β-1,6-Glucan plays a central role in the structure and remodeling of the bilaminate fungal cell wall
Figures
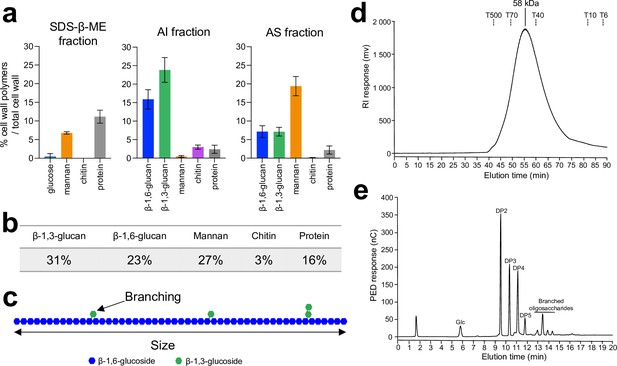
Analysis of C. albicans cell wall β-1,6-glucans.
(a) Percentages of cell wall polymers on total cell wall, distributed by fractions: sodium-dodecyl-sulfate-β-mercaptoethanol (SDS-β-ME), alkali-insoluble (AI), and alkali-soluble (AS). Cells were grown in synthetic dextrose (SD) medium at 37°C. Means and standard deviations were calculated from three independent experiments. (b) Table of the mean percentages of each polymer in the cell wall from three independent experiments. (c) Diagram of β-1,6-glucan structure. In blue are represented glucose residues linked in β-1,6 and in green glucose residues linked in β-1,3. According to nuclear magnetic resonance (NMR) analysis and high-performance anion exchange chromatography (HPAEC) after endo-β-1,6-glucanase digestion (Figure 1—figure supplement 1), based on three independent experiments, an average of 6.4% (± 0.5%) of glucose units of the main chain are substituted by a single glucose residue (88–90%) or a laminaribiose (10–12%). (d) Gel filtration analysis on a Superdex 200 column of β-1,6-glucan released by endo-β-1,3-glucanase digestion. The column was calibrated with dextrans (Tx: × kDa). (e) HPAEC analysis of the digestion products of the AI fraction treated with an endo-β-1,6-glucanase. Chromatographs in (d) and (e) are representative of three independent experiments. PED, pulsed electrochemical detector; nC, nanocoulombs; RI, refractive index; mV, millivolt; DP, degree of polymerization; Glc, glucose.
-
Figure 1—source data 1
Raw data for panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig1-data1-v1.xlsx
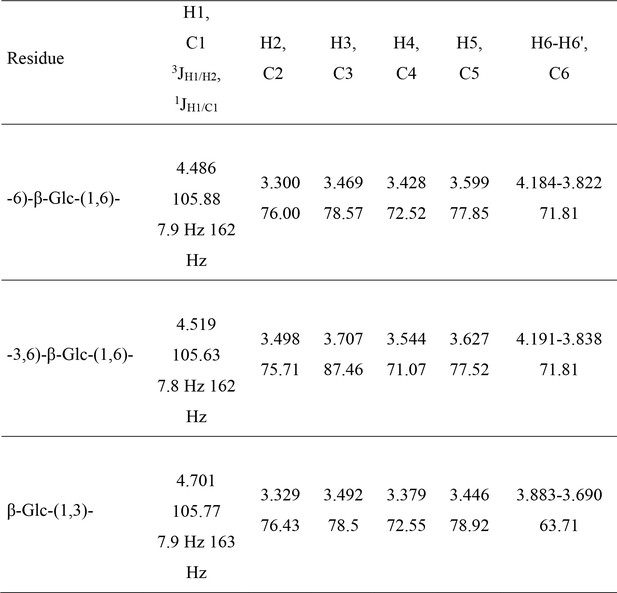
1H and 13C NMR resonance assignments, 3JH1/H2 and 1JH1/C1 coupling constants of the monosaccharide residues of cell wall β-1,6-glucan purified from the alkali-insoluble (AI) fraction.
Chemical shifts are expressed in ppm and coupling constants in Hz.
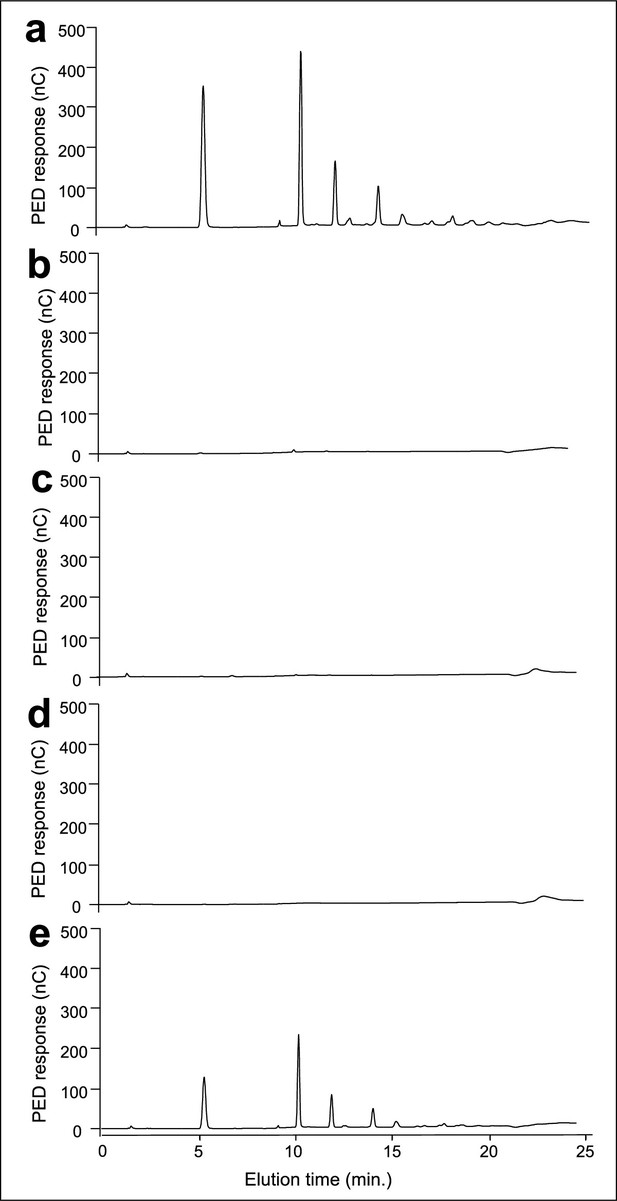
Cell disruption is essential to eliminate glycogen in alkali-insoluble (AI) and alkali-soluble (AS) fractions.
High-performance anion exchange chromatography (HPAEC) analysis of oligosaccharides released by α-amylase enzymatic digestion of AI and AS fractions. (a) Control: glycogen, (b) AI fraction obtained after biomass cell disruption, (c) AI fraction from biomass with no cell disruption, (d) AS fraction obtained after biomass cell disruption, and (e) AS fraction from biomass with no cell disruption. PED, pulsed electrochemical detector; nC, nanocoulomb.
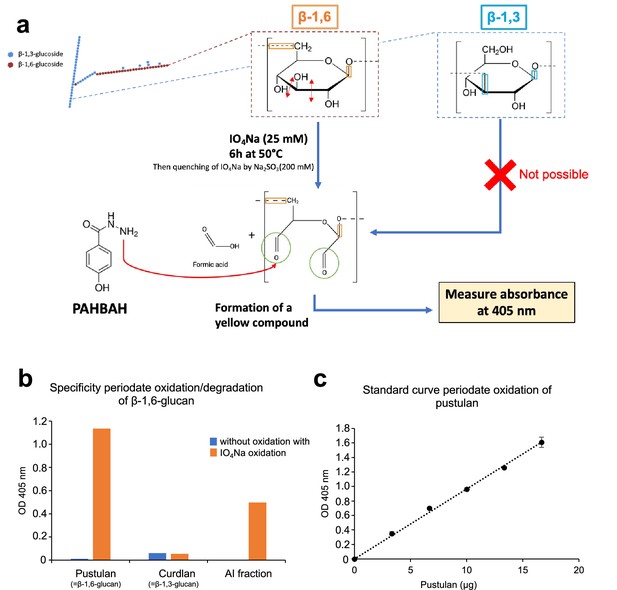
Quantification methods of β-1,6-glucans in alkali-insoluble (AI) fractions.
(a) The specific oxidation of β-1,6-glucans of the AI fraction by periodate was used for quantification. Briefly, IO4Na splits bonds between vivinal carbons bearing hydroxyl groups (only present in β-1,6-glucoside), which leads to the formation of aldehydes, which can react with 4-hydroxybenzhydrazide (PAHBAH) to form a yellow compound measurable by absorbance at OD = 405 nm. (b) Specificity of the periodate oxidation method for β-1,6-glucans (pustulan). The method is specific for β-1,6-glucans (pustulan and AI fraction) and inactive on β-1,3-glucans (curdlan). (c) Linearity of β-1,6-glucan assay after periodate oxidation. We showed that the response of the method described in (a) is proportional from 0 to 20 µg of pustulan.
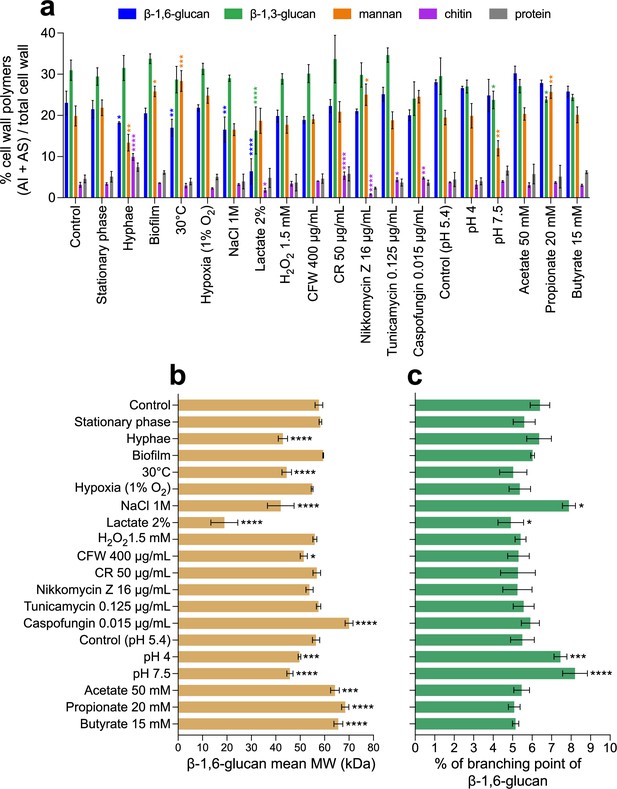
Comparative analyses of cell wall β-1,6-glucans produced in various environmental conditions.
(a) Percentages of cell wall polymers (alkali-insoluble [AI] + alkali-soluble [AS] fractions) on total cell wall. Cells were grown in liquid synthetic medium at 37°C under different conditions, as specified in ‘Materials and methods’. (b) β-1,6-Glucan mean molecular weight (MW). Average molecular weight was estimated by gel filtration chromatography on a Superdex 200 column. (c) Branching rate of β-1,6-glucans. Branching rate was determined by high-performance anion exchange chromatography (HPAEC) after digestion of the AI fraction by an endo-β-1,6-glucanase (% expressed as number of glucose units of the main chain that are substituted by a side chain). For (a–c), means and standard deviations from three independent replicate experiments are shown. All data were compared to the control conditions and were analyzed by one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 2—source data 1
Raw data for panels a, b and c.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig2-data1-v1.xlsx
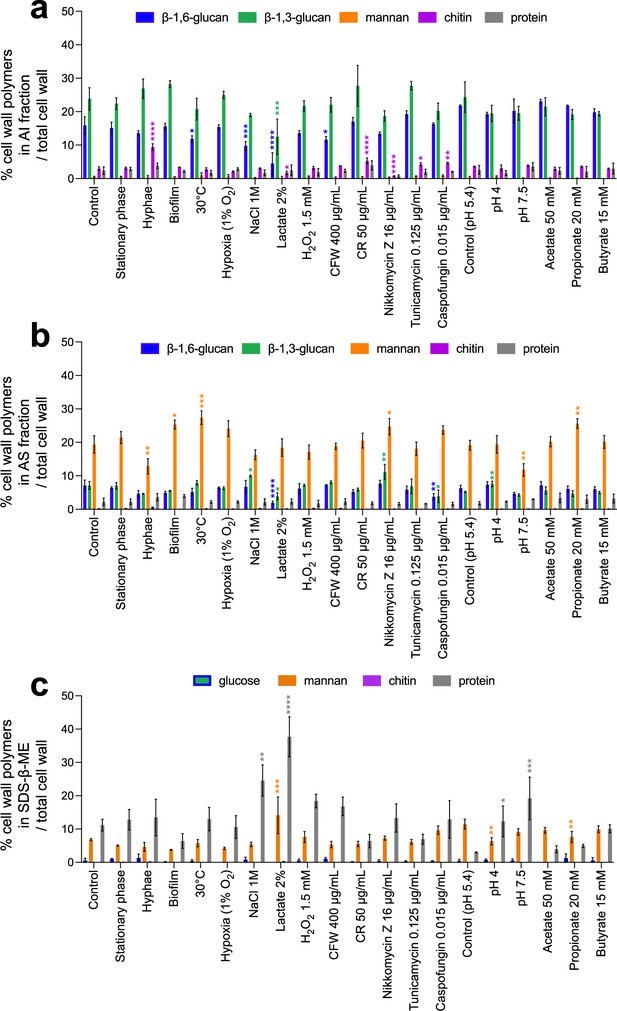
Global cell wall composition produced by C. albicans in different environmental conditions.
Results are represented as the percentage of each polymer on total cell wall in alkali-insoluble (AI) fraction (a), alkali-soluble (AS) fraction (b), and sodium-dodecyl-sulfate-β-mercaptoethanol (SDS-β-ME) fraction (c). Means and standard deviations were obtained from three independent replicate experiments. All data were compared to the control conditions and were analyzed using one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 2—figure supplement 1—source data 1
Raw data for panels a, b and c.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig2-figsupp1-data1-v1.xlsx
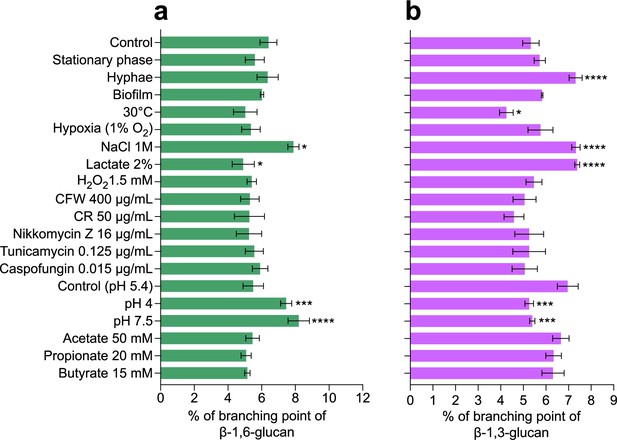
Branching rates of β-1,6-glucans and β-1,3-glucans produced by C. albicans under various environmental conditions.
(a) Branching rate of β-1,6-glucans from alkali-insoluble (AI) fraction. The branching rate was estimated by high-performance anion exchange chromatography (HPAEC) after digestion by an endo-β-1,6-glucanase. (b) Branching rate of β-1,3-glucans from AI fraction, The branching rate was estimated by HPAEC after digestion by an endo-β-1,3-glucanase. Means and standard deviations from three independent replicate experiments are shown. All data were compared to the control conditions and were analyzed using one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 2—figure supplement 2—source data 1
Raw data for panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig2-figsupp2-data1-v1.xlsx
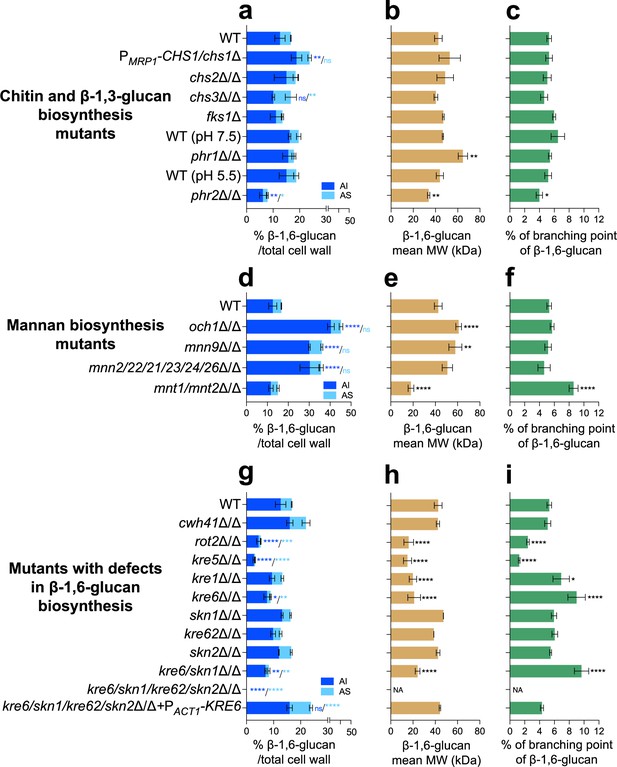
Comparative analysis of β-1,6-glucan content and structure produced by cell wall mutants.
(a, d, g) Percentages of cell wall β-1,6-glucans (alkali-insoluble [AI] and alkali-soluble [AS] fractions) on total cell wall. (b, e, h) β-1,6-Glucans mean molecular weight (MW). (c, f, i) Branching rate of β-1,6-glucans. Cells were grown in liquid synthetic dextrose (SD) medium at 30°C. Means and standard deviations from three independent replicate experiments are shown. All data were compared to the control conditions and were analyzed by one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001; ns, nonsignificant; NA, nonapplicable.
-
Figure 3—source data 1
Raw data for panels a to i.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig3-data1-v1.xlsx
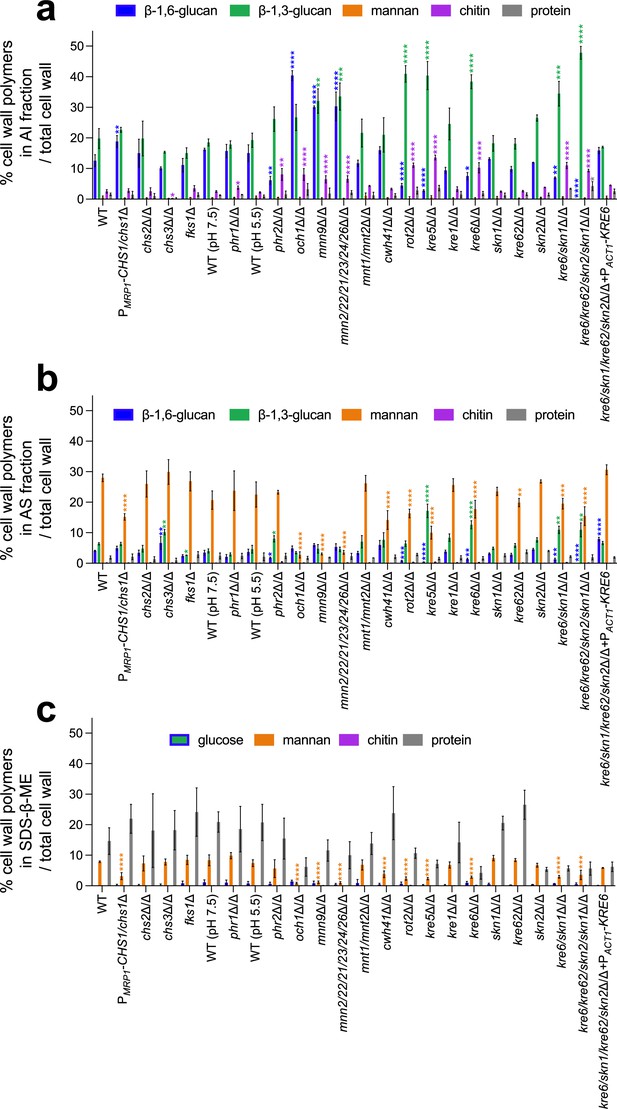
Cell wall composition of C. albicans mutants.
Results are represented as the percentage of each polymer on total cell wall in alkali-insoluble (AI) fraction (a), alkali-soluble (AS) fraction (b), and sodium-dodecyl-sulfate-β-mercaptoethanol (SDS-β-ME) fraction (c). Means and standard deviations were obtained from three independent replicate experiments. All data were compared to the parental strain SC5314 and were analyzed using one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 3—figure supplement 1—source data 1
Raw data for panels a, b and c.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig3-figsupp1-data1-v1.xlsx
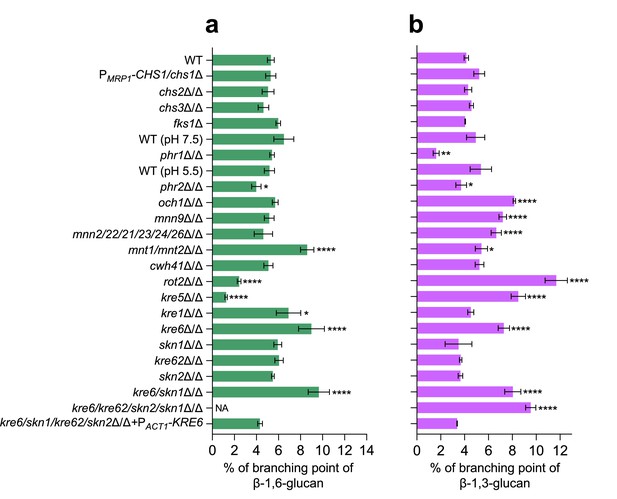
Branching rates of β-1,6-glucans and β-1,3-glucans produced by different cell wall mutants of C. albicans.
(a) Branching rate of β-1,6-glucans from alkali-insoluble (AI) fraction. The branching rate was estimated by high-performance anion exchange chromatography (HPAEC) after digestion by an endo-β-1,6-glucanase. (b) Branching rate of β-1,3-glucans from AI fraction. The branching rate was estimated by HPAEC after digestion by an endo-β-1,3-glucanase. Means and standard deviations from three independent replicate experiments are shown. All data were compared to the control conditions and were analyzed using one-way ANOVA with Dunnett’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001; NA, nonapplicable.
-
Figure 3—figure supplement 2—source data 1
Raw data for panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig3-figsupp2-data1-v1.xlsx
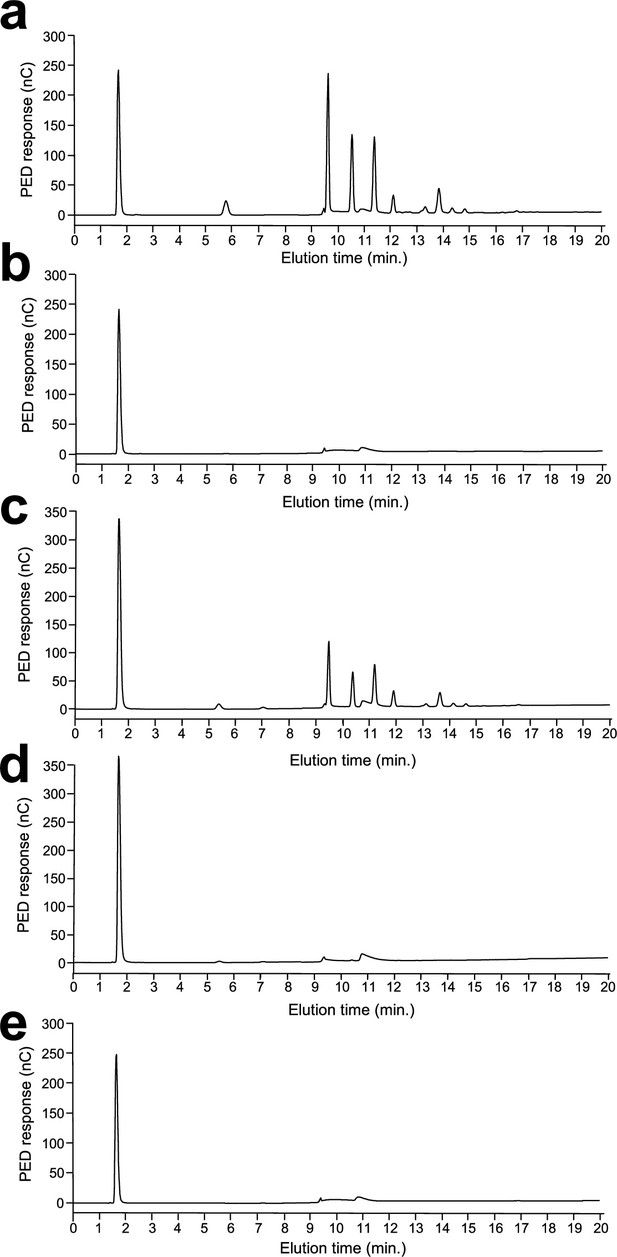
Absence of β-1,6-glucans in the cell wall of the quadruple kre6/kre62/skn2/skn1Δ/Δ mutant.
High-performance anion exchange chromatography (HPAEC) analysis of oligosaccharides released by the endo-β-(1,6)-glucanase digestions of alkali-insoluble (AI) fraction of WT (a), AI fraction of kre6/kre62/skn2/skn1Δ/Δ (b), alkali-soluble (AS) fraction of WT (c), AS fraction of kre6/kre62/skn2/skn1Δ/Δ (d), and water (e). Experiments were performed in triplicates. PED, pulsed electrochemical detector; nC, nanocoulomb.
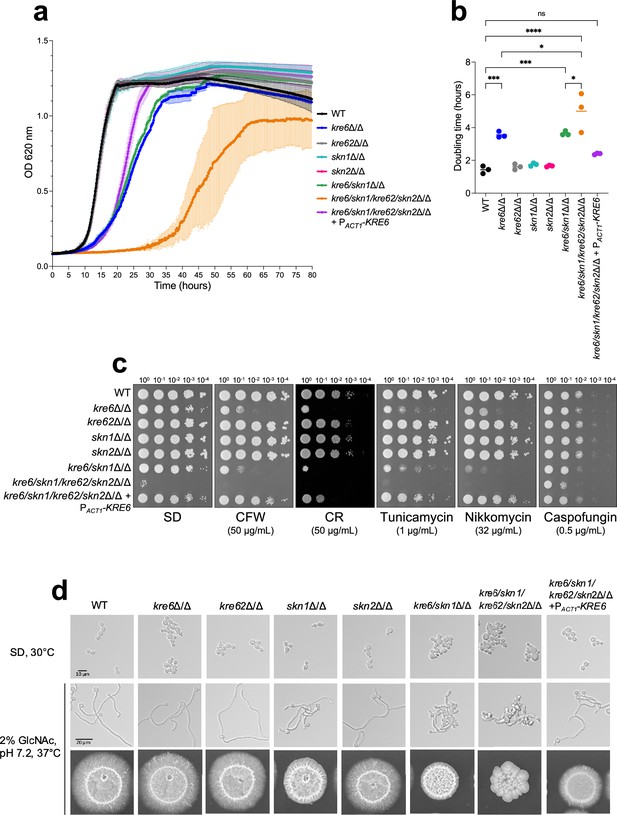
Phenotypic characterization of KRE6 family mutants: growth kinetics, drug susceptibility, and filamentation.
(a) Kinetic curve of all strains grown in liquid synthetic dextrose (SD) medium, 30°C. Optical density at 620 nm was measured every 10 min during 80 hr by TECAN SUNRISE. Means and standard deviations were calculated from three independent experiments. (b) Doubling time of each strain was determined from three independent replicates. Statistical analyses were performed with one-way ANOVA with Tukey’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001; ns, nonsignificant. (c) Spotting test of tenfold serial dilution of yeast cells of all strains on SD medium, 30°C, 48 hr, with cell wall disturbing agents (CR, Congo Red; CFW, Calcofluor White) or drugs (nikkomycin, tunicamycin, caspofungin). These results are representative of three independent experiments. Pictures were taken with a Phenobooth (Singer Instruments). (d) Filamentation assay of all strains. Top row: growth in liquid SD medium at 30°C; middle panels: growth in liquid YNB medium + 2% GlcNAc, buffered at pH 7.2 at 37°C during 6 hr. Pictures were taken using an Olympus IX 83 microscope, ×40 objective. Bottom row: cells were grown on agar YNB + 2% GlcNAc, buffered at pH 7.2, at 37°C for 6 days.
-
Figure 4—source data 1
Raw data for panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig4-data1-v1.xlsx
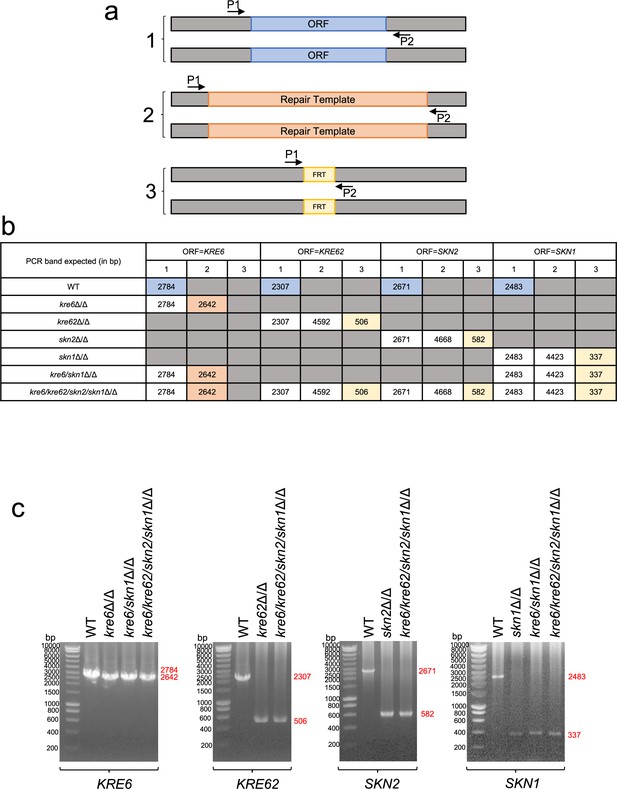
Control PCR control of mutants obtained in this study.
(a) Principle of PCR check done in (b) and (c). P1 and P2 are PCR diagnostic primers. ORF = KRE6, KRE62, SKN2, or SKN1. Repair Templates contain either HygB or a SAT1-Flipper cassette for selection. FRT correspond to the scar after SAT1-Flipper cassette excision. (b) Table of expected PCR bands according to mutant. (c) PCR bands obtained.
-
Figure 4—figure supplement 1—source data 1
PDF file containing original gels PCR for Figure 4—figure supplement 1c, indicating the relevant bands.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig4-figsupp1-data1-v1.zip
-
Figure 4—figure supplement 1—source data 2
Original files for gels PCR analysis displayed in Figure 4—figure supplement 1c.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig4-figsupp1-data2-v1.zip
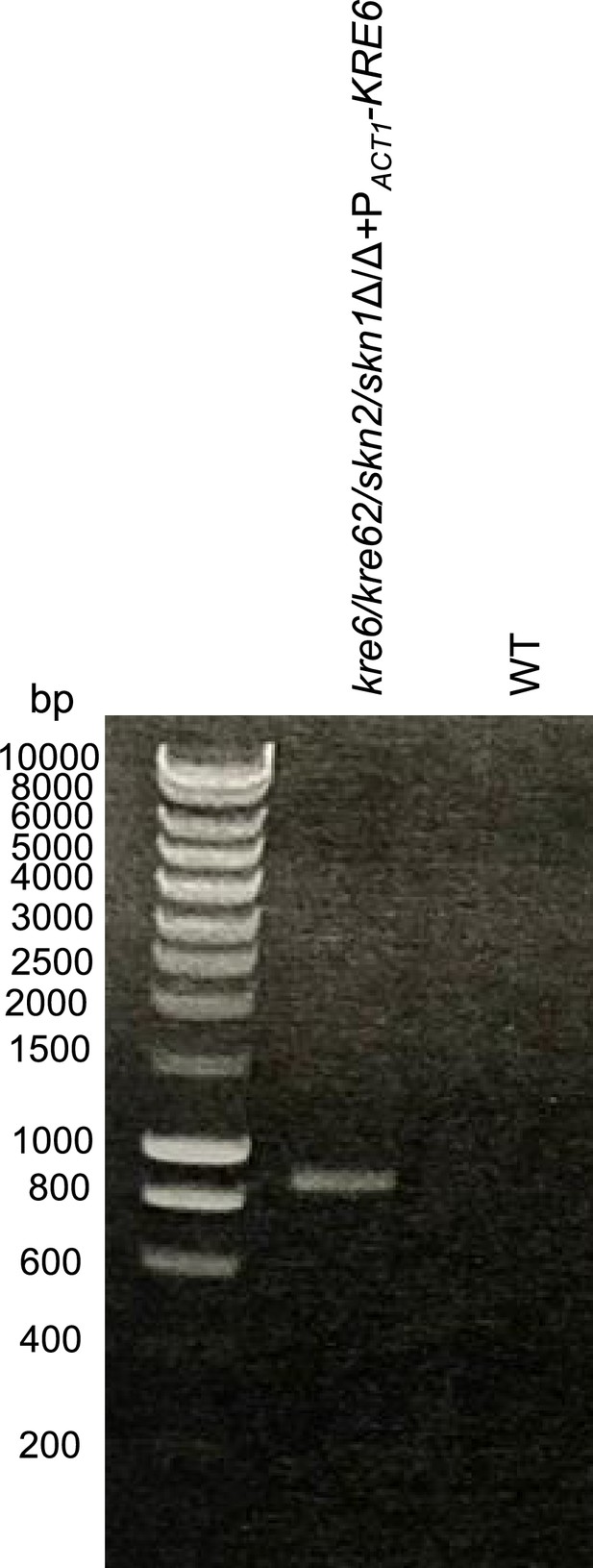
Control PCR of the quadruple mutant complemented for KRE6 (kre6/kre62/skn2/skn1Δ/Δ+PACT-KRE6).
KRE6 was reintegrated at the RPS1 locus under the control of ACT1 promoter, using SAT1 as a selection marker.
-
Figure 4—figure supplement 2—source data 1
PDF file containing original gel PCR for Figure 4—figure supplement 2, indicating the relevant bands.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig4-figsupp2-data1-v1.zip
-
Figure 4—figure supplement 2—source data 2
Original files for gel PCR analysis displayed in Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig4-figsupp2-data2-v1.zip
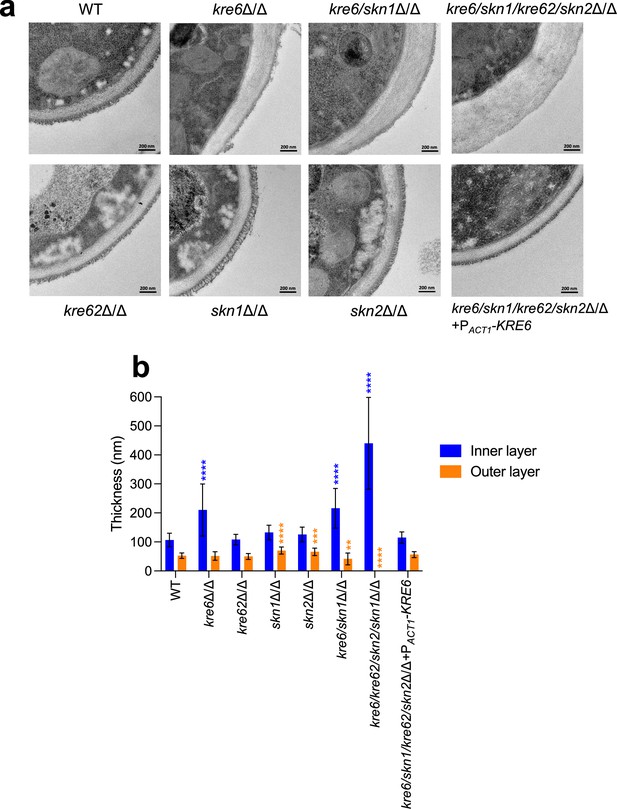
Cell wall electron microscopy observations of KRE6 simple and multiple mutants.
(a) Representative transmission electron microscopy images of the cell wall of each strains. After culture, cells were fixed and high-pressure frozen and freeze substituted with Spurr resin. Sections were cut and stained and then pictures were taken with a Tecnai Spirit 120Kv TEM microscope. Scale bar = 200 nm. (b) Measurement of the inner and outer cell wall layers of the mutants. Means and standard deviations are represented. 37–40 measurements were performed randomly on 7–13 cells. Statistical analyses were performed with one-way ANOVA with Tukey’s multiple comparisons test: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 5—source data 1
Raw data for panel b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig5-data1-v1.xlsx
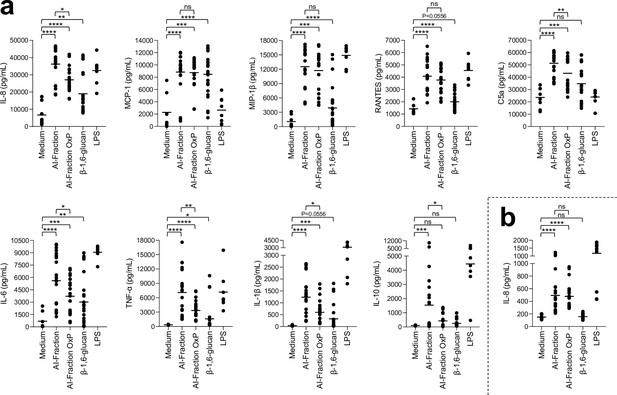
Stimulation of peripheral blood mononuclear cells (PBMCs) and neutrophils in vitro by cell wall fractions and purified β-1,6-glucans from C. albicans.
Cytokines, chemokines, or acute-phase proteins (IL-8, MCP-1, IL-6, MIP-1β, IL-1β, TNF-α, RANTES, C5a, IL-10) concentrations in culture supernatants of PBMCs (a) and neutrophils (b) stimulated by cell wall fractions of C. albicans (AI-Fraction, AI-Fraction OxP, and β-1,6-glucan) at 25 µg/mL or LPS (positive control, 0.1 µg/mL). PBMCs and neutrophils were isolated from healthy human donors (n = 8). Three independent batches of each fractions were used. Means are represented and data were analyzed using nonparametric Friedman test with Dunn’s multiple comparisons: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001; ns, nonsignificant.
-
Figure 6—source data 1
Raw data of panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-data1-v1.xlsx
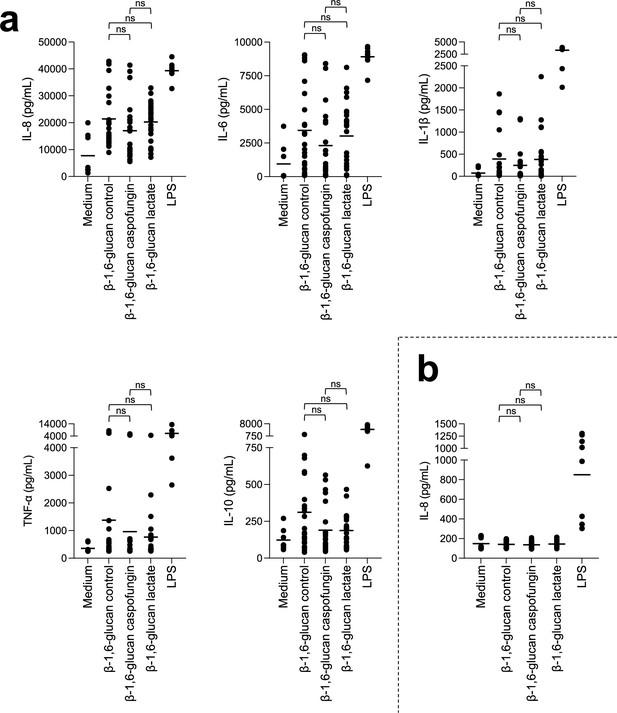
Stimulation of peripheral blood mononuclear cells (PBMCs) and neutrophils in vitro by β-1,6-glucan with different size from C. albicans.
Cytokines, chemokines, or acute-phase proteins (IL-8, MCP-1, IL-6, MIP-1β, IL-1β, TNF-α, RANTES, C5a, IL-10) concentrations in culture supernatants of PBMCs (a) or neutrophils (b) stimulated by different β-1,6-glucans from C. albicans at 25 µg/mL or LPS (positive control, 0.1 µg/mL). β-1,6-Glucans were isolated from cell wall alkali-insoluble (AI) fraction of C. albicans grown either at 37°C in synthetic dextrose (SD) medium (control, β-1,6-glucan size = 58 kDa), or in the presence of caspofungin at sublethal concentration 0.015 µg/mL (β-1,6-glucan size = 70 kDa) or in the presence of 2% lactate as sole carbon source (β-1,6-glucan size = 19 kDa). PBMCs and neutrophils were isolated from healthy human donors (n = 8). Three independent batches of the different fractions were used. Means are represented and data were analyzed using nonparametric Friedman test with Dunn’s multiple comparisons: ns, nonsignificant.
-
Figure 6—figure supplement 1—source data 1
Raw data of panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp1-data1-v1.xlsx
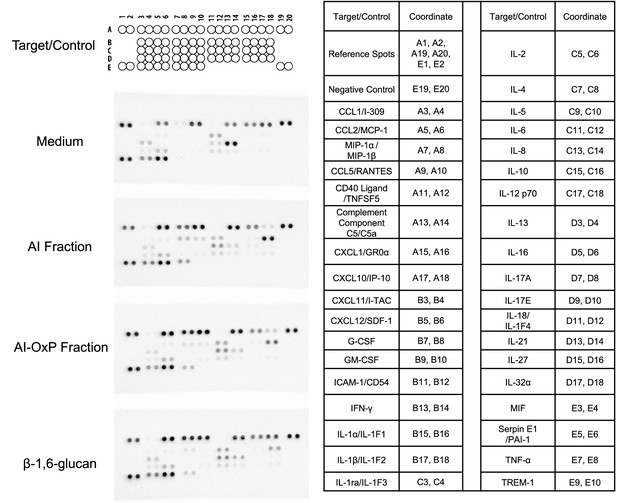
Human proteome profiler done with culture supernatant from peripheral blood mononuclear cells (PBMCs) stimulated with C. albicans cell wall fractions.
Left: membrane blots obtained after incubation with supernatants from PBMCs cultures stimulated by different C. albicans cell wall fractions: AI, AI-OxP, or β-1,6-glucans. Incubation with the culture medium was used as a control (top). Right: coordinate of each protein (cytokines, chemokines, acute-phase proteins) detected on the membranes. The experiment was performed once using a pool of 24 supernatants from the stimulation of PBMCs isolated from eight healthy donors, each stimulated with three independent batches of fractions.
-
Figure 6—figure supplement 2—source data 1
PDF file containing original blots for Figure 6—figure supplement 2, indicating the relevant bands.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp2-data1-v1.zip
-
Figure 6—figure supplement 2—source data 2
Original files for blots analysis displayed in Figure 6—figure supplement 2.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp2-data2-v1.zip
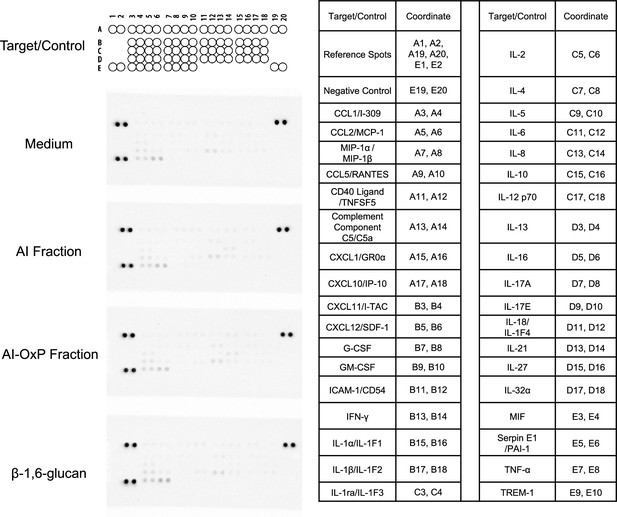
Human proteome profiler done with culture supernatant from neutrophils stimulated with C. albicans cell wall fractions.
Left: membrane blots obtained after incubation with supernatants from neutrophils cultures stimulated by different C. albicans cell wall fractions: AI, AI-OxP, or β-1,6-glucans; incubation with the culture medium was used as a control (top). Right: coordinate of each protein (cytokines, chemokines, acute-phase proteins) detected on the membranes. The experiment was performed once using a pool of 24 supernatants from the stimulation of neutrophils isolated from eight healthy donors, each stimulated with three independent batches of fractions.
-
Figure 6—figure supplement 3—source data 1
PDF file containing original blots for Figure 6—figure supplement 3, indicating the relevant bands.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp3-data1-v1.zip
-
Figure 6—figure supplement 3—source data 2
Original files for blots analysis displayed in Figure 6—figure supplement 3.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp3-data2-v1.zip
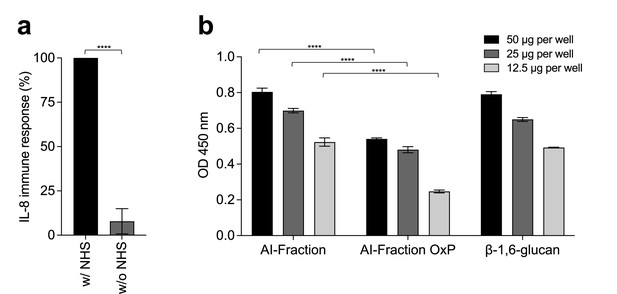
β-1,6-Glucan from C. albicans activates complement system.
(a) Normal human serum (NHS) enhances the immunostimulatory capacity of β-1,6-glucan from C. albicans. Peripheral blood mononuclear cells (PBMCs) isolated from healthy human donors (n = 2) were stimulated with three independent batches of β-1,6-glucan at 25 µg/mL with (w/) or without (w/o) NHS (10%). Immune response was analyzed by measuring IL-8 released into the culture medium. Means are represented and data were analyzed with an unpaired parametric t-test: ****p<0.0001. (b) Complement factor C3b binds to β-1,6-glucan purified from C. albicans cell wall. Three cell wall fractions (AI, AI-OxP, and β-1,6-glucan) from C. albicans were coated on microtiter plates at 50 µg, 25 µg, or 12.5 µg per well. Human normal serum, diluted in Gelatin-Veronal Buffer (GVB), was added to activate complement pathways. The amount of deposited C3b on each fraction (=level on complement activation) was determined by using anti-human C3b and peroxidase-conjugated anti-mouse IgG antibodies. 3,3’,5,5’-Tetramethylbenzidine (TMB) was used as the peroxidase substrate and the reaction was stopped with 4% H2SO4 and optical density (OD) was measured at 450 nm. The experiment was done with three independent batches of each cell wall fractions. Blank value was subtracted from the values presented. Statistical analyses were performed with one-way ANOVA with Tukey’s multiple comparisons test: ****p<0.0001.
-
Figure 6—figure supplement 4—source data 1
Raw data of panels a and b.
- https://cdn.elifesciences.org/articles/100569/elife-100569-fig6-figsupp4-data1-v1.xlsx
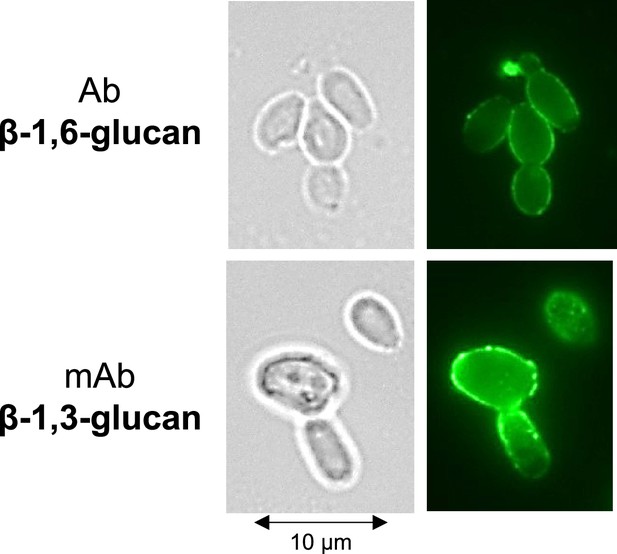
Exposure of β-1,6-glucans and β-1,3-glucans at the cell surface of C. albicans SC5314.
Cells were cultured in synthetic dextrose (SD) at 37°C. β-Glucan exposure was detected by a polyclonal rabbit anti-β-1,6-glucan serum (top panel) and or monoclonal anti-β-1,3-glucan antibody (bottom panel).
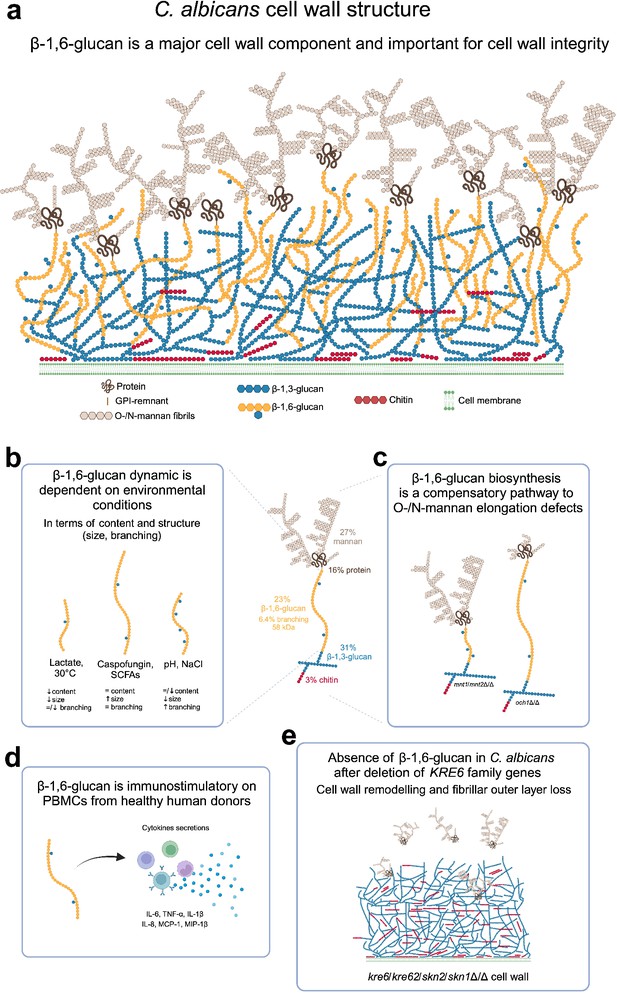
β-1,6-Glucan in C. albicans is a major and dynamic cell wall polymer.
(a) Scheme of the cell wall of C. albicans. The proportion of each cell wall polymer was representative of the results obtained on C. albicans SC5314 grown in liquid synthetic dextrose (SD) medium at 37°C. (b) Scheme representing the dynamic of β-1,6-glucan under different environmental factors. (c) β-1,6-Glucan is a compensatory pathway for mannan elongation defect. (d) β-1,6-Glucan is a PAMP. (e) Scheme of the cell wall of KRE6 family deficient mutant. Created with BioRender.com.
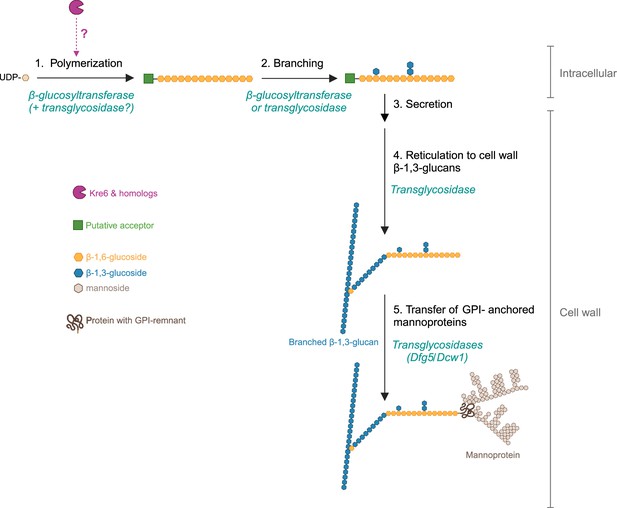
A model for β-1,6-glucan biosynthetic pathway and putative role of Kre6 family members in this process in yeast.
The cellular location of β-1,6-glucan synthesis in yeast is still unknown. We assume that synthesis begins intracellularly with the polymerization of linear β-1,6-glucan chain (step 1), which requires a β-glucosyltransferase and UDP-glucose as a donor (Aimanianda et al., 2009; Vink et al., 2004) and a putative acceptor (sugar, protein, lipid). Our data (Figure 3g, Figure 3—figure supplement 3) suggest that Kre6 and its homologs (Kre62, Skn1, Skn2) act at this stage, but their function remains unknown. Two proposed hypotheses are (1) Kre6 family members are β-glucosyltransferases and (2) they have glycosylhydrolase and transglycosidase activity essential for polymerization. Step 2 is the branching of nascent β-1,6-glucan where glucosides and laminaribiosides are added to form side chains. The enzymes (β-glucosyltransferase or transglycosidase) involved in this branching remain unknown. According to our data, members of the Kre6 family are not involved in branching (Figure 3i). Next, the polysaccharide is secreted (step 3) and then cross-linked to β-1,3-glucans in the cell wall space by an unknown transglycosidase (step 4). The transfer of GPI-anchored proteins onto β-1,6-glucan (step 5), leading to the formation of the outer layer of the cell wall, appears to be driven by Dfg5/Dcw1 (Vogt et al., 2020). The chronology between these two cross-links (steps 4 and 5) has not been established. Created with BioRender.com.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Other | Yeast Nitrogen Base with amino acids (YNB) | BD | 239210 | Medium used in this study. See section ‘Strains and growth conditions’. |
| Chemical compound, drug | Lactate | ChemCruz | sc-220120B | |
| Chemical compound, drug | H2O2 | Prolabo | 23612.294 | |
| Chemical compound, drug | MES | Sigma-Aldrich | M3058-100G | |
| Chemical compound, drug | MOPS | Sigma-Aldrich | M1254-1KG | |
| Chemical compound, drug | Acetate | Sigma-Aldrich | S2889-1KG | |
| Chemical compound, drug | Propionate | Sigma | P1880-100G | |
| Chemical compound, drug | Butyrate | Sigma-Aldrich | B5887-250MG | |
| Chemical compound, drug | Calcofluor white (CFW) | Sigma-Aldrich | 910090-20mL | |
| Chemical compound, drug | Congo Red (CR) | Sigma | C6277-25G | |
| Chemical compound, drug | Tunicamycin | Sigma | 17765-5MG | From Streptomyces spp. |
| Chemical compound, drug | Nikkomycin Z | Sigma | N-8028 | From Streptomyces tendae |
| Chemical compound, drug | Caspofungin | Sigma | SML0425-5MG | |
| Other | Yeast Nitrogen Base without amino acids | BD | 291940 | Medium used in this study for biofilm production. See section ‘Strains and growth conditions’. |
| Other | Breathseal sealing membranes | Greiner Bio-one | 676051 | Used in this study for biofilm production. See section ‘Strains and growth conditions’. |
| Commercial assay or kit | Pierce BCA Protein Assay Kit | Thermo Scientific | 23225 | |
| Chemical compound, drug | Trifluoroacetic acid (TFA) | Sigma | 8.08260.0101 | |
| Chemical compound, drug | Hydrochloric acid (HCl) | Fisher | H/1200/PC15 | |
| Chemical compound, drug | 4-Hydroxybenzhydrazide (PAHBAH) | Sigma | H9882-100G | |
| Chemical compound, drug | Sodium (meta)periodate (m-IO4Na) | Sigma-Aldrich | S1878-25G | |
| Chemical compound, drug | Sodium sulfite | Sigma-Aldrich | S0505-250G | |
| Biological sample | Pustulan | Elicityl OligoTech | GLU900-1g | From Lasallia pustulata |
| Peptide, recombinant protein | LamA | PMID:9168619 | From Thermotoga neapolitana expressed in Escherichia coli | |
| Peptide, recombinant protein | Endo-β-1,6-glucanase | PMID:20852022 | From Schizosaccharomyces pombe expressed in Pichia pastoris | |
| Chemical compound, drug | Sodium azide (NaN3) | Sigma-Aldrich | S-8032 | |
| Chemical compound, drug | D2O | Eurisotop | 17247446 | |
| Other | CarboPAC-PA1 column | Thermo Scientific | 035391 | See sections ‘Cell wall polymers quantification’, ‘Quantification of β-1,6-glucan in AI and AS fractions,’ and ‘Structural characterization of β-1,6-glucan’. |
| Biological sample | Laminarin | Sigma | L9634-5G | From Laminaria digitata |
| Other | Superdex 200 column | Cytiva | 28990944 | See sections ‘Structural characterization of β-1,6-glucan’ and ‘PBMC and neutrophil isolation, stimulation by parietal fractions and cytokine quantification’. |
| Biological sample | Dextran 6 kDa | Sigma-Aldrich | 31388-25G | From Leuconostoc spp. |
| Chemical compound, drug | Dextran 10, 40, 70, and 500 kDa | Pharmacia Fine Chemicals | ||
| Chemical compound, drug | Glutaraldehyde | Sigma-Aldrich | G6257-100ML | |
| Other | 6 mm type A and B specimen carriers | Leica | 16770181 and 16770182 | See section ‘Transmission electron microscopy’. |
| Chemical compound, drug | Spurr resin | EMS | 15000 | |
| Antibody | Polyclonal anti-β-1,6-glucan (rabbit polyclonal) | ProteoGenix | This study | Pustulan-conjugated BSA used as the antigen IF (1:100) |
| Chemical compound, drug | Curdlan | Wako | 034-09901 | |
| Chemical compound, drug | Paraformaldehyde | Electron Microscopy Sciences | 15714 | |
| Biological sample | Goat serum | Sigma | G9023 | |
| Other | Diagnostic Microscope Slides, 8-well 6 mm | Thermo Scientific | ER-301B-CE24 | See section ‘Immunolabeling’. |
| Chemical compound, drug | Poly-l-lysine | Sigma-Aldrich | P4707 | |
| Antibody | Monoclonal anti-β-1,3-glucan, named 5 H5 (mouse monoclonal) | PMID:31022215 | IF (1:250) | |
| Antibody | AlexaFluor 488 goat anti-mouse IgG (goat polyclonal) | Molecular Probes | A11029 | IF (1:200) |
| Antibody | Fluorescein goat anti-rabbit IgG (goat polyclonal) | Invitrogen | F2765 | IF (1:200) |
| Chemical compound, drug | Fluoromount-G | Invitrogen | 00-4958-02 | |
| Chemical compound, drug | Ficoll 400 | Eurobio | CMSMSL01-0U | |
| Chemical compound, drug | RPMI + GlutaMAX medium | Gibco | 61870044 | |
| Biological sample | NHS | Zenbio | HSER-ABP100ML | From human pooled donor |
| Biological sample | LPS | Sigma | L2630 | From Escherichia coli |
| Commercial assay or kit | EasySep Direct Human Neutrophil Isolation Kit | STEMCELL Technologies | 18001 | |
| Commercial assay or kit | Proteome Profiler Human Cytokines Array Kit | R&D Systems | ARY005B | |
| Commercial assay or kit | DuoSet ELISA kits | R&D Systems | DY201, DY278, DY206, DY217B, DY208, DY2037, DY271, DY279, DY210 | |
| Chemical compound, drug | BH4Na | Aldrich | 213462-100G | |
| Chemical compound, drug | Glycerol | Riedel-deHaën | 15523 | |
| Chemical compound, drug | AcOH | Honeywell | 27221-1L | |
| Other | Sep-Pak classic C18 cartridge | Waters | WAT051910 | See section ‘PBMC and neutrophil isolation, stimulation by parietal fractions and cytokine quantification’. |
| Peptide, recombinant protein | BSA | Sigma-Aldrich | A3059-100G | |
| Antibody | Anti-human C3b (mouse monoclonal) | Thermo Fisher | MA1-82814 | ELISA (1:1000) |
| Antibody | Peroxidase-conjugated secondary anti-mouse IgG (goat polyclonal) | Sigma-Aldrich | A4416 | ELISA (1:1000) |
| Chemical compound, drug | TMB | BioFX | TMBW-1000-01 | |
| Recombinant DNA reagent | pV1093 | PMID:25977940 | ||
| Recombinant DNA reagent | pSFS2A | PMID:5474295 | ||
| Recombinant DNA reagent | pV1090 | PMID:25977940 | ||
| Recombinant DNA reagent | pYM70 | PMID:20737428 | ||
| Peptide, recombinant protein | Phusion High-Fidelity DNA polymerase | NEB | M0530S | |
| Peptide, recombinant protein | Q5 polymerase | NEB | M0491S | |
| Commercial assay or kit | NucleoSpin Gel and PCR Clean-up | Macherey-Nagel | 740609.250 | |
| Chemical compound, drug | Nourseothricin | Jena Biosciences | AB-102 | |
| Commercial assay or kit | MasterPure Yeast DNA Purification Kit | Biosearch Technologies | MPY80200 | |
| Peptide, recombinant protein | BP clonase | Invitrogen | P/N 56480 | |
| Peptide, recombinant protein | LR clonase | Invitrogen | P/N 56485 | |
| Sequence-based reagent | pDONR207 | PMID:23049891 | ||
| Sequence-based reagent | Cip10-PACT1-SAT1 | PMID:29982705 | ||
| Strain | SC5314 | PMID:6394964 | Clinical blood isolate | |
| Strain | CAF2-1 | PMID:8349105 | URA3/ura3Δ::imm434 | |
| Strain | CAI4 | PMID:8349105 | ura3Δ::imm434/ura3Δ::imm434 | |
| Strain | BWP17 | PMID:10074081 | ura3Δ::imm434/ura3Δ::imm434 his1::hisG/his1::hisG arg4::hisG/arg4::hisG | |
| Strain | SN152 | PMID:15701792 | arg4Δ/arg4Δ leu2Δ/leu2Δ his1Δ/his1Δ URA3/ura3Δ::imm434 IRO1/iro1Δ::imm434 | |
| Strain | cwh41Δ/Δ | PMID:17933909 | Same as CAI4 but cwh41Δ::dp1200/cwh41Δ::dp1200 RPS1/rps1Δ::CIp10 | |
| Strain | rot2Δ/Δ | PMID:17933909 | Same as CAI4 but rot2Δ::dp1200/rot2Δ::dp1200 RPS1/rps1Δ::CIp10 | |
| Strain | kre5Δ/Δ | PMID:20543849 | Same as SN152 but kre5Δ::leu2/kre5Δ::his1 | |
| Strain | kre6Δ/Δ | This study | kre6Δ::HygB/kre6Δ::HygB | |
| Strain | kre62Δ/Δ | This study | kre62Δ::FRT/kre62Δ::FRT | |
| Strain | skn2Δ/Δ | This study | skn2Δ::FRT/skn2Δ::FRT | |
| Strain | skn1Δ/Δ | This study | skn1Δ::FRT/skn1Δ::FRT | |
| Strain | kre6/skn1Δ/Δ | This study | kre6Δ::HygB/kre6Δ::HygB skn1Δ::FRT/skn1Δ::FRT | |
| Strain | kre6/kre62/skn2/skn1Δ/Δ | This study | kre6Δ::HygB/kre6Δ::HygB kre62Δ::FRT/kre62Δ::FRT skn2Δ::FRT/skn2Δ::FRT skn1Δ::FRT/skn1Δ::FRT | |
| Strain | kre6/kre62/skn2/skn1Δ/Δ+PACT1-KRE6 | This study | kre6Δ::HygB/kre6Δ::HygB kre62Δ::FRT/kre62Δ::FRT skn2Δ::FRT/skn2Δ::FRT skn1Δ::FRT/skn1Δ::FRT RPS1/RPS1::CIpSAT1-PACT1-KRE6 | |
| Strain | kre1Δ/Δ | Provided by Mathias Richard | Same as BWP17 but kre1Δ::arg4/kre1Δ::his1 | |
| Strain | PMRP1-CHS1/chs1Δ | PMID:11251855 | Same as CAI4 but chs1Δ::hisG/chs1Δ:pSK-URA3-PMRP1-CHS1 | |
| Strain | chs2Δ/Δ | PMID:8636047 | Same as CAF2-1 but chs2Δ::hisG/chs2Δ::hisG-URA3-hisG | |
| Strain | chs3Δ/Δ | PMID:7479842 | Same as CAF2-1 but chs3-2::hisG/chs3-3::hisG-URA3-hisG | |
| Strain | mnt1/mnt2Δ/Δ | PMID:15519997 | Same as CAF2-1 but mnt1-mnt2Δ::hisG/mnt1-mnt2Δ::hisG-URA3-hisG | |
| Strain | mnn2/22/21/23/24/26Δ/Δ | PMID:23633946 | Same as CAF2-1 but mnn2Δ::dpl200/mnn2Δ::dpl200 mnn22Δ::dpl200/mnn2Δ::dpl200 mnn23Δ::dpl200/mnn23Δ::dpl200 mnn24Δ::dpl200/mnn24Δ::dpl200 mnn26Δ::dpl200/mnn26Δ::dpl200 mnn21Δ::dpl200/mnn21Δ::dpl200 | |
| Strain | mnn9Δ/Δ | PMID:10601199 | Same as CAF2-1 mnn9Δ::hisG/mnn9Δ::hisG Δura3Δ::imm434/ura3Δ::imm434 | |
| Strain | och1Δ/Δ | PMID:20543849 | Same as SN152 but och1Δ::leu2/och1Δ::his1 | |
| Strain | fks1Δ | PMID:30370375 | SC5314, but fks1/fks1Δ | |
| Strain | phr1Δ/Δ | PMID:7823929 | Same as BWP17 but phr1Δ::hisG/phr1Δ | |
| Strain | phr2Δ/Δ | PMID:9315654 | Same as BWP17 but phr2Δ::hisG/phr2Δ::hisG-URA3-hisG | |
| Sequence-based reagent | SNR52/F | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/R | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SNR52/N | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/N | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | CaCas9/F | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | CaCas9/R | PMID:27340698 | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SNR52/R/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SNR52/R/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SNR52/R/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SNR52/R/KRE6 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/F/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/F/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/F/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | sgRNA/F/KRE6 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/F/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/F/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/F/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | HygR/F/KRE6 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/R/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/R/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | SAT1FLP/R/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | hygR/R/KRE6 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_F/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_F/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_F/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_R/SKN1 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_R/SKN2 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | Flanking_R/KRE62 | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | screenKre6/F | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | screenKre6/R | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | CIpUL | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | CIpSAT | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | KRE6-FWD | This study | PCR primers | See sequence in Supplementary file 2 |
| Sequence-based reagent | KRE6-REV | This study | PCR primers | See sequence in Supplementary file 2 |
Additional files
-
Supplementary file 1
C. albicans strains used in this study.
- https://cdn.elifesciences.org/articles/100569/elife-100569-supp1-v1.docx
-
Supplementary file 2
Primers used in this study.
- https://cdn.elifesciences.org/articles/100569/elife-100569-supp2-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/100569/elife-100569-mdarchecklist1-v1.docx





