Drosophila hamlet mediates epithelial tissue assembly of the reproductive system
Figures
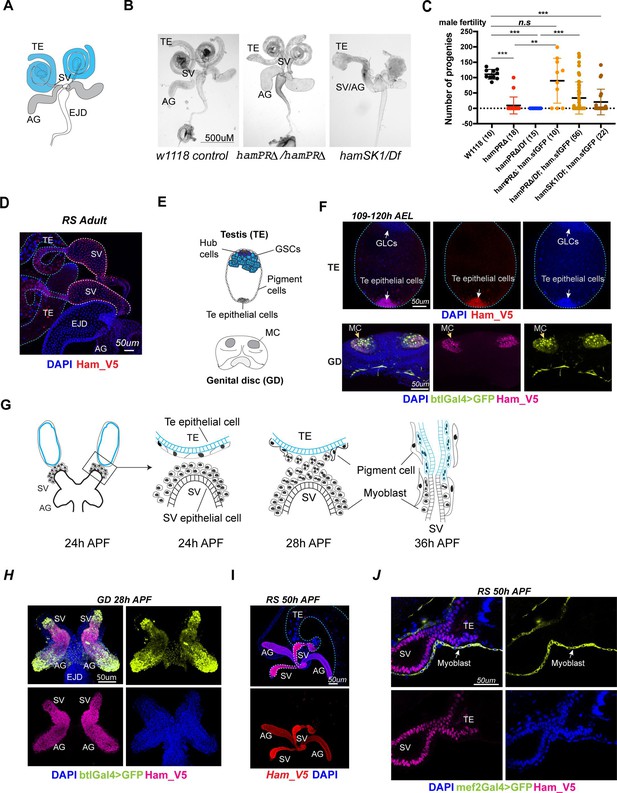
ham is expressed and required for Drosophila male reproductive system (RS) development.
(A) The adult male RS. TE, testis; SV, seminal vesicle; AG, accessory gland; EJD, ejaculatory duct. (B) White-field images of adult male RS, showing defective morphology in the ham mutants. (C) Quantification of male fertility. Ham_sfGFP with the ham genomic region was used to rescue the sterility. Statistical significance was calculated using unpaired t-test (***p < 0.001; **p < 0.01; n.s., non-significant). (D) Images of the adult RS stained with Ham_V5 in red. DNA is stained with DAPI in blue. The blue and white dashed lines highlight the TE and SV, respectively. (E) Illustration of the developing male RS at the larval stage. The testis (TE) terminal epithelial cells localize at the opposite side of the germline stem cell (GSC) niche. The genital disc (GD) contains mesenchymal cells (MC) that will develop into the SV and accessory gland (ag). (F) Images of the larval TE (top) and GD (bottom) stained with Ham_V5 in red or magenta, the reporter in lime-green (btlGal4 driving UAS-GFP) and DAPI in blue. AEL, after egg laying. (G) Illustration of the TE and SV fusion process. APF, after pupal formation. (H) Images of the pupal GD stained with Ham_V5 in magenta, the reporter in lime-green (btlGal4 driving UAS-GFP) and DAPI in blue. (I) Images of the pupal RS with Ham_V5 in red and DAPI in blue. The blue and white dashed lines highlight the TE and SV, respectively. (J) Higher magnification image of the joint site between the SV and TE, with myoblasts and myotubes labeled by mef2Gal4>GFP in lime-green.
-
Figure 1—source data 1
Raw counting data for the fertility test in Figure 1C.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig1-data1-v1.xlsx

Ham gene locus and mutant alleles.
(A) Illustration of the ham genomic locus and predicted protein isoforms, highlighting the mutation sites for the three mutant alleles and the protein domains of each isoform. All Ham protein isoforms maintain a functionally unknown motif PTZ00121 and the C-terminal zinc finger (ZF) clusters, while the long isoforms B/D and G also comprise the PR domain, the N-terminal ZF clusters. Isoform H has a truncated PR domain, and the shortest isoform E lacks the PR domain and the N-terminal ZF clusters. (B) Flanking sequences of the sgRNA target site from wt and four alleles with indels. Red letters indicate the sgRNA sequence. The corresponding amino acid sequence is shown above. ‘*’ indicates the stop codon. (C) An image from western-blot, showing the two Ham protein bands in the control samples and absence of the Ham band(s) in the indicated mutants. ‘*’ indicates an unspecific band, used as the loading control. (D) A table showing viability and sensory organ (SO) defects in four ham alleles. Note: the –6nt allele has a 6-nucleotide in-frame deletion. (E) Mechano-sensory organ phenotypes in the three indicated alleles. Red arrowheads indicate double sockets and double shafts phenotype.
-
Figure 1—figure supplement 1—source data 1
This zip archive contains the raw unedited western blot shown in Figure 1—figure supplement 1C.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig1-figsupp1-data1-v1.zip
-
Figure 1—figure supplement 1—source data 2
This zip archive contains the original western blot shown in Figure 1-figure supplement 1C with relevant band labeled.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig1-figsupp1-data2-v1.zip

Ham expression and function in the female reproductive system (RS).
(A) Bright-field images of the adult female RS from the indicated genotypes, showing accumulated eggs in the ham mutant ovaries. (B) Quantification of the number of progenies produced by wild-type and ham mutant females after crossed with wild-type males. Statistical significance was calculated using unpaired t-test (***p < 0.001). (C) Images of pupal and adult ovaries and oviducts, showing high expression of the Ham_V5 protein in the connecting sites of the ovaries and oviducts.
-
Figure 1—figure supplement 2—source data 1
Quantification of female fertility shown in Figure 1—figure supplement 2B.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig1-figsupp2-data1-v1.xlsx
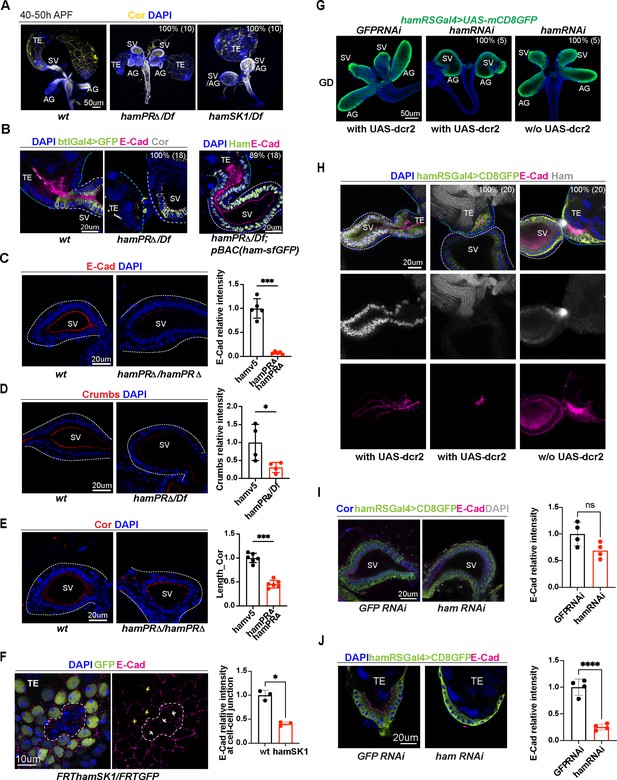
Ham promotes differentiation of the TE terminal and seminal vesicle (SV) epithelial cells.
(A) Images of the male reproductive system (RS) at the pupal stage, showing morphological defects of the TE and SV in the ham mutants. Epithelial cells are labeled with Coracle (Cor). The phenotype penetrance (%) and number of animals (n) counted are indicated as % (n). (B) High magnification images of the connection site of the TE and SV, showing that the ham mutant RS failed to form a continuous tube. Ham-sfGFP restored the normal morphology. Epithelial cells are marked by E-Cad in red, Cor in yellow, and the epithelium on the SV side also marked by btlGal4 driving UAS-mCD8GFP. DNA is labeled by DAPI in blue. (C) Images and quantification of the E-Cad signal in the wt and hamPRΔ mutant SV. n = 5. (D) Images and quantification of the Crumbs signal in the wt and hamPRΔ /Df mutant SV. n = 4. (E) Images and quantification of the length of Cor signal in the wt and hamPRΔ mutant SV. n = 6. (F) Images of ham mutant mosaic clones in the TE terminal epithelial cells. Note, the mutant cells are GFP negative and within the area surrounded by the white dashed line. The white arrows indicate junctions between mutant cells, while the yellow arrows highlight the junctions between wild-type cells. n = 3. (G) Images of pupal male genital disc (GD), showing that hamRNAi-mediated knockdown reproduces ham mutant phenotypes. (H) Images of pupal male RS, stained with E-Cad in red, Ham in yellow, GFP in green and DAPI in blue. UAS-dcr2 enhanced knockdown efficiency and led to more severe morphological defects. (I) Images and quantification of E-Cad signal in red and Cor length in blue in control and ham RNAi SV. n = 4. (J) Images and quantification of E-Cad signal in control and ham RNAi TE terminal epithelium. n = 4. In the quantification graphs, statistical significance was calculated using unpaired t-test except for TE mosaic clones in F which used paired t-test (**** p < 0.0001, ***p < 0.001; **p < 0.01; *p < 0.05; ns, non-significant).
-
Figure 2—source data 1
Quantification of the E-Cad signal in the wt and hamPRΔ mutant SV shown in Figure 2C.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data1-v1.xlsx
-
Figure 2—source data 2
Quantification of the Crumbs signal in the wt and hamPRΔ /Df mutant SV shown in Figure 2D.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data2-v1.xlsx
-
Figure 2—source data 3
Quantification of the length of Cor signal in the wt and hamPRΔ mutant SV shown in Figure 2E.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data3-v1.xlsx
-
Figure 2—source data 4
Quantification of the E-Cad signal at the border between two wt and between two hamSK1 mutant cells in the TE terminal epithelium shown in Figure 2F.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data4-v1.xlsx
-
Figure 2—source data 5
Quantification of E-Cad signal in GFP and ham.RNAi SV shown in Figure 2I.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data5-v1.xlsx
-
Figure 2—source data 6
Quantification of E-Cad signal in GFP and ham.RNAi TE terminal epithelium shown in Figure 2J.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-data6-v1.xlsx

Split channel images for images in Figure 2.

Ham controls the formation of TE epithelial cells.
(A) Images of the testes from the early L3 larval stage, showing the absence of or mis-polarized TE terminal epithelial cell cluster in the ham mutant. E-Cad marks epithelial cells in green and DAPI marks the nuclei. The white arrows indicate mis-polarized epithelial cells. (B) Illustration of the precursor cell clusters of the male reproductive system (RS) in the embryonic stages. PGC, primordium germ cells; SGPs, somatic gonadal precursors; msSGP, male-specific SGPs. (C–E) Images of stage S13 and S15 wt embryos, showing Ham staining in all SGPs. Abd-B and Sox100B label msSGPs, VASA labels PGCs. Images of the wt and ham mutant embryos at S13 (F) and S15 (G), showing the reduction or absence of msSGPs in the hamSK1 mutant. Vasa in magenta is a germ cell marker, and Sox100B labels msSGPs in lime-green.

An epithelial-specific Gal4 line in the ham locus.
(A) The ham gene locus, indicating the fragments in the Gal4 transgenes. The one (hamRSGal4) that drives reporter gene expression in the reproductive system (RS) epithelial cells is highlighted in orange. (B) Images of pupal genital disc and TE in animals with hamRSGal4 and UAS-mCD8GFP. Ham_V5 is stained in magenta, and DAPI in blue. (C) Images showing the presence of myotubes on the surface of the testis. The bar graph on the right shows the relative abundance of myotubes on the testis in wt (100%) and ham mutant condition quantified from five animals each. (D) Images showing the junction between the TE and seminal vesicle (SV), and myotubes are present even in the ham mutant. The white arrows indicate myotubes with two or three nuclei at the junction.
-
Figure 2—figure supplement 3—source data 1
Quantification of myotube number on the testis in wt and hamPRΔ/Df shown in Figure 2—figure supplement 3C.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig2-figsupp3-data1-v1.xlsx
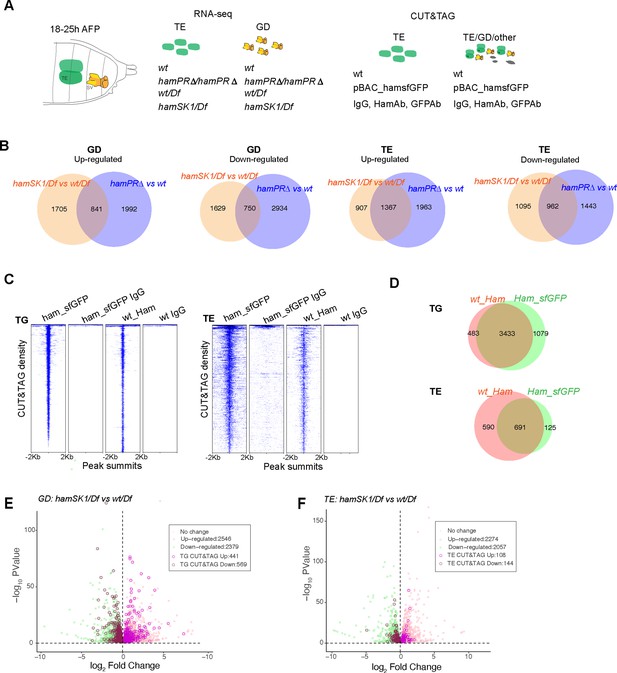
Identification of Ham-regulated genes in the developing TE and genital disc (GD).
(A) Illustration of the sample types and antibodies used in the RNA-seq and CUT&TAG experiments. (B) Venn diagram of dysregulated genes between the two mutant conditions in the GD (left) and TE (right) samples. (C) Heatmaps of Ham CUT&TAG peak density and IgG control, ranked from high to low and centered at peak summits. (D) Venn diagrams of CUT&TAG peaks from the two types of samples. Volcano plots of dysregulated genes in GD (E) and TE (F) in the hamSK1/Df mutant, and the ones with at least one Ham CUT&TAG peak are highlighted.

Ham regulatory activity and potential co-factors.
(A) Principle component analysis of the RNA-seq datasets. (B) Principle component analysis of the Cut&Tag datasets. Note: some samples had poor sequencing depth, which were excluded from the PCA and later analyses. (C) The shot locus, showing the CUT&TAG tracks from the TE and GD (TG) mixed samples and the enhancer fragments in the luciferase assays. (D) Normalized luciferase activity to the empty reporter and then to the control vector that expresses GFP and RFP proteins. Each plot represents the average from three biological replicates with each biological replicate having four technical replicates. Statistical significance was calculated using unpaired t-test (****p < 0.0001; ***p < 0.001; **p < 0.01; *p < 0.05). (E) Significantly enriched motifs from Ham binding peaks in upregulated genes (left) and downregulated genes (right). The dashed rectangles highlight MAD and ZNF263 motifs.
-
Figure 3—figure supplement 1—source data 1
Raw data of luciferase activity shown in Figure 3—figure supplement 1D.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig3-figsupp1-data1-v1.xlsx
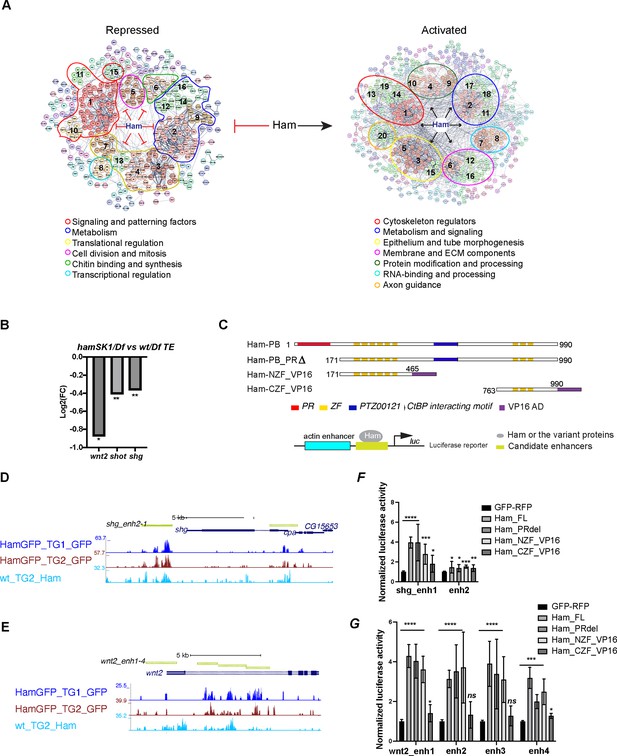
Ham activates expression of epithelial differentiation genes.
(A) Ham gene network analysis from STRING. The position of MCL clusters were manually adjusted to place genes with similar function in close proximity. The functional categories were determined by gene function tools in STRING. (B) A bar plot of the log2 fold-change value from RNA-seq for wnt2, shot, and shg, showing downregulation of these genes in the hamSK1/Df mutant. (C) Schemes of the Ham_FL and other truncation proteins, and illustration of the luciferase reporter assay. (D, E) A genome browser snapshot of the shg and wnt2 loci, showing the CUT&TAG tracks from the TE and GD (TG) mixed samples and the enhancer fragments in the luciferase assays. (F, G) Normalized luciferase activity to the empty reporter and then to the control vector that expresses GFP and RFP proteins. Each plot represents the average from three biological replicates with each biological replicate having four technical replicates. Statistical significance was calculated using unpaired t-test (****p < 0.0001; ***p < 0.001; **p < 0.01; *p < 0.05; ns, non-significant).
-
Figure 4—source data 1
Raw data of luciferase activity shown in Figure 4F.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig4-data1-v1.xlsx
-
Figure 4—source data 2
Raw data of luciferase activity shown in Figure 4G.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig4-data2-v1.xlsx
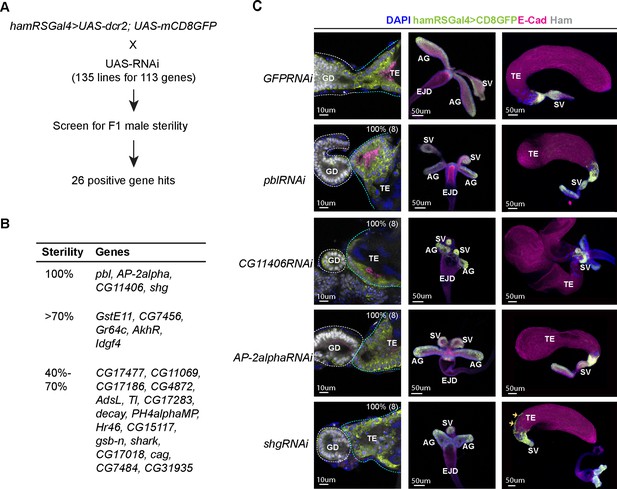
A candidate RNAi screen identified Ham-downstream effectors in reproductive system (RS) epithelial tissue fusion.
(A) Scheme for the RNAi screen. (B) Summary of positive hits. The extent of sterility is determined by the number of progenies from the F1 male crossing to the wild-type female (see Methods). 100% means no progeny. >70%, the number of progenies being fewer than 30% of the number of progenies produced by the control cross (the same Gal4 driver × UAS-GFPRNAi). 40–70%, the number of progenies being between 30% and 60% of the number of progenies produced by the control. (C) Images of the control and gene-specific RNAi knockdown RS. The left panels show the fusion site stained with E-Cad in magenta, GFP in lime-green, Ham in gray and DAPI in blue, the middle panels are genital disc (GD), and the right ones are TE. The orange arrows indicate mobilized epithelial cells in the shg knockdown TE.
-
Figure 5—source data 1
Results for a candidate RNAi screen shown in Figure 5A.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig5-data1-v1.xlsx
-
Figure 5—source data 2
The extent of sterility for positive hits from the RNAi screen shown in Figure 5B.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig5-data2-v1.xlsx

Split channel images for images in Figure 5.

Split channel images for images in Figure 5.
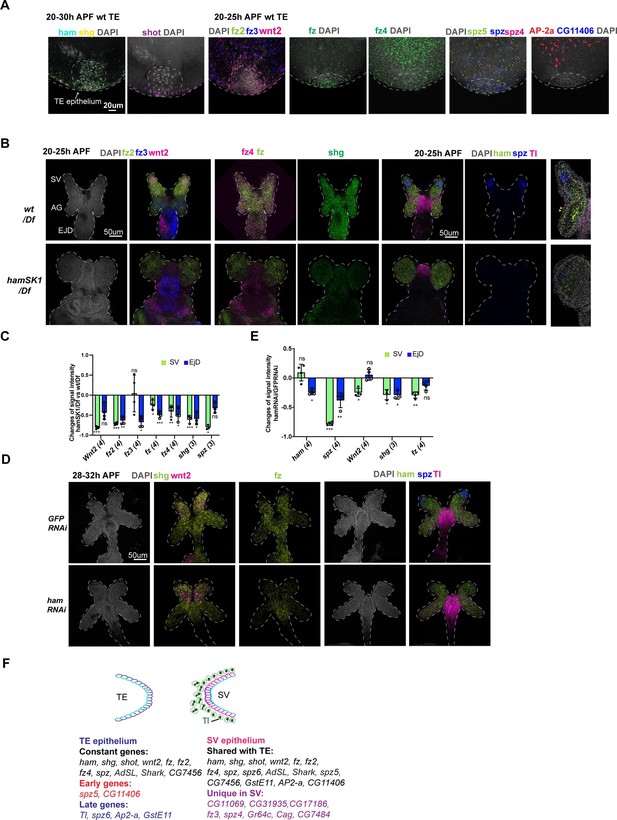
Spatial–temporal expression dynamics of Ham downstream genes.
(A) Images from multiplexed in situ hybridization (the SCRINSHOT method), showing mRNA signal of the indicated genes in wt TE samples. DNA is marked by DAPI in gray. (B) Images from SCRINSHOT, showing mRNA signals of indicated genes in wt/Df and hamSK1/Df mutant GD. The last panel is an enlarged view from the sample with Tl, ham, and spz probes. The yellow arrowheads indicate myoblast cells with Tl signal. (C) Quantification of signal intensity for genes shown in B in two areas, the SV and EjD. Statistical significance was calculated using unpaired t-test (***p < 0.001; **p < 0.01; *p < 0.05; ns, non-significant). (D) Images from SCRINSHOT, showing mRNA signals of indicated genes in GFP and ham RNAi GD samples. (E) Quantification of signal intensity for genes shown in D in two areas, the SV and EjD. Statistical significance was calculated using unpaired t-test (***p < 0.001; **p < 0.01; *p < 0.05; ns, non-significant). (F) A summary of genes expressed in the TE terminal and SV epithelium. Highlighted genes show temporal or spatial dynamic expression in the two tissues.
-
Figure 6—source data 1
Quantification of SCRINSHOT signal intensity changes for indicated genes between hamSK1/Df and wt/Df in the areas of SV and EjD, as shown in Figure 6C.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig6-data1-v1.xlsx
-
Figure 6—source data 2
Quantification of SCRINSHOT signal intensity changes for indicated genes between hamRNAi and GFPRNAi in the areas of SV and EjD, as shown in Figure 6E.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig6-data2-v1.xlsx

Expression of genes tested by SCRINSHOT in RNA-seq data.
(A) Log2FC (hamSK1/Df versus wt/Df and hamPRΔ versus wt) in TE. (B) Log2FC (hamSK1/Df versus wt/Df and hamPRΔ versus wt) in genital disc (GD).

Split channel images for images in Figure 6.

Additional genes tested in SCRINSHOT.
(A) Images from SCRINSHOT, showing mRNA signal of the indicated genes in wt TE samples. DNA is marked by DAPI in gray. (B) Images from SCRINSHOT, showing mRNA signal of the indicated genes in wt TE samples at a later pupal stage (28–32 APF). DNA is marked by DAPI in gray. (C) Images from SCRINSHOT, showing mRNA signals of indicated genes in wt/Df and hamSK1/Df mutant GDs. (D) Images from SCRINSHOT, showing mRNA signals of indicated genes in GFP and ham RNAi GD samples. (E, F) Quantification of signal intensity for genes shown in C and D in two areas, the SV and EjD. (G) Quantification of signal intensity for genes that changed expression in ham RNAi TE. Statistical significance was calculated using unpaired t-test (***p < 0.001; **p < 0.01; *p < 0.05; ns, non-significant).
-
Figure 6—figure supplement 3—source data 1
Quantification of SCRINSHOT signal intensity changes for indicated genes between hamSK1/Df and wt/Df in the areas of SV and EjD, as shown in Figure 6—figure supplement 3E.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig6-figsupp3-data1-v1.xlsx
-
Figure 6—figure supplement 3—source data 2
Quantification of SCRINSHOT signal intensity changes for indicated genes between hamRNAi and GFPRNAi in the areas of SV and EjD, as shown in Figure 6—figure supplement 3F.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig6-figsupp3-data2-v1.xlsx
-
Figure 6—figure supplement 3—source data 3
Quantification of signal intensity for genes that changed expression in ham RNAi TE shown in Figure 6—figure supplement 3G.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig6-figsupp3-data3-v1.xlsx

Split channel images for images in Figure 6—figure supplement 3.

Additional split channel images for the images in Figure 6—figure supplement 3.
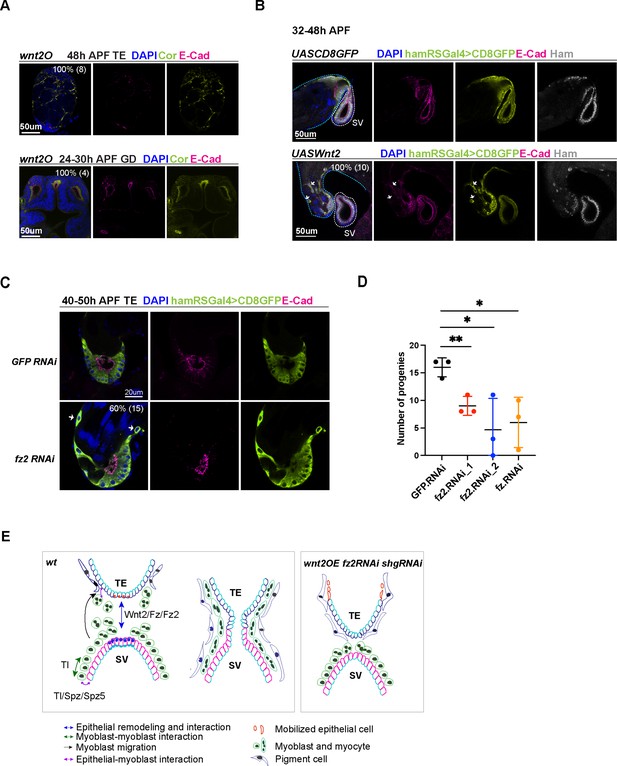
Wnt2 and its ligand genes are required for male fertility and reproductive system (RS) development.
(A) Images of wnt2o mutant TE and GD, showing disrupted morphology in these tissues. (B) Images of the TE and SV fusion site from the control and wnt2 overexpression animals. The epithelial cells of the TE and SV are labeled with hamRSGal4 driving UAS-mCD8GFP in lime-green and E-Cad in magenta and Ham in gray. The white arrows indicate mobilized GFP positive cells in the middle of the TE, and these cells have a lower level of E-Cad. The blue and white dashed lines highlight the TE and SV, respectively. (C) Images of the TE and SV fusion site from the control and fz2 RNAi animals. The white arrows indicate mobilized GFP positive cells in the middle of the TE, and these cells have a lower level of E-Cad. (D) Graphs of progeny number from male fertility tests for fz and fz2 RNAi lines in comparison to GFP RNAi. n = 3. Statistical significance was calculated using unpaired t-test (**, p< 0.01; *, p<0.05). (E) Illustration of cell–cell interactions in the developing RS, and a speculated model for Tl and Wnt2 signaling functions. Wnt2 signaling may be involved in epithelial cell interaction between the two ends. Tl signaling and Tl may be involved in the interaction between epithelial cells and myoblasts/myotubes. In three conditions (shgRNAi, wnt2OE, and fz2RNAi), some TE terminal epithelial cells (illustrated in orange) move up to the distal end of the testis.
-
Figure 7—source data 1
Number of progenies from male fertility tests for fz and fz2.RNAi lines in comparison to GFP.RNAi shown in Figure 7D.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig7-data1-v1.xlsx

Male fertility test for Toll-9.
(A) Graphs of progeny number from male fertility test for hamRSGal4 driving Toll-9 RNAi in comparison to GFP RNAi. The number of tested males = 3. Statistical significance was calculated using unpaired t-test (***, p<0.001). (B) White-field image of Toll-9 RNAi adult male reproductive system (RS), showing morphological defects of the TE and the phenotype penetrance (11% of 37 animals). (C) Images of adult RS from control GFP RNAi andToll-9 RNAi males. hamRSGal4 >UAS-mCD8GFP is used to label the testis terminal epithelium and the SV, and DAPI stains DNA. The panels below are higher magnification views on the selected SV area, highlighted with white dashed squares. White arrows indicate sperms in the control SV, which are absent from the Toll-RNAi sample. The bar graph shows the percentage of SV with sperms in GFP and Toll-9 RNAi animals (N = 10).
-
Figure 7—figure supplement 1—source data 1
Number of progenies from male fertility test for hamRSGal4 driving Toll-9 RNAi in comparison to GFP RNAi shown in Figure 7—figure supplement 1A.
- https://cdn.elifesciences.org/articles/104164/elife-104164-fig7-figsupp1-data1-v1.xlsx
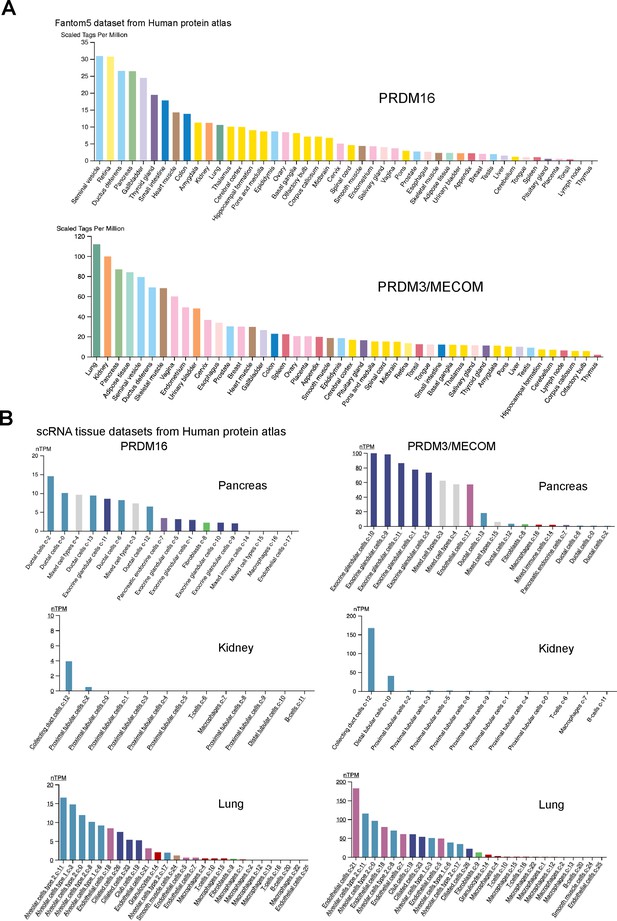
PRDM16 and MECOM/PRDM3 expression in human tissues.
(A) RNA expression data across human tissues through Cap analysis of gene expression (Fantom5 dataset from the Human Protein Atlas). (B) Expression in single-cell RNA-seq clusters in the indicated human tissues (from the Human Protein Atlas).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (D. melanogaster) | w1118 | Bloomington Drosophila Stock Center | BDSC:3605 FLYBase: FBst0003605; RRID:BDSC_3605 | FlyBase symbol: w1118 |
| Genetic reagent (D. melanogaster) | hamPR Δ FRT40A/Cyo | This paper | A new hamlet mutant fly line | |
| Genetic reagent (D. melanogaster) | hamSK1 FRT40A/Cyo | This paper | A new hamlet mutant fly line | |
| Genetic reagent (D. melanogaster) | ham1,FRT40A/Cyo | Moore et al., 2002 DOI: 10.1126/science.1072387 | ||
| Genetic reagent (D. melanogaster) | pBAC_ham_sfGFP | Bloomington Drosophila Stock Center | BDSC:83660 FLYBase: FBst0083660; RRID:BDSC_83660 | FlyBase symbol: y1 w*; PBac {ham-GFP.FPTB}VK00033/TM6C, Sb1 |
| Genetic reagent (D. melanogaster) | hamRSGal4 | Bloomington Drosophila Stock Center | BDSC:48361 FLYBase: FBst0048361; RRID:BDSC_48361 | FlyBase symbol: w1118;P {GMR80G06-GAL4}attP2 |
| Genetic reagent (D. melanogaster) | hamRNAi | Bloomington Drosophila Stock Center | BDSC:26728 FlyBase: FBst0026728; RRID:BDSC_26728 | FlyBase symbol: y1 v1;P{TRiP.JF02270}attP2 |
| Genetic reagent (D. melanogaster) | UAS-GFP | Bloomington Drosophila Stock Center | BDSC:4775 FlyBase: FBst0004775; RRID:BDSC_4775 | FlyBase symbol: w1118; P{UAS-GFP.nls}14 |
| Genetic reagent (D. melanogaster) | GFP.RNAi | Bloomington Drosophila Stock Center | BDSC:9330 FlyBase: FBst0009330; RRID:BDSC_9330 | FlyBase symbol: w1118;P{UAS-GFP.RNAi.R}142 |
| Genetic reagent (D. melanogaster) | Wnt2O | Bloomington Drosophila Stock Center | BDSC:6958 FlyBase: FBst0006958; RRID:BDSC_6958 | FlyBase symbol: Wnt2O/CyO, amosRoi-1 |
| Genetic reagent (D. melanogaster) | UASWnt2 | Bloomington Drosophila Stock Center | BDSC:6961 FlyBase: FBst0006961; RRID:BDSC_6961 | FlyBase symbol: |
| Genetic reagent (D. melanogaster) | fz2.RNAi | Bloomington Drosophila Stock Center | BDSC:27568 FlyBase: FBst0027568; RRID:BDSC_27568 | FlyBase symbol: y1 v1;P{TRiP.JF02722}attP2 |
| Genetic reagent (D. melanogaster) | fz2.RNAi | Bloomington Drosophila Stock Center | BDSC: 67863 FlyBase: FBst0067863; RRID:BDSC_67863 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS05675}attP40 |
| Genetic reagent (D. melanogaster) | fz.RNAi | Vienna Drosophila Resource Center | VDRC: 43075 FlyBase: FBst0464905; | FlyBase symbol: w1118;P{GD4614}v43075 |
| Genetic reagent (D. melanogaster) | Toll-9.RNAi | Bloomington Drosophila Stock Center | BDSC: 34853 FlyBase: FBst0034853; RRID:BDSC_34853 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS00171}attP2 |
| Genetic reagent (D. melanogaster) | pbl.RNAi | Bloomington Drosophila Stock Center | BDSC: 28343 FlyBase: FBst0028343; RRID:BDSC_28343 | FlyBase symbol: y1 v1;P{TRiP.JF02979}attP2 |
| Genetic reagent (D. melanogaster) | CG11406.RNAi | Bloomington Drosophila Stock Center | BDSC: 61185 FlyBase: FBst0061185; RRID:BDSC_61185 | FlyBase symbol: y1 v1;P{TRiP.HMJ22905}attP40 |
| Genetic reagent (D. melanogaster) | AP-2alpha.RNAi | Bloomington Drosophila Stock Center | BDSC: 32866 FlyBase: FBst0032866; RRID:BDSC_32866 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS00653}attP2 |
| Genetic reagent (D. melanogaster) | shg.RNAi | Bloomington Drosophila Stock Center | BDSC: 32904 FlyBase: FBst0032904; RRID:BDSC_32904 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS00693}attP2 |
| Genetic reagent (D. melanogaster) | GstE11.RNAi | Bloomington Drosophila Stock Center | BDSC: 82975 FlyBase: FBst0082975; RRID:BDSC_82975 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMC06653}attP40 |
| Genetic reagent (D. melanogaster) | CG7456.RNAi | Bloomington Drosophila Stock Center | BDSC: 51866 FlyBase: FBst0051866; RRID:BDSC_51866 | FlyBase symbol: y1 v1;P{TRiP.HMC03440}attP40 |
| Genetic reagent (D. melanogaster) | CG7456.RNAi | Bloomington Drosophila Stock Center | BDSC: 53679 FlyBase: FBst0053679; RRID:BDSC_53679 | FlyBase symbol: y1 v1;P{TRiP.HMJ21592}attP40 |
| Genetic reagent (D. melanogaster) | Gr64c.RNAi | Bloomington Drosophila Stock Center | BDSC: 36734 FlyBase: FBst0036734; RRID:BDSC_36734 | FlyBase symbol: y1 sc* v1 sev21;P{TRiP.HMS01625}attP40 |
| Genetic reagent (D. melanogaster) | AkhR.RNAi | Bloomington Drosophila Stock Center | BDSC: 29577 FlyBase: FBst0029577; RRID:BDSC_29577 | FlyBase symbol: y1v1;P{TRiP.JF03256}attP2 |
| Genetic reagent (D. melanogaster) | AkhR.RNAi | Bloomington Drosophila Stock Center | BDSC: 51710 FlyBase: FBst0051710; RRID:BDSC_51710 | FlyBase symbol: y1v1; P{TRiP.HMC03228}attP40 |
| Genetic reagent (D. melanogaster) | Idgf4.RNAi | Bloomington Drosophila Stock Center | BDSC: 55381 FlyBase: FBst0055381; RRID:BDSC_55381 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMC04069}attP40 |
| Genetic reagent (D. melanogaster) | CG17477.RNAi | Bloomington Drosophila Stock Center | BDSC: 42821 FlyBase: FBst0042821; RRID:BDSC_42821 | FlyBase symbol: y1 v1;P{TRiP.HMS02503}attP40 |
| Genetic reagent (D. melanogaster) | CG11069.RNAi | Bloomington Drosophila Stock Center | BDSC: 77158 FlyBase: FBst0077158; RRID:BDSC_77158 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS05930}attP40 |
| Genetic reagent (D. melanogaster) | CG17186.RNAi | Bloomington Drosophila Stock Center | BDSC: 27079 FlyBase: FBst0027079; RRID:BDSC_27079 | FlyBase symbol: y1 v1;P{TRiP.JF02425}attP2 |
| Genetic reagent (D. melanogaster) | CG4872.RNAi | Bloomington Drosophila Stock Center | BDSC: 64486 FlyBase: FBst0064486; RRID:BDSC_64486 | FlyBase symbol: y1 v1;P{TRiP.HMC05503}attP40 |
| Genetic reagent (D. melanogaster) | AdSL.RNAi | Bloomington Drosophila Stock Center | BDSC: 34347 FlyBase: FBst0034347; RRID:BDSC_34347 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS01336}attP2 |
| Genetic reagent (D. melanogaster) | Tl.RNAi | Bloomington Drosophila Stock Center | BDSC: 31044 FlyBase: FBst0031044; RRID:BDSC_31044 | FlyBase symbol: y1 v1;P{TRiP.JF01491}attP2 |
| Genetic reagent (D. melanogaster) | CG17283.RNAi | Bloomington Drosophila Stock Center | BDSC: 63723 FlyBase: FBst0063723; RRID:BDSC_63723 | FlyBase symbol: y1 v1;P{TRiP.HMJ30291}attP40/CyO |
| Genetic reagent (D. melanogaster) | decay.RNAi | Bloomington Drosophila Stock Center | BDSC:65879 FlyBase: FBst0065879; RRID:BDSC_65879 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMC06141}attP2 |
| Genetic reagent (D. melanogaster) | PH4alphaMP.RNAi | Bloomington Drosophila Stock Center | BDSC: 65190 FlyBase: FBst0065190; RRID:BDSC_65190 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMC06065}attP40 |
| Genetic reagent (D. melanogaster) | Hr46.RNAi | Bloomington Drosophila Stock Center | BDSC: 27253 FlyBase: FBst0027253; RRID:BDSC_27253 | FlyBase symbol: y1 v1;P{TRiP.JF02542}attP2 |
| Genetic reagent (D. melanogaster) | CG15117.RNAi | Bloomington Drosophila Stock Center | BDSC: 33693 FlyBase: FBst0033693; RRID:BDSC_33693 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS00562}attP2 |
| Genetic reagent (D. melanogaster) | gsb-n.RNAi | Bloomington Drosophila Stock Center | BDSC: 28078 FlyBase: FBst0028078; RRID:BDSC_28078 | FlyBase symbol: y1 v1;P{TRiP.JF02915}attP2 |
| Genetic reagent (D. melanogaster) | shark.RNAi | Bloomington Drosophila Stock Center | BDSC: 25788 FlyBase: FBst0025788; RRID:BDSC_25788 | FlyBase symbol: y1 v1;P{TRiP.JF01794}attP2 |
| Genetic reagent (D. melanogaster) | CG17018.RNAi | Bloomington Drosophila Stock Center | BDSC: 57024 FlyBase: FBst0057024; RRID:BDSC_57024 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMS04468}attP40 |
| Genetic reagent (D. melanogaster) | cag.RNAi | Bloomington Drosophila Stock Center | BDSC: 63554 FlyBase: FBst0063554; RRID:BDSC_63554 | FlyBase symbol: y1 v1;P{TRiP.HMJ30120}attP40/CyO |
| Genetic reagent (D. melanogaster) | CG7484.RNAi | Bloomington Drosophila Stock Center | BDSC: 61869 FlyBase: FBst0061869; RRID:BDSC_61869 | FlyBase symbol: y1 v1;P{TRiP.HMJ23358}attP40 |
| Genetic reagent (D. melanogaster) | CG31935.RNAi | Bloomington Drosophila Stock Center | BDSC: 64495 FlyBase: FBst0064495; RRID:BDSC_64495 | FlyBase symbol: y1 sc* v1 sev21; P{TRiP.HMC05513}attP40 |
| Antibody | Rabbit anti-Ham | Gift from Eric Lai’s lab | IF (1:300), WB (1:2000) Cut&Tag (1:150) | |
| Antibody | Rabbit anti-GFP | Invitrogen | Cat# G10362 | Cut&Tag (1:150) |
| Antibody | IgG from rabbit serum | Sigma-Aldrich | Cat# I5006-10MG | Cut&Tag (1:150) |
| Antibody | guineapig anti-rabbit | Antibodies online | Cat# ABIN101961 | Secondary antibody for Cut&Tag (1:100) |
| Antibody | Mouse anti-V5 | Invitrogen | Cat# R960-25 | IF (1:500) |
| Antibody | chicken anti-GFP | Abcam | Cat# ab13970 | IF (1:2000) |
| Antibody | Rat anti-vasa | Developmental Studies Hybridoma Bank, DSHB | Cat# anti-vasa-c | IF (1:50) |
| Antibody | Mouse anti-ABD-B | DSHB | Cat# 1A2E9-c | IF (1:50) |
| Antibody | Mouse anti-SXL | DSHB | Cat# M18-c | IF (1:50) |
| Antibody | Rat anti-E-Cad | DSHB | Cat# DCAD2 | IF (1:50) |
| Antibody | Mouse anti-Crb | DSHB | Cat# Cq4 | IF (1:10) |
| Antibody | Mouse anti-Cor | DSHB | Cat# C615.16 | IF (1:100) |
| antibody | Secondary antibodies conjugated to Cy3 or Cy5 or Alexa Fluor-488 | Jackson ImmunoResearch | IF (1:1000) | |
| Antibody | mouse anti-Rabbit IgG light chain specific | Jackson ImmunoResearch | Cat# 211-032-171 | WB (1:40,000) |
| Sequence-based reagent | hamPRΔ_CRISPR gRNA | This paper | gRNA sequence for CRISPR/Cas9 | TCAGGACGATTGTATAGGCG |
| Sequence-based reagent | hamPRΔ check_Forward | This paper | PCR primers | AACACTTTTG AGGGTTGTGTCTTG |
| Sequence-based reagent | hamPRΔ check_Reverse | This paper | PCR primers | ACAGGACTCTTGGGTCGCCC |
| Sequence-based reagent | DWnt2_Luc1_F | This paper | PCR primers | TTGGCGCGCCACTCCTCCCTCCATTTGGCCAC |
| Sequence-based reagent | DWnt2_Luc1_R | This paper | PCR primers | TTGGCGCGCCACCTTCTTCTGCTCCAGCCAACG |
| Sequence-based reagent | DWnt2_Luc2_F | This paper | PCR primers | TTGGCGCGCCGCTTGAAAAGGATCCGTGATC |
| Sequence-based reagent | DWnt2_Luc2_R | This paper | PCR primers | TTGGCGCGCCCCATCATTGATGGAAGGTC |
| Sequence-based reagent | DWnt2_Luc3_F | This paper | PCR primers | TTGGCGCGCCCTCGCAGTGTCGTACATATGTC |
| Sequence-based reagent | DWnt2_Luc3_R | This paper | PCR primers | TTGGCGCGCCTGAGATCACATGCAATGACGC |
| Sequence-based reagent | DWnt2_Luc4_F | This paper | PCR primers | TTGGCGCGCCGCGTCATTGCATGTGATCTC |
| Sequence-based reagent | DWnt2_Luc4_R | This paper | PCR primers | TTGGCGCGCCGCGGCAATAATCATGCGATC |
| Sequence-based reagent | shot_Luc1_F | This paper | PCR primers | TTGGCGCGCCAGTGCATGGATGCTAGGCGAAC |
| Sequence-based reagent | shot_Luc1_R | This paper | PCR primers | TTGGCGCGCCAACCAGCTTGTGCCTGCTACG |
| Sequence-based reagent | shot_Luc2_F | This paper | PCR primers | TTGGCGCGCCAGGTGTATAGAGCGGTGCATG |
| Sequence-based reagent | shot_Luc2_R | This paper | PCR primers | TTGGCGCGCCCAGTGCGCTTTATATCGCTCG |
| Sequence-based reagent | shg_Luc1_F | This paper | PCR primers | TTGGCGCGCCCAACTCGAACTCGACTCAGTGG |
| Sequence-based reagent | shg_Luc1_R | This paper | PCR primers | TTGGCGCGCCGTGTCGCTGGAACTCTTCCTTG |
| Sequence-based reagent | shg_Luc2_F | This paper | PCR primers | TTGGCGCGCCCTGCCACGAATGTTGACGATCC |
| Sequence-based reagent | shg_Luc2_R | This paper | PCR primers | TTGGCGCGCCCGATCTGACGACGACAGATGTC |
| Sequence-based reagent | ham_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/ATCCTGTATATCAAGATGATTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTGGTGAAGTAGCTCCTG |
| Sequence-based reagent | ham_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/ATGGTAGTTTCTGATCAAAGTCCTCT ATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTTCACCCTTTAGAAAGCCAA |
| Sequence-based reagent | ham_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CGAAGTACTCCTCCCTTTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTCATGAACTTCCTTATGTCAT |
| Sequence-based reagent | wnt2_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AGCTCATTAGTGACGCAATTTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTGCGGAGATTAACATAAATGC |
| Sequence-based reagent | wnt2_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TATCGCCAACCAGTCGAATCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTTTGATACTTCAGCATGAGTC |
| Sequence-based reagent | wnt2_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GACATAGCCCTAGTTCTAGTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTCTTAATCAACGCATTTCCTAT |
| Sequence-based reagent | shg_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/ATCGAATCGAACTCGTACATCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTCAACCGCTCAATAATTATCA |
| Sequence-based reagent | shg_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGATTATCGCATACGATTGGTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTACTACGTTATATGCCCGCA |
| Sequence-based reagent | shg_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TTACGTTGATGAATCGGATGTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTCTGATAAACTCCTCCTTGG |
| Sequence-based reagent | shot_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCGCGGTAATCCTTGCATCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTTATCAATAGTTGGTGCATAG |
| Sequence-based reagent | shot_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TCCTTTAGGGTCTTCATTACTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTCAAGATAGTGTTTCCTCTAGA |
| Sequence-based reagent | shot_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TATGCCATCGATGAAGACATTCCTCTATGATTACTGACTGCGTCTATTTAGTGGAGCCGCCCCTATCTTCTTTATGTATCGAATTTCGTATTGAG |
| Sequence-based reagent | pbl_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCGTAAACGTATAGAGCTTCTCATGATTACTGACTGCGTCTATTTAGAATATAAATGAGGTTGATGATGAGGACTTATCTGTCTCCTCATCACTGA |
| Sequence-based reagent | pbl_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/ACCGGATCGTATTCAAAGATATATGATTACTGACTGCGTCTATTTAGAATATAAATGAGGTTGATGATGAGGACTCTATCCGAGATGTCAATCACTA |
| Sequence-based reagent | pbl_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CGAATAAACTATTTGAGACAGACAATGATTACTGACTGCGTCTATTTAGAATATAAATGAGGTTGATGATGAGGACTCGGACTGGTGGTACAAT |
| Sequence-based reagent | AP-2alpha_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GCACTTGAGTTGTTGTTACTATCTATGATTACTGACTGCGTCTATTTACTCCTACACTCTTCTTAAGTACCTTTCCTTCTTCGTGTTGTACACG |
| Sequence-based reagent | AP-2alpha_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TATGAATAACAGCTTACAGACATCTATGATTACTGACTGCGTCTATTTACTCCTACACTCTTCTTAAGTACCTTTCCATGGCCGAGGAGAAA |
| Sequence-based reagent | AP-2alpha_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GCGACTGCAAATCATTCTTTACTAT GATTACTGACTGCGTCTATTTACTCCTACA CTCTTCTTAAGTACCTTTCGATTAACGTG CACAGGATTAC |
| Sequence-based reagent | CG11406_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AATTTGCCGCTATTGATGAGTGAT CCTCTATGATTACTGACTGCGTCTATTTAG ATAGTGGTCCACTGTCCTGGGTAGTCGTACTGCTGG |
| Sequence-based reagent | CG11406_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CCCGGCACCTGATACTGATCCTC TATGATTACTGACTGCGTCTATTTAGATAGTG GTCCACTGTCCTGGAACAGAAAGTCAATGTGATTATAG |
| Sequence-based reagent | CG11406_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CTTATTCCTGTTTCCTCGTTCTGAT CCTCTATGATTACTGACTGCGTCTATTTAGA TAGTGGTCCACTGTCCTGGTCCACGTTTGGTGGAA |
| Sequence-based reagent | AkhR_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CAAGGATTTGCCTCAAAGATAATCCTCT ATGATTACTGACTGCGTCTATTTACTGAGGAGA ATGATCATCGTGATGTATCTGAATGCAACTGCAT |
| Sequence-based reagent | AkhR_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TCGCTTAATTGATCCAGTTGTCCTCTATG ATTACTGACTGCGTCTATTTACTGAGGAGAATGAT CATCGTGATTCCAGTTGAACATTAGCAGA |
| Sequence-based reagent | AkhR_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/AGATGCATTAGCATGATATCAATACTCCT CTATGATTACTGACTGCGTCTATTTACTGAGGAG AATGATCATCGTGATGATCGGCGATGGCC |
| Sequence-based reagent | CG17186_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CAACATCTGGCTCTTCCTCTATGATTACT GACTGCGTCTATTTAGATAGAGGCTCTAGATTT CATCTACGTATCTATATCCTGTATATCCTCCA |
| Sequence-based reagent | CG17186_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GATCTATTTGAACCACCTCGTCTATGAT TACTGACTGCGTCTATTTAGATAGAGGCTCTAG ATTTCATCTACGTATATTCTATCATTTCAAAGCCGG |
| Sequence-based reagent | CG17186_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CAAGAGCTTCAGTTTCCTCTCTATGAT TACTGACTGCGTCTATTTAGATAGAGGCTCTAG ATTTCATCTACGTTCCAAGTATTCTACTGGCT |
| Sequence-based reagent | CG4872_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CTGCTCATATTGGCAATGAAATCTCTAT GATTACTGACTGCGTCTATTTAGAAACTATCCC AAGGAGCTAGAAATGTAGTAAGCTGGCGTAGAG |
| Sequence-based reagent | CG4872_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGTGCGATTGTAAAGATTAAGCTCTATG ATTACTGACTGCGTCTATTTAGAAACTATCCCAA GGAGCTAGAAATCAGTGCCTTTGCCTG |
| Sequence-based reagent | CG4872_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CTCTCGGTGTTCAATTTCTGCTCTATGA TTACTGACTGCGTCTATTTAGAAACTATCCCAAG GAGCTAGAAATATACAGTAATCTGGGTTCCG |
| Sequence-based reagent | decay_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AAGAATTTGTCGAATGTGGAGATCCTCT ATGATTACTGACTGCGTCTATTTACTCAACCAGA TGAAGGAAATGCGTCAACGTTACGGAATGAG |
| Sequence-based reagent | decay_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GTTTGTTCTTGAGCGTCTTATCCTCTATG ATTACTGACTGCGTCTATTTACTCAACCAGATGAA GGAAATGCCTGGATGAAGAAGAGTTTGG |
| Sequence-based reagent | decay_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TCGTTGATCTCGGAGAAGATCCTCTATG ATTACTGACTGCGTCTATTTACTCAACCAGATGA AGGAAATGCCAACCTCTTTGAGCGTG |
| Sequence-based reagent | PH4alphaMP_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CCTCCATAAAGTCTAGGGATATCCTCTAT GATTACTGACTGCGTCTATTTACAATGTATGGATT CGTGAACGCCGATGATATTCCACCACTG |
| Sequence-based reagent | PH4alphaMP_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/AAGTCGCGTAAGAGCTATCATCCTCTATG ATTACTGACTGCGTCTATTTACAATGTATGGATTC GTGAACGCCATCCAATTGATAAGTCTGCTG |
| Sequence-based reagent | PH4alphaMP_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CTAAGATTTCTTTGCTTGGATGATCCTCTATGATTACTGACTGCGTCTATTTACAATGTATGGATTCGTGAACGCCTCTTCCTTAAGCGACAC |
| Sequence-based reagent | GstE11_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AGACAGTGCTCCAAGCCTCTATGATCCTCTATGATTACTGACTGCGTCTATTTACTGGTGGATGCCCGTCAAGTAATCACCTTCAGCC |
| Sequence-based reagent | GstE11_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGACAATGAAACGGATTCGCTCTATGATCCTCTATGATTACTGACTGCGTCTATTTACTGGTGGATGCCCGTCGAAATAGATCACCGGCT |
| Sequence-based reagent | GstE11_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/ATATCCAGACCTTCCTGATTGCTCTATGATCCTCTATGATTACTGACTGCGTCTATTTACTGGTGGATGCCCGTTTAACCAGTCCCACCAAC |
| Sequence-based reagent | Cog4_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GGATTCCAGTACTCCGATAGCCTCTATGATTACTGACTGCGTCTATTTACAATCAGTTCCTGAACTACAGTCAGATGACATCTCTTTGTTATTCTCC |
| Sequence-based reagent | Cog4_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TGCACTATGAAGAACTGGACCTCTATGATTACTGACTGCGTCTATTTACAATCAGTTCCTGAACTACAGTCAGAGTTAAGTAGTCCATCCAAT |
| Sequence-based reagent | Cog4_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CTTCTCATAGCGTTCCATTATATCCCTCTATGATTACTGACTGCGTCTATTTACAATCAGTTCCTGAACTACAGTCACAGATCGGAGTTCTTCATGAT |
| Sequence-based reagent | Gr64c_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AAACTAAGAGATCCTATGGTATAGTGATCCTCTATGATTACTGACTGCGTCTATTTAGGCAAGGGATACTTTCTGCTGATGCAGATCACGCTG |
| Sequence-based reagent | Gr64c_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GTTCTTGAATTGGAGATACGAGTGATCCTCTATGATTACTGACTGCGTCTATTTAGGCAAGGGATACTTTCTGCTCGTGACACTTATGACTAAATGC |
| Sequence-based reagent | Gr64c_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GAAAGCTATGATAAATCTGCACTGATCCTCTATGATTACTGACTGCGTCTATTTAGGCAAGGGATACTTTCTGCTCATAGTTGCCCTTGGATT |
| Sequence-based reagent | Idgf4_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CGTAGTCCAAGTTATTGATAATAGATGATCCTCTATGATTACTGACTGCGTCTATTTAGACTCAACGCAAACCTGTGGTAGGTGTGCAGGTTCA |
| Sequence-based reagent | Idgf4_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/AACAAAGTAGATGATGAGATCCATGATCCTCTATGATTACTGACTGCGTCTATTTAGACTCAACGCAAACCTGTGAACTGTTGCCGTCATAGT |
| Sequence-based reagent | Idgf4_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GCTCTACAACTTGCTCTTAACATGATCCTCTATGATTACTGACTGCGTCTATTTAGACTCAACGCAAACCTGTGAAAGAATCAGAAGGATGAGGA |
| Sequence-based reagent | CG17477_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CCTGAGTTAGTGCGTTAAAGATCCTCTATGATTACTGACTGCGTCTATTTAGATAATTACGGCTGGTCACTGTATGTGGGTAGTTCAACCG |
| Sequence-based reagent | CG17477_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GGCAGACATCATGGACTATCCTCTATGATTACTGACTGCGTCTATTTAGATAATTACGGCTGGTCACTGTCAGTTCCAGATCCTCGTA |
| Sequence-based reagent | CG17477_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TAAGCACAAATATGGCAAGGATCCTCTATGATTACTGACTGCGTCTATTTAGATAATTACGGCTGGTCACTGTTCCAATATTTGCCTGACGA |
| Sequence-based reagent | CG11069_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AAATCGGAAATAAAGATATGGTAGCCTCTATGATTACTGACTGCGTCTATTTACAACTCTACTGATTGCCTACAACTGAGAACTTCTTGCGCC |
| Sequence-based reagent | CG11069_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CTCTTATGCGCCACTTGCCTCTATGATTACTGACTGCGTCTATTTACAACTCTACTGATTGCCTACAACTCACTGATATTAAGATACTCCACT |
| Sequence-based reagent | CG11069_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TGACCAACTGTATTCCAATAGCCTCTATGATTACTGACTGCGTCTATTTACAACTCTACTGATTGCCTACAACTCAATAGCATGACAGGATCTC |
| Sequence-based reagent | AdSL_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CGTCTTGAACGTTCTTCAGTGATCCTCTATGATTACTGACTGCGTCTATTTACTCGAGTCGACGTTCTTGATCTAGATCTAGATGTTCAGTTTGA |
| Sequence-based reagent | AdSL_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/AGCCAGTAGCAACTGTTTTGATCCTCTATGATTACTGACTGCGTCTATTTACTCGAGTCGACGTTCTTGATCATAAGATATTCGGCTGCTTAA |
| Sequence-based reagent | AdSL_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CGAGACATTGCTGGTTAGTGATCCTCTATGATTACTGACTGCGTCTATTTACTCGAGTCGACGTTCTTGATGTGATAATCCATTTAGCAACAG |
| Sequence-based reagent | Tl_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GATCTGCACGTAGTCTTTGCTCTATGATTACTGACTGCGTCTATTTACTACAACAATAATCTGGTGCATGTGTATCCGTCAGATTGCACAT |
| Sequence-based reagent | Tl_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TTAGTTCGAGATGACTCAGATTCCTCTATGATTACTGACTGCGTCTATTTACTACAACAATAATCTGGTGCATGTGCATTTCCTCGATATTTGCCC |
| Sequence-based reagent | Tl_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TCCGAGTATATGATCACAATAATGCTCTATGATTACTGACTGCGTCTATTTACTACAACAATAATCTGGTGCATGTGCTTCTCCACATCTCCGATG |
| Sequence-based reagent | CG17283_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TTGGTCCTCACCTTTAATTTCATCCTCTATGATTACTGACTGCGTCTATTTACAAGACCGAGTCAATCGTTAGAGACTGTCCTGCCTCTG |
| Sequence-based reagent | CG17283_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/ATACGTAGTGGCAGCGATCCTCTATGATTACTGACTGCGTCTATTTACAAGACCGAGTCAATCGTTAGAGTTAATACAGATAAGTCTAAGTGGA |
| Sequence-based reagent | CG17283_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GGTCACGAAAGTGGTTCATCCTCTATGATTACTGACTGCGTCTATTTACAAGACCGAGTCAATCGTTAGACCTAGTATACCATCGAAATTTGA |
| Sequence-based reagent | Hr46_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GGTATATTGTTCAGCAGTGTATGATTACTGACTGCGTCTATTTAGAACATAATCGAATTTGCTAAGCTCATACGGAAATATCGCGGAAATTG |
| Sequence-based reagent | Hr46_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TCACTTTAACTGCTACTCAGATGATTACTGACTGCGTCTATTTAGAACATAATCGAATTTGCTAAGCTCATACGTATTCATTCGATTTCTCGC |
| Sequence-based reagent | Hr46_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GGCAAAGACAACCTAGGATGATTACTGACTGCGTCTATTTAGAACATAATCGAATTTGCTAAGCTCATACCACATACTTTAGTTACAACGTAA |
| Sequence-based reagent | CG15117_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GTGGTTATGTCATTATAGGATGCTATGATTACTGACTGCGTCTATTTACAAACCCATTATCATGTCTGAATATGGTGATCCCTCAAATTGTCC |
| Sequence-based reagent | CG15117_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GATGTAGGTGAAGAGATCTTCCTATGATTACTGACTGCGTCTATTTACAAACCCATTATCATGTCTGAATATGGCATTAGGATATCAGATCGGC |
| Sequence-based reagent | CG15117_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/AGCTATGGAGAAGTGCAGCTATGATTACTGACTGCGTCTATTTACAAACCCATTATCATGTCTGAATATGGCTTTGTTGACCAGGATCAG |
| Sequence-based reagent | gsb-n_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CCTCCAAGAATTCCATTTATGATGATTACTGACTGCGTCTATTTACAAATATAATTGTGCAATGGTATCGACTCCACTTATATCTGAGTCTCGC |
| Sequence-based reagent | gsb-n_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TCGAGAATCTAAGCGACGATGATTACTGACTGCGTCTATTTACAAATATAATTGTGCAATGGTATCGACTCGAATGCAAAGAGCTTATAGCAA |
| Sequence-based reagent | gsb-n_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/CGTTGCCAGCCTTAATGATGATTACTGACTGCGTCTATTTACAAATATAATTGTGCAATGGTATCGACTCATGTTCATAATGCATGTATGGAAA |
| Sequence-based reagent | shark_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TTCTCTGGTCAGAGTGCTCCTCTATGATTACTGACTGCGTCTATTTATTTCGTATTTCGCAAGCTGATCTTTTAGTATGGCTACTGCCTC |
| Sequence-based reagent | shark_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/AATTCTTCTACTGCCGTTTGTCCTCTATGATTACTGACTGCGTCTATTTATTTCGTATTTCGCAAGCTGATCTAGCTTGTAGTTGAGCAGG |
| Sequence-based reagent | shark_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/AACCGTTTGCACCAATTCTCCTCTATGATTACTGACTGCGTCTATTTATTTCGTATTTCGCAAGCTGATCTAATGGAAACTGGCTTAAATGTG |
| Sequence-based reagent | Marf1_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GATTCGTCCAGCGAGTAGCTATGATTACTGACTGCGTCTATTTAATATATGTAATGGAACATTCCGATCCCAAGTAGAAAGCTGCGTATATGT |
| Sequence-based reagent | Marf1_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TGAATAGGTCTTTATGTGCTGCTATGATTACTGACTGCGTCTATTTAATATATGTAATGGAACATTCCGATCCCGTTGGTCCGTTGTTAATAC |
| Sequence-based reagent | Marf1_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TTGGAGTCCTTCCATTAAGGCTATGATTACTGACTGCGTCTATTTAATATATGTAATGGAACATTCCGATCCCGAATACTACAGTAAGGAACACTATG |
| Sequence-based reagent | cag_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/CCCTCTTTAATCGACTGTTTGCCTCTATGATTACTGACTGCGTCTATTTAAGTCTTCATCGCTCACTAGTAATGTTAAGTTCTCCACTCAACAGA |
| Sequence-based reagent | cag_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/ACGGGAGAACCAATCAAAGCCTCTATGATTACTGACTGCGTCTATTTAAGTCTTCATCGCTCACTAGTAATGGGTATATTCTCGGTACGGAC |
| Sequence-based reagent | cag_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GCTTGGGTCAGGGAAGCCTCTATGATTACTGACTGCGTCTATTTAAGTCTTCATCGCTCACTAGTAATGGTATTTATATTGCACCTAAATTGCAT |
| Sequence-based reagent | CG7484_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCTGTGTTCCACTTGGTTATCTCTATGAT TACTGACTGCGTCTATTTAGATCAAATACGTAAGAGGACTGGATGAAGAACTCCTCGACAGTG |
| Sequence-based reagent | CG7484_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/ACTGAGGCTTGATGGTATCCTCTATGATTACTGACTGCGTCTATTTAGATCAAATACGTAAGAGGACTGGATAAAGTGCAGCATTGCTTAC |
| Sequence-based reagent | CG7484_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GCGGATAGGCCCGCTCTATGATTACTGACTGCGTCTATTTAGATCAAATACGTAAGAGGACTGGATCTTTGAATAAAGGCCTGAATCT |
| Sequence-based reagent | Rab3GAP1_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GAATGCAGTAGTGGAATTACTCCTCTATGATTACTGACTGCGTCTATTTAGACCGACTGATAAGCAAGGATTACAGCTAGTTTGAGTATTGCT |
| Sequence-based reagent | Rab3GAP1_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGTAGATCTGGACGAAGATTCCTCTATGATTACTGACTGCGTCTATTTAGACCGACTGATAAGCAAGGATTAGTATAGTAGTTCCATTTGGGAT |
| Sequence-based reagent | Rab3GAP1_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GTGCAACAGTTGTCATTGTCCTCTATGATTACTGACTGCGTCTATTTAGACCGACTGATAAGCAAGGATTATCAGTCGTCGATTTACAATC |
| Sequence-based reagent | fz2_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AGGAAAGTGTTTGTGGAAGTATGATCCTCTATGATTACTGACTGCGTCTATTTAGCGTTTGTTCTACCTGCCGTTTCTCAGTCTATTTCGTTG |
| Sequence-based reagent | fz2_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/TACAGGTAACATCCGATAACTATGTATGATCCTCTATGATTACTGACTGCGTCTATTTAGCGTTTGTTCTACCTGCCTCAAAGTAGGCTGCTTCG |
| Sequence-based reagent | fz2_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TCAGTCGATTGTGTCTCATTTTATGATCCTCTATGATTACTGACTGCGTCTATTTAGCGTTTGTTCTACCTGCCCCCAGGATCAGGACCT |
| Sequence-based reagent | fz_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/ACTAAGAACTACGACTGCGCTATGATTACTGACTGCGTCTATTTACTTGTTCTACGAGTACTACAACTTTGATAGATGATGAACTTTATACAAAGCC |
| Sequence-based reagent | fz_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/AGATCGATATGGTGATGGGCTATGATTACTGACTGCGTCTATTTACTTGTTCTACGAGTACTACAACTTTGAGTCATGTTATATGGTATATTCTTGC |
| Sequence-based reagent | fz_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GCTAAACTCTAAGTAACTTTCGTTACTATGATTACTGACTGCGTCTATTTACTTGTTCTACGAGTACTACAACTTTGAGCATAGGGAACGTCTATGT |
| Sequence-based reagent | fz4_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCATCGAGCCAAAGATCCCTATGATTACTGACTGCGTCTATTTACAATCCTCTACTATAACTTAATGCGGTCAAATAGCCGGAGATCAGAT |
| Sequence-based reagent | fz4_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGAAGAATACGAATTCGATACGCTATGATTACTGACTGCGTCTATTTACAATCCTCTACTATAACTTAATGCGGTCTGATTTGTTGTTCTTTCGTTC |
| Sequence-based reagent | fz4_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GTCTAGAGTAGCGTCTCATTGCTATGATTACTGACTGCGTCTATTTACAATCCTCTACTATAACTTAATGCGGTGAAATGGTACCTAATCCAATTCC |
| Sequence-based reagent | fz3_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/GGTTATGTAGGAGGCCAGGATCCTCTATGATTACTGACTGCGTCTATTTACTCCAACTCTTTCGTATGCCACACAGGCTTAAATAGGAGGT |
| Sequence-based reagent | fz3_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GTGGTTTACGTATTTCTCCGGATCCTCTATGATTACTGACTGCGTCTATTTACTCCAACTCTTTCGTATGCCATTTATATTACTGTGCGACTCTCA |
| Sequence-based reagent | fz3_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/GTCTTCTTCTGAGAGCTCGGATCCTCTATGATTACTGACTGCGTCTATTTACTCCAACTCTTTCGTATGCCACTAGAATGAGGGTCTCAGAC |
| Sequence-based reagent | spz_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TGTTACTGTTGCCTCTTATTTTCTATGATTACTGACTGCGTCTATTTACGGTTATAAACGATATCAATATCGCGCCTTTATACTGGTAGCTGG |
| Sequence-based reagent | spz_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CCACCGATCTTAAGTGTTTATAGTCTATGATTACTGACTGCGTCTATTTACGGTTATAAACGATATCAATATCGCGAAGCCCGATACCATCTG |
| Sequence-based reagent | spz_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TCATTCCTCAAAGGACGAGTCTATGATTACTGACTGCGTCTATTTACGGTTATAAACGATATCAATATCGCGAGGGATTGTGCTCTTTAGTG |
| Sequence-based reagent | spz4_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCGTACTATTGGTATTTCCAGGTCTATGATTACTGACTGCGTCTATTTACAGACGATTTGCGATTGTATTGAAATCGCAGTTCCTTTAGTAGTACTTA |
| Sequence-based reagent | spz4_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CAGGAGGGAATCTAATGGGTCTATGATTACTGACTGCGTCTATTTACAGACGATTTGCGATTGTATTGAAATCCTAATACTACTGTACAATTGTTCA |
| Sequence-based reagent | spz4_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/ATCGTTATTAGTCTAGTGATCGTCTATGATTACTGACTGCGTCTATTTACAGACGATTTGCGATTGTATTGAAATCAAATACCCGATAACCTCG |
| Sequence-based reagent | spz6_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/TCCTCTAAAGGCTAGAACTTAGATGATCCTCTATGATTACTGACTGCGTCTATTTAGACGATACCCTACCCAAGGCTTTCACCTCAAAGTTGTGT |
| Sequence-based reagent | spz6_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/GCTCGTTTGCTTCTTGTAGATGATCCTCTATGATTACTGACTGCGTCTATTTAGACGATACCCTACCCAAGGCACTAGACGATAATCCATCTCC |
| Sequence-based reagent | spz6_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/TTGTTTAGATCTCCGCAAATGATGATCCTCTATGATTACTGACTGCGTCTATTTAGACGATACCCTACCCAAGGGATTCTTGGGTATCTGACCC |
| Sequence-based reagent | spz5_PL1 | This paper | SCRINSHOT padlock probe | /5Phos/AGGGTAAGACTAATGCTACTATGCCTCTATGATTACTGACTGCGTCTATTTACCCACACATGAAACACTCAATATCGATATCCATTAGATCGCTCAC |
| Sequence-based reagent | spz5_PL2 | This paper | SCRINSHOT padlock probe | /5Phos/CGGCTTGTAGATTTGTTTGCCTCTATGATTACTGACTGCGTCTATTTACCCACACATGAAACACTCAATATCAATTCGGAAACCTAGTGTG |
| Sequence-based reagent | spz5_PL3 | This paper | SCRINSHOT padlock probe | /5Phos/AGCGAGACATCGCGCCTCTATGATTACTGACTGCGTCTATTTACCCACACATGAAACACTCAATATCGATATCCAAGAGATCCATGTTG |
| Sequence-based reagent | ham_det1_FAM | This paper | SCRINSHOT detection probe | GTAGCTCCUGATCCTGTAUATCAAGAT/36-FAM/ |
| Sequence-based reagent | ham_det2_FAM | This paper | SCRINSHOT detection probe | CCTTUAGAAAGCCAAAUGGTAGTTTC/36-FAM/ |
| Sequence-based reagent | ham_det3_FAM | This paper | SCRINSHOT detection probe | CTTCCTTAUGTCATCGAAGUACTCC/36-FAM/ |
| Sequence-based reagent | wnt2_det1_Cy3 | This paper | SCRINSHOT detection probe | GATTAACAUAAATGCAGCUCATTAGTGAC/3Cy3Sp/ |
| Sequence-based reagent | wnt2_det2_Cy3 | This paper | SCRINSHOT detection probe | CTTCAGCAUGAGTCTAUCGCC/3Cy3Sp/ |
| Sequence-based reagent | wnt2_det3_Cy3 | This paper | SCRINSHOT detection probe | AACGCATTUCCTATGACAUAGCC/3Cy3Sp/ |
| Sequence-based reagent | shg_det1_Cy5 | This paper | SCRINSHOT detection probe | CCGCTCAAUAATTATCAAUCGAATCGAA/3Cy5Sp/ |
| Sequence-based reagent | shg_det2_Cy5 | This paper | SCRINSHOT detection probe | CGCACGAUTATCGCAUACGAT/3Cy5Sp/ |
| Sequence-based reagent | shg_det3_Cy5 | This paper | SCRINSHOT detection probe | ACTCCTCCUTGGTTACGUTGAT/3Cy5Sp/ |
| Sequence-based reagent | shot_det1_Cy3 | This paper | SCRINSHOT detection probe | TAGTTGGUGCATAGUCGCG/3Cy3Sp/ |
| Sequence-based reagent | shot_det2_Cy3 | This paper | SCRINSHOT detection probe | GTTTCCTCUAGATCCTTUAGGGTCTT/3Cy3Sp/ |
| Sequence-based reagent | shot_det3_Cy3 | This paper | SCRINSHOT detection probe | GAATTTCGUATTGAGTAUGCCATCGA/3Cy3Sp/ |
| Sequence-based reagent | pbl_det | This paper | SCRINSHOT detection probe | GAATAUAAATGAGGUTGATGAUGAGGACT /FITC/ |
| Sequence-based reagent | AP-2alpha_det | This paper | SCRINSHOT detection probe | CTCCUACACTCUTCTTAAGUACCTTTC /CY3/ |
| Sequence-based reagent | CG11406_det | This paper | SCRINSHOT detection probe | GATAGUGGTCCACTGUCCTG /CY5/ |
| Sequence-based reagent | AkhR_det | This paper | SCRINSHOT detection probe | CUGAGGAGAAUGATCATCGUGAT /FITC/ |
| Sequence-based reagent | CG17186_det | This paper | SCRINSHOT detection probe | GAUAGAGGCTCUAGATTTCAUCTACG /CY3/ |
| Sequence-based reagent | CG4872_det | This paper | SCRINSHOT detection probe | GAAACTAUCCCAAGGAGCUAGAAAT /CY5/ |
| Sequence-based reagent | decay_det | This paper | SCRINSHOT detection probe | CUCAACCAGAUGAAGGAAAUGC /FITC/ |
| Sequence-based reagent | PH4alphaMP_det | This paper | SCRINSHOT detection probe | CAATGTAUGGATTCGUGAACGC /CY3/ |
| Sequence-based reagent | GstE11_det | This paper | SCRINSHOT detection probe | CTGGTGGAUGCCCGT /CY5/ |
| Sequence-based reagent | Cog4_det | This paper | SCRINSHOT detection probe | CAATCAGUTCCTGAACUACAGTCA /FITC/ |
| Sequence-based reagent | Gr64c_det | This paper | SCRINSHOT detection probe | GGCAAGGGAUACTTTCUGCT /CY3/ |
| Sequence-based reagent | Idgf4_det | This paper | SCRINSHOT detection probe | GACUCAACGCAAACCUGTG /CY5/ |
| Sequence-based reagent | CG17477_det | This paper | SCRINSHOT detection probe | GATAAUTACGGCUGGTCACUGT /CY3/ |
| Sequence-based reagent | CG11069_det | This paper | SCRINSHOT detection probe | CAACTCTACUGATTGCCUACAACT /FITC/ |
| Sequence-based reagent | AdSL_det | This paper | SCRINSHOT detection probe | CTCGAGUCGACGTTCTUGAT /FITC/ |
| Sequence-based reagent | Tl_det | This paper | SCRINSHOT detection probe | CTACAACAAUAATCTGGUGCATGTG /CY3/ |
| Sequence-based reagent | CG17283_det | This paper | SCRINSHOT detection probe | CAAGACCGAGUCAATCGTUAGA /FITC/ |
| Sequence-based reagent | Hr46_det | This paper | SCRINSHOT detection probe | GAACATAAUCGAATTTGCUAAGCTCAUAC /CY3/ |
| Sequence-based reagent | CG15117_det | This paper | SCRINSHOT detection probe | CAAACCCAUTATCATGTCUGAATAUGG /CY5/ |
| Sequence-based reagent | gsb-n_det | This paper | SCRINSHOT detection probe | CAAATATAAUTGTGCAAUGGTATCGACUC /FITC/ |
| Sequence-based reagent | shark_det | This paper | SCRINSHOT detection probe | TTTCGTATUTCGCAAGCUGATCT /CY3/ |
| Sequence-based reagent | Marf1_det | This paper | SCRINSHOT detection probe | ATATATGUAATGGAACAUTCCGAUCCC /CY5/ |
| Sequence-based reagent | cag_det | This paper | SCRINSHOT detection probe | AGTCTTCAUCGCTCACUAGTAATG /CY3/ |
| Sequence-based reagent | CG7484_det | This paper | SCRINSHOT detection probe | GATCAAAUACGUAAGAGGACUGGAT /FITC/ |
| Sequence-based reagent | Rab3GAP1_det | This paper | SCRINSHOT detection probe | GACCGACUGAUAAGCAAGGAUTA /CY3/ |
| Sequence-based reagent | fz2_det | This paper | SCRINSHOT detection probe | GCGTTTGUTCTACCUGCC /FITC/ |
| Sequence-based reagent | fz_det | This paper | SCRINSHOT detection probe | CTTGTTCUACGAGTACUACAACTUTGA /FITC/ |
| Sequence-based reagent | fz4_det | This paper | SCRINSHOT detection probe | CAATCCTCUACTATAACUTAATGCGGT /CY3/ |
| Sequence-based reagent | fz3_det | This paper | SCRINSHOT detection probe | CTCCAACUCTTTCGTAUGCCA /CY5/ |
| Sequence-based reagent | spz _det | This paper | SCRINSHOT detection probe | CGGTTAUAAACGATAUCAATAUCGCG /CY5/ |
| Sequence-based reagent | spz4_det | This paper | SCRINSHOT detection probe | CAGACGATUTGCGATUGTATUGAAAT /CY3/ |
| Sequence-based reagent | spz6_det | This paper | SCRINSHOT detection probe | GACGAUACCCUACCCAAGG /CY5/ |
| Sequence-based reagent | spz5_det | This paper | SCRINSHOT detection probe | CCCACACAUGAAACACUCAATATC /FITC/ |
| Commercial assay or kit | SuperSignal West Femto Maximum Sensitivity Substrate | Thermo Fisher | Cat# 34096 | |
| Commercial assay or kit | Effectene Transfection kit | QIAGEN | Cat# 301427 | |
| Commercial assay or kit | Dual-Glo Luciferase Assay System | Promega | Cat# E2940 |
Additional files
-
Supplementary file 1
Clusters and genes from STRING network analysis.
- https://cdn.elifesciences.org/articles/104164/elife-104164-supp1-v1.xlsx
-
Supplementary file 2
Genes that showed male sterility in the RNAi screen.
- https://cdn.elifesciences.org/articles/104164/elife-104164-supp2-v1.xlsx
-
Supplementary file 3
Tested genes and their expression in SCRINSHOT.
- https://cdn.elifesciences.org/articles/104164/elife-104164-supp3-v1.xlsx
-
Supplementary file 4
Differentially expressed genes from RNA-seq data.
- https://cdn.elifesciences.org/articles/104164/elife-104164-supp4-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/104164/elife-104164-mdarchecklist1-v1.pdf





