The fate of pyruvate dictates cell growth by modulating cellular redox potential
Figures
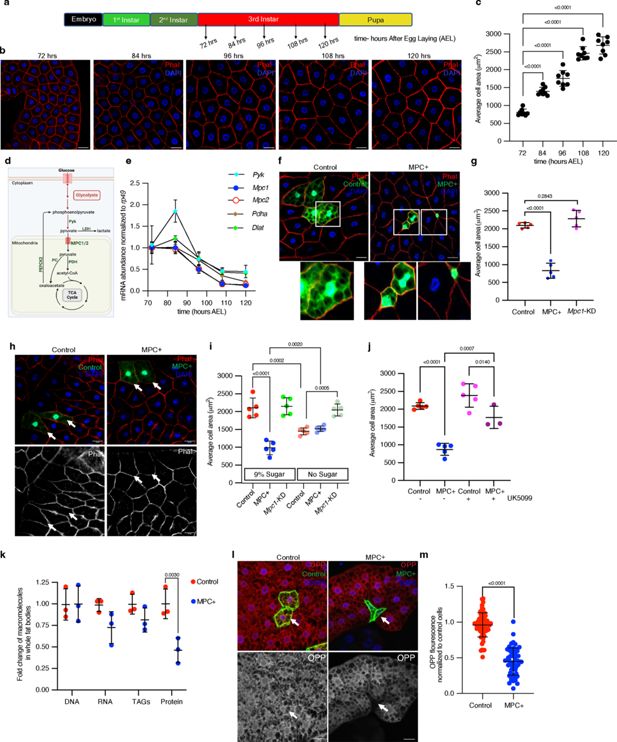
Increased mitochondrial pyruvate transport reduces size of Drosophila fat body cells.
(a) A schematic representation of Drosophila developmental stages with specified time points (hours after egg laying (AEL) at 25°C) at which larvae were dissected to collect fat bodies. (b) Representative images of larval fat bodies at the indicated times (hours AEL) stained with rhodamine phalloidin to visualize cell membranes and DAPI to visualize DNA. The scale bar represents 25 μm. (c) Quantification of fat body cell area based on rhodamine phalloidin stained cell membranes at the indicated time points. Data are presented as mean ± standard deviation (s.d.) from six biological replicates, with each replicate averaging the size of 50 randomly selected cells from fat bodies dissected from five male larvae. (d) A schematic of pyruvate metabolism. In the cytoplasm, pyruvate is a product of glycolysis, synthesized by pyruvate kinase (Pyk) or from lactate via lactate dehydrogenase (LDH). Pyruvate is transported into mitochondria by the mitochondrial pyruvate carrier (MPC) complex. Within mitochondria, pyruvate is converted into acetyl-CoA by pyruvate dehydrogenase (PDH) or into oxaloacetate by pyruvate carboxylase (PC), both of which are substrates for the TCA cycle. PEPCK2 converts oxaloacetate into phosphoenolpyruvate. (e) Pyk, Mpc1, Mpc2, Pdha, and Dlat transcripts were quantified from larval fat bodies collected at the indicated times. (f) Representative confocal microscope images of phalloidin- and DAPI-stained fat bodies with flip-out Gal4 clones expressing MPC1-P2A-MPC2 (MPC+), marked with GFP at 120 hr AEL. The images at the bottom show magnified insets of GFP-positive cells in control and MPC-expressing clones. (g) Quantification of GFP-positive clonal cell area with the indicated genetic manipulations—control, MPC+, and Mpc1-KD. Data are presented as mean ± s.d. from six biological replicates, with each set averaging the size of 20 clonal cells from fat bodies collected from five male larvae. (h) Images showing control and MPC+ clones from larvae fed on no sugar diet. (i) Quantification of the area of control, MPC+ and Mpc1-KD fat body clonal cells from larvae fed on a diet containing either 9% sugar or no sugar. (j) Quantification of the area of MPC+ fat body clonal cells from larvae fed a diet supplemented with or without 20 μM UK5099. (k) Fold change of the abundances of the indicated macromolecules in fat bodies expressing MPC (MPC+) in all fat body cells using CG-Gal4. The abundance of each individual macromolecule is normalized to that of the respective macromolecular abundance in GFP-expressing, control fat bodies. Data are represented as mean ± s.d. from three biological replicates with fat bodies collected from 10 larvae at 120 hr AEL. (l) Representative images of fat body clones stained with O-propargyl-puromycin (OPP, 20 μM for 30 min), showing control and MPC expressing GFP-positive cells. The top panels show GFP-positive clones and OPP staining in red, while the bottom panels show respective OPP channel. Arrows indicate cells with the specified genetic manipulation. The scale bar represents 20 μm. (m) Quantification of OPP fluorescence intensity of control or MPC+ fat body cells compared to neighboring non-clonal cells. Data are presented as mean ± s.d. from six biological replicates, with each set averaging the size of 20 clonal cells from fat bodies collected from five male larvae. Unpaired t-tests or one-way ANOVA tests were performed to evaluate the statistical significance of the data, with p-values mentioned in the graphs where significance was noted. Panel d was created with BioRender.com.
-
Figure 1—source data 1
Source data related to Figure 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig1-data1-v1.xlsx

Transcriptomic changes in fat body during larval development.
(a) A heatmap displaying changes in mRNA abundance of genes encoding mitochondrial proteins and enzymes involved in various metabolic pathways in wild-type w1118 fat bodies collected at the specified time points. Right. Clustered hallmarks are indicated by a color gradient corresponding to relative gene expression levels. (b) A heatmap illustrating transcriptomic changes in signaling pathways in w1118 fat body cells during Drosophila larval development. (c) Quantification of Pepck and Pepck2 transcripts from larval fat bodies collected at the indicated times.
-
Figure 1—figure supplement 1—source data 1
Source data related to Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig1-figsupp1-data1-v1.xlsx

Increased mitochondrial pyruvate transport reduces size of Drosophila fat body cells.
(a) Schematic representation of experimental time course using temperature-sensitive Gal80ts20 to control Gal4 expression. At 18°C, Gal80ts20 is active, inhibiting Gal4 expression; at 29°C, Gal80ts20 is inactive, allowing Gal4 expression. The temperature switch occurs toward the end of the second instar stage so that MPC expression construct is induced temporally only during the third instar stage. Fat bodies were collected at the specified time points. (b) Quantification of Mpc1 and Mpc2 transcripts from fat bodies of CG-Gal4>MPC+ versus CG-Gal4>GFP or control larvae, dissected at the indicated times after the induction of Gal4 activity. (c) Quantification of fat body cell area in MPC+ (CG-Gal4>MPC+) versus control (CG-Gal4>GFP) larvae at the indicated times after Gal4 induction. Data are presented as mean ± s.d. from six biological replicates, with each replicate representing the average size from 50 randomly selected cells from fat bodies dissected from five male larvae. (d) Schematics illustrating the use of the Ay-Gal4 cassette to generate clones with MPC expression. Clones are generated by leaky expression of hs-flp1.22-flippase, which binds to specific FRT sites in chromatin, leading to the excision of the spacer ‘yellow’ gene and positioning of the actin5C promoter next to the Gal4 coding sequence. This is followed by Gal4 expression in the cell and subsequent expression of UAS-GFP or UAS-MPC1-P2A-MPC2 constructs. Due to the leaky and random expression of the flippase, GFP-marked mosaicism occurs in only 1–5% of fat body cells. Representative images of fat bodies with MPC expression and control clones, showing increased abundance of (e) MPC1 and (f) MPC2 proteins. Arrows indicate cells with the specified genetic manipulation. The scale bar represents 20 μm. (g) Representative images of fat bodies with MPC+ and control clones dissected from larvae fed on a diet supplemented with 20 μM UK5099. Arrows indicate cells with the specified genetic manipulation. The scale bar represents 20 μm. The abundances of (h) DNA, (i) RNA, (j) triacylglycerides (TAGs), and (k) protein in fat bodies where MPC is expressed in all fat body cells using CG-Gal4 driver. The abundance of each individual macromolecule is normalized to that of the control. Data are represented as mean ± s.d. from three biological replicates with fat bodies collected from ten larvae at 120 hr AEL. (l) Representative images of MPC+ and control fat body clones showing lipid droplets stained with LipidTox in red. The area in square is presented in high-magnification insets below to show lipid distribution in the clonal cells. The scale bar represents 25 μm. (m) Quantification of lipid droplets relative to cell size in MPC+ versus control cells. (n) The data are presented as the number of lipid droplets per cell area and the fraction of cell area covered by lipid droplets. (o) Representative EdU-stained confocal images of MPC+ and control fat body cells, showing replicating DNA. Arrows indicate cells with the specified genetic manipulation. The scale bar represents 20 μm. (p) Quantification of total EdU fluorescence intensity in GFP-positive cells compared to neighboring GFP-negative cells in MPC+ versus control clones. Data are presented as mean ± s.d. Unpaired t-tests were performed to evaluate the statistical significance of the data, with p-values noted in the graph if significance was observed. Panel d was created with BioRender.com.
-
Figure 1—figure supplement 2—source data 1
Source data related to Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig1-figsupp2-data1-v1.xlsx
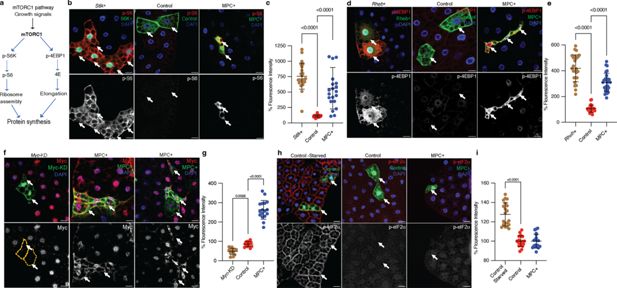
mTORC1 and Myc pathways are hyperactivated in MPC expressing fat body cells.
(a) Schematic of the mTORC1 pathway. The mTORC1 pathway is activated by pro-growth signals such as insulin, leading to the phosphorylation of S6 kinase (S6K) and 4EBP. S6K phosphorylates ribosomal protein S6, while 4EBP phosphorylation inactivates 4EBP and releases elongation factor eIF4E. These events increase ribosomal assembly and elongation rates, thereby enhancing protein synthesis. (b) Representative images of fat body clones with S6k overexpression, control clones, and MPC+ clones, stained for phosphorylated S6 (p-S6) in red (top panels) and white (bottom panels). Arrows indicate clonal cells with the specified genetic manipulation. The scale bar represents 20 μm. (c) Quantification of total p-S6 fluorescence intensity in GFP-positive cells compared to neighboring GFP-negative cells in MPC+ versus control clones. Data are presented as mean ± s.d. with statistical significance as scored with one-way ANOVA test. (d) Representative images of fat body clones with Rheb over-expression, control clones, and MPC+ clones stained for phosphorylated 4EBP (p-4EBP) in red (top panels) and white (bottom panels). Arrows indicate clonal cells with the specified genetic manipulation. The scale bar represents 20 μm. (e) Quantification of total p-4EBP fluorescence intensity in MPC+ versus control clones with statistical significance as scored with One-way ANOVA test. (f) Representative images of fat body clones with Myc knockdown, control clones, and MPC+ clones stained for Myc protein in red (top panels) and white (bottom panels). Arrows indicate clonal cells with the specified genetic manipulation. The scale bar represents 20 μm. (g) Quantification of total Myc fluorescence intensity in MPC+ versus control clones with statistical significance as scored with one-way ANOVA test. (h) Representative images of fat body clones from starved wild type, control clones, and MPC+ clones, stained for phosphorylated eIF2α (p-eIF2α) in red (top panels) and white (bottom panels). Arrows indicate clonal cells with the specified genetic manipulation. The scale bar represents 20 μm. (i) Quantification of total p-eIF2α fluorescence intensity in MPC+ versus control clones with statistical significance as scored with one-way ANOVA test. Data are presented as mean ± s.d. One-way ANOVA tests were performed to evaluate the statistical significance of the data, with p-values noted in the graph if significance was observed.
-
Figure 2—source data 1
Source data related to Figure 2.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig2-data1-v1.xlsx

mTORC1 and Myc pathways are hyperactivated in MPC-expressing fat body cells.
(a) Representative images of fat body clones with PI3K-DN (dominant-negative) expression, control clones, and MPC+ clones stained for dFoxo in red (top panels) and white (bottom panels). Arrows indicate clonal cells with the specified genetic manipulations. The scale bar represents 20 μm. (b) Transcriptomic analysis of MPC+ fat bodies compared to control, represented as Hallmarks from GSEA (gene set enrichment analysis). Quantification of the area of MPC+ cells with or without PI3K overexpression (c), and with or without Tsc1 knockdown (d). Data are presented as mean ± s.d. from five biological replicates, where each data point represents the average size of 20 clones from fat bodies collected from five male larvae. Quantification of cell area in MPC+ clones with Myc over-expression (e) or Myc knockdown (f). Data are presented as mean ± s.d. from five biological replicates, where each data point represents the average size of 20 clones from fat bodies collected from five male larvae.
-
Figure 2—figure supplement 1—source data 1
Source data related to Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig2-figsupp1-data1-v1.xlsx
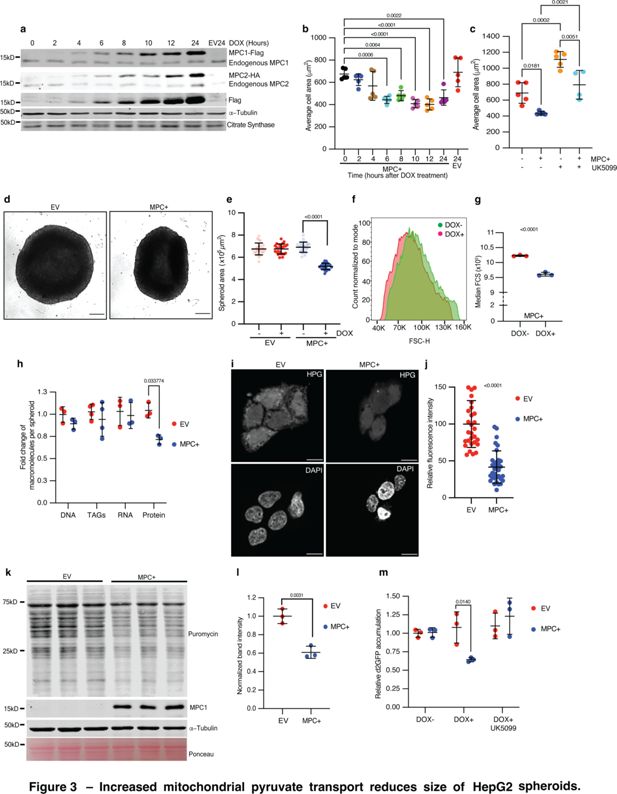
Increased mitochondrial pyruvate transport reduces size of HepG2 spheroids.
(a) Western blots showing inducible expression of MPC1 and MPC2 at 2-hr intervals following treatment with 1 μg/ml doxycycline. Citrate synthase and tubulin were used as loading controls. Both endogenous and epitope tag bands are shown. (b) Quantification of the 2D area of HepG2 cells with MPC expression (MPC+) or empty vector (EV) fixed and stained with rhodamine phalloidin at the indicated times after doxycycline treatment. Data are presented as mean ± s.d. from five biological replicates, with each replicate representing the average size of 25 randomly selected cells. (c) Quantification of the 2D area of MPC+ or EV HepG2 cells treated with 10 μM UK5099. Data are presented as mean ± s.d. from five biological replicates, with each replicate representing the average size of 25 randomly selected cells. (d) Representative brightfield images of HepG2 spheroids with empty vector (EV) or MPC expression (MPC+) treated with 1 μg/ml doxycycline for 6 days. The scale bar represents 200 μm. (e) Quantification of spheroid area from images of MPC+ or EV HepG2 spheroids, with or without doxycycline treatment. Data are presented as mean ± s.d. from 30 technical replicates. (f) Forward scatter (FSC) of cells dissociated from MPC+ (red) or EV spheroids with cell count normalized to mode. (g) Median FSC of MPC+ HepG2 spheroids treated with or without 1 μg/ml doxycycline. Data are presented as mean ± s.d. from three biological replicates. (h) Fold change in macromolecules—DNA, TAGs, RNA, and protein—fractionated from EV or MPC+ HepG2 spheroids normalized to that in EV HepG2 cells. (i) Representative images showing L-homopropargylglycine (HPG)-labeled newly synthesized proteins in EV or MPC+ HepG2 cells. The top panels show HPG staining, and the lower panels show nuclei stained with DAPI. The scale bar represents 20 μm. (j) Quantification of HPG fluorescence intensity is presented as mean ± s.d. from 35 cells, for both EV and MPC+ cells. (k) Western blot analysis of nascent protein synthesis using a puromycin incorporation assay (20 μg/ml puromycin for 30 min) in either EV or MPC+ HepG2 spheroids (protein lysates from 16 spheroids loaded in each lane). (l) Quantification of 70 kD band intensity in puromycin blot normalized with tubulin band intensity and represented as mean ± s.d. from three independent experiments. (m) Relative accumulation of destabilized GFP (d2GFP) in EV or MPC+ HepG2 spheroids treated with or without 1 μg/ml doxycycline ± 10 μM UK5099. Data are presented as mean ± s.d. from three biological replicates. Unpaired t-tests, one-way, or two-way ANOVA tests were performed to evaluate the statistical significance of the data, with p-values noted in the graph if significance was observed.
-
Figure 3—source data 1
Source data related to Figure 3.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-data1-v1.xlsx
-
Figure 3—source data 2
Original files for western blot analysis displayed in Figure 3a.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-data2-v1.zip
-
Figure 3—source data 3
PDF files containing original western blots for Figure 3a, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-data3-v1.zip
-
Figure 3—source data 4
Original files for western blot analysis displayed in Figure 3k.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-data4-v1.zip
-
Figure 3—source data 5
PDF files containing original western blots for Figure 3k, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-data5-v1.zip

MPC overexpression reduces size of HepG2 cells in 2D and spheroid culture.
(a) Quantification of cell volume from confocal images of HepG2 cells expressing MPC (MPC+) or an empty vector (EV). Data is presented as mean ± s.d. from 22 EV cells and 25 MPC+ cells. (b) Quantification of the number of cells per spheroid in EV or MPC+ HepG2 cells with or without doxycycline treatment. Data is presented as mean ± s.d. from 18 technical replicates. (c) Cell cycle profiles of MPC+ HepG2 spheroids treated with or without doxycycline treatment. Data is presented as mean ± s.d. from three biological replicates. (d) Annexin/Propidium Iodide (PI) analysis of MPC+ HepG2 cells cultured with or without doxycycline. Data is presented as mean ± s.d. from three biological replicates. Concentrations of (e) DNA, (f) RNA, (g) triacylglycerides (TAGs) and (h) protein measured for 18 MPC+ and 18 EV HepG2 spheroids. Data is shown as mean ± s.d. from three biological replicates. Unpaired t-tests and one-way ANOVA tests were performed to evaluate the statistical significance of the data, with p-values mentioned in the graph if significance is noted.
-
Figure 3—figure supplement 1—source data 1
Source data related to Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig3-figsupp1-data1-v1.xlsx
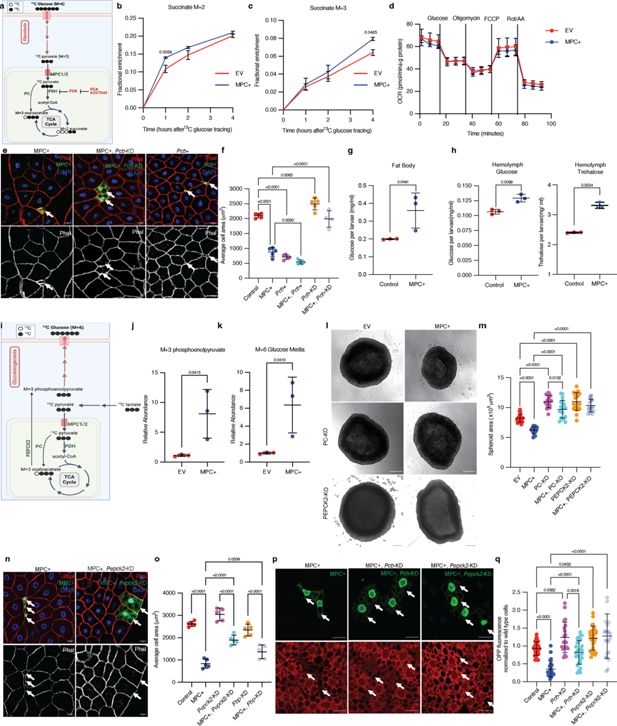
Increased mitochondrial pyruvate metabolism promotes gluconeogenesis via pyruvate carboxylase to suppress protein synthesis.
(a) Schematic illustration of the 13C-glucose tracing strategy used to measure the activity of pyruvate dehydrogenase (PDH) and pyruvate carboxylase (PC). TCA metabolites labeled with two heavy carbons (13C or M+2 TCA pool) result from PDH activity, whereas M+3 TCA metabolites result from PC activity. PDH is inhibited by PDK-mediated phosphorylation. DCA and AZD7545 are inhibitors of PDK. (b) Fractional enrichment of M+2 succinate in empty vector (EV; red) and MPC-expressing (MPC+; blue) HepG2 cells at the indicated times after 13C-glucose tracing. MPC expression was induced for 24 hr by treatment with 1 µg/ml doxycycline and media was changed to 12C-glucose. The change in M+2 succinate is significant (by two-way ANOVA test) at 1 hr after 13C glucose incubation. (c) Fractional enrichment of M+3 succinate in EV (red) and MPC+ (blue) HepG2 at the indicated times after 13C-glucose tracing. MPC expression was induced for 24 hr by treatment with 1 μg/ml doxycycline. The change in M+3 succinate is significant (by two-way ANOVA test) at 4 hr after 13C-glucose incubation. (d) Rate of oxygen consumption (OCR) in EV and MPC+ HepG2 cells. (e) Representative images of phalloidin- and DAPI-stained fat body cells. Arrows indicate GFP-positive clones with MPC expression (MPC+), Pcb knockdown with MPC expression (MPC+, Pcb-KD), or Pcb overexpression (Pcb+). The scale bar represents 20 µm. (f) Quantification of the area of GFP-positive clones with control, MPC+, Pcb over-expression (Pcb+), Pcb and MPC co-expression (MPC+, Pcb+), Pcb knockdown (Pcb-KD), and Pcb knockdown with MPC expression (MPC+, Pcb-KD) shown as mean ± s.d. of five biological replicates, with each group representing the analysis of 20 the indicated clonal cells. Concentration of glucose in the fat body (g) and concentration of glucose and trehalose in hemolymph (h) of larva with fat body-specific expression MPC or control. Data is presented as mean ± s.d. of three biological replicates analyzed by unpaired t-tests. (i) Schematic illustration of the strategy to analyze gluconeogenesis from 13C-lactate. Cells convert 13C-lactate into 13C-pyruvate, which is transported into mitochondria by the MPC. PC converts 13C-pyruvate (M+3) into oxaloacetate (M+3). PEPCK2 converts oxaloacetate (M+3) into phosphoenolpyruvate (M+3), which is converted into M+6 glucose and excreted from cells. (j) Relative abundances of M+3 phosphoenolpyruvate (PEP) in EV and MPC+ HepG2 cells and (k) M+6 glucose in their respective media following treatment with 20 mM 13C-lactate for 4 hr. Data is presented as mean ± s.d. of three biological replicates, each with an average of three technical replicates. (l) Representative brightfield images of EV, MPC+, and PC knockout (KO) or PEPCK2 KO with or without MPC expression HepG2 spheroids. The scale bar represents 200 µm. (m) Quantification of spheroid area is presented as mean ± s.d. of 30 technical replicates. (n) Representative images of phalloidin- and DAPI-stained fat body cells. Arrows indicate GFP-positive clones with MPC expression (MPC+), and Pepck2 knockdown with MPC expression (MPC+, Pepck2-KD). The scale bar represents 20 µm. (o) Quantification of the area of GFP-positive clones with MPC+, Pepck2 knockdown (Pepck2-KD), Pepck2 knockdown with MPC+ (MPC+, Pepck2-KD), Fbp knockdown (Fbp-KD), Fbp knockdown with MPC+ (MPC+, Fbp-KD). Data is presented as mean ± s.d. of five biological replicates, with each group analyzing 20 clonal cells of the mentioned genetic manipulations. (p) Representative images of fat body clones stained with OPP (red). Arrows indicate GFP-positive clones with MPC expression (MPC+), Pcb knockdown with MPC expression (MPC+, Pcb-KD), or Pepck2 knockdown with MPC expression (MPC+, Pepck2-KD). The scale bar represents 20 µm. (q) Quantification of OPP intensity in the indicated clones compared with adjacent wild-type cells. Data is presented as mean ± s.d. Unpaired t-tests, one-way ANOVA tests, or two-way ANOVA tests were performed to evaluate the statistical significance of the data, with p-values mentioned in the graph if significance is noted. Panel a was created with BioRender.com. Panel i was created with BioRender.com.
-
Figure 4—source data 1
Source data related to Figure 4.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-data1-v1.xlsx

Surge of mitochondrial pyruvate reduced glycolysis and promotes gluconeogenesis via pyruvate carboxylase.
Fractional enrichment and metabolite counts of M+3 3-phosphoglyceric acid (a), M+3 pyruvate (b), and M+3 alanine (c) in empty vector (EV; red) and MPC expressing (MPC+; blue) HepG2 cells at the indicated times after 13C glucose tracing. Metabolite counts of M+2 succinate (d) and M+3 succinate (e) in EV (red) and MPC+ (blue) HepG2 cells. Fractional enrichment and metabolite counts of M+2 fumarate (f), M+2 malate (g) M+3 fumarate (h), and M+3 malate (i) in EV (red) and MPC+ (blue) HepG2 cells. A two-way ANOVA test showed significant differences at 4 hr after 13C glucose tracing.
-
Figure 4—figure supplement 1—source data 1
Source data related to Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp1-data1-v1.xlsx

Surge of mitochondrial pyruvate promotes gluconeogenesis via pyruvate carboxylase to suppress protein synthesis.
qRT-PCR showing (a) Mpc1, (b) Mpc2, and (c) pcb transcripts from fat bodies of control, MPC+, Pcb over-expression (Pcb+), Pcb and MPC co-expression (MPC+, Pcb+), Pcb knock down (Pcb-KD) and Pcb knock down with MPC expression (MPC+, Pcb-KD) normalized to rp49 transcript abundance. (d) Quantification of the area of GFP-positive fat body clones with control, MPC expression (MPC+), Pcb knockout (Pcb KO), or MPC expression with Pcb knockout (MPC+, Pcb KO). Data is presented as mean ± s.d. of five biological replicates, with each group analyzing 20 clonal cells of the indicated genetic manipulations. (e) Quantification of the areas of HepG2 cells with MPC+ with or without pyruvate carboxylase (PC) knockout using two independent sgRNA guides (PCg5 and PCg6). Western blots show the efficiency of PC knockout. Data is presented as mean ± s.d. of five biological replicates, each representing 25 randomly selected cells for each indicated genotype. qRT-PCR showing (f) Mpc1, (g) Mpc2, (h) Pdha, and (i) Dlat transcripts from fat bodies of control, MPC+, Pdha knockdown (Pdha-KD), Pdha knockdown with MPC expression (MPC+, Pdha-KD), Dlat knockdown (Dlat-KD), Dlat knockdown with MPC expression (MPC+, Dlat-KD) normalized to rp49 transcript abundance. (j) Representative images of phalloidin- and DAPI-stained fat body cells. Arrows indicate GFP-positive clones with MPC expression (MPC+), Pdha knockdown (Pdha-KD), or MPC expression with Pdk knockdown (MPC+, Pdk-KD). The scale bar represents 20 µm. Right. (k) Quantification of the areas of GFP-positive clones with control, MPC+, Pdha knockdown (Pdha-KD), Pdha knockdown with MPC expression (MPC+, Pdha-KD), Dlat knockdown (Dlat-KD), Dlat knockdown with MPC expression (MPC+, Dlat-KD); Pdk knockdown (Pdk-KD), and Pdk knockdown with MPC expression (MPC+, Pdk-KD). Data is presented as mean ± s.d. of five biological replicates, with 20 clonal cells analyzed for each of the indicated genetic manipulations. (l) Quantification of the areas of MPC+ HepG2 cells treated with the PDK inhibitors AZD7545 (10 µM) or dichloroacetate (1 mM). Data is presented as mean ± s.d. of five biological replicates, with 25 randomly selected cells analyzed for each of the indicated groups. (m) Quantification of the areas of MPC+, PEPCK2 knockout HepG2 cells. Western blots show the efficiency of PEPCK2 knockout. Data is presented as mean ± s.d. of five biological replicates with 25 randomly selected cells analyzed for each of the indicated groups. qRT-PCR showing (n) Mpc1, (o) Mpc2, (p) Pepck2, and (q) Fbp transcripts from fat bodies of MPC+, Pepck2 knockdown (Pepck2-KD), Pepck2 knockdown with MPC+ (MPC+, Pepck2-KD), Fbp knockdown (Fbp-KD), Fbp knockdown with MPC+ (MPC+, Fbp-KD) normalized to rp49 transcript abundance. (r) Quantification of the area of GFP-positive control; MPC+; Pepck2 KO; and MPC+, Pepck2 KO fat body clonal cells. Data is presented as mean ± s.d. of five biological replicates, with 20 clonal cells analyzed for each of the indicated genetic manipulations. Unpaired t-tests, one-way ANOVA tests, or two-way ANOVA tests were performed to evaluate the statistical significance of the data, and p-values are mentioned in the graph if significance is noted.
-
Figure 4—figure supplement 2—source data 1
Source data related to Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp2-data1-v1.xlsx
-
Figure 4—figure supplement 2—source data 2
PDF files containing original western blots for Figure 4—figure supplement 2e, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp2-data2-v1.zip
-
Figure 4—figure supplement 2—source data 3
Original files for western blot analysis displayed in Figure 4—figure supplement 2e.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp2-data3-v1.zip
-
Figure 4—figure supplement 2—source data 4
PDF files containing original western blots for Figure 4—figure supplement 2m, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp2-data4-v1.zip
-
Figure 4—figure supplement 2—source data 5
Original files for western blot analysis displayed in Figure 4—figure supplement 2m.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig4-figsupp2-data5-v1.zip
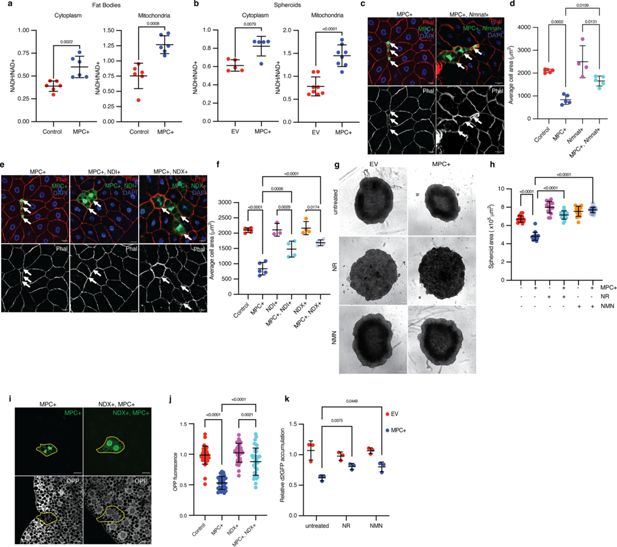
Redox imbalance impedes protein synthesis and cell growth from elevated pyruvate metabolism in mitochondria.
(a) NADH/NAD+ ratios from the cytoplasmic and mitochondrial fractions of control and MPC-expressing (MPC+) fat bodies. Data is presented as mean ± s.d. of six biological replicates. (b) NADH/NAD+ ratios from the cytoplasmic and mitochondrial fractions of empty vector (EV) and MPC-expressing (MPC+) HepG2 spheroids. Data is presented as mean ± s.d. of six biological replicates. (c) Representative images of phalloidin- and DAPI-stained fat bodies. Arrows indicate GFP-positive cells with either MPC+ or MPC and Nmnat co-expression (MPC+, Nmnat+). The scale bar represents 20 µm. (d) Quantification of the areas of GFP-positive cells with control, MPC+, Nmnat overexpression (Nmnat+), MPC and Nmnat co-expression (MPC+, Nmnat+). Data is presented as mean ± s.d. of five biological replicates, with 20 clonal cells analyzed for each of the indicated genetic manipulations. (e) Representative images of Phalloidin and DAPI-stained fat body tissues. Arrows indicate GFP-positive cells showing MPC expression (MPC+); NDI and MPC expression (MPC+, NDI+); and NDX and MPC expression (MPC+, NDX+). The scale bar represents 20 µm. (f) Quantification of the area of GFP-positive cells with control, MPC+, NDI expression (NDI+), NDI and MPC expression (MPC+, NDI+), NDX expression (NDX+), NDX and MPC co-expression (MPC+, NDX+). Data is presented as mean ± s.d. of five biological replicates, with 20 clonal cells analyzed for each of the indicated genetic manipulations. (g) Representative bright field images of EV or MPC+HepG2 spheroids cultured with NAD+ supplements (100 nM nicotinamide riboside or 1 µM NMN) as indicated. The scale bar represents 200 µm. (h) Quantification of spheroid area is presented as mean ± s.d. of 30 technical replicates. (i) Representative images MPC+ or MPC+, NDI+ GFP-positive clones and of fat body cells stained with OPP (bottom). Clonal cells are mapped with dotted lines. The scale bar represents 20 µm. (j) Fold change in OPP intensity of 35 GFP-positive cells compared with adjacent wild-type cells. Data is presented as mean ± s.d. (k) Relative accumulation of destabilized GFP (d2GFP) in spheroids of EV or MPC+ HepG2 cells treated with NAD+ supplements (100 nM nicotinamide riboside or 1 µM NMN) as indicated. Unpaired t-tests, one-way ANOVA tests, or two-way ANOVA tests were performed to assess the statistical significance of the data, with p-values indicated in the graph where significance was observed.
-
Figure 5—source data 1
Source data related to Figure 5.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-data1-v1.xlsx

Redox imbalance impedes protein synthesis and cell growth from elevated pyruvate metabolism in mitochondria.
(a) A schematic illustration of mitochondrial pyruvate metabolism and gluconeogenesis. Following are the abbreviations: PEP: phosphoenolpyruvate, F1,6BP: fructose-1, 6-bisphosphate, F6P: fructose-6-phosphate, G6P: glucose-6-phosphate, F6Pase: fructose bisphosphatase, and G6PC: glucose-6-phosphatase (b) Western blot analysis shows the expression levels of phosphorylated PDH (p-PDH), total PDH, G6PC, PEPCK2, PC, and tubulin in MPC-expressing (MPC+) HepG2 cells treated with 1 µg/ml doxycycline at the indicated time points. (c) Schematic illustrating the regulation of PDH, PC, and gluconeogenesis by a network of cofactors and substrates. Following are the abbreviations: 1,3 BPG: 1,3-bisphosphoglycerate, G3P: glucose-3-phosphate, PGK: phosphoglycerate kinase, and GAPDH: glyceraldehyde-3-phosphate dehydrogenase. (d) Fold enrichment of acetyl-CoA in EV and MPC+ HepG2 cells. Data is presented as mean ± s.d. of three biological replicates using unpaired t-tests. (e) The ratio of ATP to ADP in EV and MPC+ HepG2 cells. Data is presented as mean ± s.d. of three biological replicates using unpaired t-tests. Cellular concentrations of NADH (f) and NAD+ (g) and NADH/NAD+ ratios (h) in EV and MPC+ HepG2 cells 24 hr after treatment with 1 µg/ml doxycycline. Data is presented as mean ± s.d. of three biological replicates using unpaired t-tests. (i) Quantification of the areas of MPC+ HepG2 cells cultured with 2 nM gramicidin. Data is presented as mean ± s.d. of five biological replicates, with 25 randomly selected cells analyzed for each of the indicated groups. (j) Representative bright field images of EV or MPC+ HepG2 spheroids with NDI expression (NDI+). The scale bar represents 200 µm. (k) Quantification of spheroid areas is presented as mean ± s.d. of 30 technical replicates. Western blots show the efficiency of MPC expression. (l) Quantification of the area of MPC-expressing HepG2 cells cultured with or without 100 nM duroquinone. Data is presented as average ± s.d. of five biological replicates, with each group consisting of 25 randomly selected cells. Data is presented as mean ± s.d. of three biological replicates. Unpaired t-tests, one-way ANOVA, or two-way ANOVA tests were performed to assess the statistical significance, with p-values indicated in the graph where significance was noted. Panel a was created with BioRender.com. Panel c was created with BioRender.com.
-
Figure 5—figure supplement 1—source data 1
Source data related to Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-figsupp1-data1-v1.xlsx
-
Figure 5—figure supplement 1—source data 2
PDF files containing original western blots for Figure 5—figure supplement 1k, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-figsupp1-data2-v1.zip
-
Figure 5—figure supplement 1—source data 3
Original files for western blot analysis displayed in Figure 5—figure supplement 1k.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-figsupp1-data3-v1.zip
-
Figure 5—figure supplement 1—source data 4
PDF files containing original western blots for Figure 5—figure supplement 1b, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-figsupp1-data4-v1.zip
-
Figure 5—figure supplement 1—source data 5
Original files for western blot analysis displayed in Figure 5—figure supplement 1b.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig5-figsupp1-data5-v1.zip
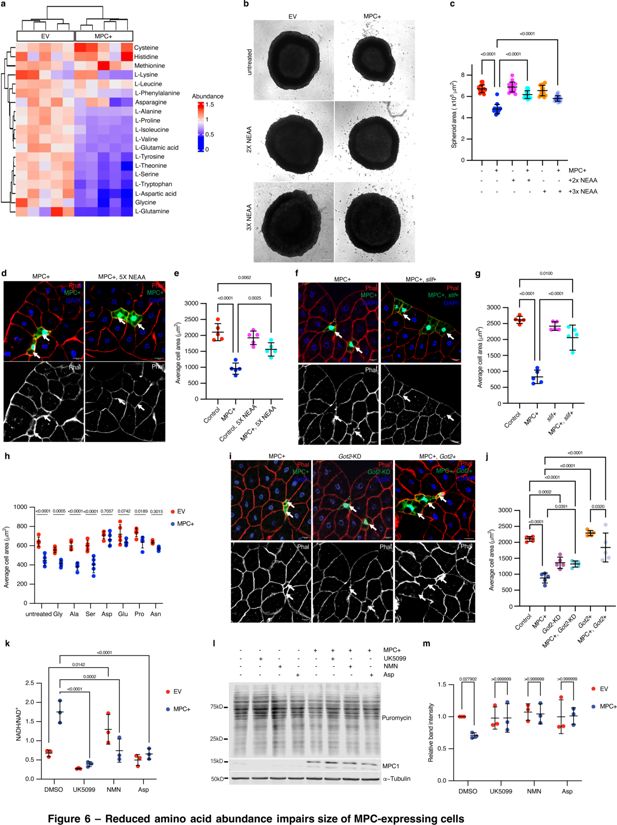
Reduced amino acid abundance impairs size of MPC-expressing cells.
(a) A heatmap of the abundances of amino acids in empty vector (EV) and MPC expressing (MPC+) HepG2 cells cultured under standard conditions. Color codes indicate relative abundances for each amino acid: blue (low), green (similar), and yellow (high). (b) Representative bright field images of EV or MPC+ HepG2 spheroids cultured with 2x or 3x the recommended concentration of non-essential amino acids cocktail (NEAA). The scale bar represents 200 µm. (c) Quantification of spheroid areas from EV and MPC+ HepG2 spheroids cultured with 2x or 3x NEAA. Data is presented as mean ± s.d. of 30 technical replicates. (d) Representative images of phalloidin- and DAPI-stained fat body cells from animals fed a standard diet or a diet supplemented with 5x NEAA. Arrows indicate GFP-positive clones with MPC expression (MPC+). The scale bar represents 20 µm. (e) Quantification of the area of MPC+ fat body clonal cells. Data is presented as mean ± s.d. of five biological replicates, with each data point representing the average size of 20 clones collected from five male larvae. (f) Representative images of phalloidin- and DAPI-stained fat body cells. Arrows indicate GFP-positive MPC+ clones and MPC+ and slimfast over-expression (MPC+, Slif+). The scale bar represents 20 µm. (g) Quantification of the area of GFP-positive clones with control, MPC expression, Slimfast overexpression (Slif+) and MPC+ clones with Slimfast overexpression (MPC+, Slif+). Data presented as mean ± s.d. of five biological replicates, with each 20 clonal cells analyzed for each of the indicated genetic manipulations. (h) Quantification of the cell areas of EV or MPC+ HepG2 cells cultured under standard conditions or with excess of the indicated amino acid—10 mM glycine, 5 mM alanine, 5 mM serine, 5 mM asparagine, 5 mM aspartic acid, 5 mM glutamic acid, or 5 mM proline. Data is presented as mean ± s.d. of five biological replicates. (i) Representative images of phalloidin- and DAPI-stained fat body cells. Arrows indicated GFP-positive cells with MPC expression (MPC+), Got2 knockdown (Got2 KD), and Got2 overexpression with MPC expression (MPC+, Got2+). (j) Quantification of the area of GFP-positive clones with control, MPC expression (MPC+), Got2 knockdown (Got2-KD), Got2 knockdown with MPC expression (MPC+, Got2-KD), Got2 overexpression (Got2+), and Got2 overexpression with MPC expression (MPC+, Got2+). Data is presented as mean ± s.d. of five biological replicates, with each 20 clonal cells analyzed for each of the specified genetic manipulations. (k) NADH/NAD+ ratio in cells treated with 10 µM UK5099, 1 µM NMN, or 5 mM aspartate. Data is presented as mean ± s.d. of three biological replicates. (l) Western blot analysis of puromycin-labeled (20 µg/ml puromycin for 30 min) nascent protein in EV or MPC+ HepG2 cells cultured with 10 µM UK5099, 1 µM NMN, or 5 mM aspartate. (m) Quantification of intensities of puromycin labeling in EV and MPC+ cell lysates. Unpaired t-tests and one-way ANOVA tests were performed to evaluate the statistical significance of the data, and p-values are noted in the graph if significance is observed.
-
Figure 6—source data 1
Source data related to Figure 6.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-data1-v1.xlsx
-
Figure 6—source data 2
PDF files containing original western blots for Figure 6i, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-data2-v1.zip
-
Figure 6—source data 3
Original files for western blot analysis displayed in Figure 6i.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-data3-v1.zip

Reduced aspartate/glutamate abundance impairs size of MPC-expressing cells.
(a) Representative bright field images of empty vector (EV) or MPC over-expressing (MPC+) HepG2 spheroids cultured with either 5 mM aspartate (Asp) or 5 mM glutamate (Glu). The scale bar represents 200 µm. (b) Quantification of spheroid area is presented as mean ± s.d. of 30 technical replicates. (c) Representative bright field images of EV or MPC+ HepG2 spheroids with SLC1A3 overexpression (SLC1A3+). The scale bar represents 200 µm. (d) Quantification of spheroid area is presented as mean ± s.d. of 30 technical replicates. Western blots show the efficiency of MPC expression. (e) Quantification of d2GFP in EV or MPC+ HepG2 spheroids cultured under standard conditions or with three times the recommended concentration of non-essential amino acids cocktail (3X NEAA). qRT-PCR showing (f) Mpc1, (g) Mpc2, and (h) Got2 transcripts from fat bodies with control, MPC expression (MPC+), Got2 knockdown (Got2-KD), Got2 knockdown with MPC expression (MPC+, Got2-KD), Got2 overexpression (Got2+), and Got2 overexpression with MPC expression (MPC+, Got2+) normalized to rp49 transcript abundance. (i) Quantification of the areas of EV and MPC+ HepG2 cells and HepG2 cells GOT2 knock out (GOT2-KO) with or without MPC expression. Data is presented as mean ± s.d. of five biological replicates, with 25 randomly selected cells analyzed per replicate. Western blots show the efficiency of GOT2 knockout using the two sgRNAs described. (j) Representative bright field images of EV, MPC+, GOT2+, and MPC+, GOT2+ HepG2 spheroids. The scale bar represents 200 µm. (k) Quantification of spheroid areas is presented as mean ± s.d. of 30 technical replicates. Western blots show the efficiency of GOT2 overexpression. Unpaired t-tests and one-way ANOVA tests were performed to evaluate the statistical significance of the data, and p-values are noted in the graph if significance is observed.
-
Figure 6—figure supplement 1—source data 1
Source data related to Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-figsupp1-data1-v1.xlsx
-
Figure 6—figure supplement 1—source data 2
PDF files containing original western blots for Figure 6—figure supplement 1k, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-figsupp1-data2-v1.zip
-
Figure 6—figure supplement 1—source data 3
Original files for western blot analysis displayed in Figure 6—figure supplement 1k.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-figsupp1-data3-v1.zip
-
Figure 6—figure supplement 1—source data 4
PDF files containing original western blots for Figure 6—figure supplement 1i, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-figsupp1-data4-v1.zip
-
Figure 6—figure supplement 1—source data 5
Original files for western blot analysis displayed in Figure 6—figure supplement 1i.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig6-figsupp1-data5-v1.zip
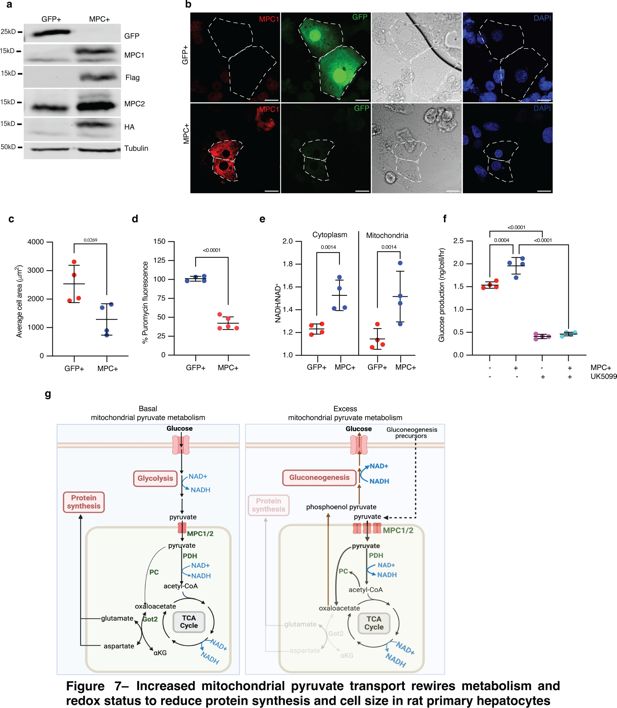
Increased mitochondrial pyruvate transport rewires metabolism and redox status to reduce protein synthesis and cell size in rat primary hepatocytes.
(a) Western blot analysis of MPC expression in cultured rat primary hepatocytes. (b) Representative confocal images of rat primary hepatocytes expressing exogenous GFP or MPC. DIC images and DAPI staining of nuclei are also shown. Cell boundaries are marked with dotted lines. The scale bar represents 20 µm. (c) Quantification of the areas of GFP and MPC expressing primary hepatocytes. Data is presented as mean ± s.d. of six biological replicates, with 20 hepatocytes analyzed for GFP and MPC+. (d) Fluorescence intensity of puromycin-labeled nascent proteins in GFP and MPC+ primary hepatocytes. Data is presented as mean ± s.d. of six biological replicates, with 20 hepatocytes analyzed for each condition. (e) Quantification of glucose in the culture media of primary hepatocytes transfected with GFP or MPC constructs, conditioned with 20 mM lactate and 2 mM pyruvate for 4 hr. 10 µM UK5099 was used to inhibit MPC’s downstream impact on gluconeogenesis. Data is presented as mean ± s.d. of three biological replicates. (f) NADH/NAD+ ratios in the cytoplasmic and mitochondrial fractions of GFP or MPC+ primary hepatocytes. Data is presented as mean ± s.d. of three biological replicates. (g) Schematics illustrating the metabolic consequences of excess mitochondrial pyruvate in hepatocytes. Under normal conditions, mitochondrial pyruvate fuels the TCA cycle, maintaining redox balance and generating sufficient amino acids for cellular homeostasis. However, when mitochondrial pyruvate transport is increased and excess pyruvate is metabolized, both mitochondrial and cytoplasmic redox states are altered. This excess pyruvate enhances the activities of pyruvate carboxylase (PC), pyruvate dehydrogenase (PDH), and the TCA cycle, leading to an elevated NADH/NAD+ ratio. The oxaloacetate produced by PC is converted into phosphoenolpyruvate via PEPCK2, promoting gluconeogenesis. This shift reduces the availability of aspartate and related amino acids necessary for protein synthesis, ultimately resulting in a reduction in cell size without impacting the canonical cell growth signaling pathways. Panel g was created with BioRender.com.
-
Figure 7—source data 1
Source data related to Figure 7.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig7-data1-v1.xlsx
-
Figure 7—source data 2
PDF files containing original western blots for Figure 7a, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig7-data2-v1.zip
-
Figure 7—source data 3
Original files for western blot analysis displayed in Figure 7a.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig7-data3-v1.zip
-
Figure 7—source data 4
Original files for western blot analysis displayed in Figure 7a.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig7-data4-v1.zip
-
Figure 7—source data 5
Original files for western blot analysis displayed in Figure 7a.
- https://cdn.elifesciences.org/articles/103705/elife-103705-fig7-data5-v1.zip
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Drosophila melanogaster) | UAS-MPC1-P2A-MPC2 | Schell et al., 2017 | BDSC, #84087 | Expresses Drosophila Mpc1 and Mpc2 cDNA separated by P2A cleavage site |
| Strain, strain background (D. melanogaster) | hs-Flp1.22 | Bloomington Drosophila Stock Center | BDSC, #77928 | |
| Strain, strain background (D. melanogaster) | Act >CD2>Gal4, UAS-GFP | Bloomington Drosophila Stock Center | BDSC, #4413 | Ay-Gal4 fly stock used to induce mosaics in fat body |
| Strain, strain background (D. melanogaster) | CG-Gal4 | Bloomington Drosophila Stock Center | BDSC, #7011 | |
| Strain, strain background (D. melanogaster) | UAS-S6kSTDETE | Bloomington Drosophila Stock Center | BDSC, #6914 | Drives constitutively active S6k expression |
| Strain, strain background (D. melanogaster) | UAS-RhebPA | Bloomington Drosophila Stock Center | BDSC, #9689 | Drives Rheb overexpression |
| Strain, strain background (D. melanogaster) | UAS-MycOE | Bloomington Drosophila Stock Center | BDSC, #9675 | Drives Myc overexpression |
| Strain, strain background (D. melanogaster) | UAS-JF01761 | Bloomington Drosophila Stock Center | BDSC, #25783 | Drives Myc dsRNA, used as UAS-MycRNAi |
| Strain, strain background (D. melanogaster) | UAS-PI3K93E.Excel | Bloomington Drosophila Stock Center | BDSC, #8287 | Drives PI3K93 overexpression |
| Strain, strain background (D. melanogaster) | UAS-Pi3K92E.A2860C | Bloomington Drosophila Stock Center | BDSC, #8288 | Drives PI3K93 dominant negative |
| Strain, strain background (D. melanogaster) | UAS-Tsc1RNAi | Bloomington Drosophila Stock Center | BDSC, #31314 | Drives Tsc1 dsRNA, used as UAS-Tsc1RNAi |
| Strain, strain background (D. melanogaster) | UAS-hPC | Bloomington Drosophila Stock Center | BDSC, #77928 | Drives human PC cDNA |
| Strain, strain background (D. melanogaster) | UAS-HMC04104 | Bloomington Drosophila Stock Center | BDSC, #56883 | Drives Pc dsRNA, used as UAS-PCRNAi |
| Strain, strain background (D. melanogaster) | UAS-HMS00200 | Bloomington Drosophila Stock Center | BDSC, #36915 | Drives Pepck2 dsRNA, used as UAS-Pepck2RNAi |
| Strain, strain background (D. melanogaster) | UAS-HMC03445 | Bloomington Drosophila Stock Center | BDSC, #51871 | Drives Fbp dsRNA, used as UAS-FbpRNAi |
| Strain, strain background (D. melanogaster) | UAS-GL00009 | Bloomington Drosophila Stock Center | BDSC, #35142 | Drives Pdk dsRNA, used as UAS-PdkRNAi |
| Strain, strain background (D. melanogaster) | UAS-NMNAT | Bloomington Drosophila Stock Center | BDSC, #39702 | Drives Nmnat cDNA |
| Strain, strain background (D. melanogaster) | UAS-CintNDX | Bloomington Drosophila Stock Center | BDSC, #93883 | Drives Cionia intestinalis NDX cDNA |
| Strain, strain background (D. melanogaster) | UAS-ScerNDI1 | Bloomington Drosophila Stock Center | BDSC, #93878 | Drives Saccharomyces cerevisiae NDI cDNA |
| Strain, strain background (D. melanogaster) | UAS-slif | Bloomington Drosophila Stock Center | BDSC, #52661 | Drives slimfast cDNA |
| Strain, strain background (D. melanogaster) | UAS-HMS05873 | Bloomington Drosophila Stock Center | BDSC, #78778 | Drives Got2 dsRNA, used as UAS-Got2RNAi |
| Strain, strain background (D. melanogaster) | UAS-Mpc1RNAi | Bricker et al., 2012 | Drives Mpc1 dsRNA | |
| Strain, strain background (D. melanogaster) | UAS-PdhaRNAi | National Institute of Genetics, Japan | NIG 7010 R-3 | Drives Pdha dsRNA |
| Strain, strain background (D. melanogaster) | UAS-DlatRNAi | National Institute of Genetics, Japan | NIG 5261 R-3 | Drives Dlat dsRNA |
| Strain, strain background (D. melanogaster) | w1118 | Bloomington Drosophila Stock Center | BDSC:3605 | Wild-type fly strain |
| Strain, strain background (D. melanogaster) | UAS-GOT2 | This paper | see materials and methods | Drives Got2 cDNA, available at Bloomington Drosophila Stock Center |
| Strain, strain background (D. melanogaster) | Pcb KO | This paper | see materials and methods | Pcb CRISPR KO, available at Bloomington Drosophila Stock Center |
| Strain, strain background (D. melanogaster) | Pepck2 KO | This paper | see materials and methods | Pepck2 CRISPR KO, available at Bloomington Drosophila Stock Center |
| Cell line (Homo sapiens) | HepG2 | ATCC | Cat# HB-8065 | a hepatocellular carcinoma cell line |
| Cell line (Rattus rattus) | Cryopreserved Rat Primary Hepatocytes | Lonza | Cat# RWCP01 | Plateable, Rat Wistar Hannover Hepatocytes, cryopreserved |
| Antibody | anti-puromycin [3RH11] (host: mouse monoclonal) | Kerafast | Cat# EQ0001 | Dilution factor 1:1000 for western blot 1:200 for immunofluorescence |
| Antibody | anti-MPC1 (Drosophila) (host: rabbit monoclonal) | gift from R. Kletzein | Dilution factor 1:200 for immunofluorescence | |
| Antibody | anti-MPC2 (Drosophila) (host: rabbit monoclonal) | gift from R. Kletzein | Dilution factor 1:200 for immunofluorescence | |
| Antibody | anti-p-S6 (host: rabbit monoclonal) | gift from A. Teleman | Dilution factor 1:200 for immunofluorescence | |
| Antibody | anti-dFoxo (host: rabbit monoclonal) | gift from P. Bellosta | Dilution factor 1:500 for immunofluorescence | |
| Antibody | anti-p-4EBP (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 2855 | Dilution factor 1:1000 for western blot 1:500 for immunofluorescence |
| Antibody | anti-p-eIF2α (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 9721 | Dilution factor 1:500 for immunofluorescence |
| Antibody | Cy3 conjugated anti-rabbit (host: donkey polyclonal) | Jacksons Immuno Research Laboratories | Cat# 711–165–152 | Dilution factor 1:400 for immunofluorescence |
| Antibody | Cy3 conjugated anti-mouse (host: donkey polyclonal) | Jacksons Immuno Research Laboratories | Cat# 115-165-166 | Dilution factor 1:400 for immunofluorescence |
| Antibody | anti-MPC1 (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 14462 | Dilution factor 1:1000 for western blot |
| Antibody | anti-MPC2 (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 46141 | Dilution factor 1:1000 for western blot |
| Antibody | anti-PDH (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 3205 | Dilution factor 1:1000 for western blot |
| Antibody | anti- p-PDH (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 31866 | Dilution factor 1:1000 for western blot |
| Antibody | anti-PC (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 66470 | Dilution factor 1:1000 for western blot |
| Antibody | anti-PEPCK2 (D3E11) (host: rabbit monoclonal) | Cell Signaling Technologies | Cat# 8565 | Dilution factor 1:1000 for western blot |
| Antibody | anti-Got2 (host: rabbit monoclonal) | Sigma | Cat# HPA018139 | Dilution factor 1:1000 for western blot |
| Antibody | anti-tubulin (DM1A) (host: mouse monoclonal) | Cell Signaling Technologies | Cat# 3873 | Dilution factor 1:1000 for western blot |
| Antibody | anti- Flag M2 (host: mouse monoclonal) | Sigma-Aldrich | Cat# F1800 | Dilution factor 1:10000 for western blot |
| Antibody | IRDye 680RD anti-mouse (host: Donkey monoclonal) | Li-COR | Cat# 926–68072 | Dilution factor 1:5000 for western blot |
| Antibody | IRDye 800RD anti-Rabbit (host: Donkey monoclonal) | Li-COR | Cat# 926–32213 | Dilution factor 1:5000 for western blot |
| Recombinant DNA reagent | pUAST-aatB | |||
| Recombinant DNA reagent | pLVX-TetOne-Zeocin | Takara | ||
| Recombinant DNA reagent | Gag-Pol | Addgene | ||
| Recombinant DNA reagent | VSVG | Addgene | ||
| Recombinant DNA reagent | pMD2.G | Addgene | ||
| Recombinant DNA reagent | pLenti-CMV-Blast | Addgene | Cat# 17486 | |
| Recombinant DNA reagent | pLVX-TetOne- HA-MPC2-P2A-T2A-MPC1-FLAG-Zeo | This paper | HA-tagged MPC2 and Flag-tagged MPC1 cDNA separated by P2A/T2A cleavage site cloned into pLenti-CMV-Blast | Drives Got2 cDNA, available at Bloomington Drosophila Stock Center |
| Recombinant DNA reagent | pLenti-CMV-PC-V5-Blast | This paper | V5-tagged PC cDNA cloned into pLenti-CMV-Blast | Drives human PC cDNA tagged with V5, available at request to lead contact |
| Recombinant DNA reagent | pLenti-CMV-GOT2-V5-Blast | This paper | V5-tagged GOT2 cDNA cloned into pLenti-CMV-Blast | Drives human GOT2 cDNA tagged with V5, available at request to lead contact |
| Recombinant DNA reagent | PB-CAG-GFPd2 | Addgene | Cat# 115665 | |
| Recombinant DNA reagent | PMXS-NDI | Addgene | Cat# 72876 | |
| Recombinant DNA reagent | pLenti-CMV-d2GFP-Blast | This paper | d2GFP cloned into pLenti-CMV-Blast | Drives human d2GFP cDNA, available at request to lead contact |
| Recombinant DNA reagent | pLenti-CMV-NDI-V5-Blast | This paper | V5-tagged NDI cloned into pLenti-CMV-Blast | Drives yeast NDI cDNA tagged with V5, available at request to lead contact |
| Recombinant DNA reagent | pLenti-CMV-SLC1A3-V5-Blast | This paper | V5-tagged SLC1A3 cDNA cloned into pLenti-CMV-Blast; see materials and methods | Drives human SLC1A3 cDNA tagged with V5, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2-blasticidin | Addgene | Cat# 83480 | |
| Recombinant DNA reagent | lentiCRISPRv2-hPCg5e-blast | This paper | sgRNA targeting human PC exon 5 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human PC exon 5, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2-hPCg6e-blast | This paper | sgRNA targeting human PC exon 6 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human PC exon 6, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2- hPEPCK2g2e-blast | This paper | sgRNA targeting human PEPCK2 exon 2 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human PEPCK2 exon 2, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2- hPEPCK2g3e-blast | This paper | sgRNA targeting human PEPCK2 exon 3 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human PEPCK2 exon 3, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2- hGOT2ga-blast | This paper | sgRNA targeting human GOT2 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human GOT2, available at request to lead contact |
| Recombinant DNA reagent | lentiCRISPRv2- hGOT2gb-blast | This paper | sgRNA b targeting human GOT2 in lentiCRISPRv2 vector; see Materials and methods | Drives sgRNA targeting human GOT2, available at request to lead contact |
| Recombinant DNA reagent | pUAST-aatB-dGOT2 | This paper | dGOT2 cDNA was cloned into pLenti-CMV-Blast | Drives human GOT2, available at request to lead contact |
| Commercial assay or kit | Annexin V/PI staining kit | Molecular Probes | Cat# V13241 | |
| Commercial assay or kit | Click-iT Plus OPP Alexa Fluor 594 kit | Molecular Probes | Cat# C10457 | |
| Commercial assay or kit | Click-iT HPG Alexa Fluor 594 kit | Molecular Probes | Cat# C10429 | |
| Commercial assay or kit | Triglycerides Reagent | Thermo Fisher Scientific | Cat# TR22421 | |
| Commercial assay or kit | BCA Protein Assay Reagent | Thermo Fisher Scientific | Cat# 23225 | |
| Commercial assay or kit | TRIzol Reagent | Thermo Fisher Scientific | Cat# 15596026 | |
| Commercial assay or kit | Amplite Fluorimetric NAD/NADH ratio assay kit | AAT Bioquest | Cat# 15263 | |
| Commercial assay or kit | Click-iT Plus EdU Alexa Fluor 594 | Molecular Probes | Cat# C10639 | |
| Commercial assay or kit | NucleoSpin RNA kit | Takara Bio USA, Inc | Cat# 740955.50 | |
| Commercial assay or kit | Autokit Glucose reagent | Wako | Cat# 997–03001 | |
| Commercial assay or kit | Amplex Red Glucose Assay Kit | Thermo Fisher Scientific | Cat# A22189 | |
| Chemical compound, drug | paraformaldehyde | Sigma-Aldrich | Cat# P6148 | |
| Chemical compound, drug | Rhodamine Phalloidin | Thermo Scientific | Cat# R418 | |
| Chemical compound, drug | DAPI-supplemented VectaShield | Vector Labs | Cat# H1200 | |
| Chemical compound, drug | Normal Goat Serum | Jackson ImmunoResearch Laboratories | Cat# 005-000-121 | |
| Chemical compound, drug | Human Plasma-Like Medium | Gibco | Cat# 765090 | |
| Chemical compound, drug | doxycycline | Sigma-Aldrich | Cat# #D5207 | |
| Chemical compound, drug | UK5099 | Sigma-Aldrich | Cat# PZ0160-5MG | |
| Chemical compound, drug | glycine | Sigma-Aldrich | Cat# G7403 | |
| Chemical compound, drug | alanine | Sigma-Aldrich | Cat# A7469 | |
| Chemical compound, drug | asparagine | Sigma-Aldrich | Cat# A4159 | |
| Chemical compound, drug | aspartic acid | Sigma-Aldrich | Cat# A7219 | |
| Chemical compound, drug | glutamic acid | Sigma-Aldrich | Cat# G8415 | |
| Chemical compound, drug | proline | Sigma-Aldrich | Cat# P5607 | |
| Chemical compound, drug | serine | Sigma-Aldrich | Cat# S4311 | |
| Chemical compound, drug | AZD7545 | MedChemExpress, | Cat# HY-16082 | |
| Chemical compound, drug | dichloroacetate | Sigma-Aldrich | Cat# 347795 | |
| Chemical compound, drug | duroquinone | Sigma-Aldrich | Cat# D223204 | |
| Chemical compound, drug | gramicidin | Sigma-Aldrich | Cat# G5002 | |
| Chemical compound, drug | nicotinamide riboside | Sigma-Aldrich | Cat# SMB00907 | |
| Chemical compound, drug | MEM non-essential amino acids | Gibco | Cat# 11140050 | |
| Chemical compound, drug | Nicotinamide mononucleotide | Sigma-Aldrich | Cat# N3501 | |
| Chemical compound, drug | Vybrant DyeCycle Violet stain | Molecular Probes | Cat# V35003 | |
| Chemical compound, drug | Eagle’s Minimal Essential Medium | ATCC | Cat# 30–2003 | |
| Chemical compound, drug | Camptothecin | MedChemExpress | Cat# HY-16560 | |
| Chemical compound, drug | Shields Sang M3 Insect Media | Sigma-Aldrich | Cat# S8398 | |
| Chemical compound, drug | puromycin | Sigma-Aldrich | Cat# P4512 | |
| Chemical compound, drug | methionine-free DMEM | Gibco | Cat# 21013024 | |
| Chemical compound, drug | LipidTOX Red | Invitrogen | Cat# H34351 | |
| Chemical compound, drug | SuperScript II Reverse Transcriptase | Molecular Probes | Cat# 18064–022 | |
| Chemical compound, drug | PowerUp SYBR Green Master Mix | Applied Biosystems | Cat# 2828831 | |
| Chemical compound, drug | 13C Glucose | Millipore Sigma | Cat# 389374 | |
| Chemical compound, drug | DMEM, no glucose, no glutamine, and no phenol red | Gibco | Cat# A1443001 | |
| Chemical compound, drug | Sodium L-Lactate-C13 solution | Millipore Sigma | Cat# 490040 | |
| Chemical compound, drug | Hepatocyte Plating Medium | Lonza | Cat# MP100 | |
| Chemical compound, drug | Hepatocyte Basal Medium | Lonza | Cat# CC-4182 | |
| Chemical compound, drug | Lipofectamine 3000 | Invitrogen | Cat# L3000001 | |
| Chemical compound, drug | Triton X-100 | Sigma-Aldrich | Cat# X100 | |
| Chemical compound, drug | FBS | Sigma-Aldrich | Cat# F0926 | |
| Chemical compound, drug | PenStrep | Thermo Fisher Scientific | Cat# 15140 | |
| Chemical compound, drug | NP-40 | Millipore | Cat# 492018 | |
| Chemical compound, drug | sodium deoxycholate | Sigma-Aldrich | Cat# D6750 | |
| Chemical compound, drug | SDS | Sigma-Aldrich | Cat# L3771 | |
| Chemical compound, drug | EDTA | Sigma-Aldrich | Cat# E9884 | |
| Chemical compound, drug | Tris-HCl | Roche | Cat# 10812846001 | |
| Software | FIJI | NIH Image | RRID:SCR_002285 | https://fiji.sc/ |
| Software | Prism | GraphPad | RRID:SCR_002798 | https://www.graphpad.com/ |
| Software | FlowJo | FlowJo | RRID:SCR_008520 | https://www.flowjo.com/solutions/flowjo/downloads |
| Other | Ultra Low Cluster, 96 well, Ultra Low Attachment Polystyrene | Costar | Cat# 7007 | Corning 96-well Clear Round Bottom Ultra-Low Attachment Microplate used to make spheroids of HepG2 |
| Other | Dumont 5 forceps | Fine Science Tools | Cat# 11254–20 | Forceps used to dissect Drosophila fat bodies |






