GGGGCC microsatellite RNA is neuritically localized, induces branching defects, and perturbs transport granule function
Figures
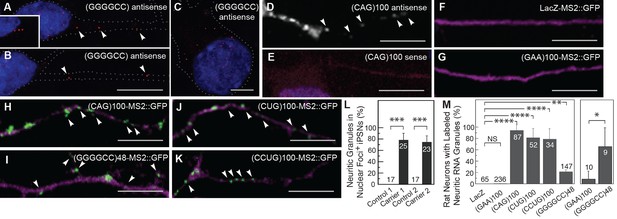
The GGGGCC repeat and other microsatellite RNA repeats with high secondary structure content are neuritically localized.
(A–B) GGGGCC repeat RNA is neuritically localized in discrete granules in human iPSNs derived from C9orf72 GGGGCC repeat expansion carriers. (A–C) GGGGCC repeat RNA (red) was detected with a (GGCCCC)4 antisense probe. Neuritic GGGGCC RNA granules (arrowheads) were observed (A) proximally, (inset in A) as linear arrays, and (B) distally. (C) GGGGCC RNA granules in the cell body. (A–C) iPSNs from carrier 2 are shown. Neurites and cell body are outlined (dotted line). (D–E) CAG repeat RNA is localized to neuritic granules in primary rat spinal cord culture. In situ hybridization of primary rat spinal cord neurons transfected with (CAG)100 RNA. The (CAG)100 RNA construct was not MS2-tagged (see Figure 1—figure supplement 2A and Materials and methods). Neuritic RNA granules (white; arrowheads) were detected with a (D) (CUG)8 antisense but not with a (E) (CAG)8 sense probe. Distributions shown are representative of two biological replicates. (F–K) Primary rat spinal cord neurons transfected with NLS-CP-GFP (green; arrowheads) and (F) LacZ-MS2, (G) (GAA)100-MS2, (H) (CAG)100-MS2, (I) (GGGGCC)48-MS2, (J) (CUG)100-MS2, or (K) (CCUG)100-MS2 (see Figure 1—figure supplement 2A and Materials and methods for construct details). Whereas (F) the control RNA LacZ-MS2, or (G) an expanded repeat RNA without secondary structure, (GAA)100-MS2, did not show GFP accumulations, (H–K) the other expanded repeat RNAs conferred punctate GFP staining indicative of RNA enrichment in neuritic RNA granules. See Figure 1—figure supplement 1E for the pattern of expression of the MS2 alone control, which lacked neuritic puncta. DsRed (magenta) was coexpressed to outline the neurons. (L) Quantitation of iPSNs with neuritic GGGGCC repeat RNA granules. Nuclear foci positive (+) neurons were defined as having ≥5 (carrier 2) or ≥1 (carrier 1) nuclear GGGGCC RNA foci. ANOVA p-value = 0.00033. (M) Primary rat spinal cord neurons were transfected as in (F–K) and the percentage of neurons with neuritic repeat RNA particles was determined. At right, neurons within the mixed culture with the large morphology characteristic of motor neurons were scored for neuritic particles. ANOVA p-value = 1.1×10–7. (L–M) Numbers indicate the total number of neurons scored from a minimum of (L) two or (M) three biological replicates. Averages ± standard deviation are given. See Materials and methods for statistical analysis. Post-hoc: ****p < 2.5×10–6, ***p < 0.00042, **p < 0.0085, *p < 0.023. (D) A single confocal section or (A–C, E–K) Z-series projections. Bars: 10 μm. DAPI: blue. See also Figure 1—figure supplement 1–2. ANOVA, analysis of variance; DAPI, 4',6-diamidino-2-phenylindole; CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; NLS, nuclear localization signal.
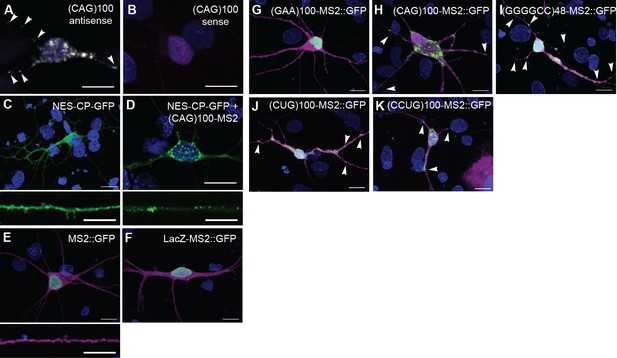
Neuritic localization of (CAG)100 RNA by in situ hybridization.
(A–B) In situ hybridization of primary rat spinal cord neurons expressing (CAG)100 RNA. The (CAG)100 construct was not MS2-tagged, but contained a leader sequence and translation reporter tags (see Figure 1—figure supplement 2A). (A) (CUG)8 antisense or (B) (CAG)8 sense probes were used for RNA detection (white; arrowheads). (B) Magenta indicates a transfected cell. (C–D) Detection of nuclear CAG repeat RNA foci. Primary rat spinal cord neurons expressing (C) CP-GFP fused to a NES (NES-CP-GFP; green), or (D) NES-CP-GFP and (CAG)100-MS2 RNA. (E-F) Primary rat spinal cord neurons were transfected with NLS-CP-GFP (green) and (E) MS2 or (F) LacZ-MS2. (E) MS2 RNA (green) was not enriched in cellular processes (0/52 neurons scored had neuritic MS2 RNA granules). (F) LacZ-MS2 was not enriched in cellular processes (0/65 neurons scored had neuritic LacZ-MS2 RNA granules), see quantitation in Figure 1M, and high magnification of neurite in Figure 1F. (G–K) Primary rat spinal cord neurons were transfected with NLS-CP-GFP (green;arrowheads) and (G) (GAA)100-MS2, (H) (CAG)100-MS2, (I) (GGGGCC)48-MS2, (J) (CUG)100-MS2, or (K) (CCUG)100-MS2. Whereas an expanded repeat RNA, (GAA)100-MS2, without secondary structure was not enriched, the other expanded repeat RNAs showed punctate GFP staining, indicative of granules. The MS2, (GAA)100-MS2, (CAG)100-MS2, (GGGGCC)48-MS2, (CUG)100-MS2, and (CCUG)100-MS2 constructs all contained a leader sequence, a translation reporter tag, and an MS2 tag (see Figure 1—figure supplement 2A and Materials and methods). Therefore, in addition to serving as a control for the MS2 tag and structured RNA, the MS2 and (GAA)100-MS2 constructs also serve as controls for the leader and translation reporter tags. (A) A single confocal section, or (B–K) confocal Z-series projections. Distributions shown are representative of (A–B) two biological replicates, or (C–K) a minimum of three biological replicates. (A–B, F–K, and top panels of C–E) Bar: 15 μm. (C–D, E) Bottom panels show neuronal processes at high magnification. Bar: 10 μm. (E–K) DsRed (magenta) was coexpressed to outline the neurons. DAPI: blue. CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; DAPI, 4',6-diamidino-2-phenylindole; NES, nuclear export signal; NLS, nuclear localization signal.
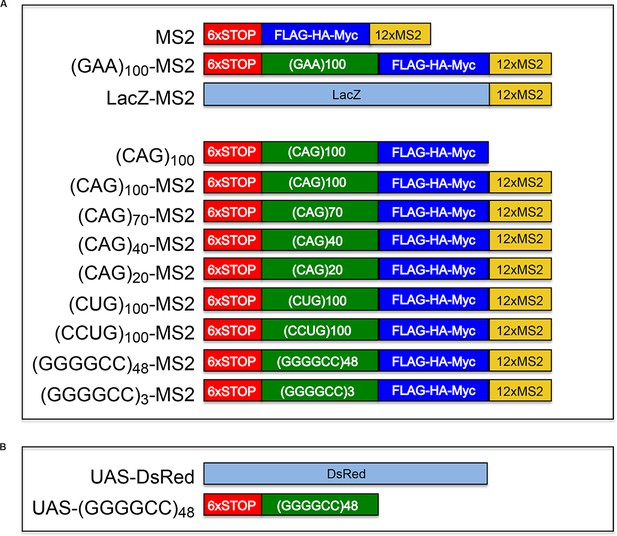
Microsatellite repeat and control expression constructs.
(A) Constructs used to express control (top three constructs) and microsatellite repeat (bottom nine constructs) RNA in transfected primary rat spinal cord neurons are shown. With the exception of (CAG)100, which was cloned into pcDNA, all constructs, including (CAG)100-MS2, were cloned into the pGW expression vector. The three control constructs were MS2, (GAA)100-MS2 and LacZ-MS2. (Red box) A leader sequence that includes 6 stop codons, 2 in each reading frame. (Green box) A repeat sequence. (Blue box) Three tags (FLAG, HA, and Myc, one in each reading frame) to detect RAN translation. (Yellow box) Twelve MS2 repeats (the CP binding site to allow detection via CP-GFP binding). (B) Constructs used to express control and microsatellite repeat RNA in Drosophila da neurons. Control UAS construct (top, UAS-DsRed) and the experimental UAS construct (bottom, UAS-(GGGGCC)48) are diagrammed. The UAS-(GGGGCC)48 construct has the same 5’ leader as the constructs described above in (A). CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein.
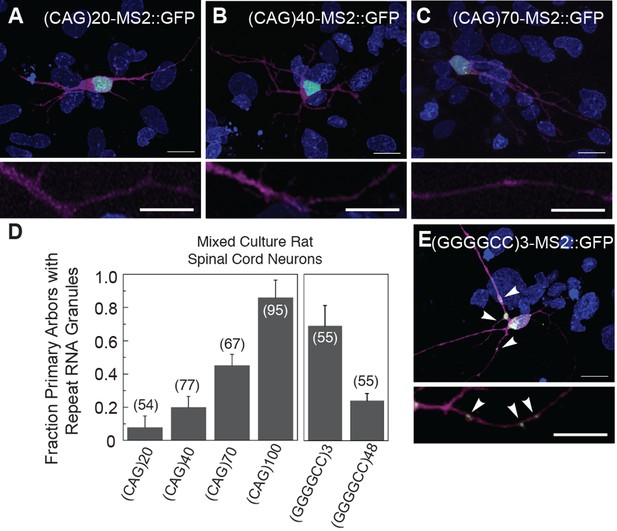
Particle formation dependence on CAG and GGGGCC RNA repeat number.
(A–C, E) Rat mixed spinal cord neurons were transfected with NLS-CP-GFP (green; arrowheads) and (A) (CAG)20-MS2, (B) (CAG)40-MS2, (C) (CAG)70-MS2, or (E) (GGGGCC)3-MS2. (CAG)100-MS2 and (GGGGCC)48-MS2 were transfected as above and are shown in Figure 1H and Figure 1—figure supplement 1H, and in Figure 1I and Figure 1—figure supplement 1I, respectively. See Figure 1—figure supplement 2A and Materials and methods for construct details. (D) Quantitation of the fraction of primary arbors containing ≥1 GFP granule. Parentheses indicate the total number of primary branches counted in three biological replicates. A minimum of 15 transfected neurons were scored for neuritic GFP granules in total. Averages ± standard deviation are given. Data are representative of a minimum of three biological replicates. Confocal Z-series projections. DsRed (magenta) was coexpressed to outline the neurons. Bottom panels: High magnification of neuronal processes. Bars: top panels, 15 μm; bottom panels,10 μm. DAPI: blue. DAPI, 4',6-diamidino-2-phenylindole; CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; NLS, nuclear localization signal.
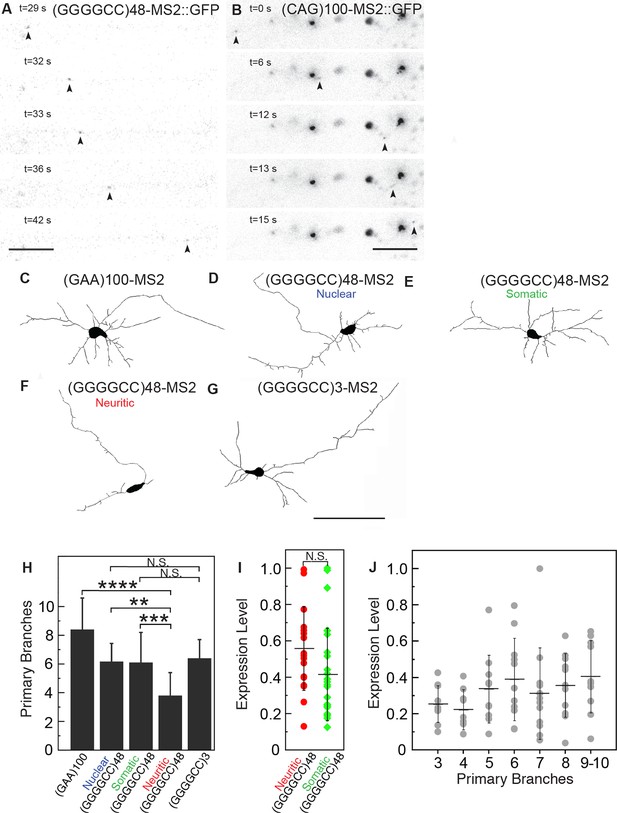
(GGGGCC)48 and (CAG)100 RNA assembles into neuritic transport particles and neuritic (GGGGCC)48 RNA correlates with branching defects.
(A–B) Rat spinal cord neurons were transfected with NLS-CP-GFP and (A) (GGGGCC)48-MS2 or (B) (CAG)100-MS2, and the trajectory of motile RNA particles along neuronal processes was captured by time-lapse microscopy. The location of an RNA particle (arrowheads) at indicated time points is shown in individual frames. (A) The uninterrupted, unidirectional anterograde particle run originates 22 μm from the cell body (which is outside of the shown frames), and ends 18 μm further away (see Video 1). (B) The uninterrupted, unidirectional anterograde particle run originates 67 μm from the cell body (which is to the left and outside of the shown frames), and ends 44 μm further away (see Videos 2 and 3). Larger stationary particles are also seen. (C–G) Cultured primary rat spinal cord neurons with neuritically localized (GGGGCC)48-MS2 RNA have fewer primary branches. (C–G) Tracings depicting the cell body and primary branches of rat mixed spinal cord neurons expressing either NLS-CP-GFP and (C) (GAA)100-MS2, (E-F) (GGGGCC)48-MS2, (G) (GGGGCC)3-MS2, or (D) NES-CP-GFP and (GGGGCC)48-MS2 RNA (see Figure 1—figure supplement 2A and Materials and methods for construct details). (H) Neurons were transfected as in C–G and the number of primary branches were scored in neurons that had (D) nuclear RNA particles, (E) somatic (but not neuritic) RNA particles, or those that had (F) neuritic RNA particles, as indicated. Repeat construct and GGGGCC repeat RNA localization affected branch number (p < 0.0001, ANOVA). For individual comparisons by post-hoc Tukey’s multiple comparisons test: ****p < 0.0001; ***p < 0.001; **p <0.01; N.S. p > 0.05. (I) Neurons with neuritic (GGGGCC)48-MS2 RNA did not have significantly higher expression level, as determined by ImageJ measurement of mean fluorescence intensity of the neural cell bodies (see Materials and methods), than neurons with somatic (GGGGCC)48-MS2 RNA. N.S. p > 0.07. (J) Expression level did not affect branch number (p = 0.2331, ANOVA). Individual comparisons did not reach significance (p value range: 0.2865 to > 0.9999). (H, J) Only neurons with cell bodies >20 μm and with >2 primary branches were included. Standard deviations are given. Bars: A, 5 μm; B, 10 μm; G, 100 μm. ANOVA, analysis of variance; CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; NES, nuclear export signal; NLS, nuclear localization signal.
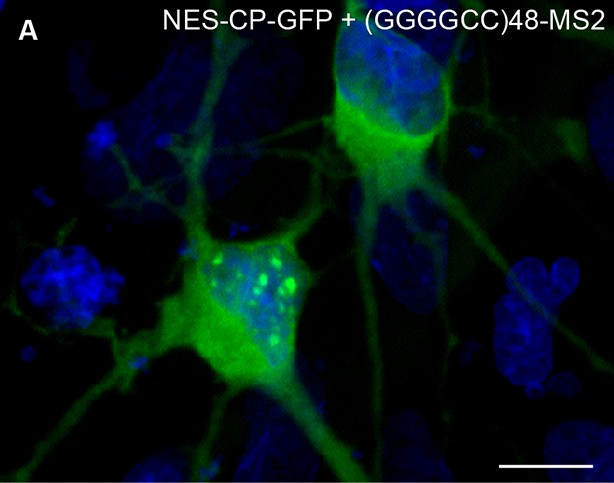
Nuclear (GGGGCC)48-MS2 foci in transfected rat primary spinal cord neurons.
(A) Primary rat spinal cord neurons transfected with NES-CP-GFP (green) and (GGGGCC)48-MS2. (A) Bottom left, a transfected neuron with nuclear (GGGGCC)48-MS2 RNA foci. Top right, a transfected neuron that lacks (GGGGCC)48-MS2 RNA foci in the nucleus. Image is representative of three biological replicates. Widefield epifluorescence micrograph. Bar: 10 μm. DAPI: blue. CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; DAPI, 4',6-diamidino-2-phenylindole; NES, nuclear export signal.
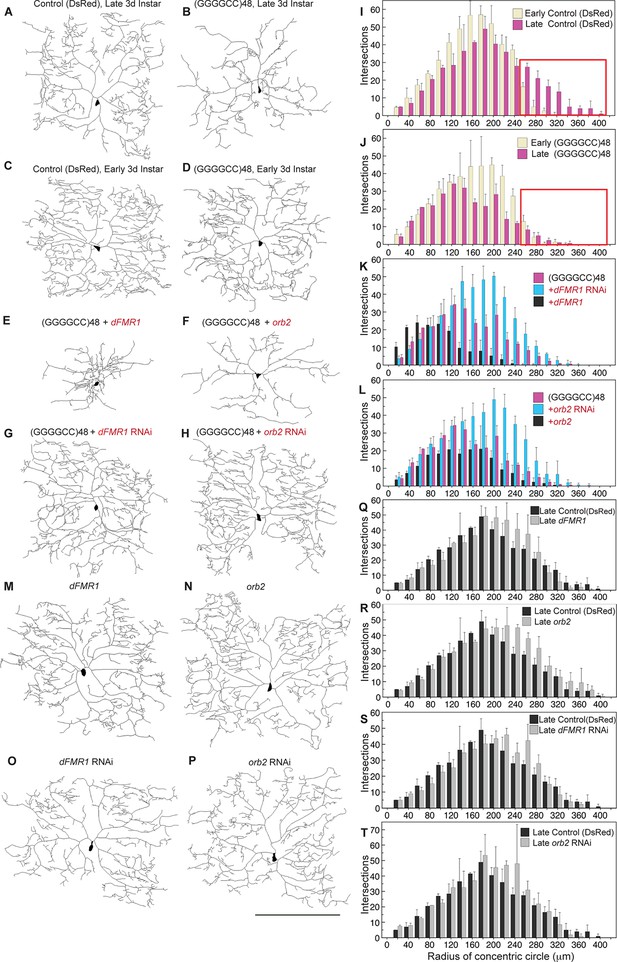
(GGGGCC)48-induced dendritic arborization defects are modulated by altered levels of transport granule components in Drosophila.
(A–H) Tracings, from confocal Z-series projections, of the cell body and dendritic arbor of class IV da neurons located in the body wall of Drosophila early or late third instar larvae. Expression of (GGGGCC)48 has a dramatic effect on the branching pattern compared with control (a transgene expressing DsRed). The effect on branching is enhanced by upregulation and suppressed by downregulation of dFMR1 or orb2. (A–D) GAL4477 (Grueber et al., 2003) driven expression of UAS-mCD8::GFP with (A, C) UAS-DsRed control (Li et al., 2008), or with (B, D) UAS-(GGGGCC)48, in early and late third instar neurons. (E–H) GAL4477driven expression of UAS-mCD8::GFP with (E) UAS-(GGGGCC)48 and UAS-dFMR1, (F) UAS-(GGGGCC)48 and UAS-orb2, (G) UAS-(GGGGCC)48 and UAS-dFMR1-RNAi, or with (H) UAS-(GGGGCC)48 and UAS-orb2-RNAi, in late third instar neurons. (I–L) Sholl analysis of traced class IV da neurons shown in (A–H), indicating the number of dendrite intersections with circles drawn at increasing radii from the cell body centroid. (I–J) Early (yellow) or late (magenta) third instar (I) UAS-DsRed control or (J) UAS-(GGGGCC)48 expressing da neurons. Distal intersections (260–400 μm from the cell body centroid) are boxed in red. (K–L) Late third instar neurons expressing UAS-(GGGGCC)48 alone (magenta), or (K) with UAS-dFMR1-RNAi (cyan) or UAS-dFMR1 (black), or (L) with UAS-orb2-RNAi (cyan) or UAS-orb2 (black). (M–T) Expression of the dFMR1 or orb2 modifier lines alone minimally alters the dendritic intersection distribution. Controls included comparison of the DsRed control (see A) to dFMR1 and orb2 lines in absence of UAS-(GGGGCC)48. (M–P) Tracings of class IV da neurons with GAL4477driven expression of UAS-mCD8::GFP with (M) UAS-dFMR1, (N) UAS-orb2, (O) UAS-dFMR1-RNAi, or (P) UAS-orb2-RNAi. (Q–T) Sholl analysis of traced class IV da neurons shown in (M–P). One dorsal neuron from the third or fourth abdominal hemisegment was scored per larvae and three to five larvae were scored per genotype (except for the late third instar control, A; n=2). (A–H, M–P) Dorsal, up; anterior, right. The UAS constructs did not contain a translation reporter or MS2 tags, see Figure 1—figure supplement 2B. Standard deviations are shown. Data are representative of three biological replicates. Bar, 300 μm. See also Figure 4—figure supplement 1.
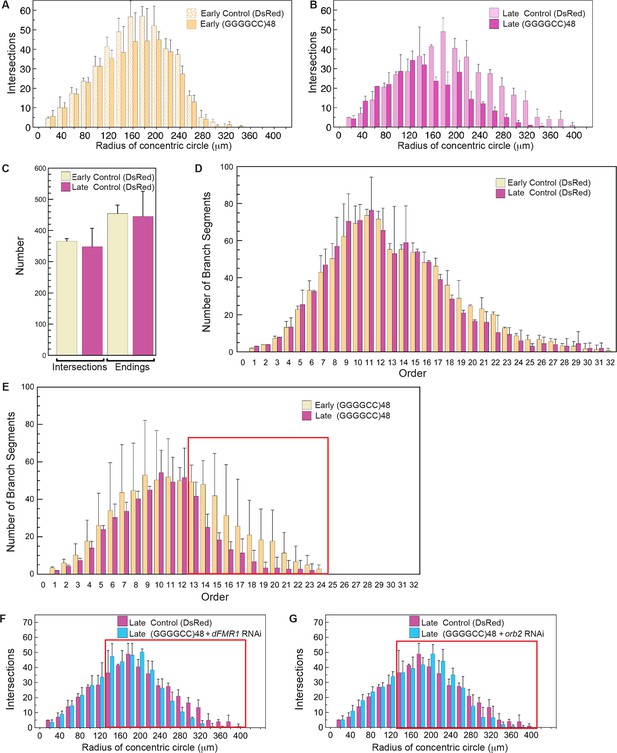
Knock down of dFMR1 or orb2 restores (GGGGCC)48-induced branching defects.
(A–G) Tracings from confocal Z-series projections of GAL4477 driven UAS-mCD8::GFP expression (Grueber et al., 2003) in Drosophila class IV da neurons were used for the analyses. (A–E) The GAL4477 driver was used to express (A–D) UAS-DsRed (control) or (A–B, E) UAS-(GGGGCC)48. (A) Mild dendritic branching defects are observed in (GGGGCC)48 da neurons at early third instar compared with the DsRed control. (B) Dramatic dendritic branching defects result from (GGGGCC)48 expression at late third larval instar compared to the DsRed control. (C–D) Branch loss is not evident in control larvae during the early to late third instar transition. The total number of (C) dendrite intersections and endings, or (D) branch segments per order were scored at early (yellow) or late (magenta) third instar stages. (E) Reduction of high order branches at late third instar (magenta) compared to early third instar (yellow) in neurons expressing (GGGGCC)48. Branch segments at high orders are boxed in red (orders 13–24). (F–G) Sholl diagrams of traced late third instar larval neurons. The GAL4477 driver was used to express (F–G) UAS-DsRed (control), (F) UAS-(GGGGCC)48 and UAS-dFMR1-RNAi, or (G) UAS-(GGGGCC)48 and UAS-orb2-RNAi. Distal intersections are restored upon (F) dFMR1 or (G) orb2 knockdown in larvae expressing UAS-(GGGGCC)48 (cyan). Comparisons with the UAS-DsRed control (magenta) are shown. Distal intersections at 140–400 μm from the cell body centroid are boxed in red. Standard deviations are shown. One dorsal neuron from the third or fourth abdominal hemisegment was scored per larvae and three to five larvae were scored per genotype (except for the late third instar control; n=2). The UAS constructs did not contain a translation reporter or MS2 tag, see Figure 1—figure supplement 2B. Data are representative of 3 biological replicates.
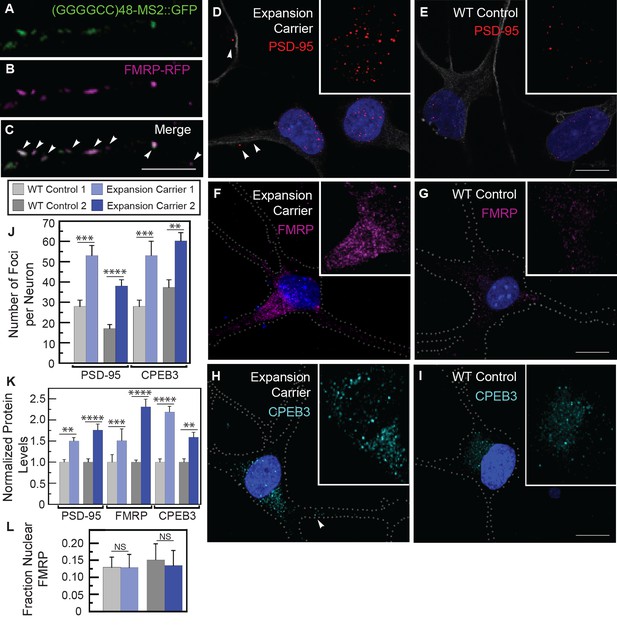
Misregulation of transport granule components in human iPSNs from carriers with a C9orf72 GGGGCC expansion.
(A–C) Neuritic particles consisting of expanded GGGGCC repeat RNA co-label for FMRP. Rat primary spinal cord neurons were transfected with NLS-CP-GFP (green), FMRP-RFP (magenta), and (GGGGCC)48-MS2, and neuronal processes were defined as regions of interest. Colocalization coefficients M1 (FMRP-RFP overlap with (GGGGCC)48-MS2 RNA) and M2 ((GGGGCC)48-MS2 RNA overlap with FMRP-RFP) were 0.64 ± 0.15 SD and 0.68 ± 0.23 SD, respectively (n=6 neurons). Colocalization coefficients for overlap between endogenous FMRP and (GGGGCC)48-MS2 were M1=0.61 ± 0.06 SD and M2=0.56 ± 0.14 SD (n=5 neurons; not shown). See Figure 1—figure supplement 2A and Materials and methods for construct details. Data are representative of three biological replicates. Confocal Z-series projections. (D–K) FMRP targets (D–E) PSD-95 and (F–G) FMRP, as well as (H–I) CPEB3, a local translation regulator, are increased in human iPSNs from C9orf72 GGGGCC expansion carriers, with a concomitant increase in PSD-95 and CPEB3 foci. High magnification of cell bodies are shown as insets. Neurites were marked with α-b III Tubulin or are outlined (dotted line), and (D) neuritic PSD-95 foci are indicated (arrowheads). (D–I) Images for GGGGCC expansion carrier 2 are shown. (J–L) Key for carriers and controls is shown at top. (J–K) Quantitation of PSD-95 and CPEB3 (J) foci, and of (K) total protein levels by immunostain in human iPSNs from carriers with a C9orf72 GGGGCC expansion. Kruskal–Wallis analysis for carrier versus control for all conditions: p < 0.0001. Post-hoc Dunn’s test, multiplicity adjusted p-values: ****p < 0.0005; ***p < 0.0015; **p < 0.018; N.S. p > 0.05. (J) From left to right, PSD-95: control, n=812 foci in 29 neurons; carrier 1, n=1590 foci in 30 neurons; control, n=851 foci in 49 neurons; carrier 2, n=1455 foci in 38 neurons. CPEB3: control, n=980 foci in 35 neurons; carrier 1, n=2067 foci in 39 neurons; control, n=735 foci in 21 neurons; carrier 2, n=1980 foci in 44 neurons. (K) From left to right, PSD-95: control, n=29; carrier 1, n=30; control, n=37; carrier 2, n=48 neurons scored. FMRP: control, n=32; carrier 1, n=30; control, n=25; carrier 2, n=27 neurons scored. CPEB3: control, n=34; carrier 1, n=37; control, n=17; carrier 2, n=49 neurons scored. (L) FMRP is not sequestered in the nuclei of carrier iPSNs. Quantitation of the fraction nuclear to total FMRP in carrier vs. control iPSNs. Data are averages from carrier 1, carrier 2, and controls ± standard error of the mean. From left to right: control, n=5; carrier 1, n=5; control, n=8; carrier 2, n=6 neurons scored. All comparisons are non-significant by ANOVA and post-hoc Sidak’s t-test. Confocal Z-series projections are shown. Bars: (E, G, I) 10 μm; (C) 5 μm. DAPI: blue. See also Figure 5—figure supplement 1. ANOVA, analysis of variance; CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; FMRP, fragile X mental retardation protein; NLS, nuclear localization signal; PSD, postsynaptic density protein; RFP, red fluorescent protein.
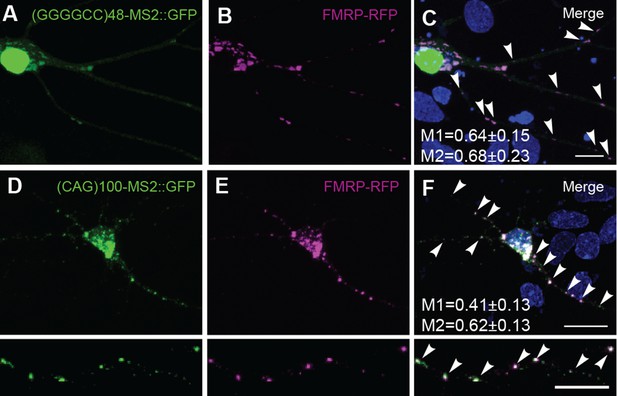
FMRP colocalizes with neuritic (GGGGCC)48-MS2 and (CAG)100-MS2 RNA.
Neuritic particles consisting of expanded GGGGCC or CAG repeat RNA co-label for FMRP. (A–F) Neurons were transfected with NLS-CP-GFP (green), FMRP-RFP (magenta), and (A–C) (GGGGCC)48-MS2, or (D–F) (CAG)100-MS2. Average ± standard deviation for colocalization coefficients M1 (FMRP overlap with RNA) and M2 (RNA overlap with FMRP), are given. (GGGGCC)48-MS2 (n=6 neurons); (CAG)100-MS2 (n=9 neurons). Neuronal processes were selected as regions of interest. See Figure 1—figure supplement 2A and Materials and methods for construct details. Data are representative of three biological replicates. Confocal Z-series projections are shown. CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; FMRP, fragile X mental retardation protein; NLS, nuclear localization signal.
Videos
Movement of a distal (GGGGCC)48-MS2 particle.
Two identical videos (60 s real-time duration each) are combined vertically; the bottom video displays a tracked particle in green, starting at 22 μm and reaching 40 μm from the cell body. Images were acquired at 1 frame/s and the video displays at 8 frames/s. The complete caption was 60 s. Selected images are shown in Figure 3A.
Movement of a distal (CAG)100-MS2 particle.
Two identical videos (40 s real-time duration each) are combined vertically; the bottom video displays the tracked particle and its path in green, starting at 67.0 μm and reaching 111.4 μm from the cell body. Images were acquired at 1 frame/s and the video displays at 8 frames/s. The complete caption was 133 s. Selected images are shown in Figure 3B.
Movement of a proximal (CAG)100-MS2 particle.
Two identical videos (97 s real-time duration each) are combined vertically; the bottom video displays the tracked particle and its path in green, starting in the cell body and reaching 6.5 μm into a neurite. Images were acquired at 1 frame/s and the video displays at 8 frames/s. The complete caption was 221 s.
Tables
Behavior of repeat RNA particles in rat spinal cord neurons.
| Mean average velocity (μm/s) | Average max velocity (μm/s) | Velocity range (μm/s) | Particles tracked in neurites | Particles tracked in cell body | Basal velocity* (μm/s) | |
|---|---|---|---|---|---|---|
| (GGGGCC)48-MS2::GFP n = 11 cells | 1.06 | 1.40 | 0.32–2.67 | 10 | 15 | 0.11 |
| (CAG)100-MS2::GFP n = 2 cells | 1.30 | 1.85 | 0.30–4.73 | 5 | 28 | 0.13 |
-
Uninterrupted unidirectional anterograde and retrograde particle runs with an average run distance of 5.3 μm ((GGGGCC)48-MS2), and 6.7 μm ((CAG)100-MS2), were analyzed. (*)The basal velocity is given as a mean average and was estimated by analyzing five particles that underwent corralled movements with an average net displacement of <0.51 μm within 20s. Data are from four (GGGGCC)48-MS2 and two (CAG)100-MS2 independent live imaging sessions. (GGGGCC)48-MS2 and (CAG)100-MS2 were co-expressed with NLS-CP-GFP. CP-GFP, MS2 RNA-binding coat protein fused with green fluorescent protein; NLS, nuclear localization signal.
Expression of transport-granule related transcripts in brains of C9orf72 patients.
| Expression in C9orf72 Cortex | Gene subset | Expected percentage | Observed percentage | Chi-squared p-value |
|---|---|---|---|---|
| Upregulated | RNA binding proteins | 7.5 | 15.10 (167/1103) | p<0.0001 |
| mRNA binding proteins | 3.4 | 7.80 (86/1103) | p<0.0001 | |
| FMRP targets | 4.2 | 4.90 (54/1103) | p=0.2568 | |
| STAT5B targets | 3.1 | 6.07 (67/1103) | p<0.0001 | |
| Downregulated | RNA binding proteins | 7.5 | 4.93 (129/2618) | p<0.0001 |
| mRNA binding proteins | 3.4 | 1.72 (45/2618) | p<0.0001 | |
| FMRP targets | 4.2 | 3.40 (89/2618) | p=0.0389 | |
| STAT5B targets | 3.1 | 1.38 (36/2618) | p<0.0001 |
-
Comparison of uniquely-identified protein coding genes that were either up- or down-regulated in cortical samples from C9orf72 patients (Donnelly et al., 2013) with transcripts associated with the regulation of RNA. Supplementary file 1 lists the RNA binding proteins upregulated and downregulated, as noted above.
Additional files
-
Supplementary file 1
Genes related to RNA granules that are misregulated in brain tissue from C9ORF72 patients.
Lists of the RNA binding proteins upregulated or downregulated in cortical samples from C9orf72 patients (Donnelly et al., 2013). Analysis is presented in Table 2.
- https://doi.org/10.7554/eLife.08881.018





