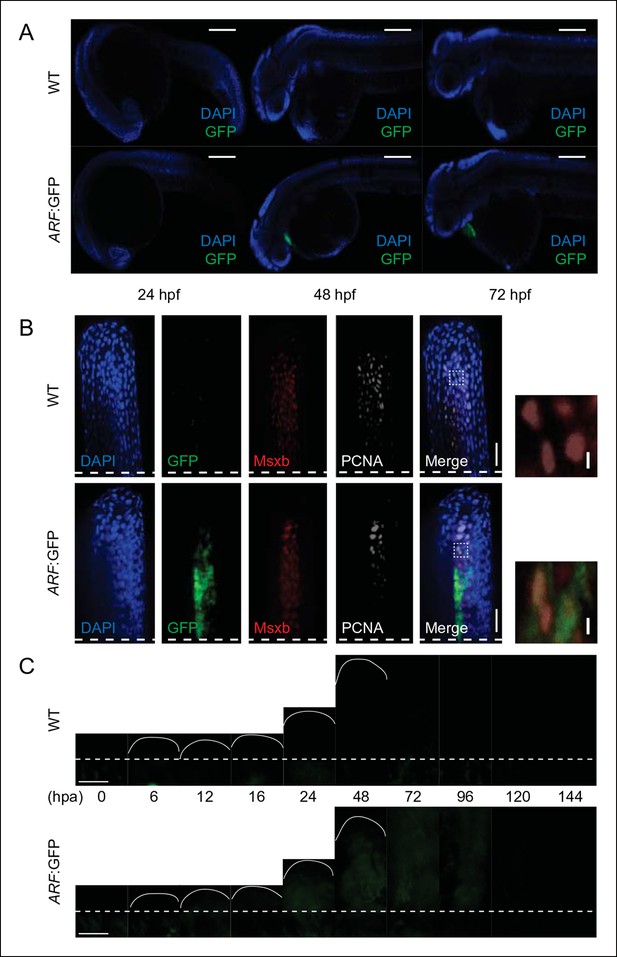
GFP reporter activity in WT or ARF:GFP zebrafish at multiple developmental and regenerative time points.
(A) Immunostaining (sagittal confocal images) for GFP at 24 hpf, 48 hpf, and 72 hpf in WT and ARF:GFP embryos. Scale bars: 200 μm. GFP expression is restricted to the hearts of transgenic fish due to presence of a separate transgene used for selection (cmlc2:GFP). (B) Confocal images from Figure 1 of coronal vibratome sections immunostained for GFP, Msxb, and PCNA at 2 dpa in WT and ARF:GFP fins. Scale bars: 50 μm. Included to the right of the figure are insets showing Msxb +, PCNA +, GFP- blastema cells in WT fins and cytoplasmic GFP expression in Msxb +, PCNA + blastema cells in ARF:GFP fins (white dashed boxes). DAPI is excluded from the inset images to improve clarity of costaining. Scale bars: 10 μm. (C) Wide-field epifluorescent images of WT and ARF:GFP fins at multiple time points during fin regeneration. GFP intensity of individual ARF:GFP images was evaluated relative to that of WT images at the same time points, and the resulting values were plotted in Figure 1E. There is a small amount of detectable autofluorescence below the amputation plane in the regenerating wild-type and transgenic fins. Scale bars: 100 μm. Dashed lines represent amputation planes. GFP: Green fluorescent protein; hpa: Hours postamputation; PCNA: Proliferating cell nuclear antigen; WT: Wild type.
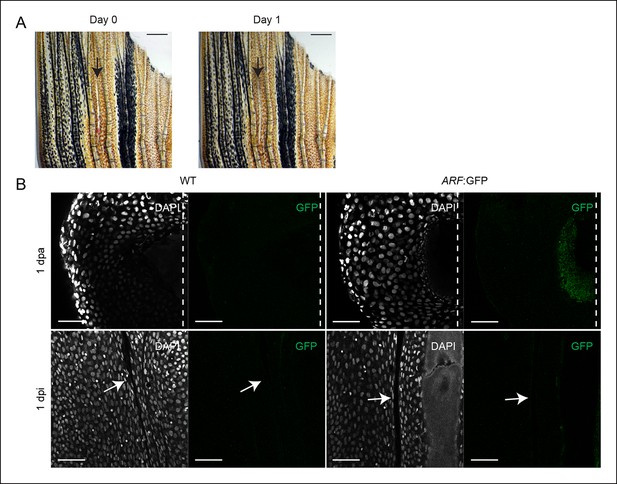
ARF is not activated during wound healing in the absence of a blastema.
(A) At day 0, dorsal fin lobes were wounded (interray laceration, 0 dpi), while ventral fin lobes were amputated (0 dpa). At day 1, GFP expression was assayed in the healing (dorsal) and regenerating (ventral) fins. Scale bars: 0.5 mm. (B) Representative images (sagittal confocal images) of cytoplasmic GFP expression in WT and ARF:GFP fins that were either amputated (1 dpa) or wounded (1 dpi). GFP is only detected in ARF:GFP fins that have been amputated (N = 5). Scale bars: 50 μm. Arrows point to the interray wound. Dashed lines represent amputation planes.GFP: Green fluorescent protein; WT: Wild type.
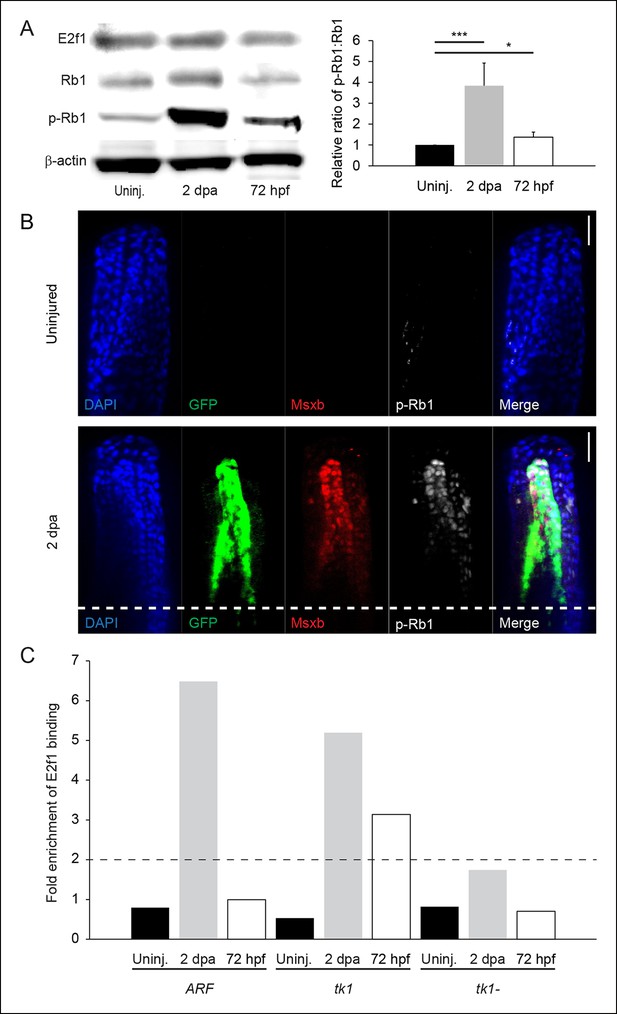
Rb1 hyperphosphorylation and E2f1 binding of the human ARF promoter in the blastema during regeneration.
(A) Representative Western blot of three experimental replicates of Rb pathway components, E2f1, Rb1, and hyperphosphorylated Rb1 (p-Rb1), before injury (uninj.), at 2 dpa, and during embryogenesis at 72 hpf (left). Quantification of p-Rb1 and Rb1 levels normalized to β-Actin and relative to uninjured tissue. Results are from three independent biological replicate experiments and are shown as mean ratios ± standard deviation. *p<0.05; ***p<0.001 (right). (B) Confocal images of coronal vibratome sections immunostained for Green fluorescent protein (GFP), Msxb, and p-Rb1 in uninjured and regenerating (2 dpa) ARF:GFP fins. Scale bars: 50 μm. Very little p-Rb1 staining is seen in the uninjured fin, but high levels of p-Rb1 staining can be seen in Msxb + cells in the blastema at 2 dpa. The white dashed line represents the amputation plane. (C) Representative ChIP qPCR data of three experimental replicates with a pool of 30 fins per experiment. Tissue was collected from ARF:GFP transgenic fish before injury (uninj.), at 2 dpa (regenerate only), and at 72 hpf. Fold enrichment of E2f1 binding was normalized to rabbit IgG. The zebrafish thymidine kinase 1 (tk1) promoter was used as a positive control for E2f1 binding. Sequences 2 kbp upstream of tk1 were used as a negative control (tk1-). Values above twofold (black dashed line) are significant (p<0.05). Figure supplement 1 shows promoter sequences for the ARF, tk1, and tk1- promoters annotated for canonical E2f binding sites. hpa: Hours postamputation.

Promoter sequences evaluated for E2f1 enrichment using an E2f1 antibody to perform a ChIP assay.
Both ARF and tk1 promoters contain E2f binding sites (bold; del Arroyo et al., 2007, Tfsitescan), but the tk1- promoter does not.
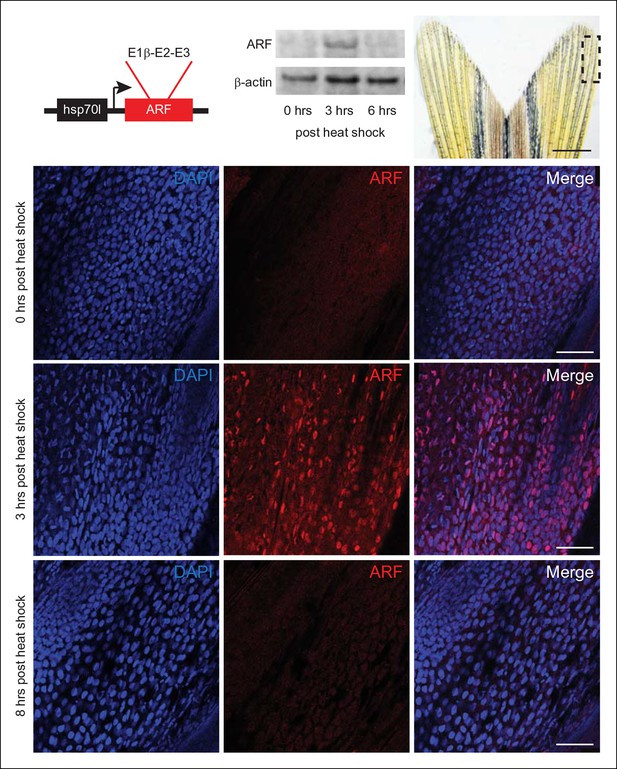
Expression of the mammalian tumor suppressor ARF in zebrafish driven by heat shock promoter.
In vivo analysis of transgenic zebrafish expressing human ARF under the control of an inducible heat shock promoter, Tg (hsp70l:ARF) (hs:ARF). Schematic of the hs:ARF transgene (top left). The ARF cassette included in the transgene is a cDNA that consists of human exons 1b, 2, and 3 of CDKN2A. Representative Western blot of 3 replicates of ARF before (0 hr) and 3 and 6 hr post heat shock induction of ARF expression (top middle). Portion of fin shown for analysis of expression in vivo (top right; dashed box). Scale bar: 1 mm. Immunostaining (sagittal confocal images) for ARF in adult hs:ARF zebrafish fins at 0, 3, and 8 hr after a single, hour long, 37°C heat shock (bottom). Scale bars: 50 μm. ARF expression is maximal at 3 hr post heat shock, and it is undetectable by 8 hr post heat shock. Figure supplement 1 shows in vitro assays.
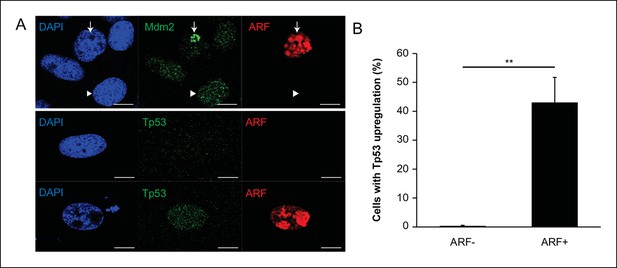
Analysis of ARF expression in zebrafish cells.
(A) Immunofluorescence for Mdm2 and ARF (top) and Tp53 (bottom) in zebrafish cells (ZKS) transfected with pcDNA-ARF. ARF and Mdm2 co-localize in the nucleolus (arrow) when ARF is expressed; in cells without ARF, Mdm2 has a diffuse nuclear staining pattern (arrow head; top). Tp53 upregulation depends on ARF expression (bottom). Scale bars: 10 μm. (B) Quantification of Tp53 upregulation in zebrafish cells (ZKS) transfected with pcDNA-ARF (N = 100, p<0.01). Results are shown as mean ± standard deviation.
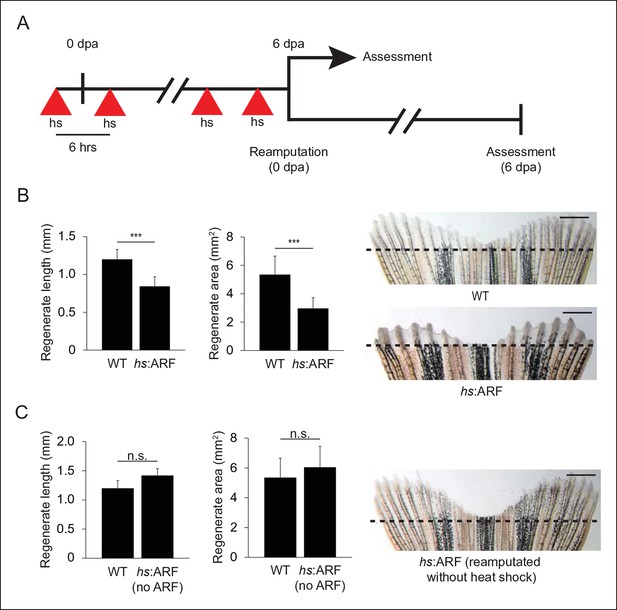
ARF suppresses fin regeneration.
(A) Schematic of heat shock regimen. An initial hour long, 37°C heat shock is delivered 3 hr prior to amputation (0 dpa) and then every 6 hrs thereafter for 6 days. Regenerates are then assessed (top) or fins are reamputated (0 dpa) and allowed to regenerate in the absence of heat shock for 6 days (bottom). (B) Quantification of regenerate length and area at 6 dpa in WT and hs:ARF fins exposed to the heat shock regimen (left; N = 40 fins representing multiple different transgene insertions, p<0.001). Representative images of fin regeneration at 6 dpa in WT and hs:ARF fins exposed to the heat shock regimen (right). (C) Quantification of regenerate length and area at 6 dpa in reamputated hs:ARF fins not exposed to heat shock (left; N = 40 fins, p>0.05). Representative image of fin regeneration at 6 dpa in a reamputated hs:ARF fin not exposed to heat shock (right). The dashed lines represent amputation planes. Scale bars: 1 mm. Results are shown as mean ± standard deviation. Figure supplement 1 shows ARF and Tp53 immunostaining at 4 dpa, and tp53 and cdkn1a expression changes with ARF expression. It also shows regeneration at 14 dpa after heat shock was discontinued at 6 dpa. hs: Heat shock; WT: Wild type. n.s.: not significant.
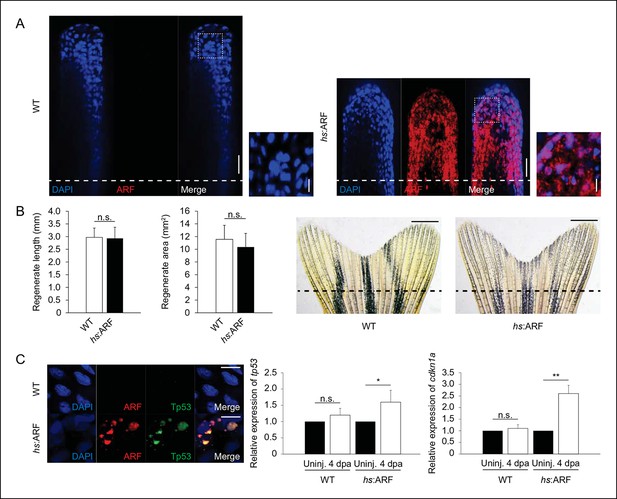
ARF expression during regeneration promotes Tp53, tp53, and cdkn1a upregulation and regeneration inhibition is reversible.
(A) Representative (sagittal confocal) images of WT and hs:ARF fins at 4 dpa. Scale bars: 50 μm. Dashed lines represent amputation planes. ARF localizes to the nucleus (inset). Scale bars: 10 μm. (B) Quantification of fin regenerate length and area in WT and hs:ARF at 14 dpa after heat shock was discontinued at 6 dpa (left; N = 10 fins, p>0.05). Representative images of WT and hs:ARF fins at 14 dpa after heat shock was discontinued at 6 dpa (right). (C) Representative images of ARF and Tp53 in WT and hs:ARF fins at 4 dpa (left). Scale bar: 10 μm. Tp53 expression is only detected in cells with ARF expression. Quantifications of relative tp53 (middle) and cdkn1a (right) transcript expression in uninjured (uninj.) WT and hs:ARF fin and regenerates at 4 dpa (N = 3 replicates). Expression was normalized to β-Actin transcripts and relative to uninjured fins within each condition. Significant increases in tp53 (N=5 fins, p<0.05) and cdkn1a (N =5 fins, p<0.01) were observed with ARF expression. Results are shown as mean ± standard deviation.hs: Heat shock; WT: Wild type. n.s.: not significant.
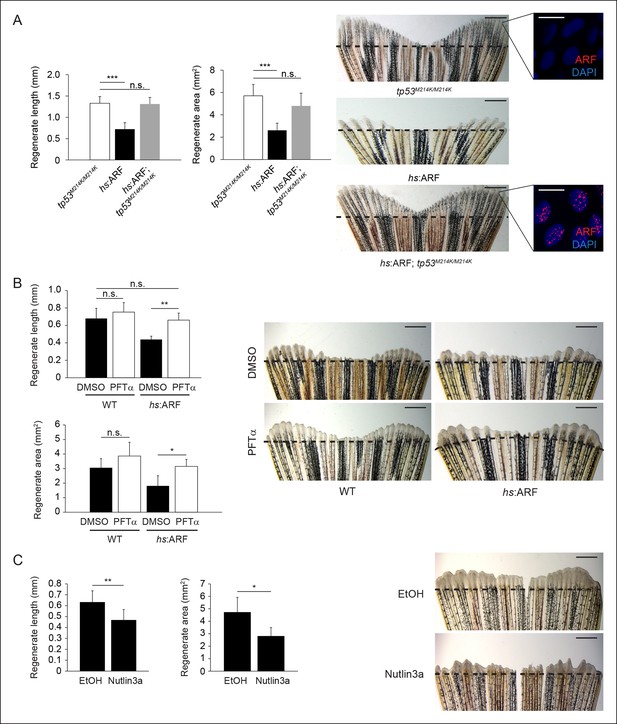
Human ARF functions through the Tp53 pathway in fish to suppress regeneration.
(A) Quantification of regenerate length and area at 6 dpa in tp53M214K/M214K, hs:ARF, and hs:ARF; tp53M214K/M214K fins exposed to the heat shock regimen as in Figure 4 (left; N = 30 fins). Representative images of fin regeneration at 6 dpa in tp53M214K/M214K, hs:ARF, and hs:ARF; tp53M214K/M214K fins exposed to heat shock (right). Scale bars: 1 mm. Immunostaining (sagittal confocal images) for ARF in tp53M214K/M214K and hs:ARF; tp53M214K/M214K fins 3 hr after a single heat shock (right inset). Scale bars: 10 μm. Fin regeneration proceeds equally well in tp53M214K/M214K and hs:ARF; tp53M214K/M214K fins exposed to heat shock, but fin regeneration inhibition is observed in hs:ARF fins exposed to heat shock. (B) Quantification of regenerate length and area at 4 dpa in wild type (WT) and hs:ARF fins exposed to heat shock and 5 μM pifithrin-α(PFTα) or 0.1% Dimethyl sulfoxide (DMSO) (vehicle) (left; N = 8 fins, p<0.01). Representative images of fin regeneration at 4 dpa in WT and hs:ARF fins exposed to heat shock and 5 μM PFTα or 0.1% DMSO (right). Scale bars: 0.5 mm. Inhibition of Tp53 activity with PFTα rescues regeneration suppression by ARF. (C) Quantification of regenerate length and area at 4 dpa in WT fins exposed to 5 μM nutlin3a or Ethanol (EtOH) (vehicle) (left; N = 8 fins, p<0.01). Representative images of fin regeneration at 4 dpa in WT fins exposed to 5 μM nutlin3a or EtOH (right). Scale bars: 0.5 mm. Inhibition of Mdm2 with nutlin3a phenocopies ARF expression by suppressing fin regeneration. The dashed lines represent amputation planes. Results are shown as mean ± standard deviation. n.s.: not significant.
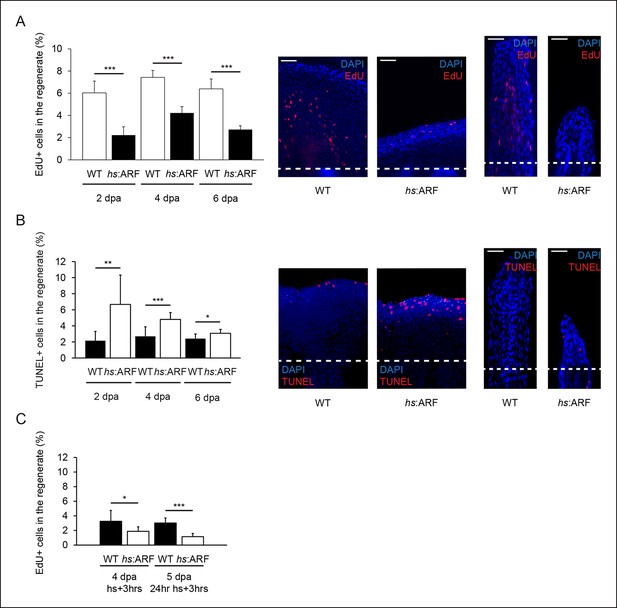
ARF suppresses fin regeneration by inducing apoptosis and cell-cycle arrest.
(A) Quantification of EdU staining at 2, 4, and 6 dpa in wild type (WT) and hs:ARF fins exposed to heat shock (left). At 2 dpa, 6.0% ± 1.1% of cells in WT regenerates were EdU + compared with approximately 2.2% ± 0.8% in Heat shock (hs):ARF regenerates. At 4 dpa, approximately 7.4% ± 0.6% of cells in WT regenerates were EdU + compared with 4.2% ± 0.6% in hs:ARF regenerates. At 6 dpa, approximately 6.4% ± 0.9% of cells in WT regenerates were EdU + compared with 2.7% ± 0.3% in hs:ARF regenerates. Significantly fewer cycling cells are detected with ARF expression (N = 10 fins, p<0.001). Representative (left – sagittal confocal, right – longitudinal) images of EdU staining at 2 dpa in WT and hs:ARF fins exposed to heat shock (right). Scale bars: left – 50 μm, right – 25 μm. Dashed lines represent amputation planes. (B) Quantification of Terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) staining at 2, 4, and 6 dpa in WT and hs:ARF fins exposed to heat shock (left). At 2 dpa, 2.2% ± 1.2% of cells in WT regenerates were TUNEL + , while 6.7% ± 3.7% of cells in hs:ARF regenerates were TUNEL + . At 4 dpa, only 2.7% ± 1.2% of cells in WT regenerates were TUNEL + compared with 4.8% ± 0.8% in hs:ARF regenerates. At 6 dpa, 2.4% ± 0.6% of cells in WT regenerates were TUNEL +, while 3.1% ± 0.5% of cell in hs:ARF regenerates were TUNEL +. Significantly more apoptosis is detected with ARF expression (N = 10 fins, p<0.001). Representative images (left – sagittal, right – longitudinal) of TUNEL staining at 2 dpa in WT and hs:ARF fins exposed to heat shock (right). Image quantification was performed on regenerates only. Dashed lines represent amputation planes. Scale bars: left – 50 μm, right – 25 μm. (C) Quantification of EdU staining in WT and hs:ARF fins 3 hr after a single heat shock or 24 hr of heat shock delivered at 4 dpa. After a single heat shock, 3.3% ± 1.5% of cells in WT regenerates were EdU + compared with 1.9% ± 0.6% in hs:ARF regenerates. After 24 hr of heat shock, 3.0% ± 0.7% of cells in WT regenerates were EdU + compared with 1.2% ± 0.4% in hs:ARF regenerates. Significantly fewer cycling cells are detected with ARF expression after blastema formation (N = 10 fins, p<0.001). Results are shown as mean ± standard deviation.
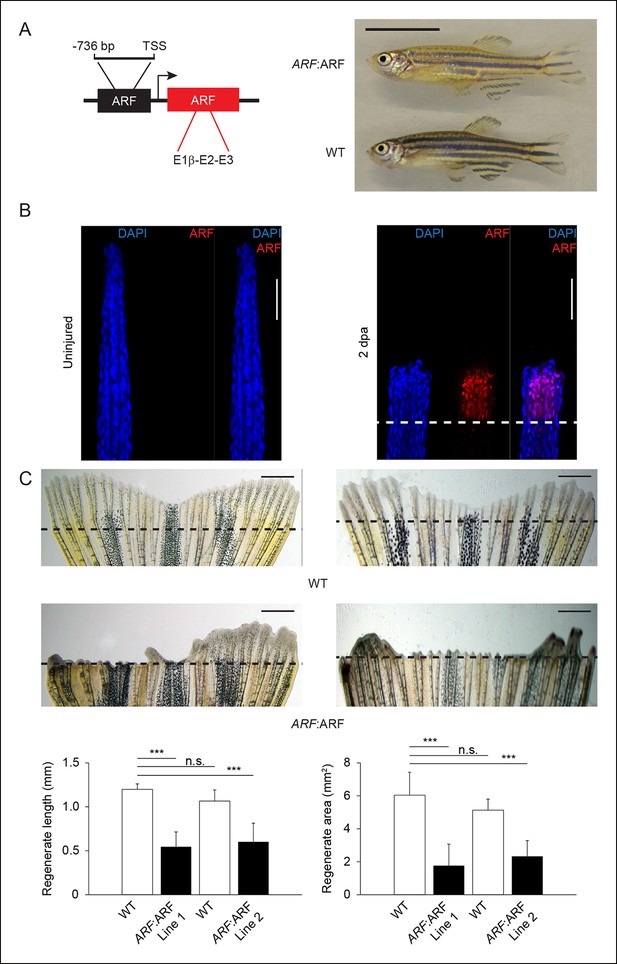
ARF senses regenerative signals and suppresses fin regeneration.
(A) Schematic of transgene expressing human ARF under the control of the human ARF promoter (left). Representative images of age- and sex-matched ARF:ARF and WT zebrafish (right; 5 months postfertilization, male). Scale bar: 1 cm. ARF:ARF fish are viable, grow to adulthood and are of normal size and patterning. (B) Immunostaining (longitudinal confocal images) for ARF in ARF:ARF transgenic fish before injury (uninjured ) and at 2 dpa. Scale bars: 50 μm. ARF is specifically expressed upon injury. The dashed line represents the amputation plane. (C) Representative images of fin regeneration at 6 dpa in WT and ARF:ARF fins (top). Scale bars: 1 mm. The dashed lines represent amputation planes. Quantification of regenerate length and area at 6 dpa in WT and ARF:ARF fins (bottom; N= 10 fins, p<0.001). The first set of bars in each graph represents the results from one transgenic line (Line 1), while the second set of bars represents the results from a second, independent transgenic line (Line 2). ARF causes marked inhibition of fin regeneration. Results are shown as mean ± standard deviation. Figure supplement 1 shows the embryonic viability of ARF transgenic lines. Figure supplement 2 shows the failure of ARF:ARF fins to completely regenerate after 15 days and even 30 days. Figure supplement 3 shows ARF immunostaining at 6 dpa, Tp53, tp53, and cdkn1a expression changes with ARF expression in WT and ARF:ARF fins at 4 dpa, fin regeneration rescue in ARF:ARF fins treated with PFTα, and EdU incorporation studies performed in WT and ARF:ARF fins. TSS: Transcriptional state site; uninj.: Uninjured; WT: Wild type. n.s.: not significant.
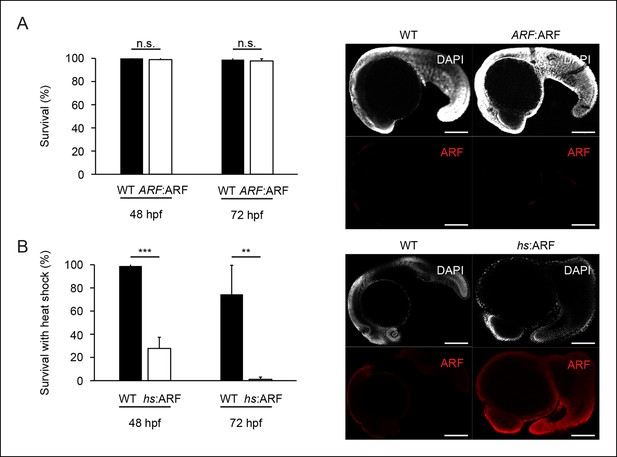
The ARF:ARF transgene does not interfere with development, whereas forced ARF expression causes embryonic lethality.
(A) Quantification of embryonic mortality at 48 hpf and 72 hpf in wild type (WT) and ARF:ARF embryos (left; N = 90, p>0.05). Representative sagittal confocal images of ARF expression at 24 hpf in WT and ARF:ARF (right). (B) Quantification of embryonic mortality at 48 hpf and 72 hpf in WT and hs:ARF embryos exposed to heat shock (left; N = 90, p<0.001). Representative sagittal confocal images of ARF expression at 27 hpf in WT and hs:ARF embryos 3 hr after a single heat shock (right). Scale bars: 200 μm. Results are shown as mean ± standard deviation . n.s.: not significant.
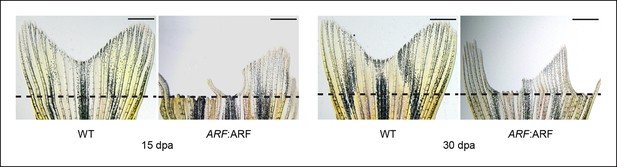
ARF:ARF fins do not completely regenerate even after 30 days.
Representative images of fin regeneration at 15 dpa and 30 dpa in wild type (WT) and ARF:ARF fins. Scale bars: 1 mm.
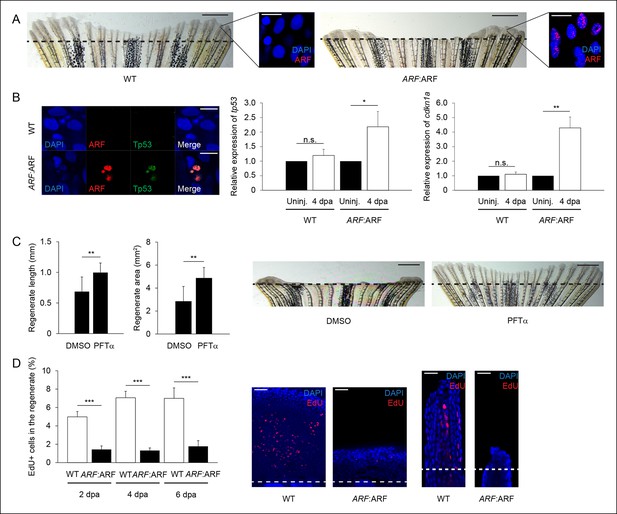
ARF:ARF expression and p53-dependent functions during regeneration.
(A) Representative images of wild type (WT) (left) and ARF:ARF (right) fins at 6 dpa. Scale bars: 1 mm. Representative images of ARF expression at 6 dpa in WT (left) and ARF:ARF (right) fins. Scale bars: 10 μm. Dashed lines represent amputation planes. (B) Representative images of ARF and Tp53 in WT and ARF:ARF fins at 4 dpa (left). Scale bar: 10 μm. Tp53 expression is only detected in cells that express ARF. Quantification of relative tp53 (middle) and cdkn1a (right) transcript expression in uninjured (uninj.) WT and ARF:ARF fin and regenerates at 4 dpa (N = 3 replicates). Expression was normalized to β-Actin transcripts and relative to fins within each condition. Significant increases in tp53 (N = 5 fins, p<0.05) and cdkn1a (N = 5 fins, p<0.01) were observed with ARF expression. (C) Quantification of regenerate length and area at 6 dpa in ARF:ARF fins treated with 0.1% Dimethyl sulfoxide (DMSO) or 5 μM Pifithrin-α (PFTα) (left; N = 8 fins/condition, p<0.01). Representative images of fin regeneration at 6 dpa in ARF:ARF fins treated with 0.1% DMSO or 5 μM PFTα (right). Scale bars: 1 mm. Dashed lines represent amputation planes. Treatment with PFTα rescues fin regeneration in ARF:ARF transgenic zebrafish. (D) Quantification of EdU staining at 2, 4, and 6 dpa in WT and ARF:ARF fins (left). At 2 dpa, 5.0% ± 0.6% of cells in WT regenerates were EdU + compared with approximately 1.4% ± 0.4% in hs:ARF regenerates. At 4 dpa, approximately 7.0% ± 0.7% of cells in WT regenerates were EdU + compared with 1.3% ± 0.3% in hs:ARF regenerates. At 6 dpa, approximately 7.0% ± 1.1% of cells in WT regenerates were EdU + compared with 1.8% ± 0.6% in hs:ARF regenerates. Significantly fewer cycling cells are detected with ARF expression (N = 10 fins, p<0.001). Representative (left – sagittal confocal, right – longitudinal) images of EdU staining at 2 dpa in WT and ARF:ARF fins (right). Scale bars: left – 50 μm, right – 25 μm. Dashed lines represent amputation planes. Results are shown as mean ± standard deviation. n.s.: not significant.
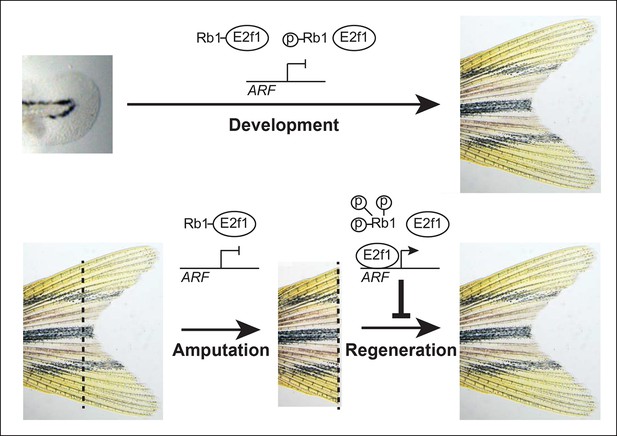
Model of ARF function in the context of Rb pathway activity during zebrafish development and fin regeneration.
ARF is not active during development during which a moderate level of mitogenic signaling causes modest phosphorylation of Rb1 (top); however, during regeneration, high mitogenic signaling induces Rb1 hyperphosphorylation and abundant free E2f1, which activates ARF and leads to inhibition of regeneration (bottom). The dashed lines represent the amputation plane.
Tables
Primary antibodies.
| Host species | Antigen | Company | Cat. No. | Dilution | Application |
|---|---|---|---|---|---|
| Mouse | Tp53 | Abcam | ab77813 | 1:50 | IHC |
| Rabbit | Mdm2 | Santa Cruz | C-18 | 1:50 | IHC |
| Rabbit | GFP | Torrey Pines | TP401 | 1:3000 | IHC |
| Chicken | GFP | Abcam | ab13970 | 1:3000 | IHC |
| Mouse | Msxb | DSHC | 4G1-c | 1:50 | IHC |
| Rabbit | PCNA | Abcam | ab2426 | 1:500 | IHC |
| Mouse | p14ARF | Cell Signaling | 2407 | 1:100/1:500 | IHC/WB |
| Rabbit | Rb1 | AnaSpec | 55432 | 1:500 | WB |
| Rabbit | p-Rb1 (S780) | Abcam | ab47763 | 1:500 | WB |
| Rabbit | Beta-actin | Millipore | EP1123Y | 1:1000 | WB |
| Rabbit | E2f1 | Abcam | ab14769 | 1:1000 | WB |
-
IHC: Immunohistochemistry; PCNA: Proliferating cell nuclear antigen; WB: Western blot.
Chromatin immunoprecipitation primers.
| Gene | Ensembl ID | Target site | Forward primer | Reverse primer |
|---|---|---|---|---|
| CDKN2A | ENSG00000147889 | TSS | 5′-GCTGAGGGTGGGAAGATG-3′ | 5′-CCTTAACTGC AGACTGGGA-3′ |
| tk1 | ENSDARG00000086561 | TSS | 5′-AGTCACTGTGCCGGTTTATT-3′ | 5′-GTCGTCTGCTTG TTGTCTTTATTT-3′ |
| tk1- | ENSDARG00000086561 | 2 kbp 5′ of TSS | 5′-CAGGCTTACGGAGACAGCAA-3′ | 5′-AGTGTTTGCTG CTGGATCAC-3′ |
-
TSS: Transcriptional state site.
Quantitative polymerase chain reactionprimers.
| Gene | RefSeq ID | Forward primer | Reverse primer |
|---|---|---|---|
| CDKN2A | NM_058195.3 | 5′-ATGGTGCGCAGGTTCTTGGTGA-3′ | 5′-CACCACCAGCGTGT CCAGGAAG-3′ |
| actb2 | NM_181601.4 | 5′-CGAGCAGGAGATGGGAACC-3′ | 5′-CAACGGAAACGCT CATTGC-3′ |
| tp53 | NM_131327.2 | 5′-CTGAAGTGGTCCGCAGATG-3′ | 5′-CGTTTGGTCCCAG TGGTGG-3′ |
| cdkn1a | NM_001128420.1 | 5′-AGCTGCATTCGTCTCGTAGC-3′ | 5′-TGAGAACTTACT GGCAGCTTCA-3′ |





