Regulatory consequences of neuronal ELAV-like protein binding to coding and non-coding RNAs in human brain
Figures
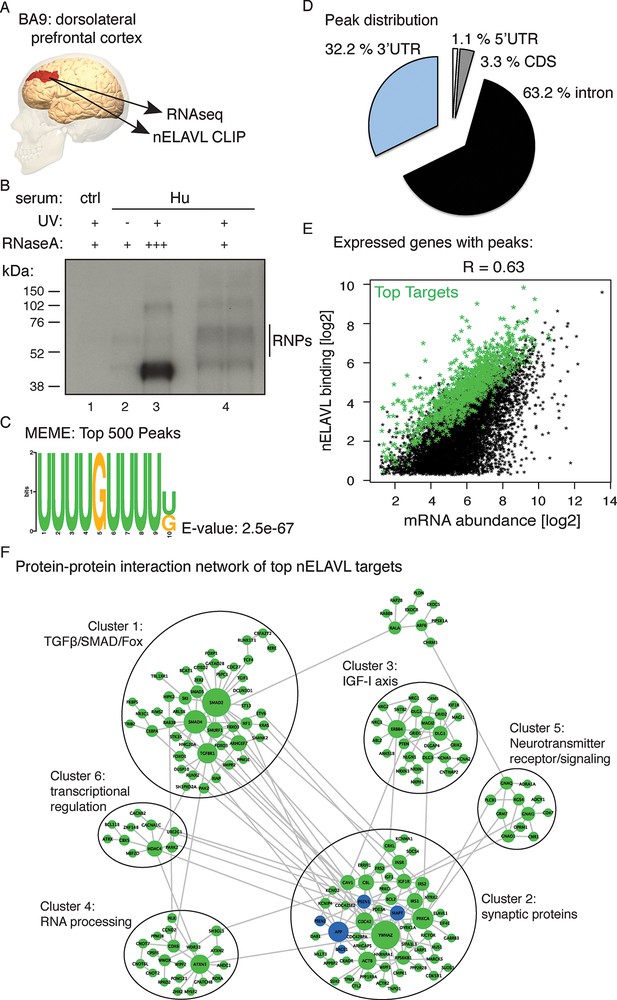
Identification of nELAVL targets in human brain.
(A) Illustration depicting the brain area analyzed by CLIP and RNAseq. The image was generated using BodyParts3D/Anatomography service by DBCLS, Japan. (B) SDS-PAGE separation of radiolabeled nELAVL-RNP complexes. nELAVL-RNP complexes from 40 mg of human brain were specifically immunoprecipitated with Hu-antiserum, compared to control serum (compare lane #4 to #1), which is dependent on UV irradiation (compare lane #4 to #2). Wide-range nELAVL-RNP complexes collapse to a single band in the presence of high RNAse concentration (lane #3). RNAse dilutions: + 19.23 Units/µl; +++ 3846 Units/µl. As in studies of mouse nELAVL (Ince-Dunn et al., 2012), higher molecular weight bands were present in nELAVL CLIP autoradiograms, which correspond at least in part to nELAVL multimers. (C) Shown is the most enriched motif in the top 500 nELAVL peaks, determined with MEME-ChiP. (D) Pie chart of the genomic peak distribution of 75,592 nELAVL peaks (p < 0.01; present in at least 5 individuals). (E) nELAVL binding correlates with mRNA abundance. nELAVL binding (CLIP tags within binding sites per transcript) was compared to mRNA abundance (RNAseq tags per transcript). Only expressed genes with peaks are shown and the correlation coefficient is indicated. The top 1000 targets were identified as genes with highest normalized nELAVL binding (binding sites were normalized for mRNA abundance and summarized per gene). (F) Subnetwork of direct protein-protein interactions of top nELAVL targets. The 1000 top nELAVL target genes and six additional genes highly associated with AD (APP, BACE1, MAPT, PICALM, PSEN1 and PSEN2) were clustered using the organic layout algorithm in yEd. Genes with no direct interactions with other target genes were excluded, leaving 172 nodes from the top nELAVL target list (green) and 5 AD associated genes (blue) in this subnetwork. The size of the nodes is proportional to the connectivity degree. Six clusters (gray circles) containing at least 10 nodes were identified, and subjected to enrichment analysis (see Supplementary file 1F).
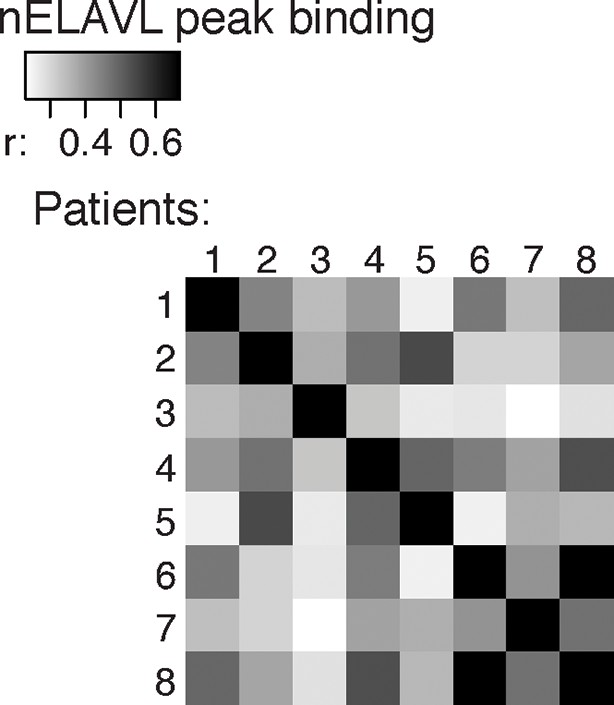
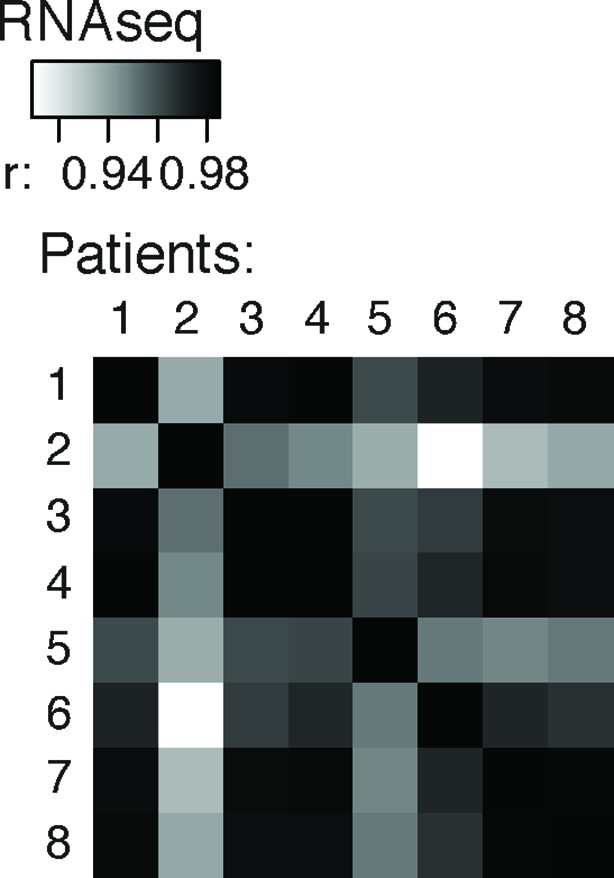
Cross-correlation plot comparing nELAVL peak binding between eight individuals (n = 75,592).
Shown are R values.
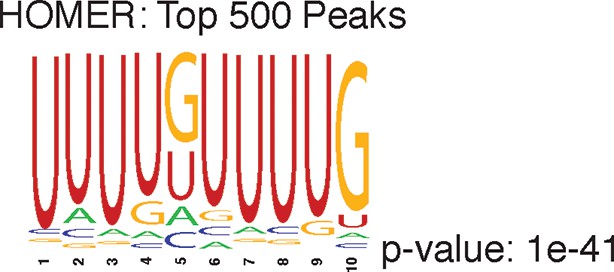

Shown is the most enriched motif in the top 500 nELAVL peaks, determined with HOMER.
https://doi.org/10.7554/eLife.10421.005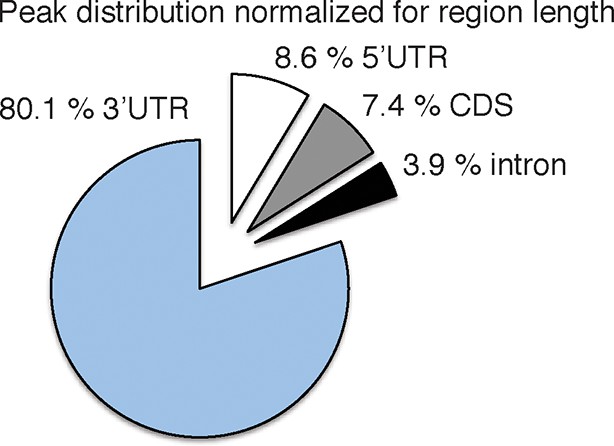
Pie chart of the genomic peak distribution of 75,592 nELAVL peaks (p < 0.01; present in at least 5 individuals), normalized for region length.
https://doi.org/10.7554/eLife.10421.006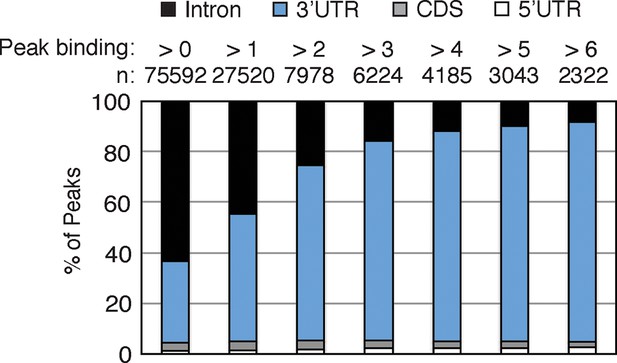
nELAVL peaks within 3’UTRs are higher than intronic binding sites.
The genomic distribution of nELAVL binding sites is plotted as a function of nELAVL peak binding. The number of nELAVL binding sites (n) within each category is indicated.
Cross-correlation plot comparing the mRNA abundance of all transcripts between eight individuals (n = 19,185).
R values are depicted.
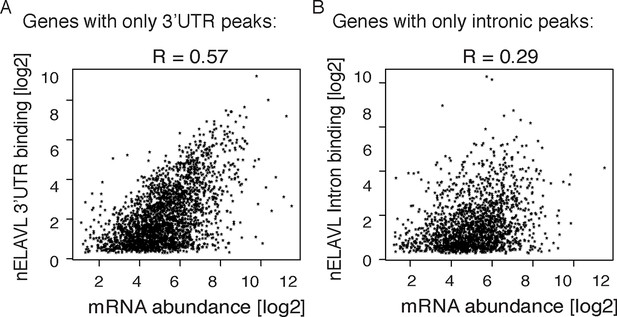
Correlations between mRNA abundance and nELAVL binding.
(A,B) mRNA abundance shows a higher correlation with nELAVL 3’UTR (A) than intronic binding (B). nELAVL binding was defined as CLIP tags within binding sites per transcript. Shown are genes with exclusively 3’UTR (A) or intronic (B) binding and R values are indicated.
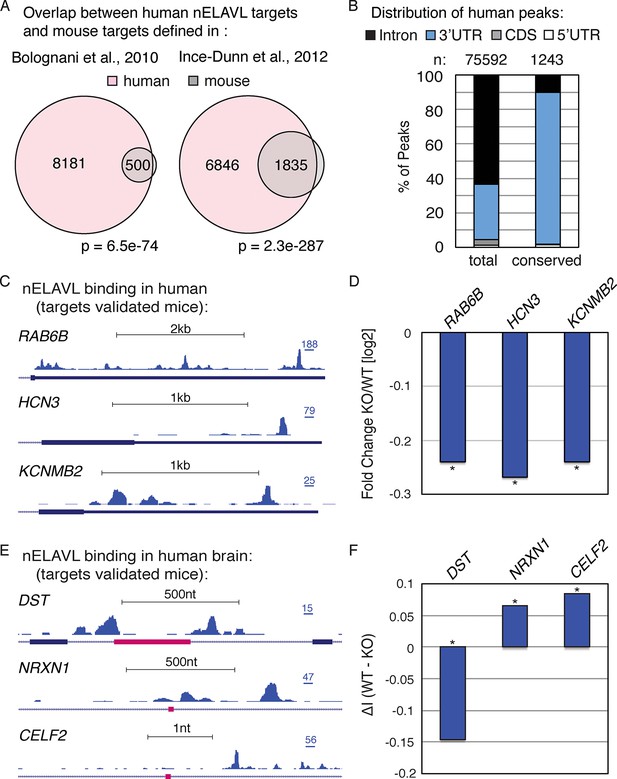
nELAVL mediated regulation is conserved in mouse and human.
(A) Overlap of nELAVL targets in human and mouse. Human nELAVL targets (n = 8681) were intersected with mouse targets identified by RIP (Bolognani et al., 2010) or HITS-CLIP (Ince-Dunn et al., 2012). 538 genes were identified as nELAVL targets by RIP and were expressed in human brain. 1978 expressed genes had HITS-CLIP nELAVL clusters that were present in at least 3 samples (biological complexity (BC) ≥ 3). Both overlaps (n = 500 and n = 1835) were highly significant (p = 6.5e-74 and p = 2.3e-287; hypergeometric test), compared to expressed transcripts (n = 14,737). (B) Only few nELAVL binding sites are conserved between mice and human, which are predominantly present within 3’UTRs. The genomic distribution of all human nELAVL binding sites (total) and nELAVL binding sites conserved in mouse is shown. The number of nELAVL binding sites (n) within each category is indicated. (C) UCSC Genome Browser images illustrating the 3’UTRs of RAB6B, HCN3, and KCNMB2 and their normalized nELAVL binding profile in human brain. The maximum PeakHeight is indicated by numbers in the right corner. (D) The mRNA levels of transcripts with nELAVL 3’UTR binding decrease in Elavl3/4 knockout (KO) mice. Shown are the mRNA expression fold changes (knockout/wildtype) of RAB6B, HCN3, and KCNMB2. *p< 0.01 (two-tailed t test; Ince-Dunn et al., 2012). (E) UCSC Genome Browser images showing pink cassette exons in the DST, NRXN1, and CELF2 genes and their normalized nELAVL binding profiles in human brain. The maximum PeakHeight is indicated by numbers in the right corner. (F) nELAVL binding adjacent to a cassette exon in the DST gene prevents exon inclusion. Downstream nELAVL binding promotes the inclusion of cassette exons in the NRXN1 and CELF2 genes. The change in alternative exon inclusion (delta inclusion (ΔI): wildtype - Elavl3/4 KO) is shown. * significantly changing (analyzed by Aspire2; Ince-Dunn et al., 2012).
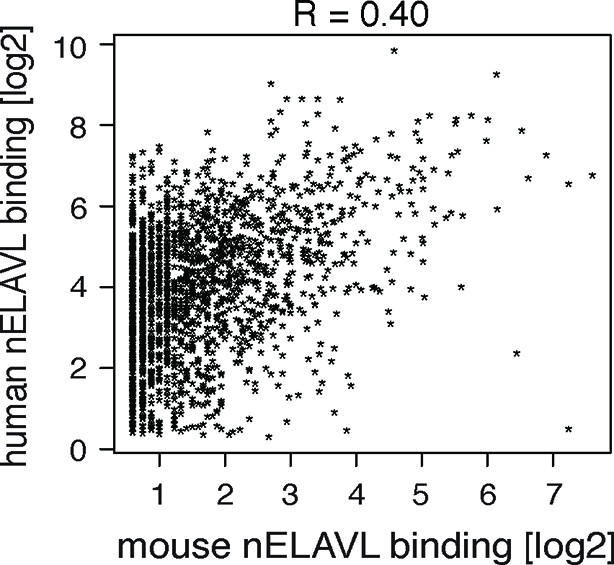
Comparison of nELAVL binding (CLIP tags within binding sites per transcript) between mice and human.
The correlation coefficient is indicated.
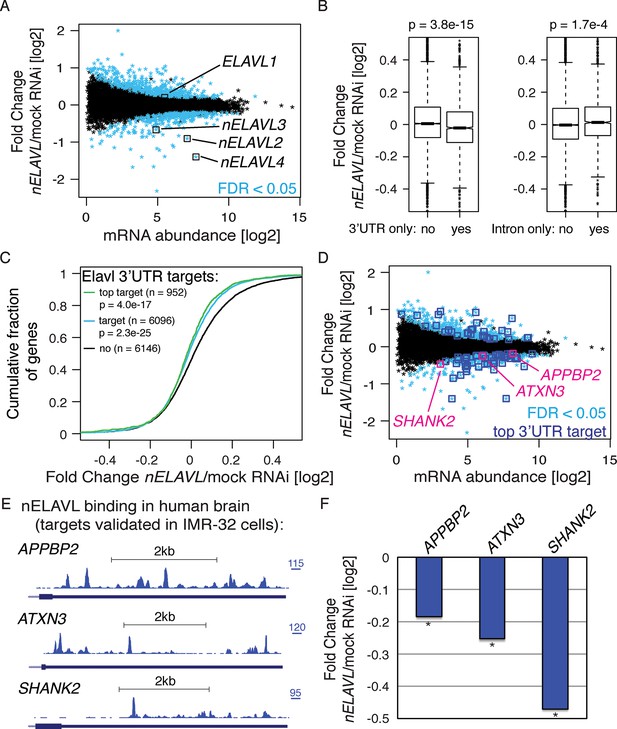
nELAVL proteins regulate mRNA abundance of human brain targets.
(A) nELAVL depletion causes mRNA level changes in IMR-32 neuroblastoma cells. The mRNA abundance change was plotted against average mRNA abundance. Significantly changing transcripts (FDR < 0.05; n = 784) are colored in blue. Shown are only expressed genes (n = 12,743), and ELAVL1/2/3/4 transcripts are indicated. (B) nELAVL with exclusively 3’UTR binding decrease upon nELAVL RNAi depletion. Box plots represent the distribution of mRNA level differences between mock and nELAVL RNAi. We compared genes with exclusively 3’UTR (n = 2346) or intronic (n = 1693) binding that were expressed in IMR-32 cells. nELAVL binding was defined as CLIP tags within binding sites per transcript. Transcripts with exclusively 3’UTR binding were less abundant upon nELAVL RNAi compared to remaining transcripts (p = 3.8e-15; two-tailed t-test). In contrast, mRNA levels of transcripts with exclusively intron binding were even slightly increased compared to remaining transcripts (p = 1.7e-4; two-tailed t-test). (C) Transcripts with nELAVL 3’UTR binding decrease upon nELAVL RNAi. Cumulative fraction curves for genes with no 3’UTR nELAVL binding in human brain, 3’UTR binding, and top 3’UTR targets. Top targets were identified as 1000 genes with highest normalized nELAVL 3’UTR binding (binding sites were normalized for mRNA abundance before summarized per gene). 952 of the top 1000 targets were expressed in IMR-32 cells. A curve displacement to the left indicates a downregulation of mRNA abundance upon nELAVL RNAi. p values were calculated with a one-sided KS test, comparing (top) targets to non-targets. (D) Many transcripts that are decreasing upon nELAVL depletion are top nELAVL 3’UTR targets. The mRNA abundance change (nELAVL/mock RNAi) of transcripts expressed in IMR-32 cells and in human brain (n = 12,242) was plotted against average mRNA abundance. Significantly changing transcripts (FDR<0.05; n = 743) are colored in blue and additionally boxed if they are top nELAVL 3’UTR targets. Transcripts shown in E/F are indicated. (E) UCSC Genome Browser images illustrating the 3’UTRs of APPBP2, ATXN3, and SHANK2 and their normalized nELAVL binding profile in human brain. The maximum PeakHeight is indicated by numbers in the right corner. (F) The mRNA abundance of top nELAVL 3’UTR targets decreases upon nELAVL RNAi. Shown are the mRNA level changes (nELAVL/mock RNAi) of APPBP2, ATXN3, and SHANK2. * FDR<0.05 (derived from edgeR).
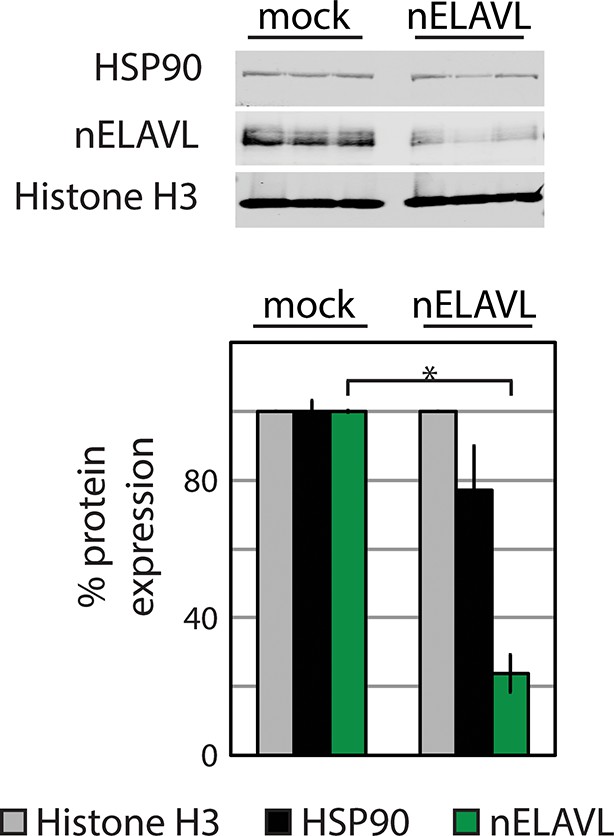
Western blot and its quantification showing protein levels of nELAVL and the housekeeping genes HSP90 and Histone H3 in mock and nELAVL RNAi-treated IMR-32 cells.
Protein expression was normalized to the house keeping gene Histone H3 and to mock RNAi treated cell. Error bars represent SEM. * p<0.01 (two-tailed t-test).
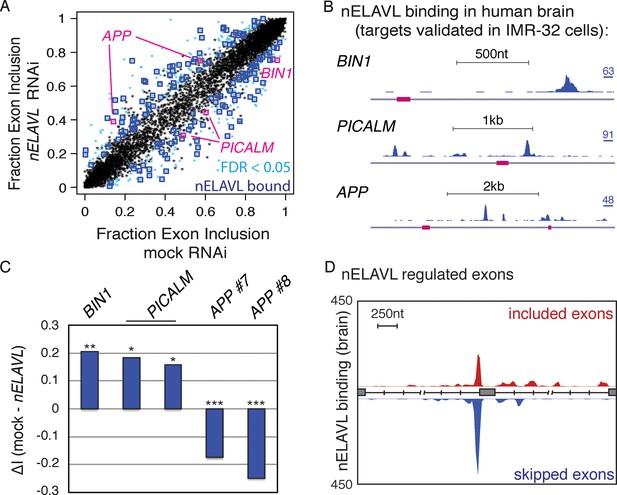
nELAVL regulates splicing of human brain targets.
(A) Analysis of splicing changes upon nELAVL RNAi. Shown is the exon inclusion fraction of cassette exons that are expressed in IMR-32 cells and in human brain (n = 7903). Significantly changing exons (FDR<0.05 and ΔI>0.1) are colored in light blue (n = 473), and additionally boxed in dark blue if adjacent (+/- 2.5 kb) to intronic nELAVL binding sites (n = 155). Significantly changing exons shown in (B/C) are boxed in pink. The two alternative events within PICALM correspond to the same alternative exon with two different 3’ splice sites. (B) UCSC Genome Browser images depicting cassette exons in pink in the BIN1, PICALM, and APP genes and their normalized nELAVL binding profiles in human brain. The maximum PeakHeight is indicated by numbers in the right corner. (C) nELAVL binding downstream of cassette exons in BIN1 and PICALM promotes exon inclusion, whereas intronic nELAVL binding of APP prevents exon inclusion downstream and upstream. The change in alternative exon inclusion (ΔI: mock – nELAVL RNAi) is shown. *FDR< 0.0005; **FDR< 1e-4; ***FDR<1e-16 (GLM likelihood ratio test). (D) Normalized nELAVL binding map of nELAVL regulated exons. Only exons that changed significantly upon nELAVL RNAi (FDR<0.05 and ΔI>0.1) and that are adjacent (+/- 2.5 kb) to intronic nELAVL binding sites (n = 155) were included. Red and blue peaks represent binding associated with nELAVL-dependent exon inclusion and exclusion, respectively.
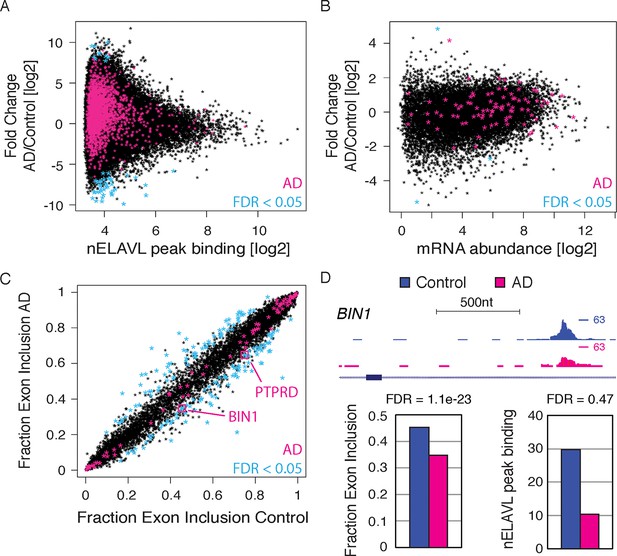
RNA regulation changes in AD.
(A) nELAVL binding changes in AD. The nELAVL peak binding change (AD/Control) was plotted against average nELAVL peak binding. Significantly changing peaks (FDR<0.05; n = 52) are colored in blue, and peaks within AD genes are colored in pink (1811 peaks within 69 genes). Shown are only peaks that are bound in control or AD brain (n = 115,393). (B) mRNA abundance changes in AD. The mRNA abundance change (AD/Control) was plotted against average mRNA abundance. Significantly changing transcripts (FDR<0.05; n = 3) are colored in blue, and AD transcripts are colored in pink (n = 89). Shown are only transcripts that are expressed in control or AD brain (n = 14,875). (C) Analysis of splicing changes in AD. Shown is the inclusion fraction of expressed cassette exons in control and AD subjects (n = 8163). Exons within AD genes are colored in pink (n = 79). Significantly changing exons (FDR<0.05 and ΔI>0.1) are colored in light blue (n = 170), and additionally boxed in pink if within AD genes (n = 2). (D) BIN1 is alternatively spliced in AD. UCSC Genome Browser image illustrating a cassette exon in the BIN1 gene and normalized nELAVL binding profiles in control and AD brain. The maximum PeakHeight is indicated by numbers in the right corner. Bar graphs depict the difference in alternative exon inclusion (ΔI: Control – AD) and nELAVL peak binding (AD/Control) in control and AD brain. Corresponding FDR values derived from edgeR are shown. The inclusion of the exon is promoted by nELAVL (see Figure 4), and exon inclusion as well as nELAVL peak binding are reduced in AD subjects.
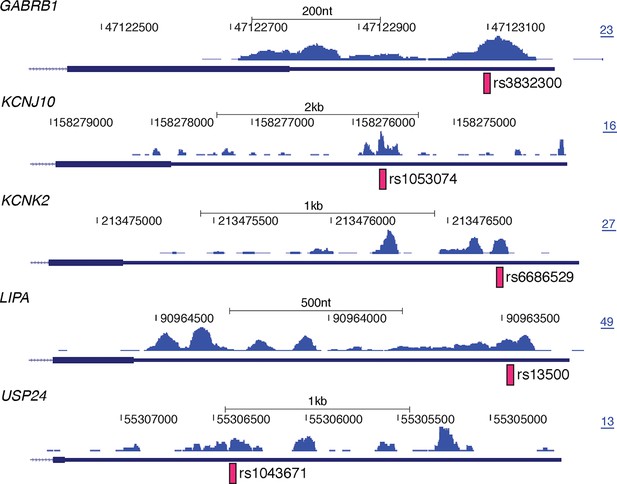

Examples of disease associated SNPs with corresponding nELAVL binding sites.
UCSC Genome Browser images depicting the last exon of GABRB1, KCNJ10, KCNK2, LIPA, and USP24 and the normalized nELAVL binding profile in human brain. Pink bars illustrate SNPs overlapping with nELAVL binding sites, and the maximum PeakHeight is indicated by numbers in the right corner.

Correlations between control and AD samples.
(A) Cross-correlation plot comparing nELAVL peak binding between eight controls and nine AD subjects (n = 247,547). Shown are R values. (B) Cross-correlation plot comparing the mRNA abundance of all transcripts between eight controls and nine AD subjects (n = 19,185). R values are depicted.
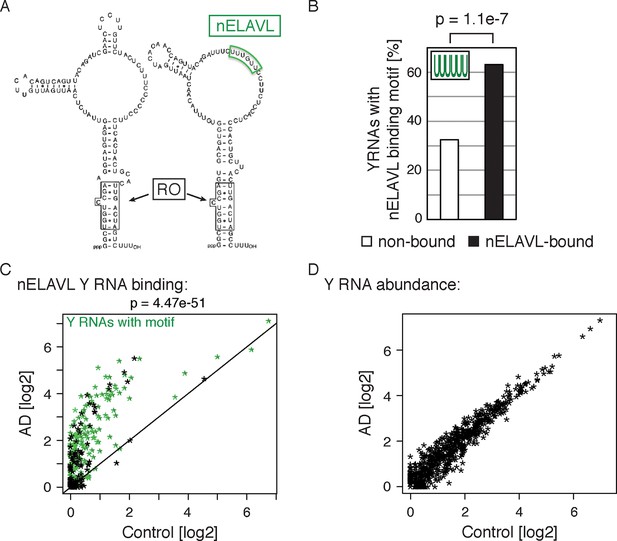
Non-coding Y RNAs are bound by nELAVL in AD.
(A) Secondary structures of Y1 and Y3. Binding sites of nELAVL and Ro are indicated. Modified from (Chen and Wolin, 2004). (B) The nELAVL binding motif (UUUUUU, allowing a G at any position) is enriched in nELAVL-bound Y RNAs compared to non-bound Y RNAs (p = 1.1e-7; Fisher’s exact test). Y RNAs were scanned for (T)6, allowing a G at any position. nELAVL-bound Y RNAs: nELAVL CLIP tags in at least two samples; n = 320. (C) nELAVL binding of Y RNAs increases in AD compared to control samples (p = 4.47e-51; paired one-sided Wilcoxon rank sum test). The axes depict nELAVL Y RNA binding (nELAVL CLIP tags per Y RNA) in control and AD subjects. Y RNAs with nELAVL binding motif are colored in green. (D) Y RNA levels do not change in AD. Y RNA abundance (RNAseq tags per Y RNA) in AD subjects was plotted against Y RNA abundance in control subjects.

Y RNAs with a motif that are not bound are not expressed.
Box plots represent the distribution of Y RNA abundance in human brain. Non-bound Y RNAs with an nELAVL binding motif show a lower expression than nELAVL-bound Y RNAs with motif (p = 0.002; two-tailed t test).
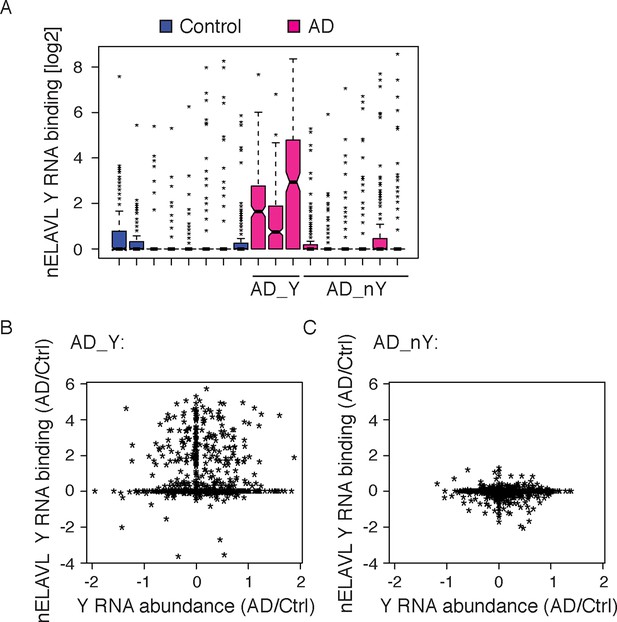
nELAVL:Y RNA binding increases in AD.
(A) A subset of AD subjects shows increased nELAVL binding to Y RNAs. Box plots represent the distribution of nELAVL Y RNA binding (tags per Y RNA) for each individual. AD subjects were grouped into AD_Y and AD_nY subjects based on nELAVL binding. Only nELAVL bound Y RNAs are included: nELAVL CLIP tags in at least two samples; n = 320. (B,C) nELAVL Y RNA binding but not Y RNA expression changes in AD_Y subjects. Comparison of Y RNA abundance and nELAVL Y RNA binding changes (AD/Control) in AD_Y (B) and AD_nY (C) subjects.
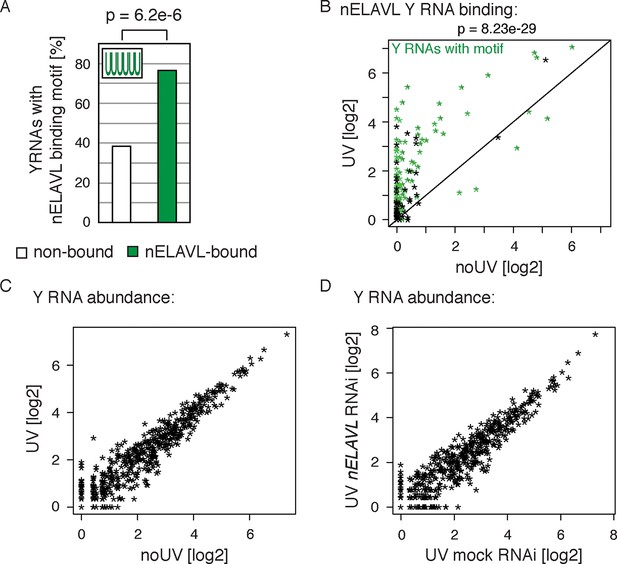
Y RNPs are remodeled during UV stress.
(A) The nELAVL binding motif (UUUUUU, allowing a G at any position) is enriched in nELAVL-bound Y RNAs compared to non-bound Y RNAs (p = 6.2e-6; Fisher’s exact test). Y RNAs were scanned for (T)6, allowing a G at any position. nELAVL-bound Y RNAs: nELAVL CLIP tags in at least two samples; n = 132. (B) nELAVL binding of Y RNAs increases during UV stress compared to non-stressed cells (p = 8.23e-29; paired one-sided Wilcoxon rank sum test). The axes depict nELAVL Y RNA binding (nELAVL CLIP tags per Y RNA) in control and UV stressed cells. Y RNAs with nELAVL binding motif are colored in green. (C) Y RNA levels do not change upon UV stress. Y RNA abundance (RNAseq tags per Y RNA) in UV stressed cells was plotted against Y RNA abundance in non-stressed control cells. (D) nELAVL is binding is not required for Y RNA stability. Comparison of Y RNA abundance between mock and nELAVL RNAi treated UV stressed cells.
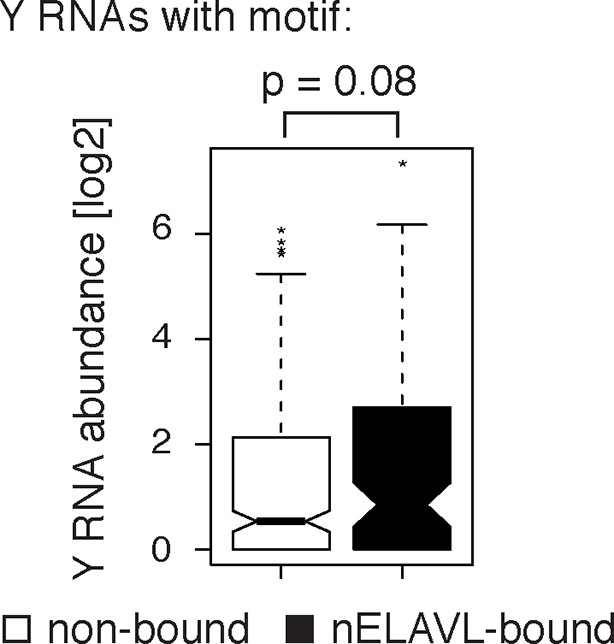
Y RNAs with a motif that are not bound are not expressed.
Box plots represent the distribution of Y RNA abundance in IMR-32 cells. Non-bound Y RNAs with an nELAVL binding motif show a lower expression than nELAVL-bound Y RNAs with motif (p = 0.08; two-tailed t-test).
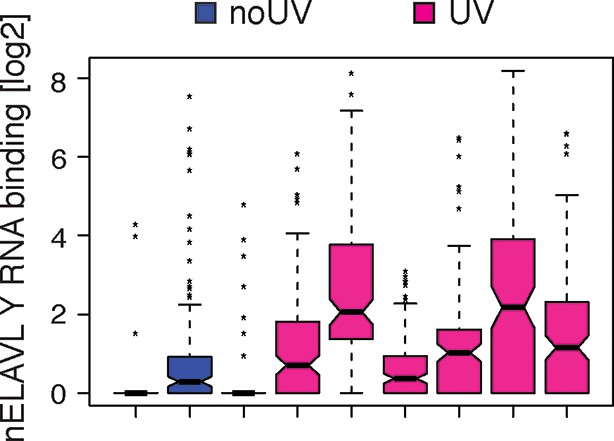
UV-stressed cells show increased nELAVL binding to Y RNAs.
Box plots represent the distribution of nELAVL binding (tags per Y RNA) for each individual. Only nELAVL bound Y RNAs are included: nELAVL CLIP tags in at least two samples; n = 132.
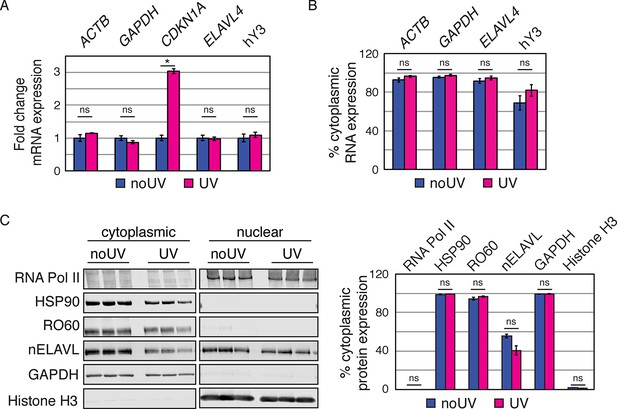
UV does not induce changes in the nucleocytoplasmic localization of Y RNP components.
(A) Validation of UV stress induction. Bar graphs depict the fold change in RNA expression in UV stressed cells compared to non-stressed control cells. CDKN1A (the most upregulated transcripts in the RNAseq dataset) but not control mRNAs (ACTB, GAPDH, ELAVL4) nor Y RNAs increase upon UV stress. RNA expression was normalized to non-UV treated cell. Error bars represent SEM. p values were calculated with a two-tailed t test (ns: not significant; * p = 6.9e-5; two-tailed t-test). (B) UV stress does not induce changes in Y RNA distribution. Bar graphs depict the percentage of cytoplasmic RNA levels (cytoplasmic RNA levels divided by the sum of cytoplasmic and nuclear RNA levels) of mRNA controls (ACTB, GAPDH, ELAVL4) and Y RNAs. Error bars represent SEM. Changes in cytoplasmic RNA levels were not significant (ns; p<0.05; two-tailed t-test). (C) UV stress does not induce changes in protein distribution. Western Blot and its quantification showing cytoplasmic and nuclear protein levels of nELAVL, RO60, as well as cytoplasmic (HSP90 and GAPDH) and nuclear (RNA PolII and Histone H3) markers. Bar graphs depict the percentage of cytoplasmic protein levels (cytoplasmic protein levels divided by the sum of cytoplasmic and nuclear protein levels). Error bars represent SEM. Changes in cytoplasmic proteins levels were not significant (ns; p<0.05; two-tailed t test).
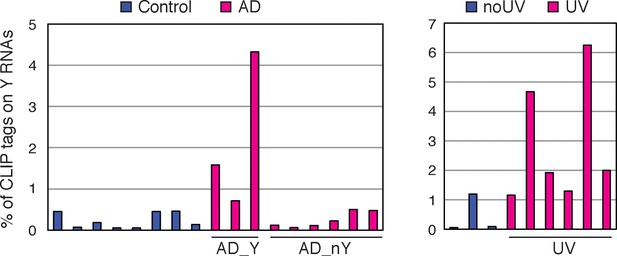
Up to 5% of nELAVL CLIP map to Y RNAs in AD_Y subjects (AD subjects with increased nELAVL/Y RNA association) and UV stressed cells when mapped with Bowtie 2 (allowing multiple alignments and reporting one).
Columns represent the percentage of nELAVL tags that mapped to Y RNAs with Bowtie 2.
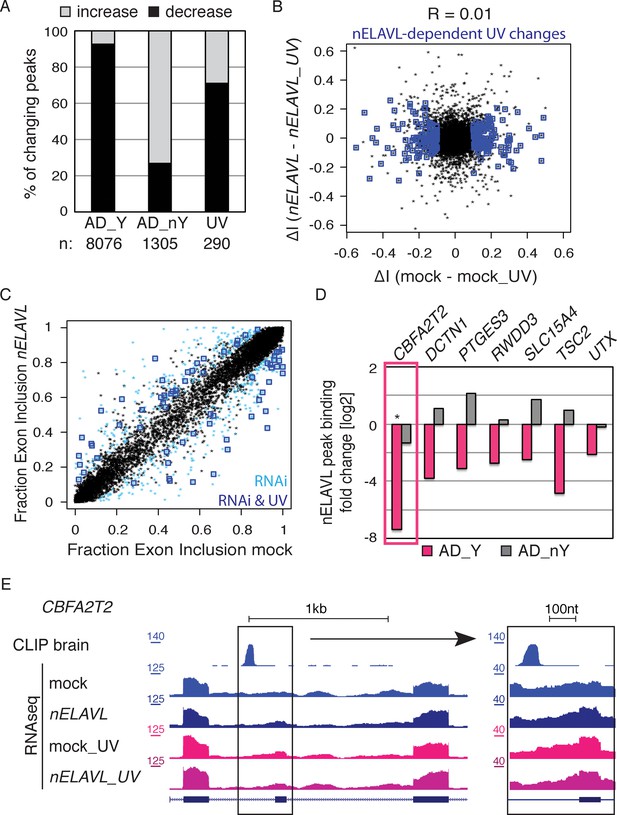
nELAVL/Y RNA correlates with loss of nELAVL-mediated splicing.
(A) Samples with high nELAVL/Y RNA association show decreased nELAVL binding on mRNA targets. Columns represent significantly changing nELAVL binding sites. Shown are changes in AD subjects with and without Y RNA association (AD_Y and AD_nY) and changes upon UV treatment. The number of nELAVL binding sites (n) within each category is indicated. (B) Identification of nELAVL-dependent UV-induced splicing changes. Comparison of the differential inclusion rate of expressed cassette exons upon UV stress between mock and nELAVL RNAi treated IMR-32 cells (n = 9397). Significant UV-induced splicing changes that do not change upon UV stress in nELAVL RNA treated cells are boxed in dark blue (FDR<0.05 and ΔI>0.1; n = 260). (C) Many exons that are alternatively spliced upon nELAVL RNAi treatment also change during UV stress in an nELAVL-dependent manner. Shown is the inclusion rate of expressed cassette exons in IMR-32 cells that were subjected to mock or nELAVL RNAi (n = 9397). nELAVL RNAi induced splicing changes are colored in light blue (n = 553), and are additionally boxed in dark blue if they are UV-induced in an nELAVL-dependent manner (n = 68). The plot is related to Figure 4A but contains additional cassette exons expressed in UV stressed cells. (D) nELAVL binding adjacent to exons that are alternatively spliced upon nELAVL RNAi and UV treatment decreases only in AD subjects with an increased Y RNA association. Displayed is the change in nELAVL peak binding. nELAVL peak binding changes were not significant except for CBFA2T2 (boxed in pink). * FDR<0.05 (derived from edgeR). (E) UCSC Genome Browser images depicting an overview and an enlarged view of a cassette exon within the CBFA2T2 gene that is alternatively spliced in nELAVL RNAi and UV-treated IMR-32 cells. The nELAVL binding track in human brain and RNAseq tracks in mock and nELAVL RNAi treated non-stressed and UV-stressed IMR-32 cells are shown.
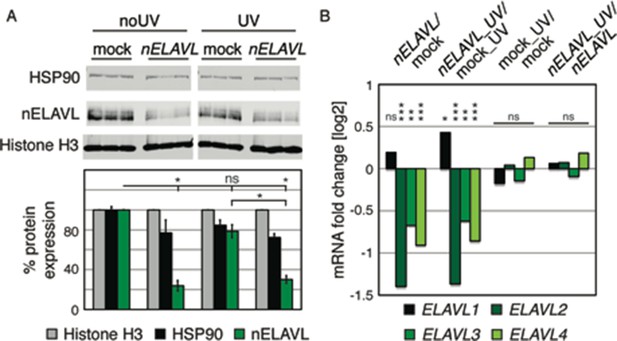
UV does not affect nELAVL RNA or protein levels.
(A) nELAVL protein levels decrease upon nELAVL RNAi treatment in non-stressed and UV-stressed IMR-32 cells but are not affected by UV stress. Western Blot and its quantification showing protein levels of nELAVL and the house keeping genes HSP90 and Histone H3 in mock and nELAVL RNAi treated non-stressed and UV-stressed IMR-32 cells. Protein expression was normalized to the house keeping gene Histone H3 and to mock RNAi treated cell. Error bars represent SEM. p-values were calculated with a two-tailed t-test (ns: not significant; *p<0.01). (B) nELAVL mRNA levels decreased upon nELAVL RNAi treatment in non-stressed and UV-stressed IMR-32 cells but were not affected by UV stress. Shown is the mRNA abundance assessed by RNAseq in response to nELAVL RNAi and UV stress. FDR values were derived from edgeR (ns: not significant; * FDR<1e-2; ** FDR<1e-4; *** FDR<1e-20).
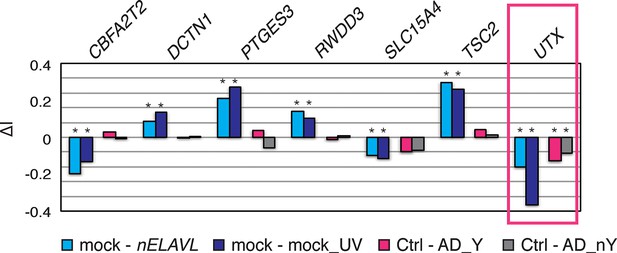
Analysis of splicing changes in nELAVL RNAi and UV treated IMR-32 cells and AD subjects with and without Y RNA association (AD_Y and AD_nY).
Shown is the differential inclusion rate of cassette exons that change similarly upon nELAVL RNAi and UV treatment and that were adjacent (+/- 2.5 kb) to intronic nELAVL binding sites in human brain. Splicing changes in AD subjects were not significant except for UTX (boxed in pink). * FDR<0.05 (GLM likelihood ratio test).
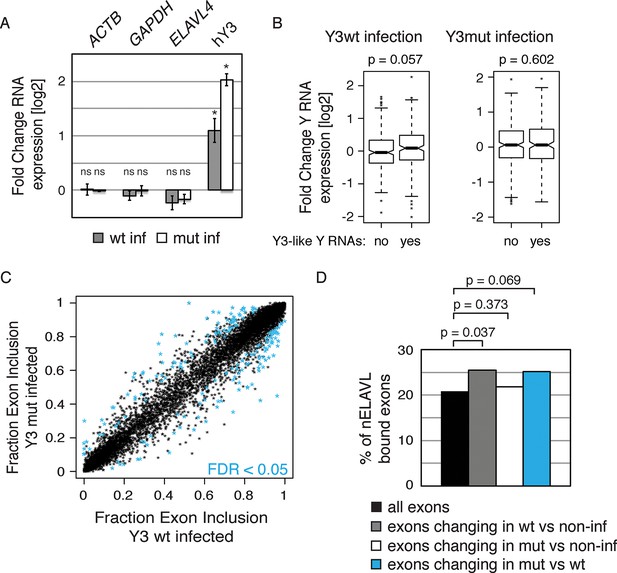
Y RNA overexpression is linked to nELAVL sequestration from mRNA targets.
(A) Validation of Y RNA overexpression. Shown are RNA expression fold changes of Y3wt or Y3mut infected IMR-32 cells compared to non-infected IMR-32 cells assessed by qPCR. Y RNAs expression increased while control mRNAs (ACTB, GAPDH, ELAVL4) were not affected. Error bars represent SEM. p values were calculated with a two-tailed t-test (ns: not significant; * p<0.05). (B) The expression of endogenous Y3-like Y RNAs increases upon Y3wt but not Y3mut infection. Box plots represent the distribution of endogenous Y3-like and non-Y3-like Y RNA expression fold changes upon Y3wt or Y3mut infection. Y3-like Y RNAs show a slight increase in abundance upon Y3wt compared to non-Y3-like Y RNAs (p = 0.057; one-tailed t-test). In contrast, the mRNA abundance of Y3-like Y RNAs does not change upon Y3mut infection, when compared to non-Y3 like Y RNAs (p = 0.602; one-tailed t-test). (C) Identification of Y3 dependent splicing changes. Shown is the exon inclusion fraction of cassette exons that are expressed in IMR-32 cells subjected to Y3wt or Y3mut infection (n = 10,189). Exons changing significantly between Y3wt and Y3mut infection (FDR<0.05 and ΔI>0.1) are colored in light blue (n = 191). (D) Exons that are alternatively spliced upon Y3wt infection are enriched for nELAVL bound exons. Bar graph representing total expressed exons (n = 10,189), exons that change in either Y3wt (n = 240; blue points in the left panel of Figure 9—figure supplement 4) or Y3mut (n = 151; blue points in the right panel of Figure 9—figure supplement 4) infected cells compared to non-infected cells, and exons that change in Y3wt compared to Y3mut infected cells (n = 191; blue points in Figure 9C). Exons that are alternatively spliced upon Y3wt infection compared to either non-infected (p = 0.037; hypergeometric test) or Y3mut infected cells (p = 0.069; hypergeometric test) are enriched for nELAVL bound exons.
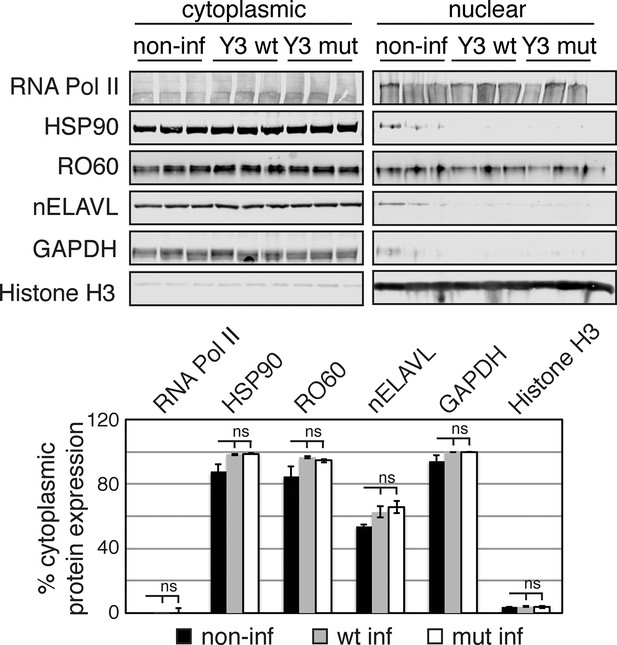
Y3 overexpression does not induce changes in protein distribution.
Western Blot and its quantification showing cytoplasmic and nuclear protein levels of nELAVL, RO60, as well as cytoplasmic (HSP90 and GAPDH) and nuclear (RNA PolII and Histone H3) markers. Bar graphs depict the percentage of cytoplasmic protein levels (cytoplasmic protein levels divided by the sum of cytoplasmic and nuclear protein levels). Error bars represent SEM. Changes in cytoplasmic proteins levels were not significant (ns; p<0.05; two-tailed t-test).
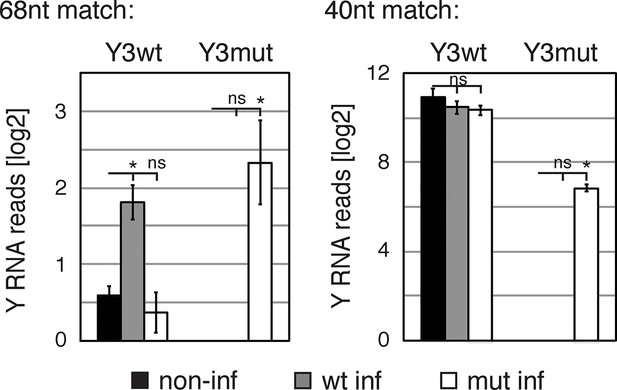
Validation of Y RNA overexpression.
Bar graphs represent log2 number of reads of Y3wt and Y3mut in non-infected, Y3wt and Y3mut infected IMR-32 cells. Read numbers were assessed by searching raw fastq files for Y3wt and Y3mut sequences, respectively. Searched sequences were either 40 (left panel) or 68 (right panel) nucleotides in length. Both sequence lengths encompassed the sequence mutated in Y3mut. While the 40nt Y3wt sequence is present in numerous Y3 RNA copies, the 68nt Y3wt sequence should only be present in the infected Y3wt RNA and the endogenous canonical hY3 RNA. Error bars represent SEM. p values were calculated with a two-tailed t-test (ns: not significant; * p<0.01).
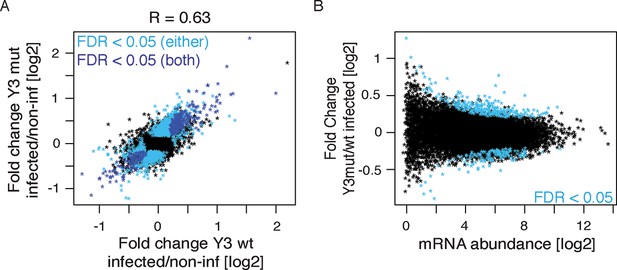
Y3 overexpression does not lead to nELAVL 3'UTR target sequestration.
(A) Correlation of mRNA abundance changes upon Y3wt and Y3mut infection compared to non-infected cell. The mRNA abundance fold change (Y3 infected/non-infected) of Y3mut infected cells was plotted against the mRNA abundance fold change of Y3wt infected cells. Transcripts that change significantly (FDR<0.05) upon either Y3wt infection (n = 502) or Y3mut infection (n = 1920) are colored in light blue. Transcripts that change in both infections and are therefore likely to be virus dependent are colored in dark blue (n = 349). Shown are only transcripts that are expressed (n = 12,659) and the R value is indicated. (B) The mRNA abundance change (Y3mut/Y3wt infection) of transcripts was plotted against average mRNA abundance. Significantly changing transcripts (FDR<0.05; n = 435) are colored in blue, and are not enriched for nELAVL 3’UTR targets.
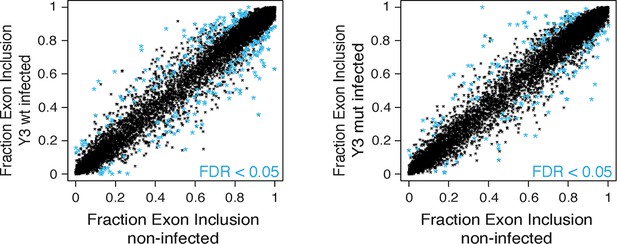
Identification of Y3wt and Y3mut dependent splicing changes.
Shown is the exon inclusion fraction of cassette exons that are expressed in IMR-32 cells subjected to Y3wt or Y3mut infection (n = 10,189). Exons changing significantly (FDR<0.05 and ΔI > 0.1) upon Y3wt (left panel; n = 240) or Y3mut (right panel; n = 151) infection are colored in light blue.
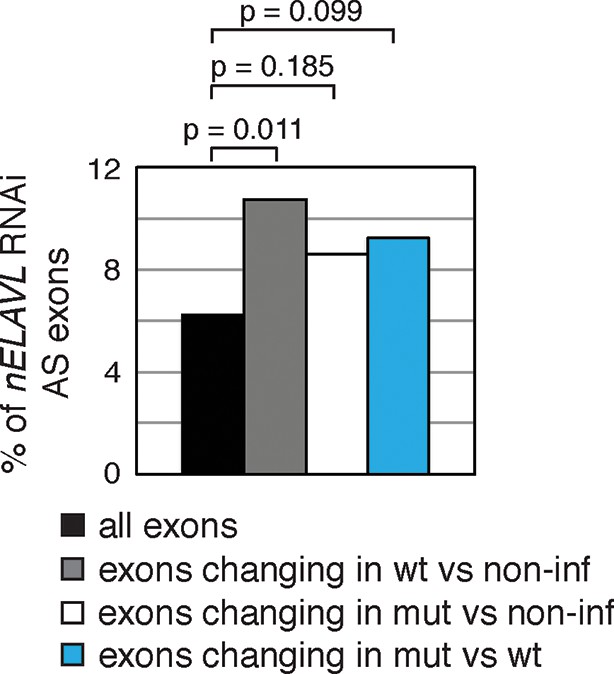
Exons that are alternatively spliced upon Y3wt infection are enriched for nELAVL RNAi dependent exons.
Bar graph representing total expressed exons (n = 10,189), exons that change in either Y3wt (n = 240; blue points in the left panel of Figure 9—figure supplement 4) or Y3mut (n = 151; blue points in the right panel of Figure 9—figure supplement 4) infected cells compared to non-infected cells, and exons that change in Y3wt compared to Y3mut infected cells (n = 191; blue points in Figure 9C). Exons that are alternatively spliced upon Y3wt infection compared to either non-infected (p = 0.011; hypergeometric test) or Y3mut infected cells (p = 0.099; hypergeometric test) are enriched for nELAVL RNAi dependent exons.
Additional files
-
Supplementary file 1
Supplementary files 1A-F.
(A) Sample summary. Shown is subject sample information including postmortem interval (PMI), age, pathology diagnosis, cognitive performance assessed by clinical dementia rating (CDR), plaque pathology assessed by plaque counts, and neuritic performance assessed by Braak staging (BB), as well as read statistics, used linker indexes, and batch information for RNAseq and CLIP data for brain and IMR-32 samples. (B) RNAseq data from human brain List containing 14,876 genes that are expressed in either control or AD brain. Shown are Entrez Gene ID and Symbol, rpkm, log cpm, and the comparison between control and AD brain using edgeR (log2 average cpm, log2 FC (fold change), p and FDR values). (C) nELAVL binding sites in human brain. The table contains 74,423 peaks in human brain that mapped to transcripts expressed in brain. Shown are PeakName, hg18 and hg19 genomic coordinates (chromosome, start, end, strand), Entrez Gene ID and Symbol, genomic region, BC (biological complexity), PH (PeakHeight), and PH normalized for mRNA abundance. (D) nELAVL targets in human brain. Shown are 8681 genes with nELAVL binding sites in human brain. Entrez Gene ID and Symbol, rpkm, log2 cpm and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of total, 3’UTR and intronic peaks are displayed. Top targets and top 3’UTR targets can be extracted by selecting the top 1000 targets according to log2 normalized nELAVL binding or log2 normalized 3’UTR nELAVL binding, respectively. (E) nELAVL top targets in human brain. List of top 1000 nELAVL targets in human brain. Entrez Gene ID and Symbol, rpkm, log2 cpm and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of total, 3’UTR and intronic peaks are displayed. Information on the validation of nELAVL-mediated regulation is included if applicable. (F) PPI Cluster analysis of top nELAVL targets in human brain. Enriched terms of nELAVL top target gene clusters in the PPI subnetwork. The Benjamini-Hochberg corrected p value is < 0.05 for all terms. For large gene clusters (≥20 genes), enriched terms containing no less than 4 genes from the cluster are shown in the table; for small gene clusters (<20 genes), enriched terms containing no less than 2 genes from the cluster are shown. Terms are ranked by increasing p value and only the top 10 terms with a p value <0.05 are shown.
- https://doi.org/10.7554/eLife.10421.035
-
Supplementary file 2
Supplementary files 2A-F.
(A) nELAVL 3’UTR targets validated in mice. Displayed are 37 genes that change significantly in their mRNA abundance in Elavl3/4 KO mice and that have 3’UTR binding in human brain. Included are mouse Entrez Gene ID and Symbol, the count of mouse 3’UTR CLIP tags, mRNA level log2 FC (fold change) in Elavl3/4 KO mice (KO/WT), and t-test results. Human Entrez Gene ID and Symbol, log2 cpm and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of 3’UTR peaks are displayed. (B) nELAVL intron targets validated in mice. Table containing exons that are alternatively spliced in Elavl3/4 KO mice and that have intronic binding adjacent to them (+/- 2.5kb) in human brain. Shown are mouse Gene Symbol, mouse exon information (mm9 and mm10 coordinates), the difference in the exon inclusion rate in Elavl3/4 KO mice compared to wilt-type (ΔI: WT-KO), adjacent mouse CLIP tags, information on experimental validation, human Entrez Gene ID and Symbol, human exon information (hg18 and hg19 coordinates), human peak information (name, hg18 and hg19 coordinates, PH (PeakHeight), PH normalized for mRNA abundance, and the position of the peak relative to the alternative exon (negative values indicate that the peak is upstream of the exon). (C) RNAseq data from IMR-32 cells. List containing 13100 genes that are expressed in at least one IMR-32 dataset (RNAi, UV stress, Y3 infection). Shown are Entrez Gene ID and Symbol, rpkm and log cpm for the different conditions, and the comparisons between the different conditions using edgeR (log2 average cpm, log2 FC (fold change), p and FDR values). If applicable, nELAVL binding information (peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) in human brain) is included. (D) nELAVL 3’UTR targets validated in IMR-32 cells. List containing 743 genes expressed in brain that are changing in their abundance upon nELAVL RNAi and their nELAVL 3’UTR binding if applicable. Included are Entrez Gene ID and Symbol, rpkm and log2 cpm in mock and nELAVL RNAi treated cells, log2 average cpm, log2 FC (fold change), p and FDR values comparing the mRNA abundance between mock and nELAVL RNAi treated cells. Additionally, rpkm and log2 cpm in human brain, and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of 3’UTR peaks (if applicable) in human brain are shown. Top 3’UTR targets are indicated in in the last column. Transcripts that decrease upon nELAVL RNAi can be obtained by selecting rows with log2FC_MockElavl <0. (E) Splicing data from IMR-32 cells. List containing 13496 alternative exons that are expressed in at least one IMR-32 dataset (RNAi, UV stress, Y3 infection). Shown are AS event information (Name, hg18 and hg19 genomic coordinates of the AS event and the alternative exon), Entrez Gene ID and Symbol, exon inclusion fraction, and the comparisons between different conditions (the change in exon inclusion (ΔI), p and FDR values). Peak information (Name, hg18 and hg19 genomic coordinates, PH (PeakHeight), PH normalized for mRNA abundance, and the peak position relative to the alternative exon peaks) of peaks adjacent (+/- 2.5kb) to alternative exons is shown. Information on exons that change in different conditions can be found in the last 7 columns. (F) nELAVL intron targets validated in IMR-32 cells. Displayed are 473 alternative splicing (AS) events that change upon nELAVL RNAi, and if applicable adjacent intronic nELAVL binding sites. Shown are AS event information (Name, hg18 and hg19 genomic coordinates of the AS event and the alternative exon), Entrez Gene ID and Symbol, alternative exon inclusion in mock and nELAVL RNAi (PSI), the change in exon inclusion (ΔI), p and FDR values comparing PSI between mock and nELAVL RNAi treated cells, and peak information if applicable. This includes Peak Name, hg18 and hg19 genomic coordinates, PH (PeakHeight), PH normalized for mRNA abundance, and the position of the peak relative to the alternative exon (negative values indicate that the peak is upstream of the exon).
- https://doi.org/10.7554/eLife.10421.036
-
Supplementary file 3
Supplementary files 3A-F.
(A) Overlap of nELAVL 3’UTR targets and genes with disease associated 3’UTR SNPs. List containing transcripts with nELAVL 3’UTR binding with disease associated 3’UTR SNPs (245 SNPs mapped to 200 genes). Included are Entrez Gene ID and Symbol, rpkm and log2 cpm, nELAVL 3’UTR binding information (peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) in human brain), SNP information (coordinates, associated disease), and, if applicable, information on overlapping peaks (name, hg18 and hg19 coordinates, PH (PeakHeight), PH normalized for mRNA abundance). (B) CLIP and RNAseq data of AD-related genes List of 96 AD-related genes. Shown are Entrez Gene ID and Symbol, rpkm and log2 cpm, and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of peaks (if applicable) in human brain. Top targets are indicated in in the last column. (C) Overlap of top nELAVL targets and GWAS-associated AD genes. Displayed are 77 top nELAVL targets with AD-associated SNPs (p< 0.001). Entrez Gene ID and Symbol, rpkm and log2 cpm, and the peak number, log2 nELAVL binding (CLIP tags within peaks were summarized per transcript), and log2 normalized nELAVL binding (CLIP tags within peaks were normalized for mRNA abundance and summarized per transcript) of peaks (if applicable) in human brain are shown. (D) nELAVL binding sites in control and AD subject brain. The table contains 113,671 nELAVL binding sites that are bound in control or AD brain and that present within expressed genes. Included are PeakName, hg18 and hg19 genomic coordinates (chromosome, start, end, strand), Entrez Gene ID and Symbol, genomic region, PH (PeakHeight), PH normalized for mRNA abundance, and BC (biological complexity) in control and AD brain, and log2 cpm (counts per million CLIP reads per peak), log2 FC (fold change), p and FDR values comparing PH between control and AD brain. (E) Splicing changes in AD Displayed are 170 alternative splicing (AS) events that change in AD. Shown are AS event information (Name, hg18 and hg19 genomic coordinates of the AS event and the alternative exon), Entrez Gene ID and Symbol, exon inclusion fraction in control and AD brain (PSI), the change in exon inclusion (ΔI), p and FDR values comparing PSI between control and AD brain. Among the differentially spliced exons were five nELAVL regulated exons (validated in IMR-32 cells), which were adjacent to 8 intronic peaks). For those exons, alternative exon inclusion in mock and nELAVL RNAi (PSI), the change in exon inclusion (ΔI), p and FDR values comparing PSI between mock and nELAVL RNAi treated cells, and information of adjacent peaks (Name, hg18 and hg19 genomic coordinates, biological complexity (BC), PH (PeakHeight), PH normalized for mRNA abundance, and the peak position relative to the alternative exon) is additionally included. (F) Y RNA expression and nELAVL binding in brain and IMR-32 cells. Table containing 864 Y RNAs. Shown are Y RNA information (Name, hg18 and hg19 Y RNA coordinates, the presence of 6mer or 5mer Hu binding motif, and type), and log2 average of YRNA sequencing (seq) and nELAVL binding (CLIP) in brain and IMR-32 cells, and the YRNA expression log2 fold change upon Y3wt and Y3mut infection. Y RNAs bound in AD and IMR-32 cells can be extracted by selecting BC_brain_CLIP≥2 and BC_IMR-32_CLIP ≥ 2, respectively.
- https://doi.org/10.7554/eLife.10421.037





