Resection is responsible for loss of transcription around a double-strand break in Saccharomyces cerevisiae
Figures
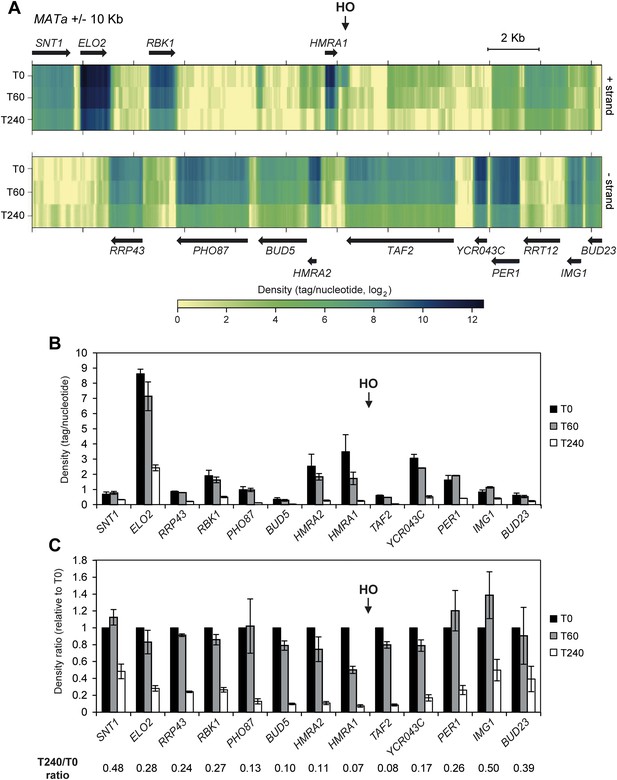
Decrease of RNA levels around a DNA double-strand break (DSB).
(A) Strand-specific RNA-seq data from the two biological replicates of wild type strain (JKM139) before (time zero, T0) and 60 (T60) or 240 (T240) min after HO induction, were uniquely mapped to the MAT locus ±10 kb. For each time point, densities (tag/nucleotide, log2 scale) from the two replicates were pooled and visualized as a heatmap with the upper and lower panels corresponding to the + and − strands, respectively. Black arrows represent annotated genes. (B) Density for genes along the MAT locus ±10 kb at T0, T60 and T240 after HO induction. Mean values ±s.d. were calculated from the two biological replicates analyzed. RRT12 showed too low signal to be significantly quantified and was not included. (C) Ratio of density for genes along the MAT locus ±10 kb at T0, T60 and T240 after HO induction. For each time point, densities were normalized on the values obtained at T0. Mean values ±s.d. were calculated as above. RNA-seq data used to construct the graphs of Figure 1B,C are available in Figure 1—source data 1.
-
Figure 1—source data 1
RNA-seq data used to construct the graphs of Figure 1B,C.
- https://doi.org/10.7554/eLife.08942.004
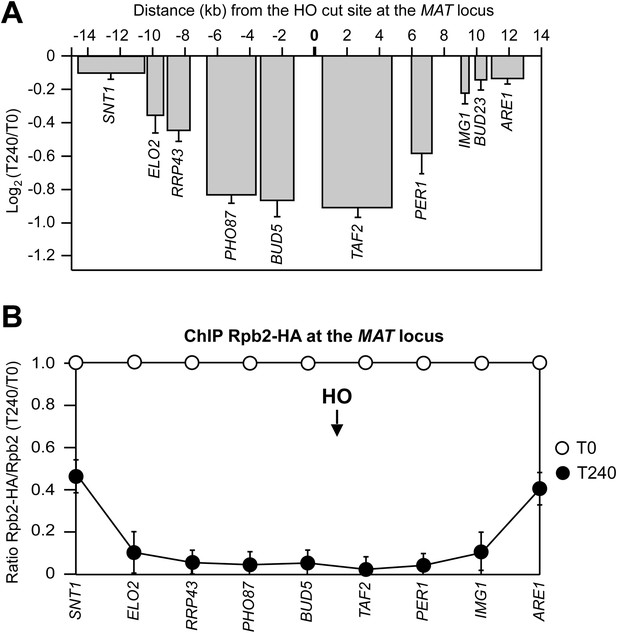
DSB-induced transcriptional inhibition at the MAT locus.
(A) YEPR exponentially growing cell cultures of the JKM139 strain, carrying the HO cut site at the MAT locus, were transferred to YEPRG at T0 to induce HO. RNA levels of genes located at different distances from the HO cut site were evaluated by quantitative reverse transcriptase PCR (qRT-PCR) at T0 and T240 after HO induction. Results are presented as ratios between T240 and T0. RNA levels were quantified using ∆∆Ct method and quantities were normalized to ACT1 RNA levels. The mean values ±s.d. are represented (n = 3). (B) Exponentially growing YEPR cell cultures of the JKM139 derivative strains expressing either a fully functional Rpb2-HA fusion protein or untagged Rpb2 were transferred to YEPRG at T0 to induce HO. Binding of Rpb2-HA at different distance from the DSB at T0 and T240 after HO induction was evaluated by ChIP and qPCR. Primers used were the same as in (A). Results are presented as ratios between Rpb2-HA and untagged Rpb2, both of which normalized against the corresponding input, at T240 relative to T0. The mean values ±s.d. are represented (n = 3). Rpb2-HA binding at the ACT1 gene was used as internal control.
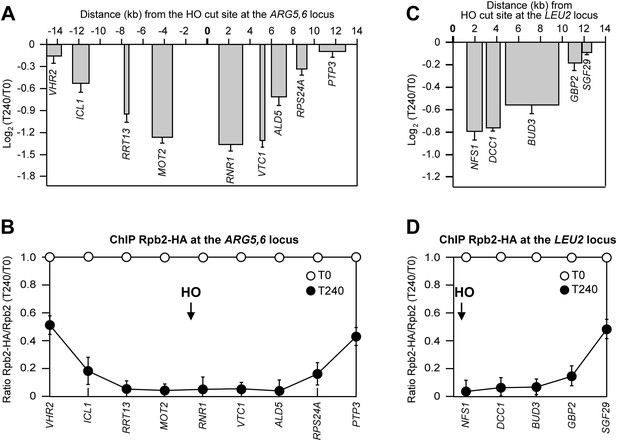
DSB-induced transcriptional inhibition at different chromosomal loci.
(A) YEPR exponentially growing cell cultures of tGI354 strain, carrying the HO site at the ARG5,6 locus and the deletion of RAD51 gene, were transferred to YEPRG at T0 to induce HO. RNA levels of genes located at different distances from the HO cut site were evaluated by qRT-PCR at T0 and T240 after HO induction, as described in Figure 2A. Results are presented as ratios between T240 and T0. The mean values ±s.d. are represented (n = 3). (B) Binding of Rpb2-HA from samples collected in (A) was evaluated by ChIP and qPCR as described in Figure 2B. Primers used were the same as in (A). The mean values ±s.d. are represented (n = 3). (C) YEPR exponentially growing cell cultures of YFP17 strain, carrying the HO cut site at the LEU2 locus, were transferred to YEPRG at T0 to induce HO. RNA levels were analyzed as in Figure 2A but normalized to the ALG9 gene transcript. Only transcription of genes located on the right side of the HO-induced DSB was analyzed because no transcription units are present on the left side of the break. The mean values ±s.d. are represented (n = 3). (D) Binding of Rpb2-HA from samples collected in (C) was evaluated by ChIP and qPCR as described in Figure 2B. Primers used were the same as in (C).
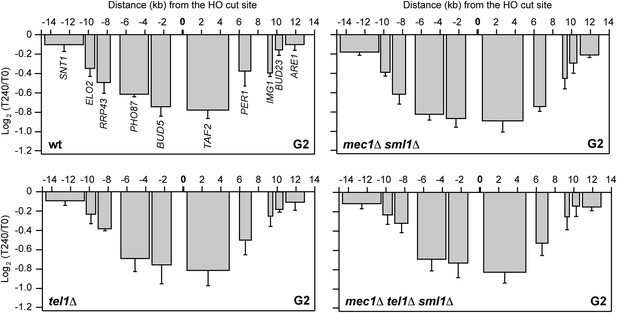
DSB-induced transcriptional inhibition does not require Mec1 and Tel1.
YEPR exponentially growing cell cultures of the JKM139 strain, carrying the HO cut site at the MAT locus, were arrested in G2 with nocodazole and transferred at T0 to YEPRG in the presence of nocodazole. RNA levels were analyzed by qRT-PCR as described in Figure 2A. The mean values ±s.d. are represented (n = 3).
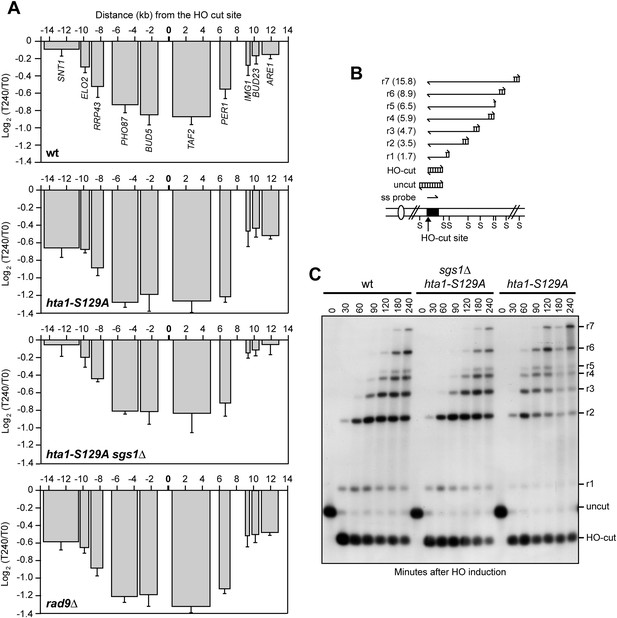
The lack of γH2A enhances transcriptional inhibition around the DSB by accelerating resection.
(A) YEPR exponentially growing cell cultures of the JKM139 derivative strains, carrying the HO cut site at the MAT locus, were transferred to YEPRG at T0. RNA levels of genes located in the surroundings of the HO cut site at the MAT locus were analyzed at T0 and T240 after HO induction by qRT-PCR as described in Figure 2A. The mean values ±s.d. are represented (n = 3). (B) Method to measure DSB resection. Gel blots of SspI-digested genomic DNA separated on alkaline agarose gel were hybridized with a single-stranded MAT probe (ss probe) that anneals to the unresected strand. 5′–3′ resection progressively eliminates SspI sites (S), producing larger SspI fragments (r1 through r7) detected by the probe. (C) DSB resection. Genomic DNA prepared from samples collected in (A) was analysed for single-stranded DNA (ssDNA) formation at the indicated times after HO induction as described in (B). The image that was used for the cropped final Figure 5C is available in Figure 5—source data 1.
-
Figure 5—source data 1
Image that was used for the cropped final Figure 5C.
- https://doi.org/10.7554/eLife.08942.009
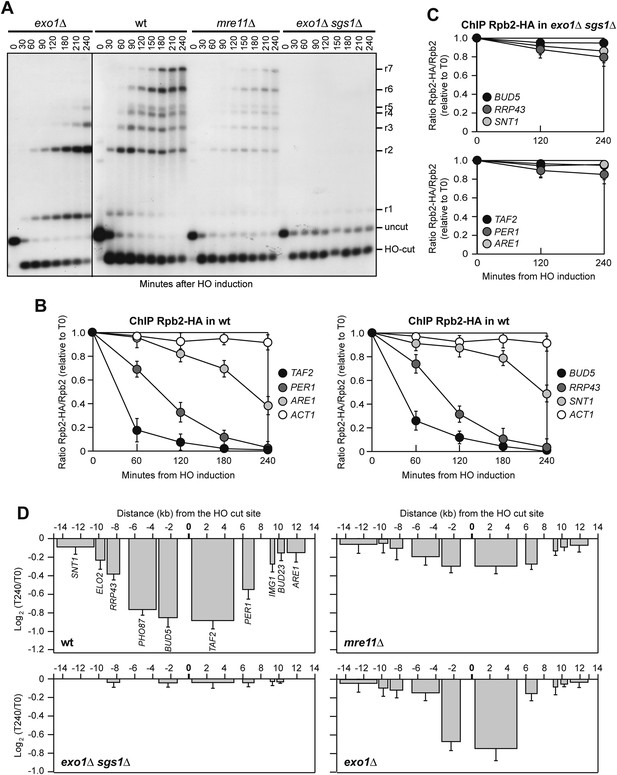
Resection mutants fail to reduce transcription around the DSB.
(A) DSB resection. YEPR exponentially growing cell cultures of JKM139 derivative strains, carrying the HO cut site at the MAT locus, were transferred to YEPRG at T0. Formation of ssDNA was determined as described in Figure 5B. (B) Binding of Rpb2-HA in wild type cells from samples collected in (A) was evaluated by ChIP and qPCR as described in Figure 2B. The mean values ±s.d. are represented (n = 3). (C) Binding of Rpb2-HA in exo1∆ sgs1∆ cells from samples collected in (A) was evaluated by ChIP and qPCR as described in Figure 2B. The mean values ±s.d. are represented (n = 3). (D) RNA levels from samples collected in (A) were analyzed at T0 and T240 after HO induction by qRT-PCR as described in Figure 2A. The mean values ±s.d. are represented (n = 3). Contiguous images that were used for the cropped final Figure 6A are available in Figure 6—source data 1.
-
Figure 6—source data 1
Contiguous images that were used for the cropped final Figure 6A.
- https://doi.org/10.7554/eLife.08942.011
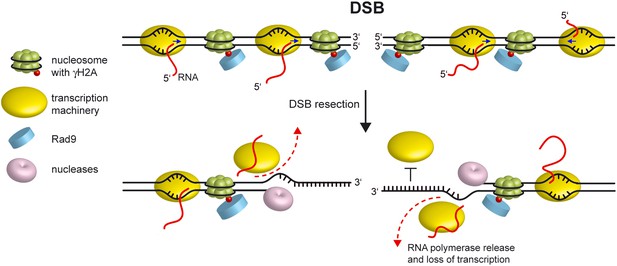
Model for loss of transcription around a DSB.
Resection of the 5′ strands at both DSB ends leads to release of the transcription machinery (dashed lines) and to subsequent transcription arrest independently of whether the degraded DNA strand acts as template or non-template. Since the RNA polymerase binds double-stranded promoter DNA, generation of ssDNA at the DSB ends prevents reinitiation events (bar-headed line). Blue arrows indicate direction of transcription.
Additional files
-
Supplementary file 1
Saccharomyces cerevisiae strains used in this study.
- https://doi.org/10.7554/eLife.08942.013
-
Supplementary file 2
Primer sequences used in this study.
- https://doi.org/10.7554/eLife.08942.014





