Deciphering the preeclampsia-specific immune microenvironment and the role of pro-inflammatory macrophages at the maternal–fetal interface
Figures
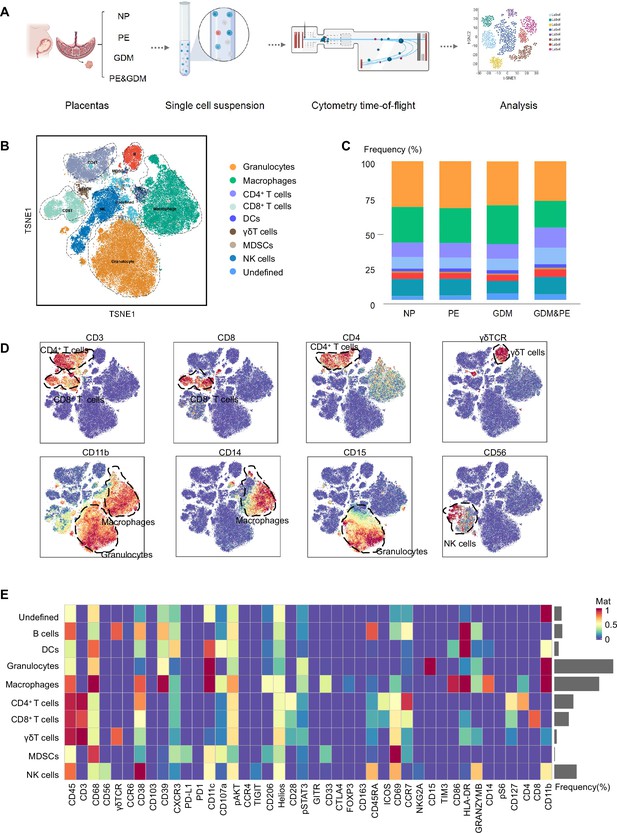
Identification and characterization of placental immune cells using cytometry by time of flight (CyTOF).
(A) Schematic of the experimental workflow in CyTOF experiment. The placentas were obtained from individuals with normal pregnancy (NP, n = 9), preeclampsia (PE, n = 8), gestational diabetes mellitus (GDM, n = 8), or GDM&PE (n = 7). (B) t-Distributed Stochastic Neighbor Embedding (t-SNE) maps showing 8 × 104 CD45+ cells (the average cell number of the all samples) from the placenta overlaid with color-coded clusters and the distributions of B cells, CD4+ T cells, CD8+ T cells, dendritic cell (DC), γδT cells, monocytes, granulocytes, myeloid-derived suppressor cell (MDSC), and natural killer (NK) cells. (C) Percentages of each cell type of CD45+ cells in placentas. (D) t-SNE maps showing the expression of CD3, CD8, CD4, γδTCR, CD14, CD15, and CD56. (E) Heatmap showing the expression levels of markers in CD45+ cell subsets. Data were compared between NP and PE, NP and GDM, and NP and GDM&PE using the Kruskal–Wallis test and represented as mean ± SEM (*p < 0.05, NS, not significant). CyTOF soure data are stored in Dryad Digital Repository, DOI: 10.5061/dryad.4qrfj6qn0.
-
Figure 1—source data 1
Raw data and detailed analysis for Figure 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig1-data1-v1.xls
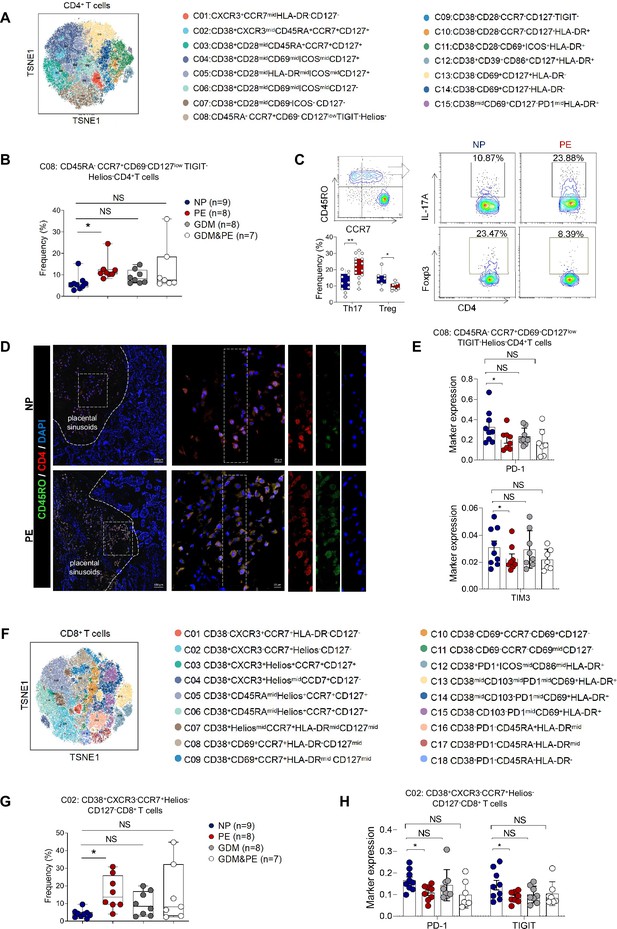
Specific altered T cell profile in the placentas of individuals with PE.
(A) Distribution of the CD4+ T cells in the group analyzed using t-Distributed Stochastic Neighbor Embedding (t-SNE). (B) Scatter dot plots showing the frequencies of cluster 8 of CD4+ T cells in the placentas of individuals with NP, PE, GDM, and GDM&PE (n = 9 in NP group; n = 8 in PE group; n = 8 in GDM group; n = 7 in GDM&PE group). (C) Expression of IL-17A and Foxp3 in CD45RO+CCR7+CD4+ T cells in placentas of individuals with NP and PE using flow cytometry (IL-17A: n = 10 in NP group, n = 15 in PE group; Foxp3: n = 7 in NP group, n = 9 in PE group). (D) Immunofluorescence co-staining of CD4 (red), CD45RO (green), and DAPI (blue) in frozen placental sections. The right panels show the fluorescence intensity of CD4 and CD45RO. Scale bar, 20 µm. (E) Scatter dot plots showing significantly altered markers of in cluster 8 of CD4+ T cells. (F) Distribution of the CD8+ T cells analyzed using t-SNE. (G) Scatter dot plots showing the frequencies of cluster 2 of CD8+ T cells in the placentas of individuals with NP, PE, GDM, and GDM&PE. (H) Scatter dot plots showing significantly altered markers in cluster 2 of CD8+ T cells. Data were compared between NP and PE, NP and GDM, and NP and GDM&PE using the Kruskal–Wallis test and represented as mean ± SEM (*p < 0.05, NS, not significant).
-
Figure 2—source data 1
Raw data and detailed analysis for Figure 2.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig2-data1-v1.zip
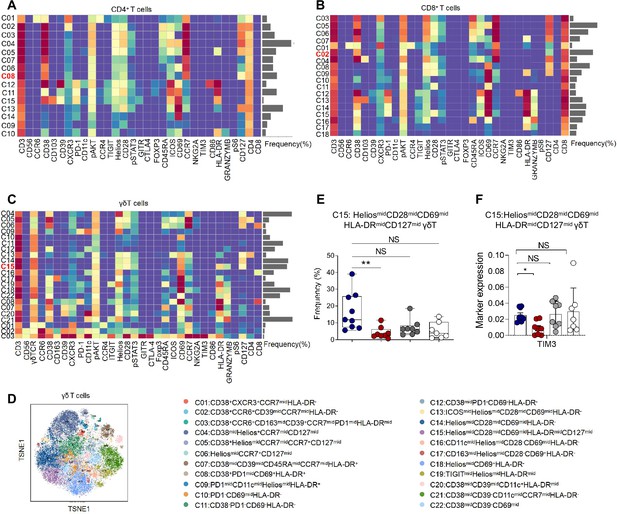
Identification of the placental T cell subsets.
(A) Heatmap showing the expression levels of markers in the CD4+ T subsets (n = 9 in NP group; n = 8 in PE group; n = 8 in GDM group; n = 7 in GDM&PE group). (B) Heatmap showing the expression levels of markers in the CD8+ T subsets. (C) Heatmap showing the expression levels of markers in the γδT subsets. (D) Distribution of the γδT cells in the group analyzed using t-Distributed Stochastic Neighbor Embedding (t-SNE). (E) Scatter dot plots showing the frequencies of cluster 15 of γδT cells in the placentas of individuals with NP, PE, GDM, and GDM&PE. (F) Scatter dot plots showing significantly altered markers in cluster 15 of γδT cells. Data were compared between NP and PE, NP and GDM, and NP and GDM&PE using the Kruskal–Wallis test and represented as mean ± SEM (*p < 0.05, **p < 0.01,NS, not significant).
-
Figure 2—figure supplement 1—source data 1
Raw data and detailed analysis for Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig2-figsupp1-data1-v1.zip
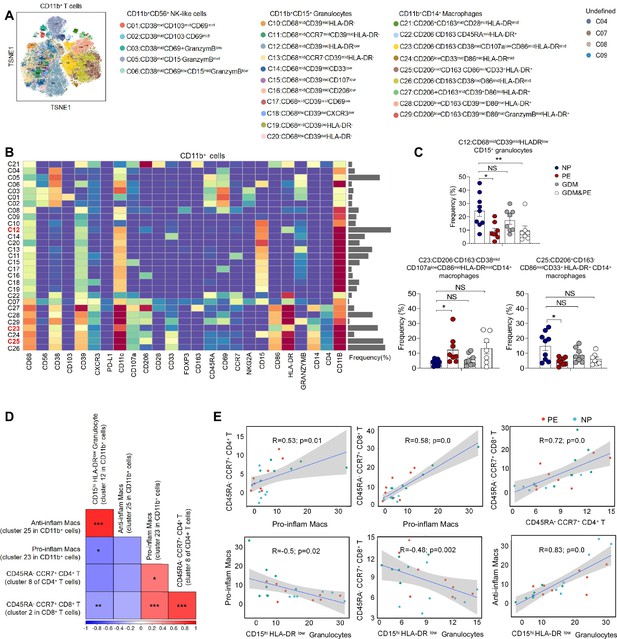
Identification of the placental CD11b+ cell subsets and the interaction between placental immune cells.
(A) Distribution of the CD11b+ immune cells in group analyzed using t-Distributed Stochastic Neighbor Embedding (t-SNE). (B) Heatmap showing the expression levels of markers in the CD11b+ cells. (C) Scatter dot plots showing the frequencies of clusters 12, 23, and 25 of CD11b+ cells in the placentas. (D) Interaction between placental immune cells showed in heatmap. (E) Scatter plots of Pearson’s correlation analysis between placental immune cells. Data were compared between NP and PE, NP and GDM, and NP and GDM&PE using the Kruskal–Wallis test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001; NS, not significant).
-
Figure 3—source data 1
Raw data and detailed analysis for Figure 3.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig3-data1-v1.zip
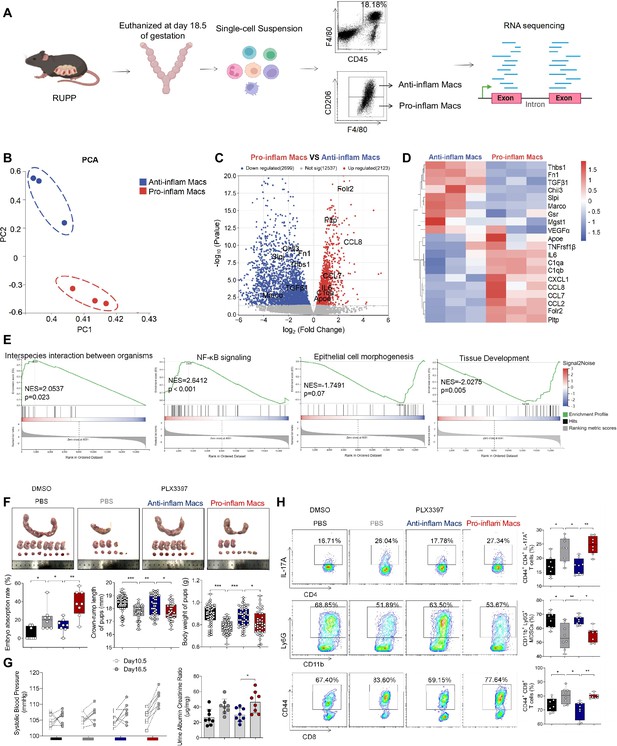
The immune imbalance at the maternal–fetal interface induced by F480+CD206− pro-inflam Macs.
(A) An illustration showcasing the RNA-seq of the CD45+F4/80+CD206− pro-inflam Macs and CD45+F4/80+CD206+ anti-inflam Macs derived from the RUPP mouse model. (B) Principal component analysis (PCA) reflected the differences between the two groups of macrophages (n = 3). (C) The volcano map shows a comparison of the content and p value of gene expression between pro-inflam and anti-inflam Macs. Differential expression genes were screened out when p < 0.05. Red dots indicate genes with increased expression in pro-inflam Macs. Blue dots indicate genes with decreased expression. (D) The volcano map shows differential expression genes between pro-inflam and anti-inflam Macs. (E) Representative pathways enriched in the identified genes as determined by gene set enrichment analysis (GSEA) (p-value <0.05). (F) Embryo abortion rate of the pregnant mice, body weight and crown-rump length of pups measured on day 18.5 of gestation. Black represents mice treated with DMSO (n = 8); gray represents mice treated with PLX3397 (n = 8); blue represents mice injected with CD45+F4/80+CD206+ anti-inflammatory macrophages (n = 8); red represents mice injected with CD45+F4/80+CD206− pro-inflammatory macrophages (n = 8). (G) Systolic blood pressure (SBP) and UACR of pregnant mice in the four groups. (H) Frequencies of CD44+CD4+IL-17A+ cells, CD44+CD8+ T cells, and CD11b+Ly6G+ granulocytes analyzed by flow cytometry. Data were compared between groups using the Kruskal–Wallis test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001). RNA transcriptome soure data are stored in Dryad Digital Repository, doi:10.5061/dryad.4qrfj6qn0.
-
Figure 4—source data 1
Raw data and detailed analysis for Figure 4.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig4-data1-v1.zip
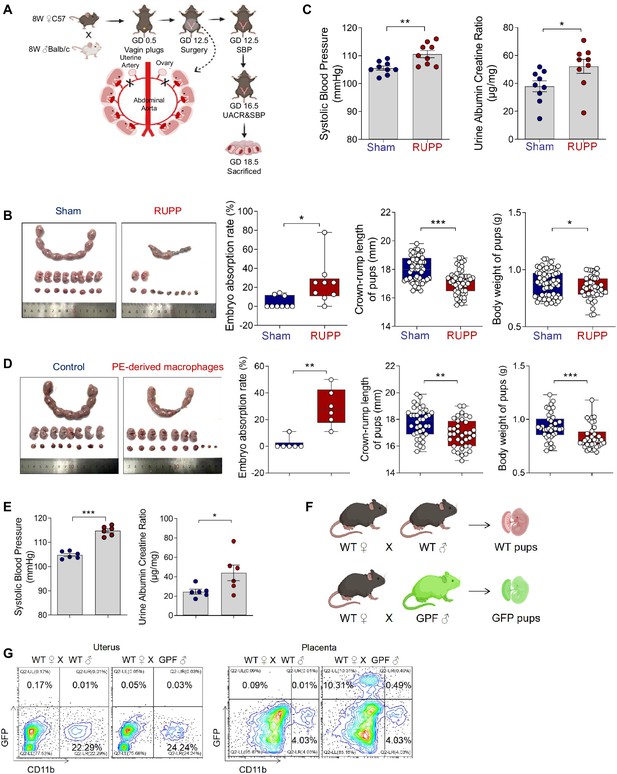
The construction of RUPP mouse model.
(A) Experimental design of mice model with PE by reducing uterine perfusion pressure (RUPP) on day 12.5 of gestation. Mice with sham operation were considered as controls. Systolic blood pressure (SBP) and urine albumin creatine ratio (UACR) were measured on days 12.5 and 16.5 of gestation, respectively. Mice were sacrificed on day 18.5 of gestation. (B) The embryo abortion rate of pregnant mice, body weight, and crown-rump length of pups measured on day 18.5 of gestation in the Sham and RUPP group. Embryo abortion rate = number of absorbed embryos/total number of embryos. Blue represents mice in the Sham group (n = 9); red represents mice in the RUPP group (n = 9). (C) SBP and UACR of pregnant mice in Sham and RUPP group. (D) The embryo abortion rate of pregnant mice, body weight, and crown-rump length of pups measured on day 18.5 of gestation in the mice injected with PBS or RUPP-derived macrophages. Embryo abortion rate = number of absorbed embryos/total number of embryos. Blue represents mice in the mice injected with PBS group (n = 6); red represents mice in the mice injected with RUPP-derived macrophages group (n = 6). (E) SBP and UACR of pregnant mice in PBS and RUPP-derived macrophages group. (F) Experimental design of GFP pregnant mice model. (G) Frequencies of GFP+CD11b+ fetal-derived macrophages and GFP−CD11b+ maternal-derived macrophages were analyzed by flow cytometry. Data were compared between the two groups using the Student’s t-test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001).
-
Figure 4—figure supplement 1—source data 1
Raw data and detailed analysis for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig4-figsupp1-data1-v1.zip
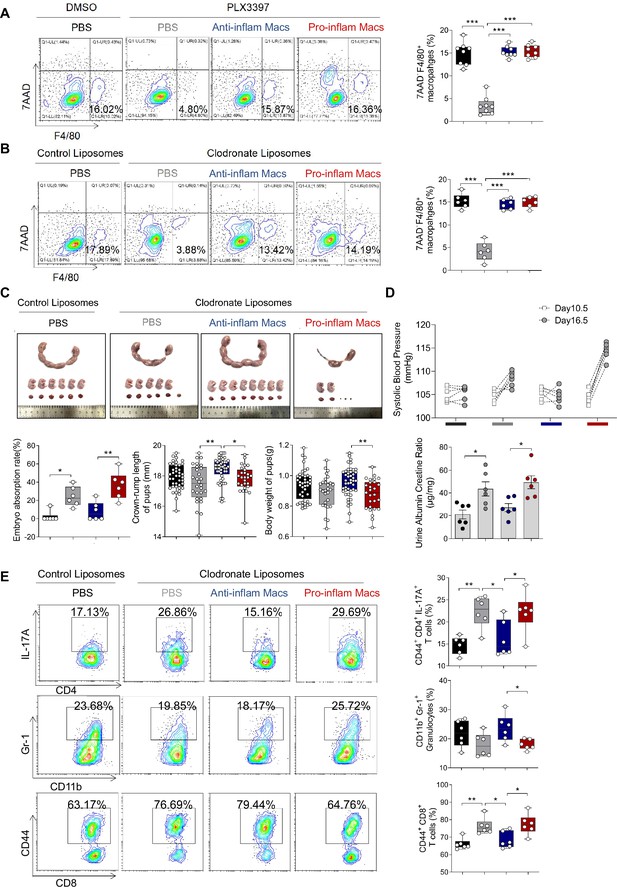
Clodronate liposomes were used to deplete the macrophages of pregnant mice to demonstrate that pro-inflam Macs lead to immune imbalance.
(A) Frequencies of F4/80+7AAD− macrophages in PLX3397-treated mouse were analyzed by flow cytometry. (B) Frequencies of F4/80+7AAD− macrophages in clodronate liposomes-treated mouse were analyzed by flow cytometry. (C) Embryo abortion rate of the pregnant mice, body weight and crown-rump length of pups measured on day 18.5 of gestation. Black represents mice treated with control liposomes (n = 6); gray represents mice treated with clodronate liposomes (n = 6); blue represents mice injected with CD45+F4/80+CD206+ anti-inflam Macs (n = 6); red represents mice injected with CD45+F4/80+CD206− pro-inflam Macs (n = 6). (D) Systolic blood pressure (SBP) and UACR of pregnant mice in the four groups. (E) Frequencies of CD44+CD4+IL-17A+ cells, CD44+CD8+ T cells, and CD11b+Gr1+ granulocytes analyzed by flow cytometry. Data were compared between groups using one-way ANOVA and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001).
-
Figure 4—figure supplement 2—source data 1
Raw data and detailed analysis for Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig4-figsupp2-data1-v1.zip
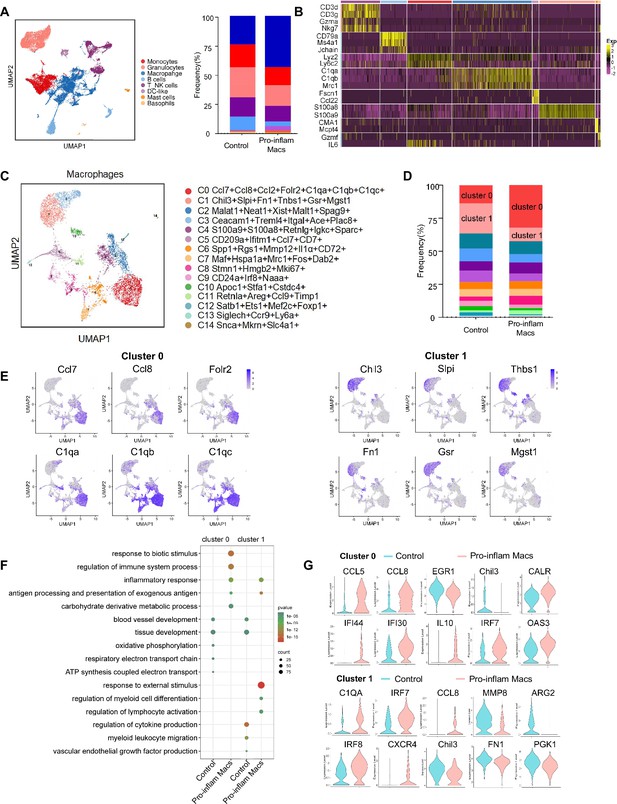
Identification of the macrophage subsets in mice injected with pro-inflam and anti-inflam Macs by using single-cell RNA sequencing (scRNA-seq).
(A) Uniform Manifold Approximation and Projection (UMAP) maps showing the eight types of mouse immune cells at the maternal–fetal interface. (B) Heatmap showing clustering analysis for markers distinguished different type of immune cells. (C) UMAP maps showing the 15 clusters of mouse macrophages was listed. (D) Bar graph showing the frequencies of clusters of macrophages in the two groups of mice was listed in the right panel. (E) UMAP maps showing the distribution of specific markers of clusters 0 and 1. (F) Dot plot depicting GO enrichment terms that were significantly enriched in the differentially expressed genes in clusters 0 and 1 from the pro-inflam Macs group and the control group. (G) Violin plot of specific differential gene expression in clusters 0 and 1 between the pro-inflam Macs and the control groups. Single-cell RNA transcriptome soure data are stored in Dryad Digital Repository, 10.5061/dryad.4qrfj6qn0.
-
Figure 5—source data 1
Raw data and detailed analysis for Figure 5.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig5-data1-v1.zip

Heatmap of different clusters of macrophages.
(A) Heatmap showing clustering analysis for markers distinguished different clusters of macrophages.
-
Figure 5—figure supplement 1—source data 1
Raw data and detailed analysis for Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig5-figsupp1-data1-v1.zip
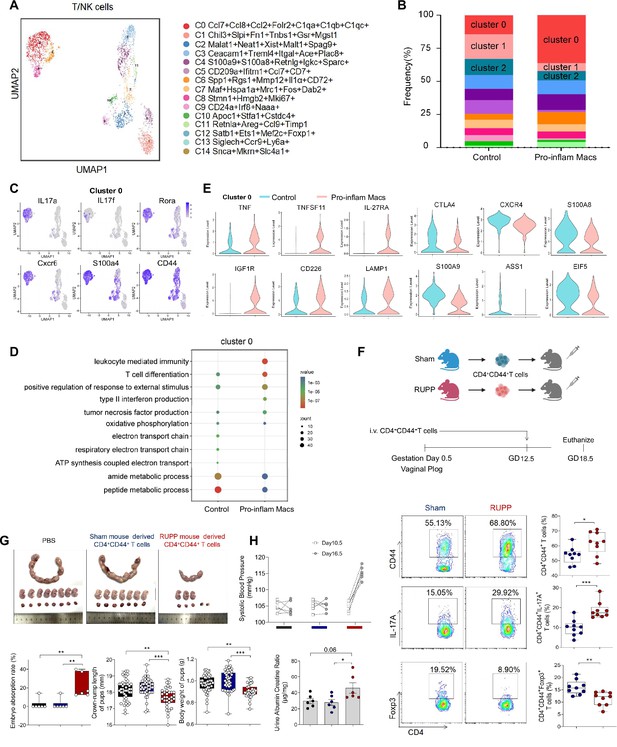
Identification of the T/NK cells subsets in mice injected pro-inflam and anti-inflam Macs using single-cell RNA sequencing (scRNA-seq).
(A) Uniform Manifold Approximation and Projection (UMAP) maps showing the 12 clusters of mouse T/NK cells was listed. Heatmap showing clustering analysis for markers distinguished 12 different clusters of T/NK cells. (B) Bar graph showing the frequencies of clusters of T/NK cells in the two groups of mice was listed in the down panel. (C) Dot plot depicting GO enrichment terms that were significantly enriched in the differentially expressed genes in cluster 0 from the pro-inflam Macs and the control groups. (D) Violin plot of specific differential gene expression in cluster 0 between the pro-inflam Macs and the control groups. (F) Frequencies of CD4+CD44+ T cells and the percentages of Foxp3+ or IL-17A+ cells in CD4+CD44+ T cells at the maternal–fetal interface in Sham and RUPP group analyzed by flow cytometry. (G) The embryo abortion rate of pregnant mice, body weight, and crown-rump length of pups measured on day 18.5 of gestation in mice injected PBS, Sham mouse-derived or RUPP mouse-derived CD4+CD44+ T cells. Black represents mice injected with PBS (n = 6); blue represents mice injected with Sham mouse-derived CD4+CD44+ T cells (n = 6); red represents mice injected with RUPP mouse-derived CD4+CD44+ T cells (n = 6). (H) Systolic blood pressure (SBP) and UACR of pregnant mice injected with PBS, Sham mouse-derived or RUPP mouse-derived CD4+CD44+ T cells. Data were compared between groups using one-way ANOVA and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001). Data were compared between the two groups using the Student’s t-test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001).
-
Figure 6—source data 1
Raw data and detailed analysis for Figure 6.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig6-data1-v1.zip
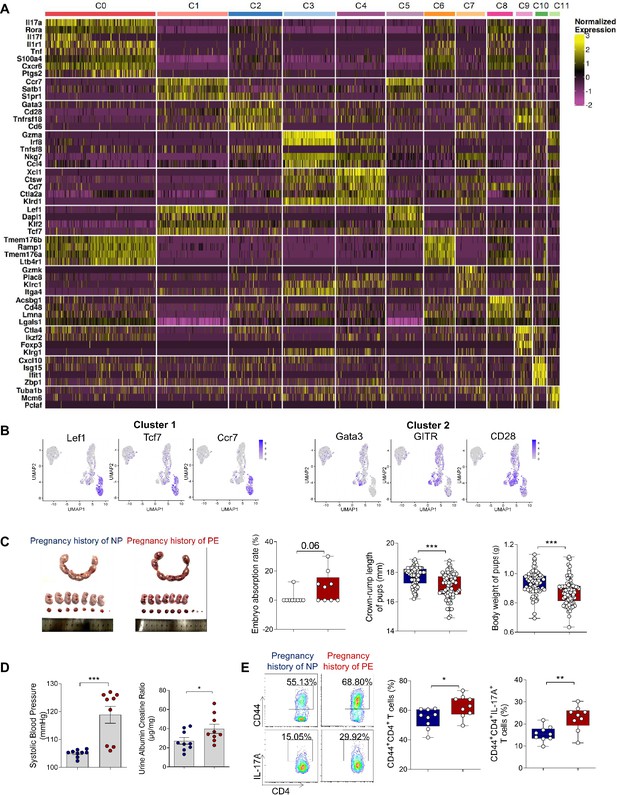
Memory-like Th17 cells may be associated with the recurrence of PE.
(A) Heatmap showing clustering analysis for markers distinguished 12 different clusters of T/NK cells. (B) Dot plot depicting gene that were significantly enriched in the differentially expressed genes in cluster 1 and cluster 2. (C) Embryo abortion rate of pregnant mice, body weight and crown-rump length of pups measured on day 18.5 of gestation. Blue represents mice with previous normal pregnancy (n = 9); red represents mice with previous pregnancy with PE (n = 9). (D) Systolic blood pressure (SBP) and UACR of second pregnant mice with a history of PE or NP in the first pregnancy measured on day 16.5 of gestation. (E) Frequencies of CD4+ CD44+ T cells and the levels of IL-17A in CD4+ CD44+ T cells in mice with a pregnancy history with NP and PE analyzed by flow cytometry. Data were compared using the Student’s t-test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001; NS, not significant). Data were compared between the two groups using the Student’s t-test and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001).
-
Figure 6—figure supplement 1—source data 1
Raw data and detailed analysis for Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig6-figsupp1-data1-v1.zip
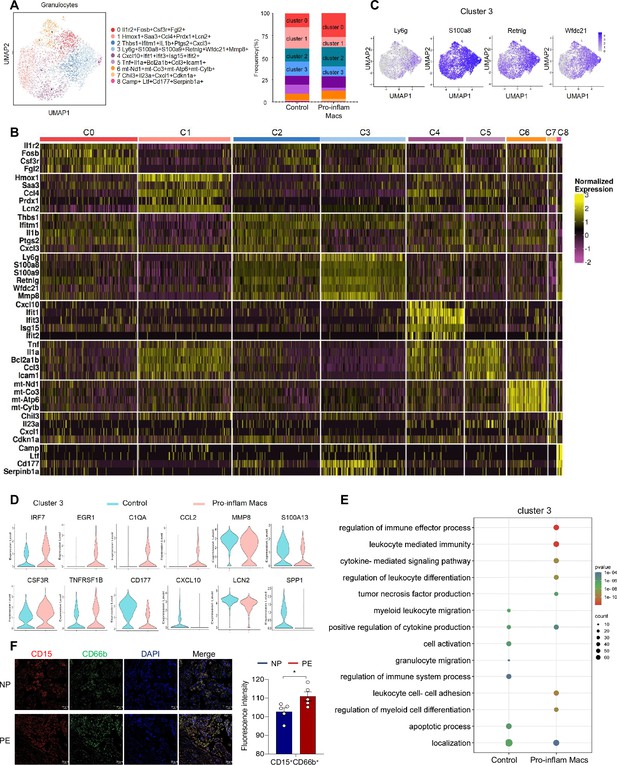
Identification of the granulocytes subsets in mice injected pro-inflam and anti-inflam Macs use single-cell RNA sequencing (scRNA-seq).
(A) Uniform Manifold Approximation and Projection (UMAP) maps showing the nine clusters of mouse granulocytes. Bar graph showing the frequencies of clusters of granulocytes in the two groups of mice. (B) Heatmap showing clustering analysis for markers distinguished nine different clusters of granulocytes cells. (C) UMAP maps showing the distribution of specific markers of cluster 3. (D) Dot plot depicting GO enrichment terms that were significantly enriched in the differentially expressed genes in cluster 3 from the pro-inflam Macs and the control groups. (E) Violin plot of specific differential gene expression in cluster 3 between the pro-inflam Macs and the control groups. (F) Immunofluorescence co-staining of CD15 (red), CD66b (green), and DAPI (blue) in frozen placental sections. Scale bar, 20 µm. Data were compared between the two groups using the Student’s t-test and represented as mean ± SEM (*p < 0.05).
-
Figure 6—figure supplement 2—source data 1
Raw data and detailed analysis for Figure 6—figure supplement 2.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig6-figsupp2-data1-v1.zip
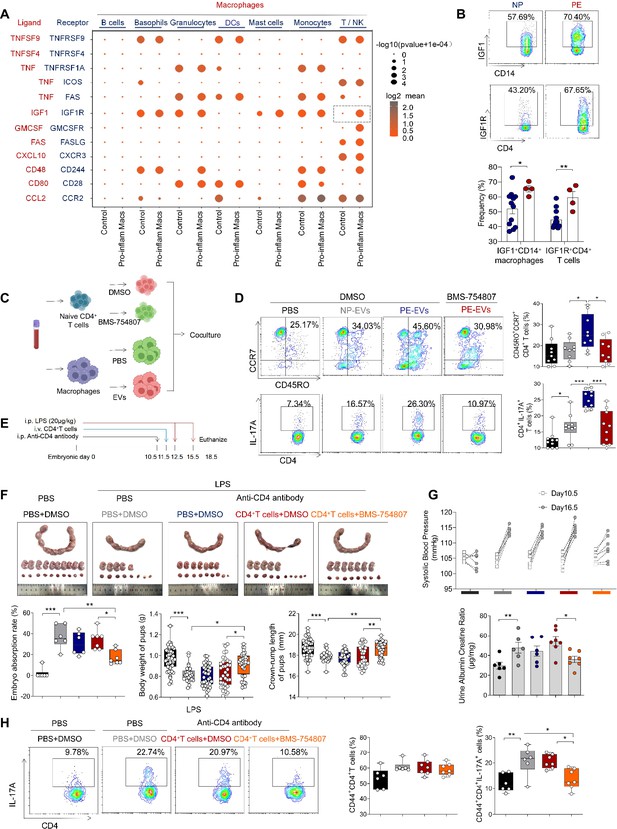
Pro-inflam Macs induce the generation of memory-like Th17 cells via IGF1–IGF1R.
(A) Signaling modules indicated by ligand–receptor pairing between macrophages and other types of immune cells at the maternal–fetal interface using CellPhoneDB. (B) Frequencies of IGF1+CD14+ and IGFIR+CD4+ cells in placentas of individuals with NP and PE (n = 12 in NP group, n = 4 in PE group). (C) Schematic of the experimental workflow to induce memory-like T cells in vitro. Macrophages, after incubating with PBS, NP-EVs or PE-EVs, were co-cultured with CD4+ naive T cells treated with DMSO or BMS-754807. Cells were isolated from human peripheral blood. (D) Frequencies of CD45RO+CCR7+Th17 cells. Black represents CD4+ naive T cells treated with DMSO; gray represents CD4+ naive T cells co-cultured with NP-EV-treated macrophages; blue represents CD4+ naive T cells co-cultured with PE-EV-treated macrophages; red represents CD4+ naive T cells treated with BMS-754807 before co-cultured with PE-EV-treated macrophages (n = 10 in each group). (E) Schematic of mice transferred CD4+ T cells treated with BMS-754807 or PBS. Anti-CD4 antibody was used to deplete CD4+ T cells in mice on day 10.5 of gestation. CD4+ T cells were transferred into mice on day 11.5 of gestation. 20 µg/kg lipopolysaccharide (LPS) was intraperitoneally injected on days 12.5 and 15.5 of gestation to induce a PE-like pregnant mice model. Mice were sacrificed on day 18.5 of gestation. (F) Embryo abortion rate of pregnant mice, body weight and crown-rump length of pups were measured on day 18.5 of gestation. Black represents the control group mice (n = 6); gray represents mice treated with LPS (20 μg/kg) to construct an animal model of PE (n = 6); blue represents anti-CD4 antibody treated PE mice (n = 6); red represents anti-CD4 antibody treated PE mice injected with CD4+ T cells with DMSO treatment (n = 7); orange represents anti-CD4 antibody treated PE mice injected with CD4+ T cells with BMS754807 treatment (n = 7). (G) Systolic blood pressure (SBP) and UACR of pregnant mice in the five groups. (H) The frequencies of CD4+ CD44+ IL-17A+ cells analyzed by flow cytometry. Data were compared between groups using one-way ANOVA and represented as mean ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001).
-
Figure 7—source data 1
Raw data and detailed analysis for Figure 7.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig7-data1-v1.zip
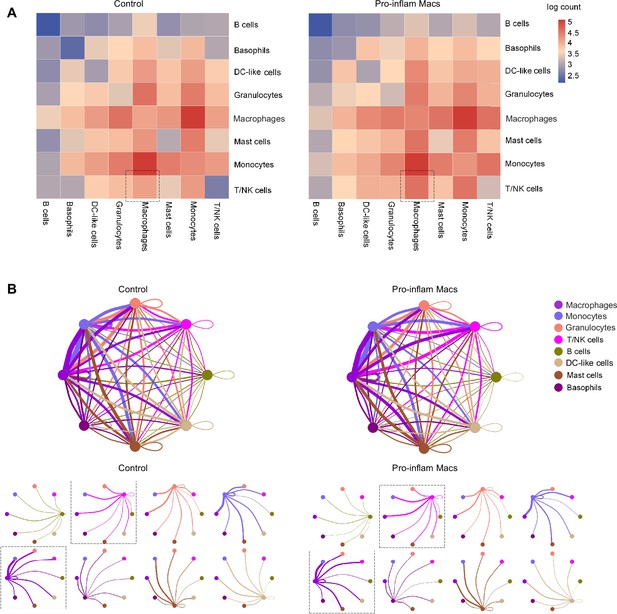
Cell–cell communications in immune cells when pro-inflam Macs accumulated at the maternal–fetal interface.
(A) Abundance of connections between different cell types at the maternal–fetal interface analyzed by CellPhoneDB. (B) Capacities for interactions between immune cells are showed. Each line indicates the ligands expressed by the cell population represented by the same color (labeled). The lines connect to cell types that express cognate receptors. Line thickness is proportional to the number of ligands when cognate receptors are present in the recipient cell type.
-
Figure 7—figure supplement 1—source data 1
Raw data and detailed analysis for Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/100002/elife-100002-fig7-figsupp1-data1-v1.zip
Tables
Details of the individuals included in the cytometry by time of flight (CyTOF).
| Parameters | NP (n = 9) | PE (n = 8) | GDM (n = 8) | GDM&PE (n = 7) | p value |
|---|---|---|---|---|---|
| Age (years) | 31.33 ± 2.108 | 31.75 ± 4.206 | 35.25 ± 3.419 | 32.43 ± 2.321 | 0.1008 |
| BMI (kg/m2) | 27.35 ± 1.740 | 30.84 ± 2.888 | 29.56 ± 2.689 | 30.16 ± 2.280 | 0.0518 |
| Gestational age (weeks) | 38.89 ± 0.558 | 37.39 ± 1.253 | 38.73 ± 0.419 | 36.22 ± 1.372 | <0.001 |
| Number of living children | 0.4444 ± 0.497 | 0.500 ± 0.500 | 1.125 ± 0.331 | 0.429 ± 0.495 | 0.0236 |
| Previous abortions | 0.7778 ± 1.030 | 1.000 ± 0.866 | 1.000 ± 1.000 | 1.714 ± 2.050 | 0.5848 |
| Mean systolic blood pressure | 109.7 ± 5.400 | 151 ± 7.225 | 106.9 ± 2.848 | 150.7 ± 9.161 | <0.0001 |
Cytometry by time of flight (CyTOF) antibody panel used for analyzing placentas from individuals with normal pregnancy (NP), preeclampsia (PE), gestational diabetes mellitus (GDM), and GDM complicated with PE (GDM&PE).
| Target | Metal tag |
|---|---|
| CD45 | HI30 |
| CD3 | UCHT1 |
| CD68 | Y1/82A |
| CD56 | NCAM16.2 |
| gd TCR | 5A6.E9 |
| CD19 | HIB19 |
| CCR6 | G034E3 |
| CD38 | HIT2 |
| CD103 | B-Ly7 |
| CD39 | A1 |
| CXCR3 | G025H7 |
| PD-L1 | 29E.2A3 |
| PD-1 | EH12.2H7 |
| CD11C | BU15 |
| CD107a | H4A3 |
| pAKT | D9E |
| CCR4 | L291H4 |
| TIGIT | A15153G |
| CD206 | 15-2 |
| Helios | 22F6 |
| CD28 | CD28.2 |
| pSTAT3 | 4/p-Stat3 |
| GITR | 110416 |
| CD33 | WM53 |
| CTLA-4 | BN13 |
| FOXP3 | PCH101 |
| CD163 | GHI/61 |
| CD45RA | HI100 |
| ICOS | C398.4A |
| CD69 | FN50 |
| CCR7 | G043H7 |
| NKG2A | 131411 |
| CD15 | W6D3 |
| TIM-3 | F38-2E2 |
| CD86 | Fun-1 |
| HLA-DR | L243 |
| Granzyme B | QA16A02 |
| CD14 | M5E2 |
| pS6 | A17020B |
| CD127 | A019D5 |
| CD4 | RPA-T4 |
| CD8 | RPA-T8 |
| CD11b | M1/70 |
Expression levels of markers identified in each immune subset in the placentas.
| B cells | CD4+ T | CD8+ T | gdTCR | NK cells | Macrophage | Granulocyte | MDSC | Dendritic cells | |
|---|---|---|---|---|---|---|---|---|---|
| CD45 | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ | ++ |
| CD3 | - | ++ | ++ | - | - | - | - | - | - |
| CD68 | + | + | + | + | + | ++ | + | ++ | ++ |
| CD56 | - | - | - | - | + | - | - | - | - |
| gd TCR | ++ | - | - | ++ | - | - | - | - | - |
| CD19 | ++ | - | - | ++ | - | - | - | - | - |
| CCR6 | + | - | - | - | - | - | - | - | - |
| CD38 | ++ | ++ | + | + | ++ | ++ | - | - | + |
| CD103 | - | - | - | - | - | - | - | - | - |
| CD39 | ++ | - | - | - | + | ++ | + | + | + |
| CXCR3 | + | + | + | + | + | + | - | + | ++ |
| PD-L1 | - | - | - | - | - | - | - | + | - |
| PD-1 | - | - | - | - | - | - | - | - | - |
| CD11C | + | - | - | - | + | ++ | ++ | + | ++ |
| CD107a | + | - | - | - | - | + | - | + | + |
| pAKT | + | + | + | + | + | ++ | + | + | + |
| CCR4 | - | - | - | - | - | - | - | - | - |
| TIGIT | - | - | - | - | - | - | - | - | - |
| CD206 | - | - | - | - | - | + | - | + | - |
| Helios | + | + | + | + | + | + | + | + | + |
| CD28 | - | + | + | + | - | - | - | - | - |
| pSTAT3 | + | + | - | + | - | + | + | - | - |
| GITR | - | - | - | - | - | - | - | - | - |
| CD33 | - | - | - | - | - | + | - | + | - |
| CTLA-4 | - | - | - | - | - | - | - | - | - |
| FOXP3 | - | - | - | - | - | - | - | - | - |
| CD163 | - | - | - | - | - | - | - | - | - |
| CD45RA | ++ | + | + | + | ++ | - | - | - | - |
| ICOS | - | + | + | - | - | - | - | - | - |
| CD69 | + | + | + | + | ++ | + | - | ++ | - |
| CCR7 | + | ++ | + | + | - | - | - | + | - |
| NKG2A | - | - | - | - | - | - | - | - | - |
| CD15 | - | - | - | - | - | - | ++ | - | - |
| TIM-3 | - | - | - | - | - | - | - | - | - |
| CD86 | - | - | - | - | - | ++ | - | - | + |
| HLA-DR | ++ | - | + | + | - | ++ | + | - | ++ |
| Granzyme B | + | - | - | + | ++ | + | + | - | - |
| CD14 | - | - | - | - | - | ++ | - | - | - |
| pS6 | - | - | - | - | - | - | - | - | - |
| CD127 | - | ++ | + | - | - | - | - | - | - |
| CD4 | - | ++ | - | - | - | - | - | - | - |
| CD8 | - | - | ++ | - | - | - | - | - | - |
| CD11b | - | - | - | + | + | ++ | ++ | - | + |
The marker profile of PE-specific immune subsets.
| Memory-like CD4+ T cells | Memory-like CD8+ T cells | gMDSCs | Pro-inflammatory macrophages | Anti-inflammatory macrophages | |
|---|---|---|---|---|---|
| Cluster 08 of CD4+ T cells | Cluster 02 of CD8+ T cells | Cluster 12 ofCD3−CD11b+ cells | Cluster 23 ofCD3−CD11b+ cells | Cluster 25 ofCD3−CD11b+ cells | |
| CD45 | ++ | ++ | ++ | ++ | ++ |
| CD3 | ++ | ++ | - | - | - |
| CD68 | - | - | + | ++ | ++ |
| CD56 | - | - | - | - | - |
| gd TCR | - | - | - | - | - |
| CD19 | - | - | - | - | - |
| CCR6 | - | - | - | - | - |
| CD38 | ++ | ++ | - | + | + |
| CD103 | - | - | - | + | + |
| CD39 | + | - | + | - | - |
| CXCR3 | + | + | - | - | - |
| PD-L1 | - | - | - | - | - |
| PD-1 | - | - | - | - | - |
| CD11C | - | - | ++ | ++ | ++ |
| CD107a | - | - | - | + | + |
| pAKT | + | + | - | - | - |
| CCR4 | - | - | - | - | - |
| TIGIT | - | - | - | - | - |
| CD206 | - | - | + | - | + |
| Helios | + | + | - | - | - |
| CD28 | + | + | - | - | - |
| pSTAT3 | - | - | - | - | - |
| GITR | - | - | - | - | - |
| CD33 | - | - | - | - | + |
| CTLA-4 | - | - | - | - | - |
| FOXP3 | - | - | - | - | - |
| CD163 | - | - | - | - | - |
| CD45RA | - | - | - | - | - |
| ICOS | + | + | - | - | - |
| CD69 | + | - | - | - | - |
| CCR7 | ++ | ++ | - | - | - |
| NKG2A | - | - | - | - | - |
| CD15 | - | - | ++ | - | - |
| TIM-3 | - | - | - | - | - |
| CD86 | - | - | - | + | + |
| HLA-DR | - | - | + | + | ++ |
| Granzyme B | - | - | + | - | - |
| CD14 | - | - | - | + | + |
| pS6 | - | - | - | - | - |
| CD127 | + | - | - | - | - |
| CD4 | ++ | - | - | - | - |
| CD8 | - | ++ | - | - | - |
| CD11b | - | - | ++ | ++ | ++ |
Details of the individual with NP or PE included in the study.
| Parameters | NP (n = 30) | PE (n = 30) | p value |
|---|---|---|---|
| Age (years) | 31.13 ± 2.9747 | 31.50 ± 4.1693 | 0.0741 |
| BMI (kg/m2) | 27.24 ± 2.8048 | 29.48 ± 3.9295 | 0.0155 |
| Gestational age (weeks) | 38.68 ± 0.6038 | 35.53 ± 3.29 | <0.0001 |
| Number of living children | 0.6000 ± 0.4900 | 0.4667 ± 0.6182 | 0.3665 |
Details of the individual with early- or late-onset preeclampsia included in the study.
| Parameters | Early-onset preeclampsia (n = 11) | Late-onset preeclampsia (n = 19) | p value |
|---|---|---|---|
| The gestational week when blood pressure begins to rise | 28.82 ± 2.8220 | 36.05 ± 1.129 | <0.0001 |
| Age (years) | 32.45 ± 4.6768 | 32.36 ± 3.8472 | 0.9568 |
| BMI (kg/m2) | 29.64 ± 2.6840 | 28.71 ± 4.4506 | 0.5388 |
| Gestational age (weeks) | 33.45 ± 3.7887 | 36.77 ± 1.2171 | 0.0014 |
| Number of living children | 0.36 ± 0.5045 | 0.36 ± 0.4956 | 0.98 |





