Vilya, a component of the recombination nodule, is required for meiotic double-strand break formation in Drosophila
Figures
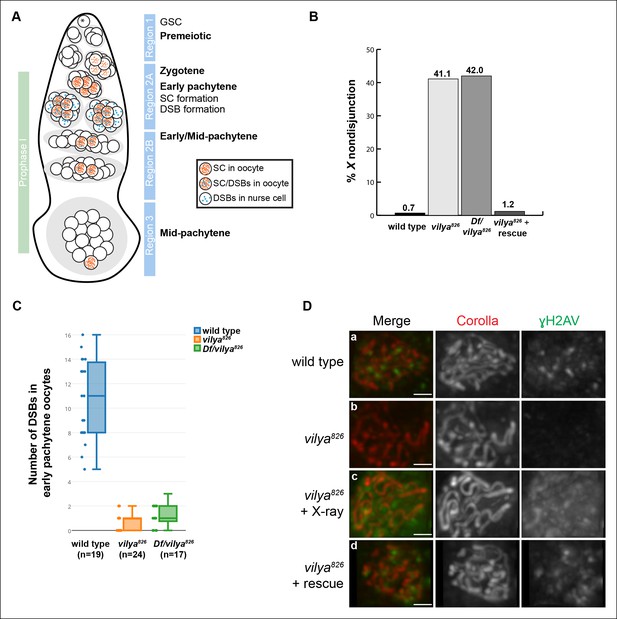
vilya encodes a RING domain-containing protein required for DSB formation.
(A) Schematic diagram of a germarium showing the timing of SC and DSB formation. (B) vilya826 homozygotes and Df/vilya826 transheterozygotes cause high levels of X chromosome nondisjunction. The high level of X nondisjunction in vilya826 is almost completely rescued by expressing vilya3XHA in the female germline. The deficiency that uncovers vilya used in the analysis was Df (1)ED6630. vilya826 + rescue refers to the genotype y w vilya826 nos-Gal4/vilya826; PUASp-vilya3XHA/+. Wild type and vilya826 nondisjunction rates are from (Collins et al., 2012). (C) vilya826 and Df/vilya826 are defective in DSB formation in early pachytene oocytes as identified by an antibody against γH2AV and compared to wild type. DSBs in region 2A nurse cells are also significantly reduced in vilya mutants (see Figure 1—figure supplement 4). (D) Region 2A oocyte nuclei stained with Corolla (red) and γH2AV (green) in wild type, vilya826, vilya826exposed to X-ray and vilya826 + vilya3XHA germline rescue construct. Images are maximum intensity projections of deconvolved z-series through the selected nuclei. Scale bar, 1 µm.
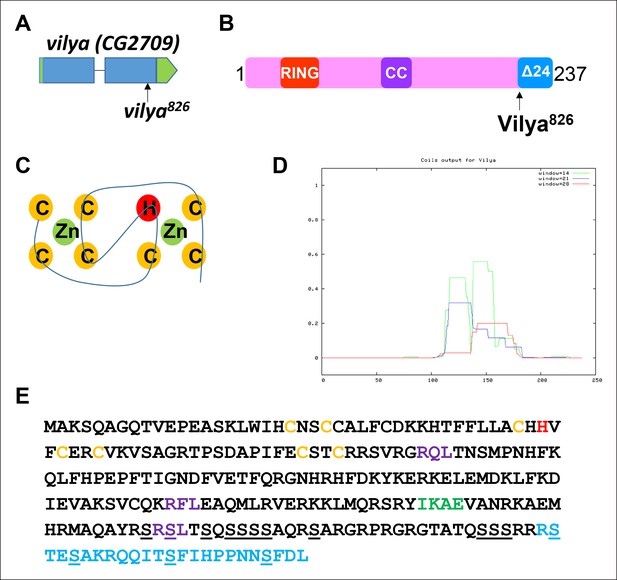
vilya, CG2709, encodes a RING domain-containing protein.
(A) mei-826 (Collins et al., 2014) was mapped to CG2709 (Materials and Methods) and renamed vilya826. (B) vilya is predicted to encode a 237 amino acid protein with a RING domain and a potential internal coiled-coil domain. vilya826 allele is predicted to truncate the protein at amino acid 213. (C) Shown is the structural RING domain consisting of Cys3HisCys4 binding to two zinc (Zn) cations. (D) Vilya is predicted to contain an internal coiled-coil region based on the COILS program (Lupas et al., 1991). (E) Vilya protein sequence is shown with cysteine and histidine residues of the RING domain (yellow and red), the mutation (R213STOP) in vilya826 (blue), a predicted SUMO-interacting motif (green), three potential RXL motifs for mediating cyclin binding (purple), and the serines in the serine-rich C-terminal domain (underlined). The last quarter of Vilya is 25% serines.
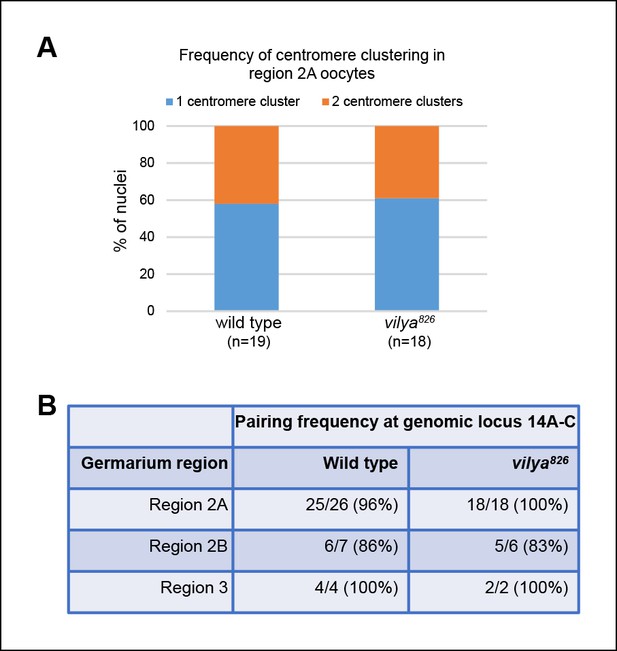
Centromere clustering and homolog pairing is not affected in vilya826.
(A) Using an antibody to the CENP-A homolog, CID, clustering of centromeres is unaffected in vilya826 compared to wild type in region 2A. 100% of region 2A oocytes analyzed (n) for both wild type and vilya826 contain two or less centromere clusters. (B) FISH analysis of an X chromosomal probe at region 14A-C indicates that homolog pairing is normal throughout pachytene in vilya826 when compared to wild type. Nuclei with either a single focus or foci separated by less than 0.75 µm were defined as paired. Those foci with centers separated by more than 0.75 µm were considered unpaired.
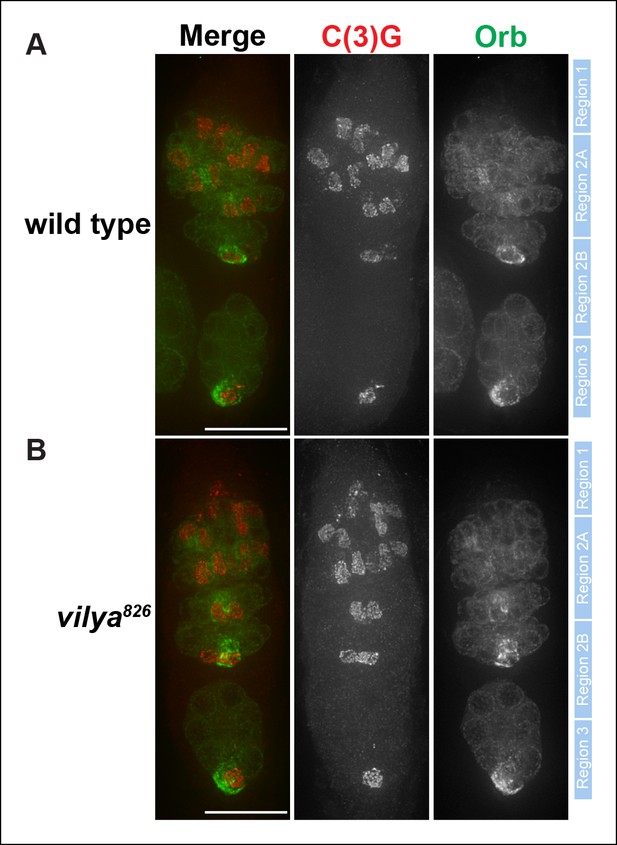
C(3)G and Orb staining appears normal in vilya826.
Immunofluorescence analysis of wild-type (A) and vilya826mutant (B) germaria showing the timing of SC formation, SC structure and oocyte determination. The SC is labeled with an antibody to C(3)G (red). By region 2B the cytoplasm of the oocyte becomes concentrated with Orb (green). The position of regions 1 through 3 are labeled to the right of each germarium. Scale bar, 15 µm.
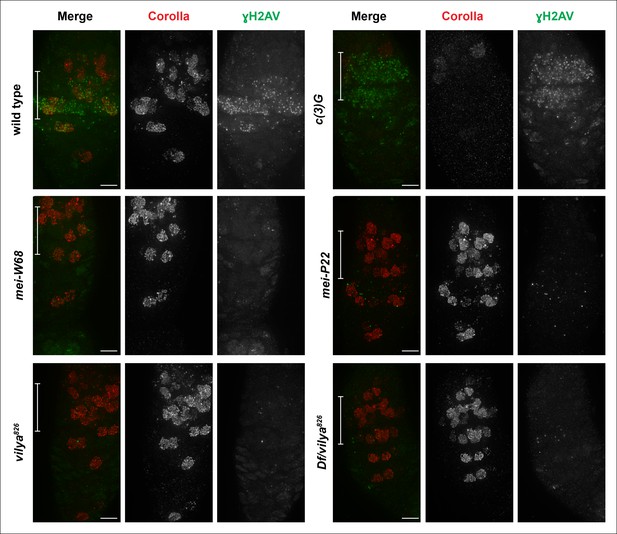
Vilya plays a direct role in DSB formation in early pachytene.
Immunofluorescence analysis comparing the induction and location of DSBs as marked by an antibody to γH2AV (green) in both surrounding nurse cells and oocyte nuclei (identified with an antibody to Corolla (red)) of wild type, c (3)G, mei-W68, mei-P22, vilya826 and Df/vilya826. In each genotype, region 2A is identified by the white bar on the merged image. γH2AV foci are readily identifiable in region 2A in both wild type and c (3)G indicating that DSBs are induced. No, or very few, DSBs can be identified with the γH2AV antibody in any region 2A nuclei (oocytes or surrounding nurse cells) in mei-W68, mei-P22 or in the vilya mutants, suggesting that vilya’s function is required for the induction of DSBs during early pachytene. Images are maximum intensity projections of deconvolved z-series through the entire germarium. Scale bar, 5 µm.
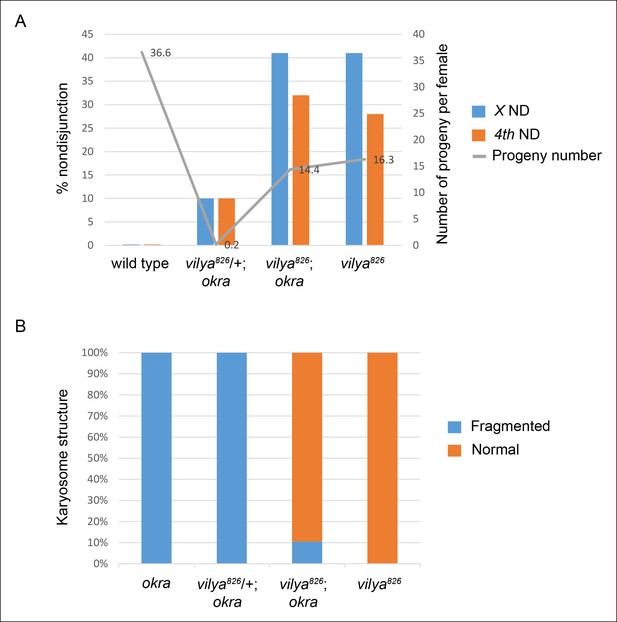
vilya826 rescues the fertility and karyosome defects of the DSB repair-deficient mutant okra.
(A) vilya826 rescues the fertility defect of the DSB repair-deficient mutant okra and displays an increase in chromosome nondisjunction in the double mutant, similar to that of the single vilya826 mutant. The fertility of vilya826 is only about 30% of the wild-type control, likely due to the high levels of chromosome missegregation. The high levels of 4th chromosome nondisjunction observed in the vilya mutant are due to the inability of the achiasmate segregation system to withstand the effects of a global reduction in recombination (Zitron and Hawley, 1989; Hawley et al., 1992). Average number of progeny per female (gray line) is shown. Number of adjusted progeny scored in the nondisjunction assay: wild type (330), vilya826/+; okra (20), vilya826; okra (1587) and vilya826 (195). Number of females tested in the fertility assay: wild type (9), vilya826/+; okra (90), vilya826; okra (110) and vilya826 (12). Wild type and vilya826 data collected independently from other genotypes. ND, nondisjunction. (B) vilya826 rescues the karyosome defect seen in the okra mutant to 89.5% of normal. Number of karyosomes analyzed: okra (38), vilya826/+; okra (20), vilya826; okra (12), and vilya826 (6). (A,B) ( + ) indicates wild-type copy of vilya present on FM7 balancer chromosome.
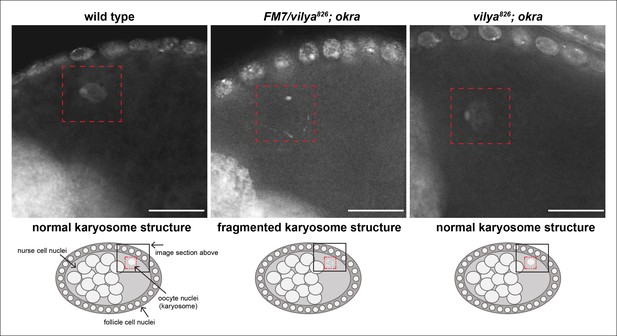
vilya826 rescues the karyosome defect of the DSB repair-deficient mutant okra.
The karyosome structure of a wild-type stage 8 egg chamber is shown for comparison purposes. A schematic of the egg chamber is shown below each image to identify the region highlighted in the image. The karyosome structure is fragmented in the DSB repair-deficient mutant okra when one copy of wild-type vilya is present (vilya826/FM7; okra). vilya826 rescues the karyosome defect seen in okra mutants (vilya826; okra), indicating that DSB formation is suppressed or abolished. Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. Stage 8 egg chambers are stained with DAPI (white) only. Karyosome is identified by a red dashed box. Scale bar, 15 µm.
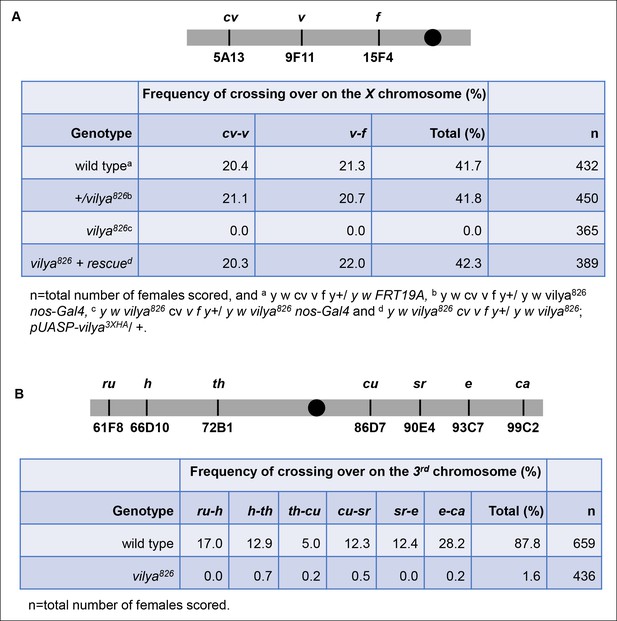
vilya826 is defective in meiotic recombination.
(A) vilya826 is defective in meiotic recombination as assayed for intervals cv-v and v-f on the X chromosome. (B) Recombination frequency across the entire 3rd chromosome in vilya826 is reduced over 50-fold compared to wild type.
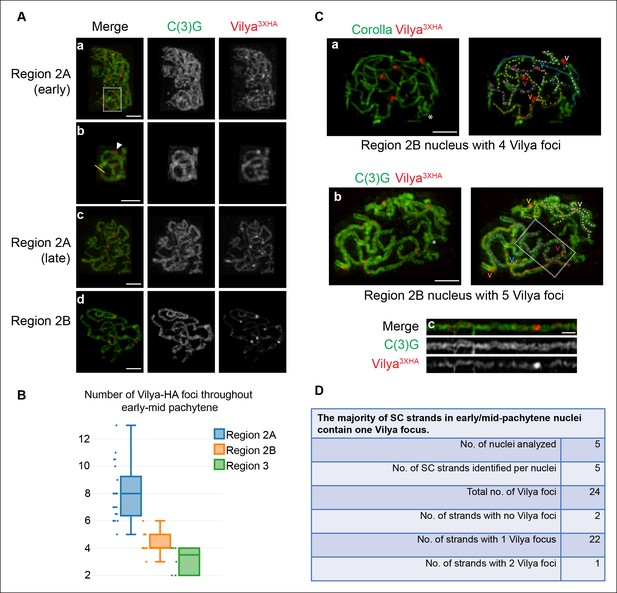
Vilya localizes to the central region of the SC in both linear elements and discrete foci.
(A) Localization of Vilya3XHA throughout early pachytene as assayed by germline expression of vilya3XHA using antibodies to the transverse filament protein C(3)G (green) and an antibody to HA (red). Images are maximum intensity projections of deconvolved z-series from a DeltaVision OMX microscope through the selected nuclei. Scale bar, 1 µm. (A-a) Early pachytene (region 2A) oocyte nucleus showing that Vilya localizes to the central region of the SC in both linear strands and discrete foci. (A-b) Higher magnification of the white dashed box in A showing Vilya3XHA clearly positioned in the central region between the two tracks of C(3)G (yellow line) and a discrete Vilya3XHA focus sitting within and above a stretch of SC (arrowhead). (A-c,d) Localization of Vilya3XHA in region 2A and 2B showing the discrete foci and SC staining. Note in region 2B the SC shortens. (B) Analysis of the number of Vilya3XHA foci throughout early/mid-pachytene. (C-a,b) Traces of SC between homologous chromosome arms in early/mid-pachytene (region 2B) nuclei expressing vilya3XHA. Images are maximum intensity projections of deconvolved z-series from a DeltaVision OMX microscope through the selected nuclei. (*) Indicates the chromosome center containing pericentric heterochromatin and is the location of the centromeres. Scale bar, 1 µm. Individual tracks of SC between homologous chromosome arms were identified and each labeled with a separate color. The corresponding Vilya3XHA foci associated with each stretch of SC between homologous chromosome arms are labeled by a (v) in the same color as the stretch of SC it is on. (C-a) The oocyte nucleus is labeled with antibodies to Corolla (green) and HA (red). This nucleus has five colored chromosome arms and four Vilya3XHA foci. Each chromosome arm has been linearized in Figure 3—figure supplement 1D. (C-b) Oocyte nucleus is labeled with antibodies to C(3)G (green) and HA (red). This nucleus has five colored chromosome arms and five Vilya3XHA foci. (C-c) The chromosome arm outlined with the white dashed line in (C-b) has been linearized. (D) The majority (92%) of Vilya3XHA foci in the five nuclei that have been identified as having five clearly identifiable chromosome arms each localize to one strand. One chromosome arm contains two foci, and two chromosome arms contain no Vilya3XHA foci.
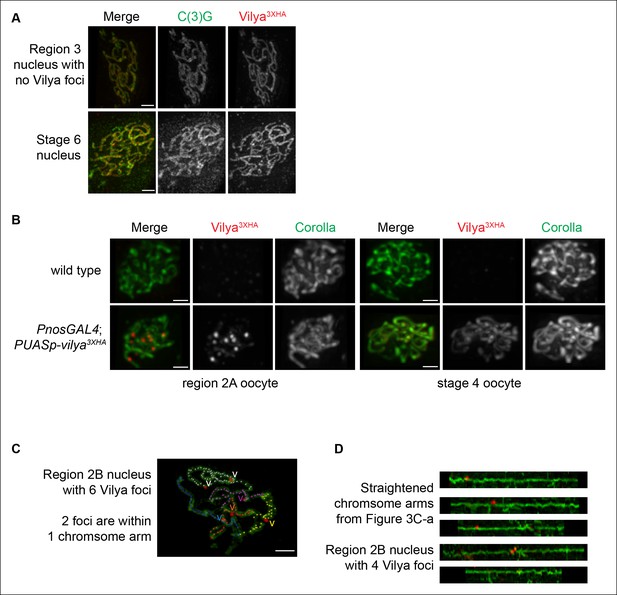
Localization of Vilya3XHA within pachytene nuclei.
(A) Immunolocalization of Vilya3XHA in a mid-pachytene (region 3) nucleus that did not have any distinct Vilya3XHA foci and a late pachytene (Stage 6) nucleus showing that once the discrete foci disappear in pachytene, the localization of Vilya3XHA is exclusively uniform throughout the central region of the SC. Nuclei are stained with antibodies to C(3)G (green) and HA (red). Images are maximum intensity projections of deconvolved z-series from a DeltaVision OMX microscope through the selected nuclei. Scale bar, 1 µm. (B) Immunofluorescence analysis showing the specificity of the anti-HA antibody to Vilya3XHA protein. A region 2A image is shown for both wild-type and vilya3XHA-expressing oocytes, as well as a stage 4 oocyte that has only the linear staining pattern. Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. See Materials and Method for details regarding image acquisition on wild-type tissue. Scale bar, 1 µm. (C) Traces of SC between homologous chromosome arms in an early/mid-pachytene region 2B nucleus expressing vilya3XHA. Image is a maximum intensity projection of a deconvolved z-series from a DeltaVision OMX microscope through the selected nucleus. Scale bar, 1 µm. Individual chromosome arms were identified and each labeled with a separate color. The corresponding Vilya3XHA foci associated with each chromosome arm are labeled by a (v) in the same color as the stretch of SC they are on. Notice in this nucleus the chromosome arm labeled in white contains two Vilya3XHA foci spaced some distance apart from one another. (D) Linearized chromosome arms from Figure 3C-a.
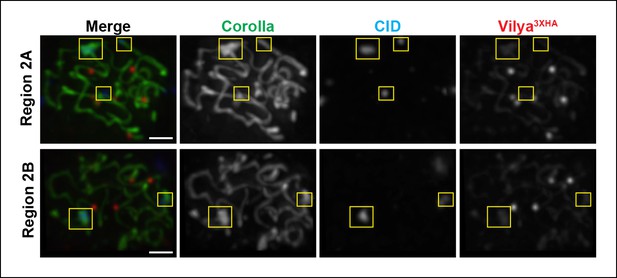
Vilya3XHA foci are not found at centromeres in early/mid-pachytene.
Immunolocalization of Vilya3XHA and CID in vilya3XHA expressing early/mid-pachytene oocytes showing the absence of foci at centromeres. Pachytene nuclei in the specified regions were labeled with antibodies to HA (mouse) (red), Corolla (green) and CID (blue). Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. Boxes mark the centromere clusters in each nucleus shown. Scale bar, 1 µm.
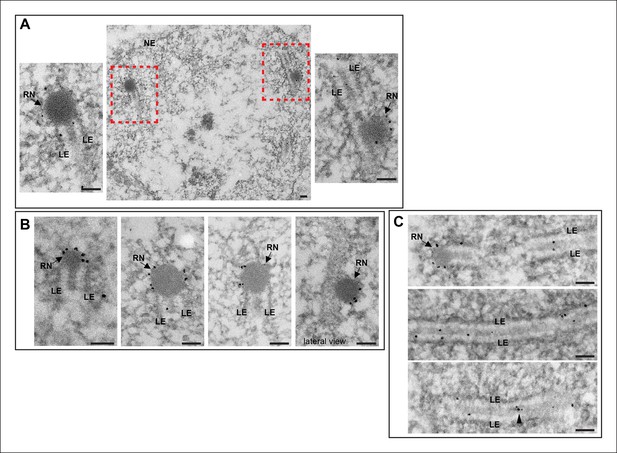
Immuno-EM of Vilya3XHA shows localization to both the RNs and to the central region of the SC.
Immuno-gold labeling of Vilya3XHA from germline-expressed PUASp-vilya3XHA ovaries. (A) A low magnification image of a section from a single nucleus with two RNs (outlined with red dashed box). A higher magnification of each RN with associated gold particles is also shown. (B) Four additional immuno-EM images showing gold particles associated with RNs. A lateral view of an RN is also shown. (C) Three immuno-EM images showing gold particles distributed throughout the central region of the SC, as well as at RNs. Arrowheads point to cluster of gold particles in what appears to be a small electron-dense region in the central region. NE, nuclear envelope; RN, recombination nodule; LE, lateral element. Scale bar, 100 nm.
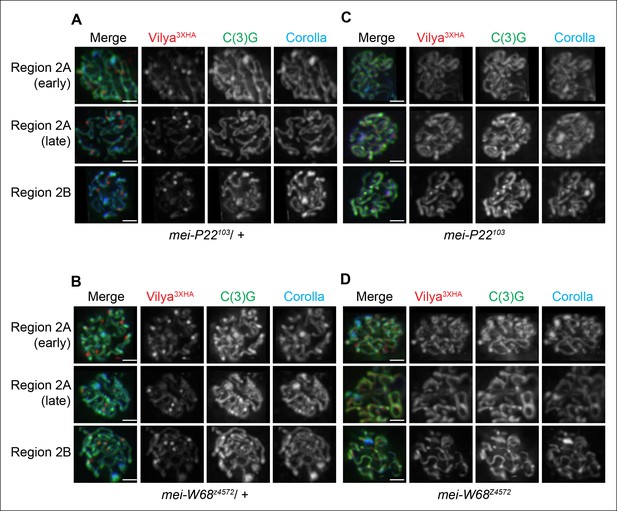
Localization of Vilya3XHA to discrete foci is dependent on the process of DSB formation.
(A-D) Immuno-localization of Vilya3XHA foci in the presence and absence of DSB formation. Pachytene nuclei in the specified regions were labeled with antibodies to HA (red), C(3)G (green) and Corolla (blue). Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. Scale bar, 1 µm. (A-B) Germline expression of PUASp-vilya3XHA in the presence of DSB formation. (A) y w nos-Gal4/w; PUASp-vilya3XHA/+; mei-P22103/+. (B) y w nos-Gal4/w; PUASp-vilya3XHA/mei-W68z4572. (C,D) Germline expression of PUASp-vilya3XHA in the absence of DSB formation. (C) y w nos-Gal4/w; PUASp-vilya3XHA/+; mei-P22103. (D) y w nos-Gal4/w; PUASp-vilya3XHA mei-W68z4572/mei-W68z4572.
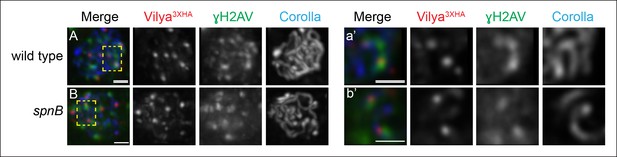
A subset of Vilya3XHA foci localize near γH2AV staining at DSB sites.
(A,B) Immunofluorescence analysis of Vilya3XHA (red) localization at the sites of programmed DSBs that are recognized with the γH2AV modification (green) and Corolla (blue) in region 2A (A) or region 3 (B) pachytene nuclei. Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. (A-a’, B-b’,) Higher magnification of a single z-section from a small region outlined in the yellow box of the corresponding image A-B, respectively, showing the close association of Vilya3XHA with γH2AV marks. Genotype for (A) wild type in this figure refers to the genotype y w nos-Gal4/+; PUASp-vilya3XHA/+; mei-P22103/+. mei-P22 is not haploinsufficient. (B) y w nos-Gal4/+; PUASp-vilya3XHA/+; spnB. Scale bar, 1 µm.
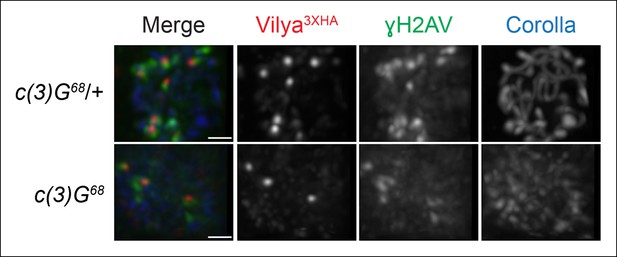
Localization of Vilya3XHA to discrete foci in early pachytene is not dependent on the SC.
Immunofluorescence analysis of vilya3XHA-expressing region 2A oocytes in the presence and absence of c(3)G. Early pachytene nuclei in region 2A were labeled with antibodies to HA (red), γH2AV (green) and Corolla (blue). Images are maximum intensity projections of deconvolved z-series from a DeltaVision microscope through the selected nuclei. Scale bar, 1 µm.
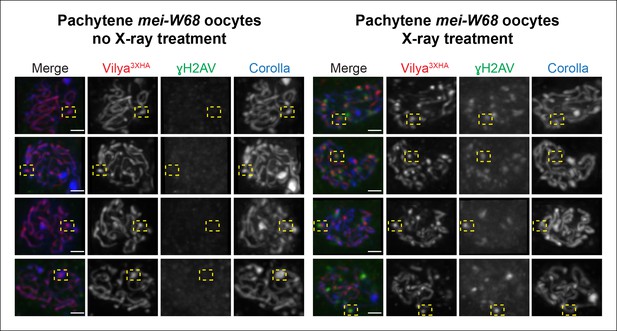
Some DSBs created by X-ray can recruit Vilya3XHA to discrete foci.
Immunofluorescence analysis of germline expression of PUASp- vilya3XHA in the absence of functional mei-W68 with and without X-ray treatment. Ovaries were stained with antibodies to HA (red), γH2AV (green) and Corolla (blue). Four examples of early/mid-pachytene oocytes are shown for each treatment. Yellow boxes in the no X-ray treatment show examples of concentrated regions of linear Vilya3XHA that are associated with dense Corolla staining. Yellow boxes in the X-ray treatment show examples of discrete Vilya3XHA foci associated with γH2AV marks that are not associated with dense Corolla staining. Scale bar, 1 µm.
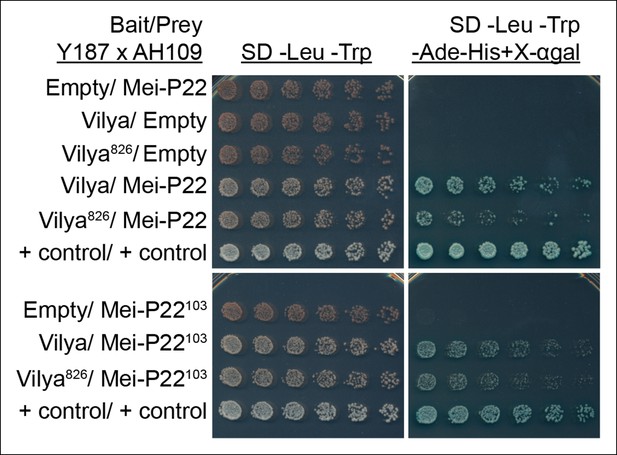

Vilya and Mei-P22 interact by yeast two-hybrid.
Yeast two-hybrid was used to test for an interaction between Vilya and Mei-P22. All diploid strains, where the OD600 was equalized before plating, grow equally well under selection for both the bait and prey plasmids (SD -Leu-Trp). Six two-fold dilutions for each diploid were plated on each selection plate. Vilya and Mei-P22 strongly interact on the reporter plate (SD -Leu-Trp-Ade-His + X-αgal). Vilya826 and Mei-P22 also interact, but it appears to be a weaker interaction than with full-length Vilya on the reporter plate. Vilya and Mei-P22103 interact, as well as Vilya826 and Mei-P22103, although this interaction is also weaker than with full-length Vilya. No interaction was detected with empty construct for any of the plasmids used. The control plasmids are pGBKT7-53 and pGADT7-T supplied by Clontech.
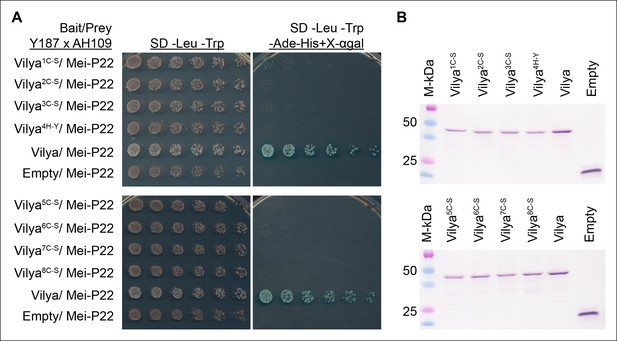

A functional RING domain is required for Vilya to interact with Mei-P22 in yeast-two hybrid assay.
(A) Mutations in any of the key residues in the RING domain ablate the ability of Vilya to interact with Mei-P22. (B) Western blot analysis showing that the RING domain mutants are expressed in the Y187 strain used as the bait in (A). GAL4-BD-cMyc-Vilya protein is predicted to be 49 kDa. GAL4-BD-cMyc protein (empty vector) is predicted to be 22 kDa.
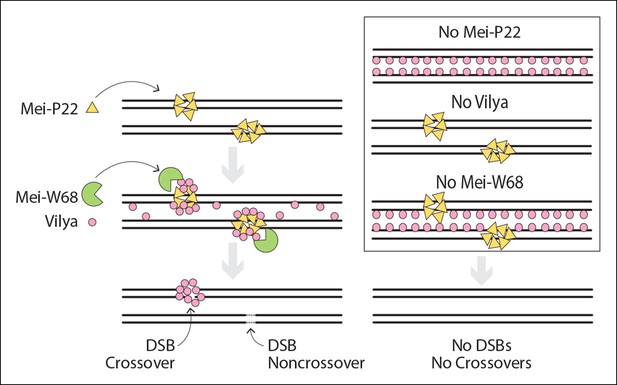
Model of DSB formation in Drosophila female oocytes.
(Left) In wild-type oocytes, Mei-P22, which localizes to discrete foci prior to the time γH2AV foci are present, is located at chromatin adjacent to the SC (Liu et al., 2002). Vilya localizes to the central region of the SC and is required along with its binding partner, Mei-P22, and Mei-W68 (the Spo11 homolog) for formation of DSBs. Although initially Vilya may localize to a vast majority of, if not all, DSBs, as the oocytes mature into early/mid-pachytene, Vilya is retained and/or recruited to form discrete foci at sites of crossing over. Failure to accumulate Vilya at DSB sites would direct that DSB to a noncrossover fate. (Right) In the absence of Mei-P22 or Mei-W68, and thus in the absence of DSB formation, Vilya fails to localize to discrete foci and is found exclusively along the central region of the SC. In the absence of Vilya, we speculate, based on the fact that Mei-P22 can localize to discrete foci in the absence of Mei-W68 (Liu et al., 2002), that Mei-P22’s localization is unaffected. However, DSB formation would fail due to the absence of vilya function. In the absence of Mei-W68, Mei-P22 is able to bind normally (Liu et al., 2002), however, due to the absence of DSBs, Vilya does not form discrete foci. In all these instances, crossovers do not form.
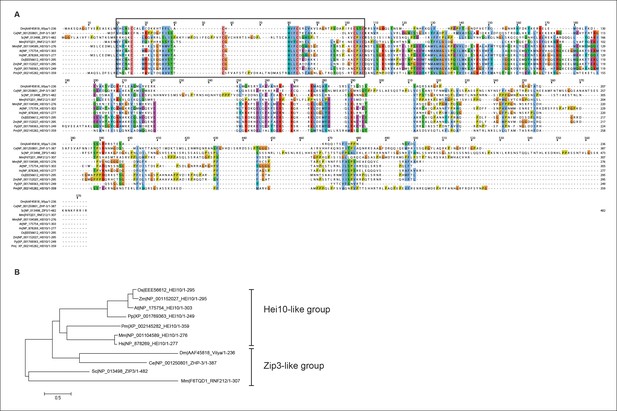
Protein alignment of Vilya to Zip3 and HEI10 homologous proteins.
(A) Protein alignment of Drosophila melanogaster (Dm) Vilya (AAF45818), Caenorhabditis elegans (Ce) ZHP-3 (NP_001250801), Saccharomyces cerevisiae (Sc) Zip3 (NP_013498), Mus musculus (Mm) RNF212 (F6TQD1) and HEI10 (NP_001104589), Arabidopsis thaliana (At) HEI10 (NP_175754), Homo sapiens (Hs) HEI10 (NP_878269), Oryza sativa (Os) HEI10 (EEE56612), Zea mays (Zm) HEI10 (NP_001152027), Physcomitrella patens (Pp) HEI10 (XP_001769363) and Penicillium marneffei (Pm) HEI10 (XP_002145282). Proteins were aligned and visualized using Muscle and ClustalX programs in Jalview (http://www.jalview.org). Black box corresponds to the region surrounding the RING domain. (B) Maximum likelihood tree constructed from the sequences above using LG/G + I model (best fit model identified with MEGA 6 (http://www.megasoftware.net)) (Hall, 2013). This maximum likelihood tree appears to have similar grouping to that reported in the BLAST similarity network by Chelysheva et al. (Chelysheva et al., 2012) showing the HEI10 proteins as one group and the Zip3-like proteins (including Zip3, ZHP-3 and RNF212) as the other group. In this analysis Vilya is positioned within the Zip3-like group.
Videos
Rotation of an early/mid-pachytene nuclei.
Movie showing X and Y rotation of the nucleus in Figure 3C-b. Each chromosome arm is marked with a separate color. The SC is labeled using an antibody to C(3)G (green), and Vilya3XHA foci are identified with an antibody to HA (red).





