Transdifferentiation mediated tumor suppression by the endoplasmic reticulum stress sensor IRE-1 in C. elegans
Figures
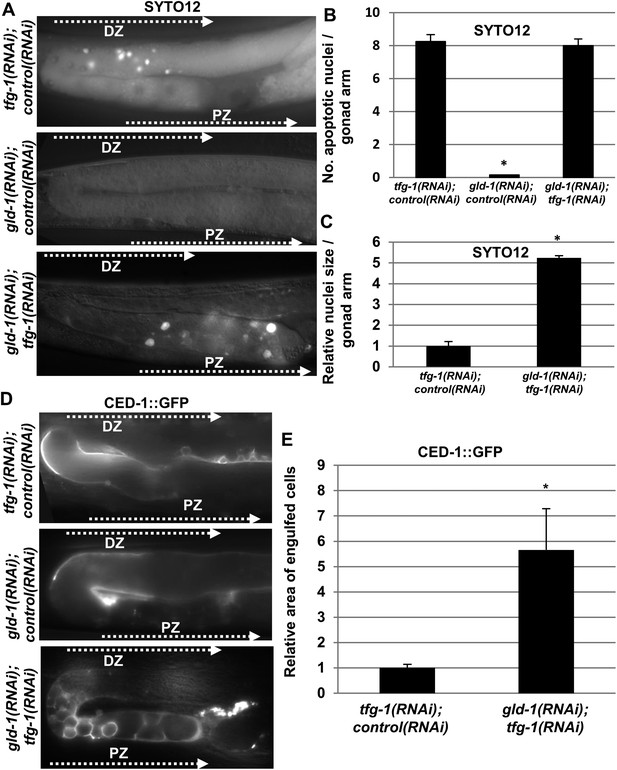
Apoptotic corpses are detected in the gonads of gld-1-deficient animals upon induction of ER stress.
(A–C) Day-3 animals treated with the indicated RNAi were stained with SYTO12 to detect apoptotic cell corpses. The average number of SYTO12-labeled apoptotic corpses per gonad is shown in B. The relative size of the SYTO12-labeled nuclei is shown in C. See Figure 1—figure supplement 1 for SYTO12-labeling of tunicamycin treated animals. (D, E) CED-1::GFP expressed in the gonadal sheath cells was used to follow engulfment of apoptotic cells within the gonad of day-3 animals. The relative average area of the engulfed cells is shown in E. Note that in non-tumorous animals the apoptotic cells are detected in the distal gonad zone (DZ), whereas in the ER stressed-tumorous animals they are detected in the proximal gonad zone (PZ). Asterisk marks Student's T-test values of p < 0.001 compared to animals treated with a mixture of control and tfg-1 RNAi. gld-1 RNAi knocked down GLD-1 protein levels to a similar extent upon treatment with control or tfg-1 RNAi (see Figure 1—figure supplement 2). At least 40 gonads of each genotype were analyzed.
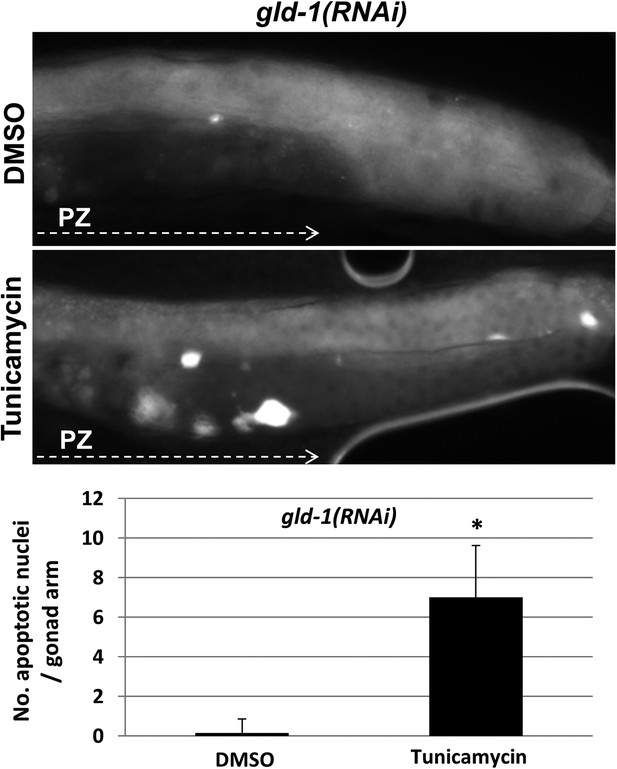
Apoptotic cell corpses are detected in the gonads of tunicamycin-treated tumorous animals.
Representative micrographs showing gonads (x400) of day-3 gld-1 RNAi-treated animals treated with either 45 μg/ml tunicamycin or DMSO as of L4 and stained with SYTO12 to detect apoptotic cell corpses. The average number of SYTO12-labeled apoptotic corpses per gonad is shown.
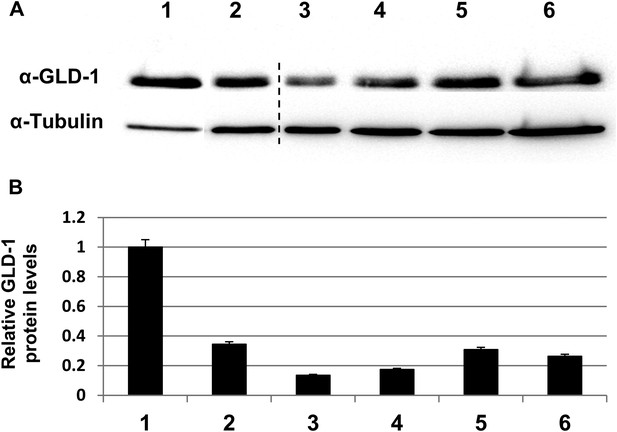
GLD-1 protein levels are efficiently reduced by gld-1 RNAi in the single, double and triple RNAi mixtures.
(A) Representative western blot of GLD-1 and tubulin in day 3 wild-type animals treated with the following RNAi combinations: 1 = control RNAi. 2 = gld-1 RNAi. 3 = gld-1 and control RNAi double mix. 4 = gld-1and tfg-1 RNAi double mix. 5 = gld-1, control and ced-3 RNAi triple mix. 6 = gld-1, tfg-1and ced-3 triple mix. Dashed line indicates removal of irrelevant lanes. (B) Bar graph shows the mean ratio of GLD-1 protein levels normalized to tubulin levels ±SEM in 3 independent experiments. Note that GLD-1levels were efficiently reduced in all RNAi conditions compared to lane 1. Note that GLD-1 levels were similarly reduced upon both conditions of double RNAi treatment (compare lanes 3–4) as well as upon both conditions of triple RNAi treatment (compare lanes 5–6). Furthermore, under all conditions, the gonads of gld-1 RNAi-treated animals appeared tumorous, and lacked oocytes and embryos.
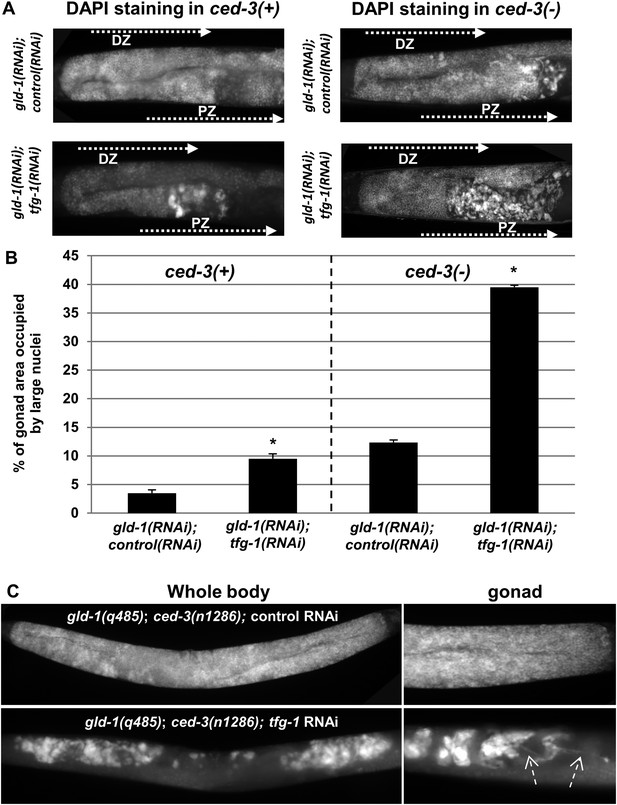
Ectopic cells with large nuclei accumulate in the gonads of gld-1; ced-3 animals.
(A) Representative micrographs (x400) of DAPI-stained gonads of day-4 animals. Animals were treated with the indicated RNAi. gld-1 RNAi was used to induce a germline tumor. ced-3 RNAi served to block apoptosis. tfg-1 RNAi was used to induce ER stress. Treatment with tfg-1 RNAi increased the levels of ectopic cells with large misshaped nuclei at the proximal zone of the gonad of gld-1 deficient animals, especially upon apoptosis inactivation. DZ marks the distal zone of the gonad. PZ marks the proximal zone of gonad. (B) Bar graph presents percentage of gonad area occupied by large nuclei in the indicated genotypes (n = at least 60 gonads per genotype). Asterisks mark Student's T-test values of p < 0.001 of tfg-1 RNAi-treated animals compared to their non-stressed controls. Note that ectopic cells with large nuclei were detected to different extents in most of the animals examined (see Figure 2—figure supplement 1). (C) The induction of ectopic cells in the gonad by tfg-1-induced ER stress was recapitulated in gld-1(q485); ced-3(n1286) double mutants. Arrows point at axon-like structures detected within the gonads of gld-1-deficient animals upon ER stress.
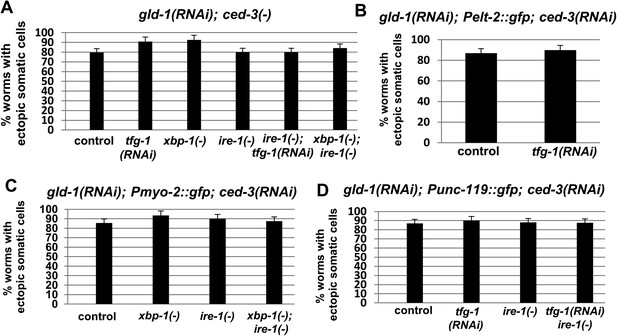
Ectopic somatic cells are detected in most of the gonads of gld-1-deficient animals.
Bar graphs present the percentage of animals treated with a mixture of gld-1 and ced-3 RNAi which contained ectopic cells in their gonads. (A) Ectopic cell were detcted by abnormally large DAPI-stained nuclei in the gonads (n = 50–60 gonads per genotype) (B–D) Ectopic cell were detcted by the expression of transgenic fluorescent somatic markers (n = 50–60 gonads per genotype). Pelt-2::gfp is an intestinal marker. Pmyo-2::gfp is a pharyngal muscle marker. Punc-119::gfp is a neuronal marker.
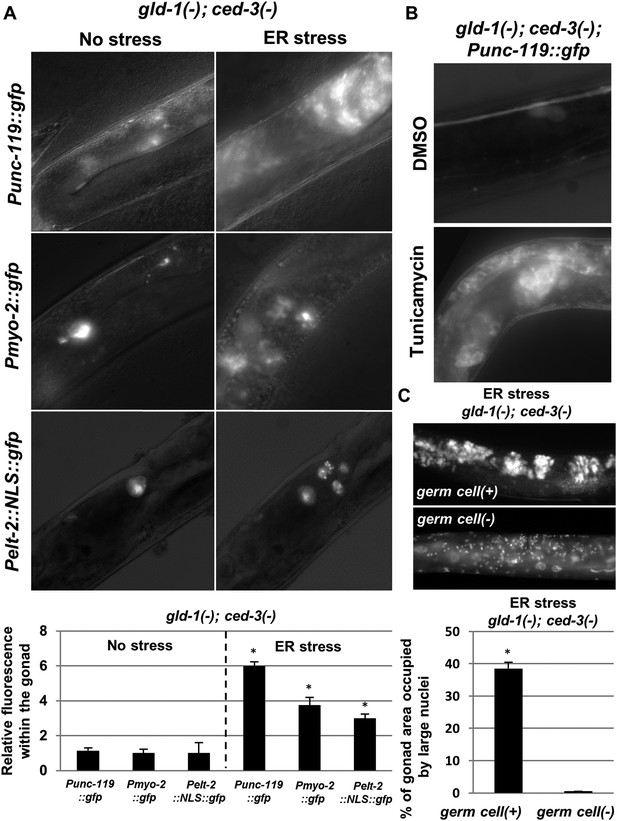
The ectopic somatic cells in the ER-stressed gonad of gld-1-deficient animals are germ cell-derived differentiated somatic cells.
(A) Representative fluorescence micrographs (x400) of somatic differentiation markers expressed in the gonads of gld-1; ced-3-deficient animals on day-4 of adulthood. gld-1 RNAi was used to sensitize the germline for transdifferentiation. ced-3 RNAi was used to prevent the clearance of the ectopic somatic cells from the gonad. Punc-119::gfp is a neuronal marker, Pmyo-2::gfp is a pharyngeal muscle marker. Pelt-2::NLS::gfp is an intestinal marker. ER stress was induced by tfg-1 RNAi or by an xbp-1 mutation. Asterisks mark Student's T-test values of p < 0.001 compared to non-stressed conditions. At least 40 animals were analyzed per genotype. (B) Representative fluorescence micrographs (x400) of Punc-119::gfp in the gonads of ced-3(n1286) day-4 animals treated with gld-1 RNAi. ER stress was induced chemically with tunicamycin and compared to DMSO treatment. (C) Representative fluorescence micrographs of DAPI-stained nuclei of ER-stressed day 4 adults treated with a mixture of gld-1, ced-3 and tfg-1 RNAi. Accumulation of abnormal somatic-like nuclei was detected in germ cell(+) animals and not in germ cell(−) glp-1(−) mutants. At least 50 gonads were analyzed per genotype. Asterisk marks Student's T-test values of p < 0.001. See Figure 3—figure supplement 1 for co-localization of the somatic marker expressing cells and the cells with the large nuclei and/or the cell under engulfment.
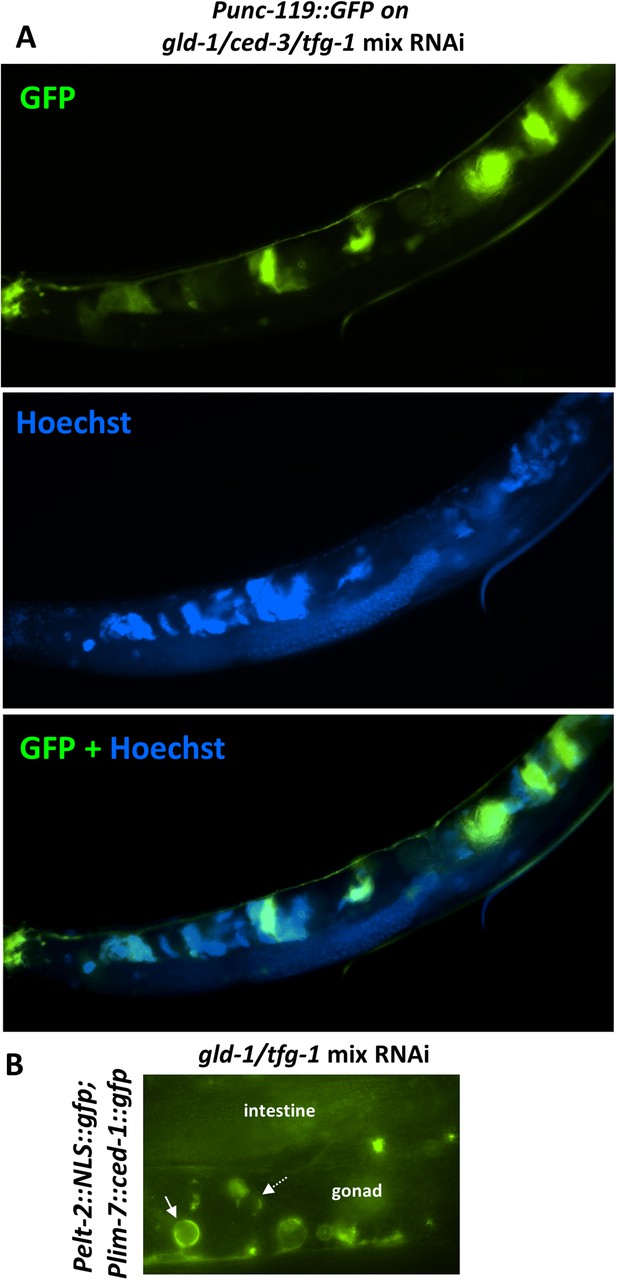
The ectopic somatic cells in the tumorous gonad have large nuclei and are engulfed by the surrounding gonad.
(A) Day 4 Punc-119::gfp transgenic animals treated with a mixture of gld-1, ced-3 and tfg-1 RNAi were stained with the nuclear dye Hoechst. Hoecst staining and GFP expression were individually captured and used to assess co-localization between the pattern of the GFP-expressing cells in the gonad and the cells harboring ectopically large nuclei. (B) Day 3 transgenic animals co-expressing the somatic marker Pelt-2::NLS::GFP and the engulfment marker Plim-7::ced-1::gfp were treated with a mixture of gld-1 and tfg-1 RNAi. Solid arrow indicates an engulfed cell expressing the somatic marker, demonstrating that the ectopic somatic cells in the tumorous gonad are engulfed and removed by the surrounding cells. Dashed arrow indicates an engulfed cell which does not express the somatic marker.
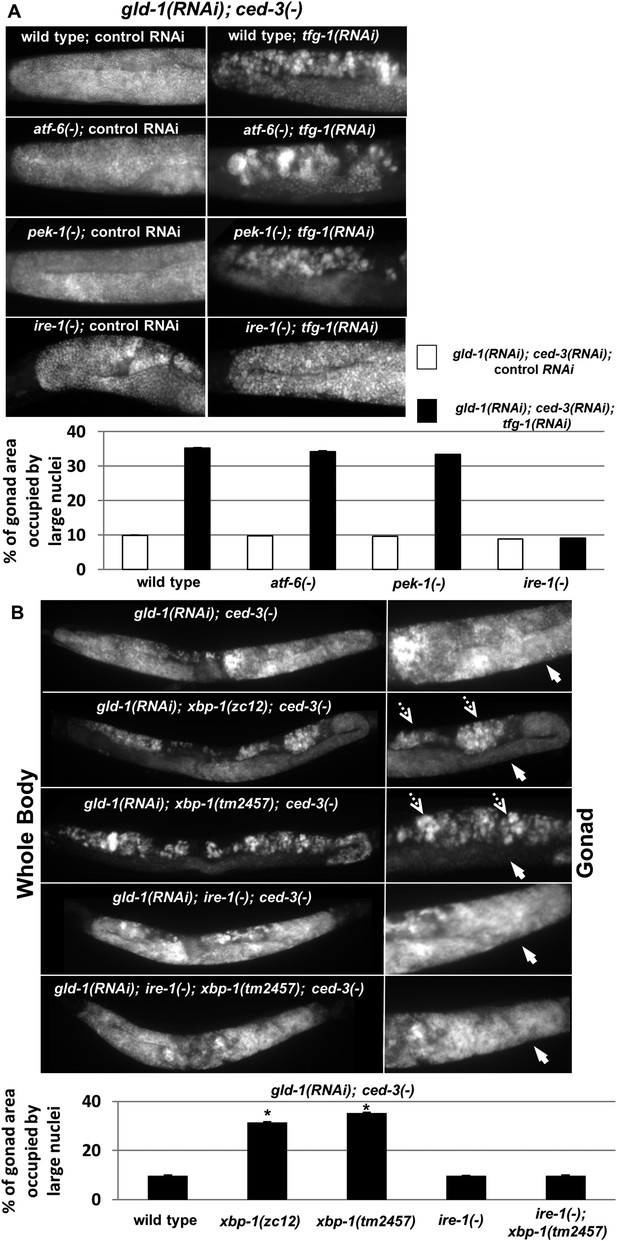
ER stress induces germline transdifferentiation in an ire-1-dependent but xbp-1-independent manner.
(A) Representative micrographs (x400) of DAPI-stained gonads of day-4 animals treated with either a mixture of control, gld-1and ced-3 RNAi or with a mixture of tfg-1, gld-1and ced-3 RNAi. Treatment with tfg-1, gld-1and ced-3 RNAi failed to induce germ cell transdifferentiation in ire-1 mutants. (B) Representative micrographs of whole body (x100) and gonads (x400) of DAPI-stained day-4 animals of the indicated genotypes treated with gld-1 and ced-3 RNAi. Solid arrows indicate mitotic germ cells. Dashed arrows indicate somatic nuclei. Bar graphs present percentage of gonad area occupied by ectopic cells. Asterisk marks Student's T-test of p < 0.001 relative to wild-type animals. Bar graphs present percentage of gonad area occupied by ectopic cells of the indicated genotypes (n = at least 70 gonads per genotype). Asterisks mark Student's T-test of p < 0.001 relative to the same animals treated with control, gld-1and ced-3 RNAi. Note that both alleles of xbp-1 similarly increased the percentage of gonad area occupied by ectopic somatic cells (p = 0.23).
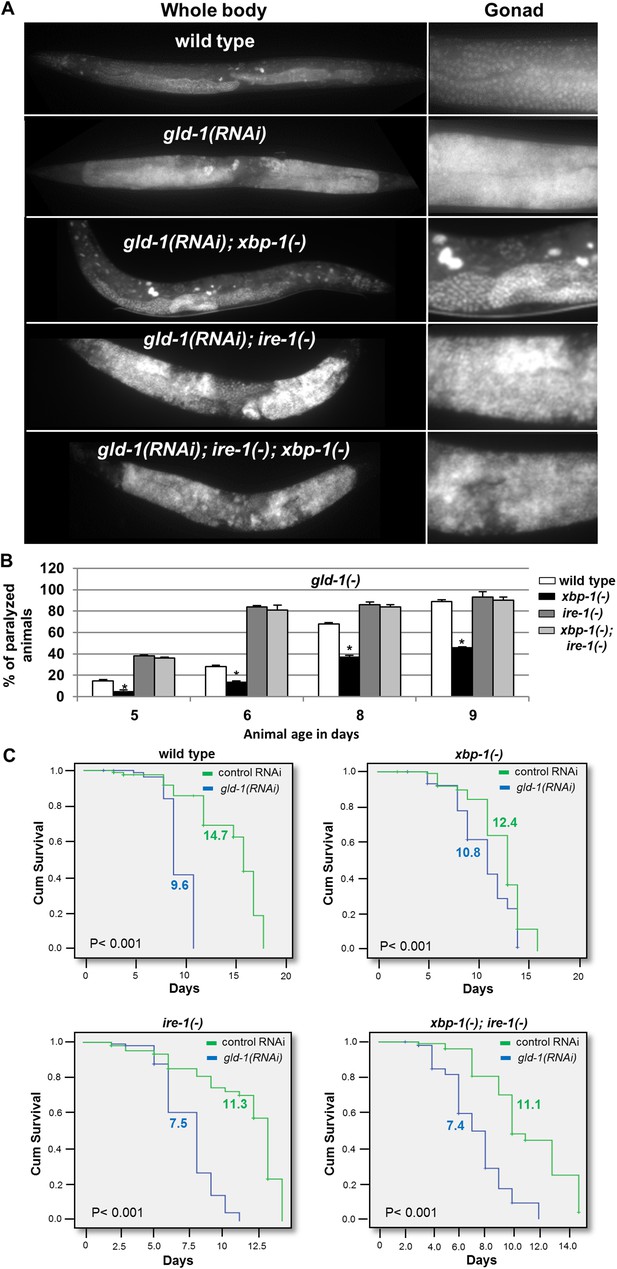
ER stress suppresses the germline tumor in an ire-1-dependent manner.
(A) Representative micrographs showing whole body (x100) and gonads (x400) of day-4 animals stained with DAPI. (B) Paralysis assay in wild type, xbp-1(tm2457), ire-1(ok799) and xbp-1(tm2457); ire-1(ok799) animals. At least 90 synchronized adult animals per genotype were placed on gld-1RNAi plates and their paralysis was scored on days 5, 6, 8 and 9. Bar graphs present percentage of paralyzed animals. At all timepoints the xbp-1 mutation significantly decreased the paralysis of the tumorous animals in an ire-1-dependent manner. Asterisks mark Student's T-test of p < 0.001 for reduced paralysis relative to wild-type animals. (C) Lifespan analysis of wild type, xbp-1(tm2457), ire-1(ok799) and xbp-1(tm2457); ire-1(ok799) animals treated with either gld-1(RNAi) to induce tumor formation or with control RNAi. Lifespan shortening was significantly suppressed in xbp-1 mutants in an ire-1-dependent manner. Mean lifespan and p-values are indicated within each graph. tfg-1 RNAi similarly supressed gld-1-RNAi-induced paralysis and lifespan shortening (see Figure 5—figure supplement 1).
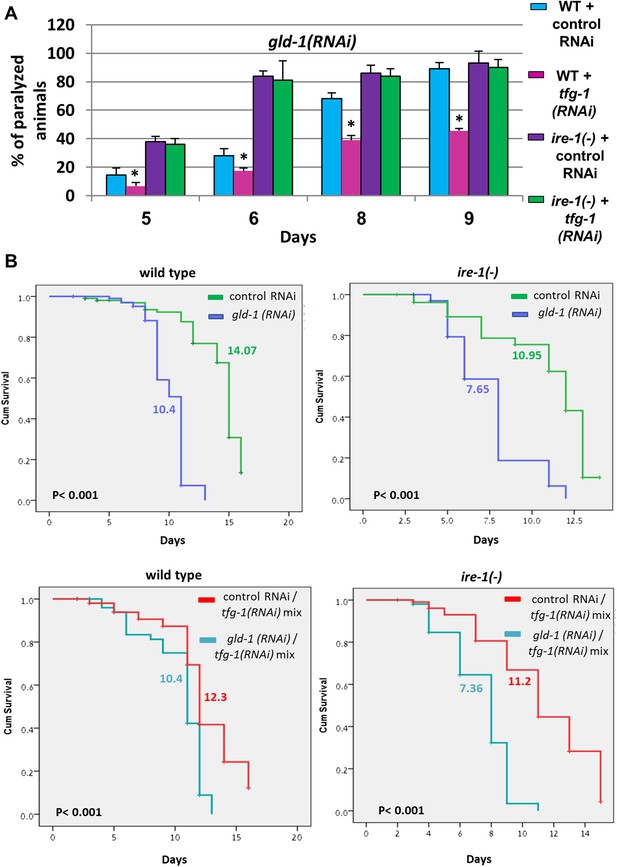
tfg-1 RNAi suppresses the germline tumor in an ire-1-dependent manner.
(A) Paralysis assay in wild type and ire-1(ok799) animals. At least 90 synchronized adult animals per genotype were placed on gld-1/control RNAi plates or gld-1/tfg-1 RNAi plates and their paralysis was scored on days 5, 6, 8 and 9. Bar graphs present percentage of paralyzed animals. At all timepoints the tfg-1 RNAi treatment significantly decreased the paralysis of the tumorous animals in an ire-1-dependent manner. Asterisks mark Student's T-test of p < 0.001 for reduced paralysis relative to wild-type animals. (B) Lifespan analysis of wild type and ire-1(ok799) animals treated with the indicated RNAi combinations. Lifespan shortening by gld-1 RNAi was significantly suppressed in animals co-treated with tfg-1 RNAI in an ire-1-dependent manner. Mean lifespan and p-values are indicated within each graph.
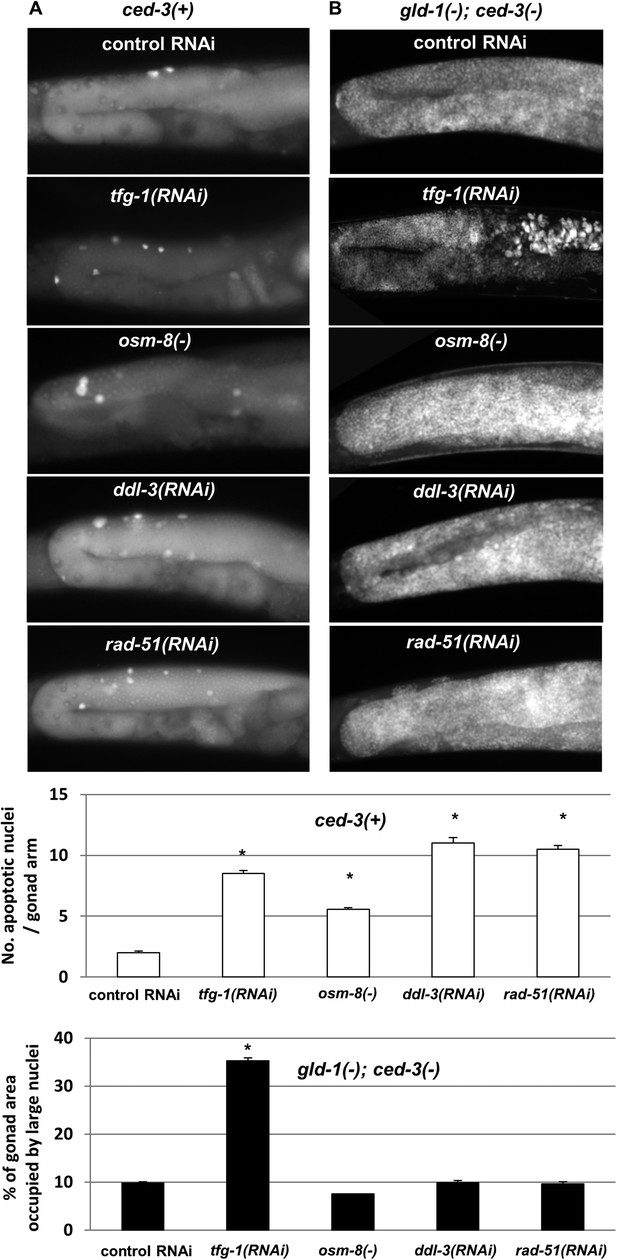
Loss of germ cell pluripotency in gld-1-deficient animals does not occur in response to all stresses.
(A) Representative micrographs (x400) of gonads of ced-3(+) day 2 animals which were stained with SYTO12 to detect apoptotic cell corpses. White bar graphs present number of apoptotic nuclei per gonad arm (n = 60 gonads per genotype). (B) Representative micrographs (x400) of gonads of gld-1(RNAi); ced-3(RNAi) day-4 animals which were stained with DAPI to detect germline and somatic nuclei within the animals' gonads. Black bar graph presents the percentage of gonad area occupied by large nuclei (n = 60 gonads per genotype). ER stress was induced by tfg-1 RNAi. Osmotic stress was induced by osm-8 inactivation. Mitochondrial stress was induced by RNAi targeting ddl-3. Genotoxic stress was induced by rad-51 RNAi. Asterisks mark Student's T-test of p < 0.001 relative to control RNAi-treated animals. Note that the stresses that did not induce germ cell transdifferentiation in gld-1(−); ced-3(−)animals also failed to suppress the germline tumor in gld-1(−); ced-3(+) animals (see Figure 6—figure supplement 1).
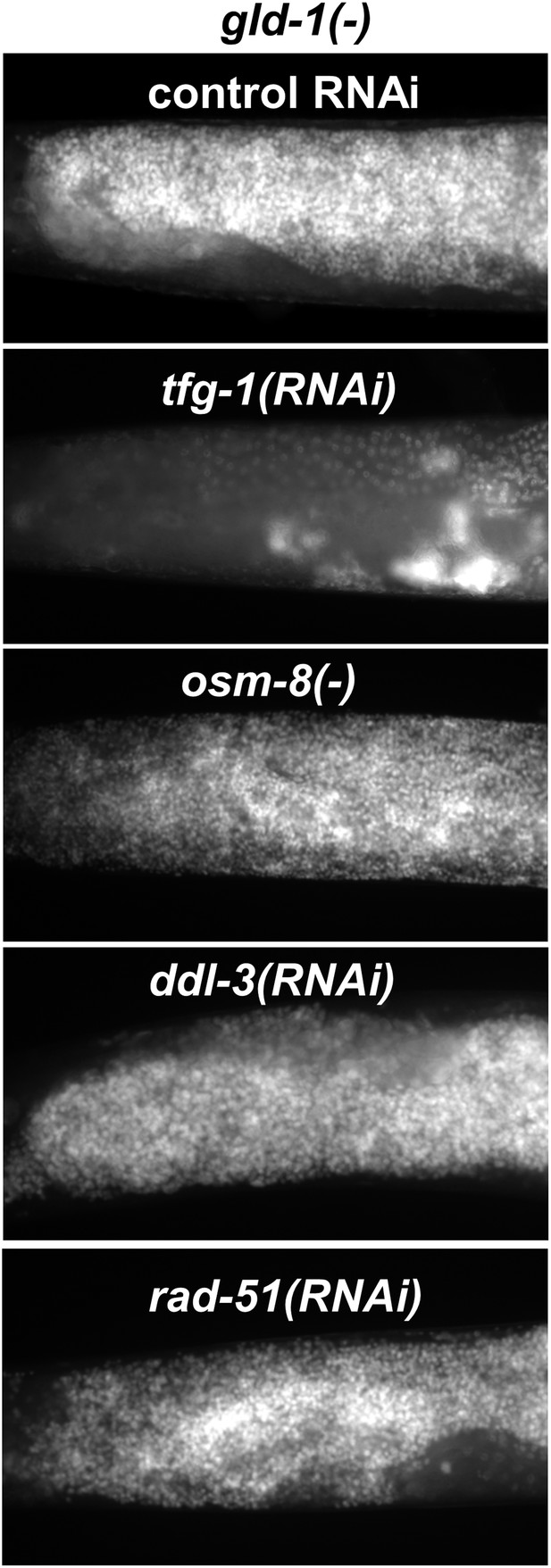
Not all stresses suppress the germline tumor in gld-1-deficient animals.
Representative micrographs (x400) of DAPI-stained gonads of gld-1(RNAi) day-4 animals (n = at least 50 gonads per genotype). Note that the ability to execute apoptosis has not been manipulated in these animals, in contrast to the animals in Figure 6B. ER stress was induced by tfg-1 RNAi. Osmotic stress was induced by osm-8 inactivation. Mitochondrial stress was induced by RNAi targeting ddl-3. Genotoxic stress was induced by rad-51 RNAi. Only tfg-1 RNAi suppressed tumor progression.
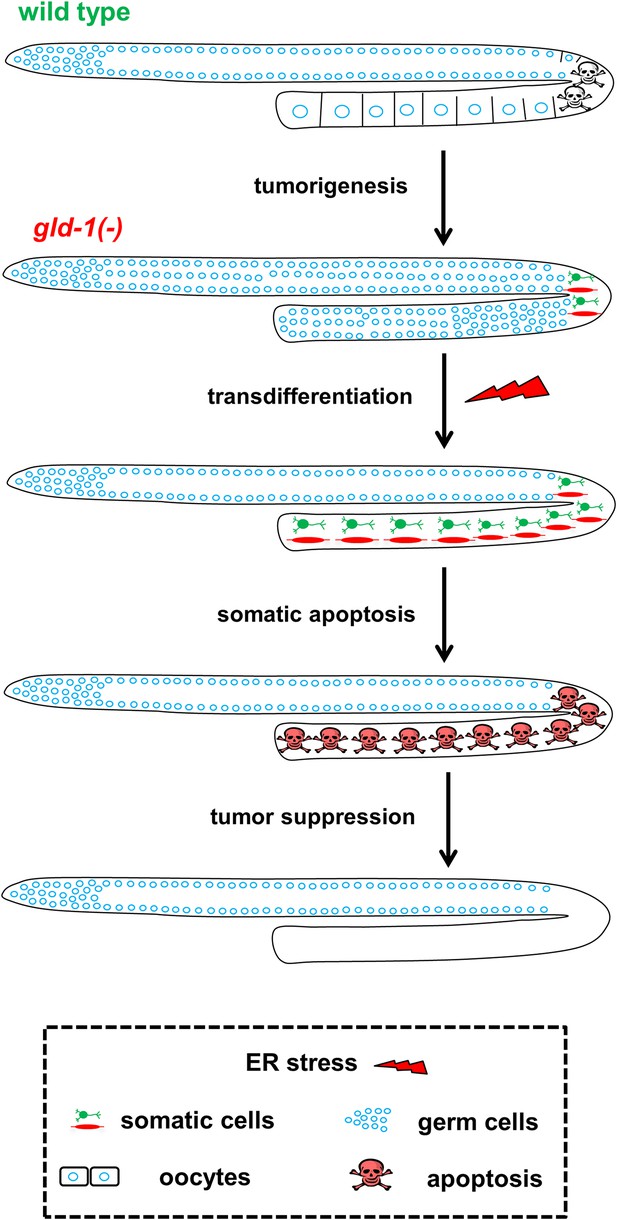
Model—tumor progression and germline fate in ER stressed tumorous gonads.
In animals with a normal germline, ER stress induces germ cell apoptosis. Inactivation of the gld-1 genes alters many aspects of germ cell fate: differentiation of germ cells into oocytes is abrogated, germ cell proliferation is enhanced, a germline tumor is formed and the germ cells lose their responsiveness to execute physiological and stress-induced apoptosis. Furthermore, in gld-1 deficient animals the germ cells are prone to generate teratoma as they become sensitized to precociously transdifferentiate into somatic cells. Under these conditions ER stress can suppress and limit the germline tumor. This suppression is achieved by enhancing germline transdifferentiation into ectopic somatic cells. Soon after the transdifferentiation, these ectopic cells undergo apoptosis, and are removed from the gonad, suppressing the germline tumor.





