On the dimensionality of odor space
Figures
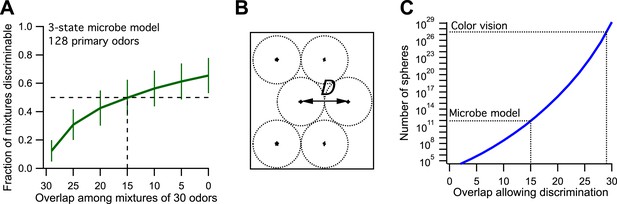
Model of olfaction in a toy microbe.
(A) This 3-state olfactory system counts how many odors in the mixture are attractants vs repellents, and converts the result into three response categories (see text). Two odor mixtures are discriminable if they cause different responses. A numerical simulation of the response to many odor mixtures yields the fraction of discriminable mixtures as a function of the number of odors, O, that they share. Mean ± SD over 1000 repeats using different random assignments of the primary odors. Horizontal dashes: criterion for critical distance (50% discriminable pairs). Vertical dashes: critical distance D = 30 − O = 15. (B) Points in odor space separated by a distance D cause different responses at least half the time. Counting how many such points exist in the space is like trying to pack spheres of diameter D to fill the space as efficiently as possible. (C) The number of such spheres in 128-dimensional space as a function of the discriminable overlap O among 30-odor mixtures, computed by the formula given in Bushdid et al. (2014). The value O = 15 from panel (A) yields ∼9 × 1011 spheres.
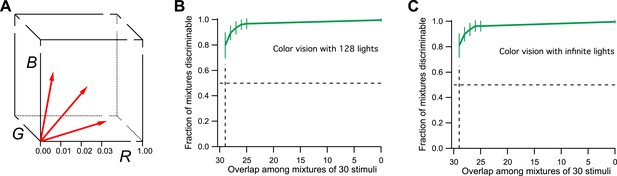
Model of human color vision.
(A) The RGB color cube with three of the 128 primary colors represented by vectors from the origin. Tick marks represent just noticeable differences, for example, along the R-axis. (B) The fraction of discriminable 30-light mixtures as a function of their overlap. Mean ± SD over 1000 repeats using different random assignments of the 128 lights. 30 lights per mixture, 20 mixture pairs per class, 26 subjects per pair. Horizontal dashes: criterion for critical distance. Vertical dashes: critical distance. (C) As in panel B but with the mixture components drawn at random from all possible directions in the space rather than from a preselected set of 128 primaries. The results are almost identical.
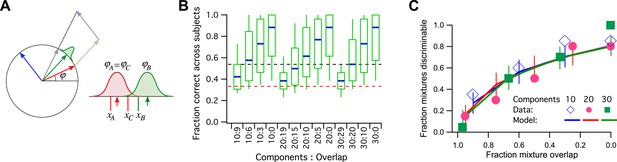
A model simulation of the human smell experiments.
(A) Left: Each primary odor gets mapped into a unit vector (e.g., red, green, blue). Mixtures of odors get mapped into the normalized sum vector (gray). Right: When a subject sniffs an odor vial, the odor angle is corrupted by Gaussian noise and a value is drawn from that distribution. Here three vials were presented, two (A and C) containing the identical odor and a third (B) a different odor. This produced response variables xA, xB, and xC. On this trial, xB and xC are closest to each other, so the subject (incorrectly) identifies A as the odd odor. (B) Discriminability of odor mixtures under this model (compare to Bushdid et al., 2014, Figure 2C). Mixtures were simulated according to the reported procedure with 10, 20, or 30 components and varying overlap. Each mixture pair was presented to 26 subjects, and the fraction of correct identification determined across subjects. Box-and-whisker plot shows the distribution of that fraction with percentiles 10, 25, 50, 75, 90. Average over 1000 repeats of the procedure with different random numbers. Red dashes: chance performance. Black dashes: criterion for discriminability (14/26 correct). (C) Fraction of discriminable mixtures as a function of their overlap (compare to Bushdid et al., 2014, Figure 3D). This is the fraction of mixture pairs in each class that exceeds 50% correct identification across subjects (above the black line in panel B). Lines are mean ± SD. Symbols are data from Bushdid et al. (2014). The model used Gaussian noise with a SD of 0.4 radians.
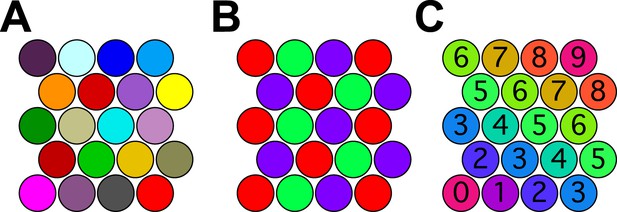
Coloring close-packed spheres in 2 dimensions so no nearest neighbors have the same color.
(A) All colors different. (B) Three colors suffice. (C) Progression of colors in one dimension only.
Tables
Number of dimensions of various spaces involved in sensory discrimination
| Toy microbe | Ring model | Human color | E. coli smell | Human smell | |
|---|---|---|---|---|---|
| Stimuli | ∞ | ∞ | ∞ | ∞ | ∞ |
| Receptors | ∞ | ∞ | 3 | 5 | ∼400 |
| Percepts | 1 | 1 | 3 | 1 | 1–20? |
-
The symbol ∞ stands for ‘very large or infinite’.
Additional files
-
Source code 1
Annotated Igor (Wavemetrics) code.
- https://doi.org/10.7554/eLife.07865.008





