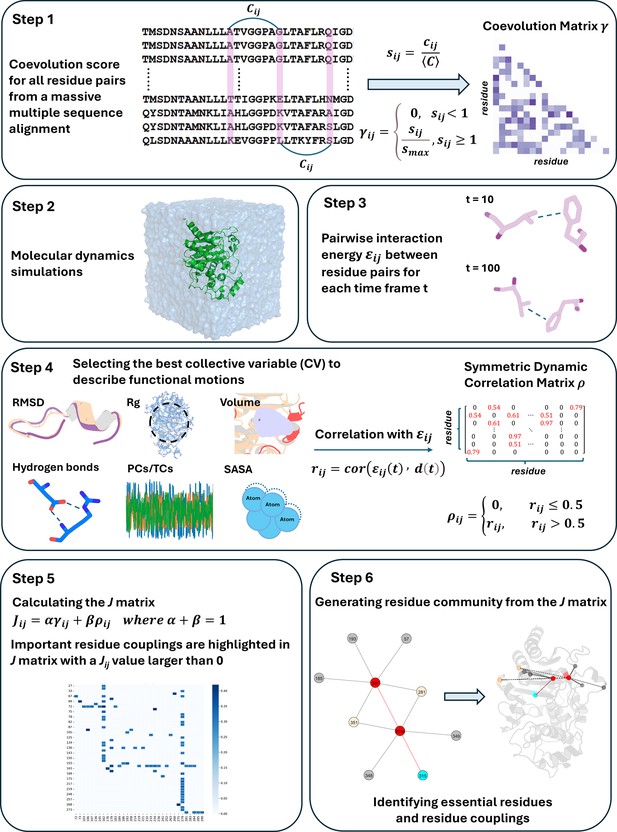
Functionally important residues from graph analysis of coevolved dynamic couplings
Figures
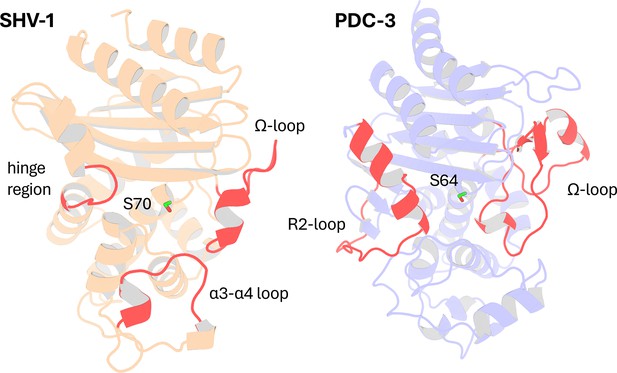
Structural Comparison of SHV-1 (PDB ID: 3N4I) and PDC-3 (PDB ID: 4HEF) β-lactamases.
Catalytic serine S70 (SHV-1) and S64 (PDC3) are highlighted using stick representation. Important loops surrounding the active site are highlighted in red. In SHV-1, highlighted loops are the α3-α4 loop (residues 101–111), the Ω-loop (residues 164–179), and the hinge region (residues 213–218). In PDC-3, highlighted loops are the Ω-loop (residues 183–226) and the R2-loop (residues 280–310).
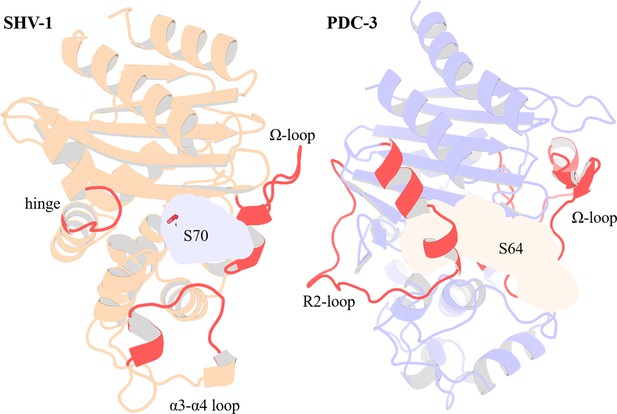
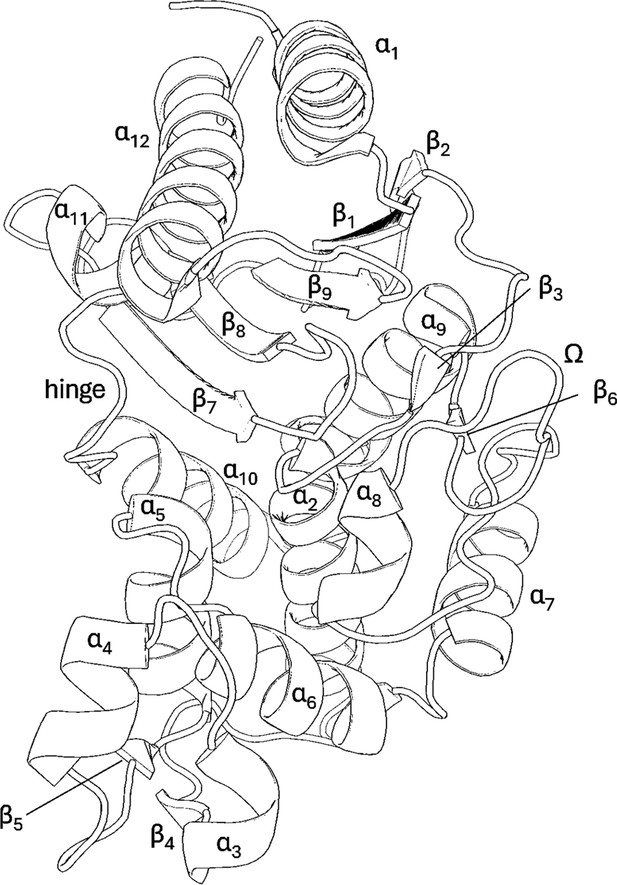
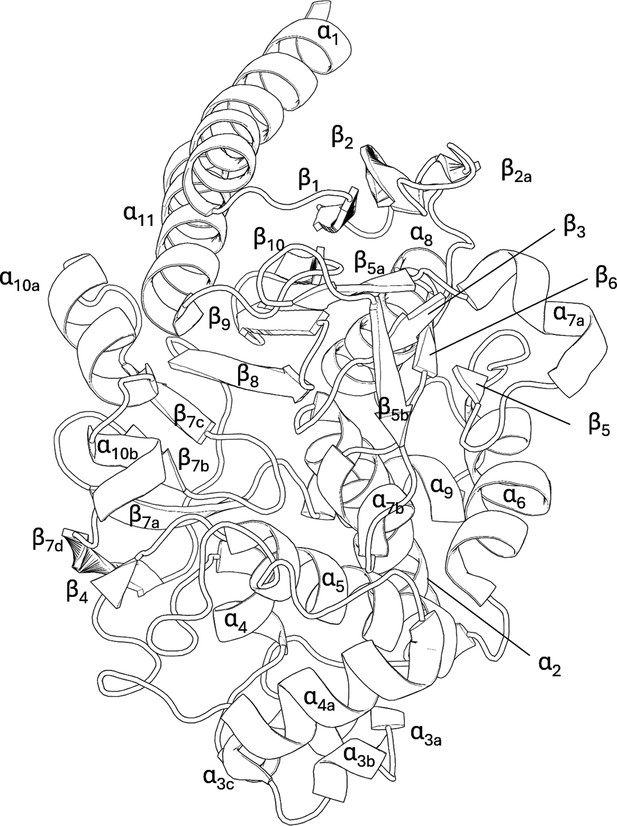
Crystal structure and binding pocket of SHV-1 (PDB ID: 3N4I) and PDC-3 (PDB ID: 4HEF).
S70 and S64 are the catalytic serine residue for SHV-1 and PDC-3, respectively. The α3-α4 loop, the Ω-loop, and the hinge region in SHV-1 are highlighted in red. The Ω-loop and the R2-loop in PDC-3 are coloured in red.
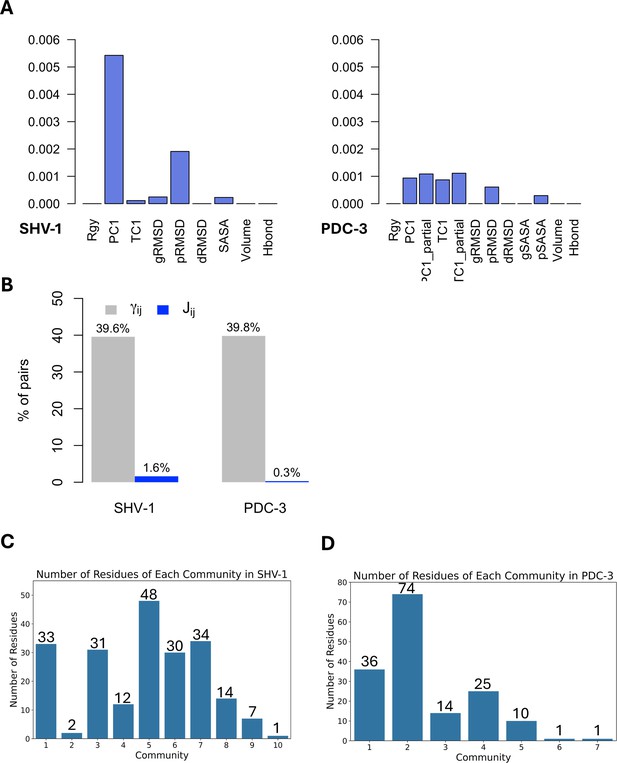
Features and residues in each community.
(A) Average J matrix score varies across different collective variables (CVs). Left: SHV-1; right: PDC-3. (B) Number of non-zero couplings detected by scaled coevolution scores () and J values calculated by DyNoPy (). (C) Number of residues of each community in SHV-1. (D) Number of residues of each community in PDC-3. A reasonable residue community should contain at least three residues.
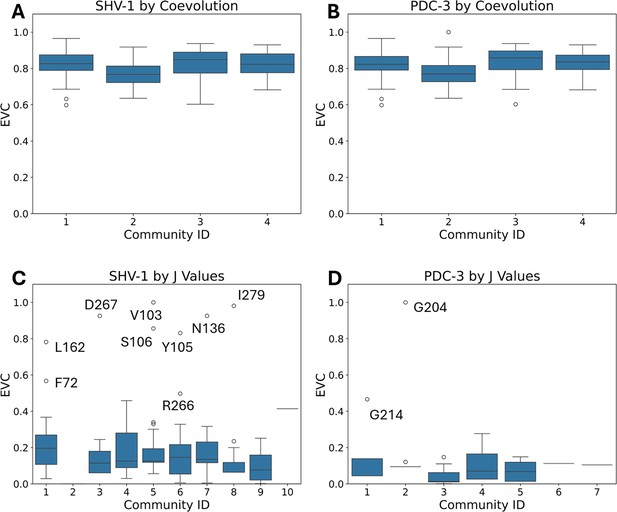
Eigenvector centrality (EVC) distribution of all the residues for each community.
(A) EVC distribution of SHV-1 for each community detected by traditional coevolution scores. (B) EVC distribution of PDC-3 of each community detected by traditional coevolution scores. (C) EVC distribution of SHV-1 of each community induced from J value calculated by DyNoPy. (D) EVC distribution of PDC-3 of each community induced from J value calculated by DyNoPy.
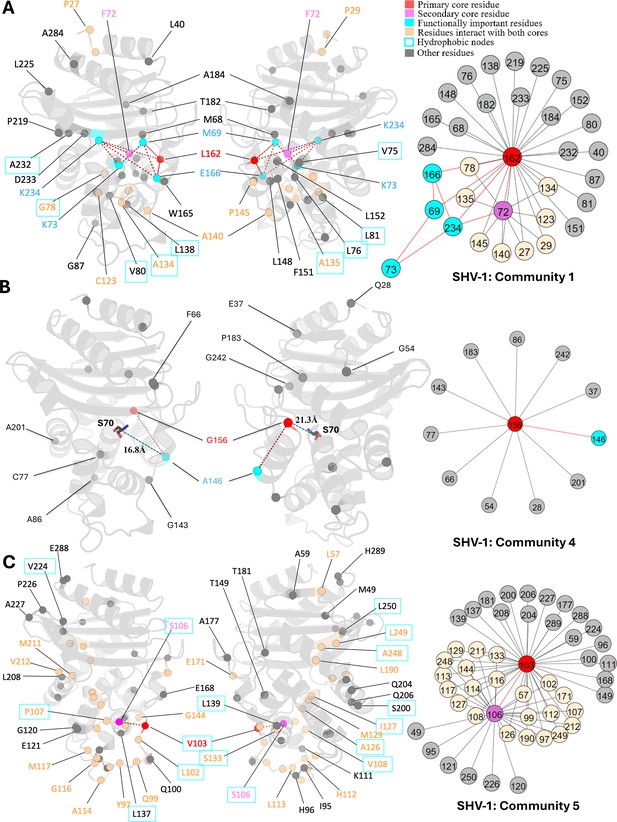
Communities 1, 4, and 5 of SHV-1 β-lactamase.
All the residues are depicted as spheres on the protein structure. The core residue for each community is highlighted in red, while purple is used to emphasize the secondary core residue. Residues that interact with both cores are coloured in light yellow. Functional important residues are marked in cyan. Hydrophobic nodes are enclosed with cyan boxes. (A) Community 1 of SHV-1, comprising 33 residues with L162 being the primary core residue. (B) Community 4 of SHV-1, containing 12 residues and is centred by G156. G156 and A146 are two functional important residues distant from the active site. G156 is 21.3 Å away from the catalytic S70. A146 is 16.8 Å away from S70. (C) Community 5 of SHV-1, embracing 48 residues and showing a strong correlation between V103 and S106.
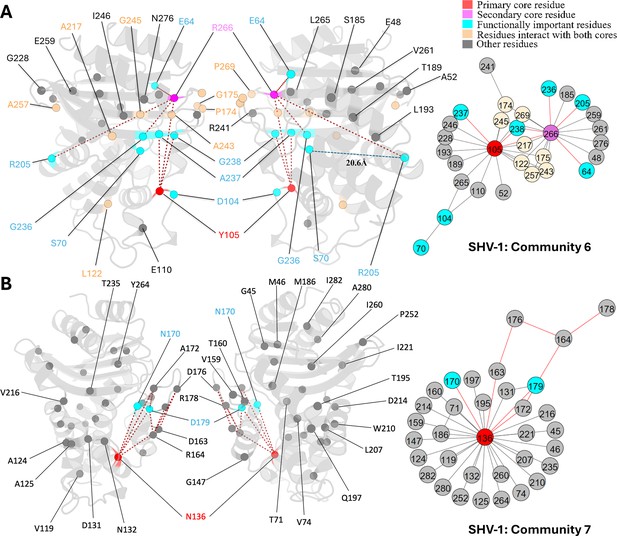
Communities 6 and 7 of SHV-1 β-lactamase.
All the residues are depicted as spheres on the protein structure. The core residue for each community is highlighted in red, while purple is used to emphasize the secondary core residue. Residues that interact with both cores are coloured in light yellow. Functional important residues are marked in cyan. (A) Community 6 of SHV-1, comprising 30 residues with Y105 being the primary core residue. R205 is a functional important residue that is 20.6 Å away from the active site S70. (B) Community 7 of SHV-1, containing 34 residues and is centred by N136.
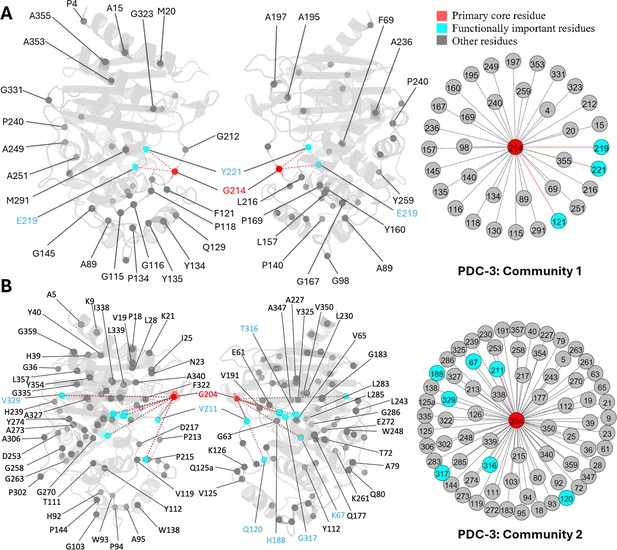
Communities 1 and 2 of PDC-3 β-lactamase.
All the residues are depicted as spheres on the protein structure. The core residue for each community is highlighted in red. Functional important residues are marked in cyan. (A) Community 1 of PDC-3, comprising 36 residues with G214 being the primary core residue. (B) Community 2 of PDC-3, containing 74 residues and is centred by G204.
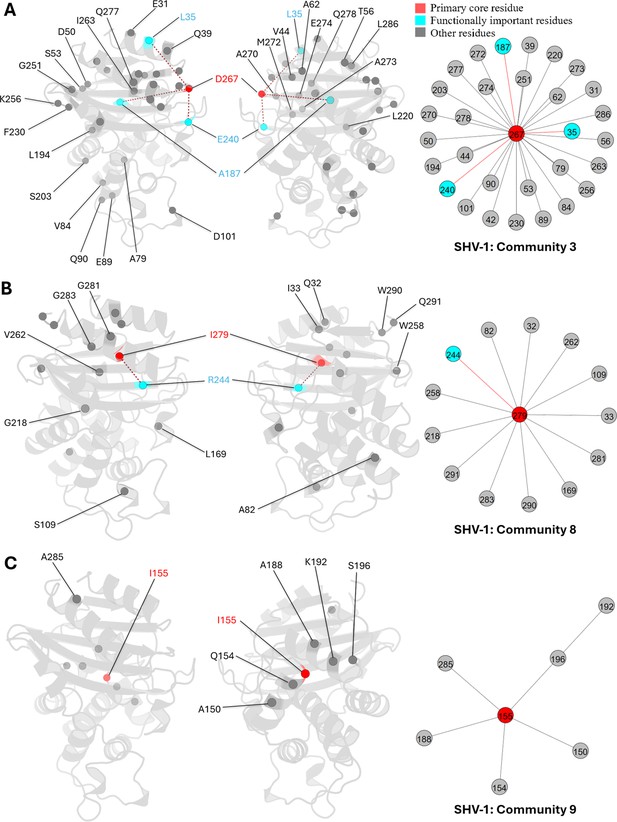
Communities 3, 8, and 9 of SHV-1 β-lactamase.
All the residues are depicted as spheres on the protein structure. The core residue for each community is D267, I279, and I155, respectively. They are highlighted in red. Functional important residues are marked in cyan. (A) Community 3 of SHV-1, comprising 31 residues. (B) Community 8 of SHV-1, containing 14 residues. (C) Community 9 of SHV-1, with 7 residues.
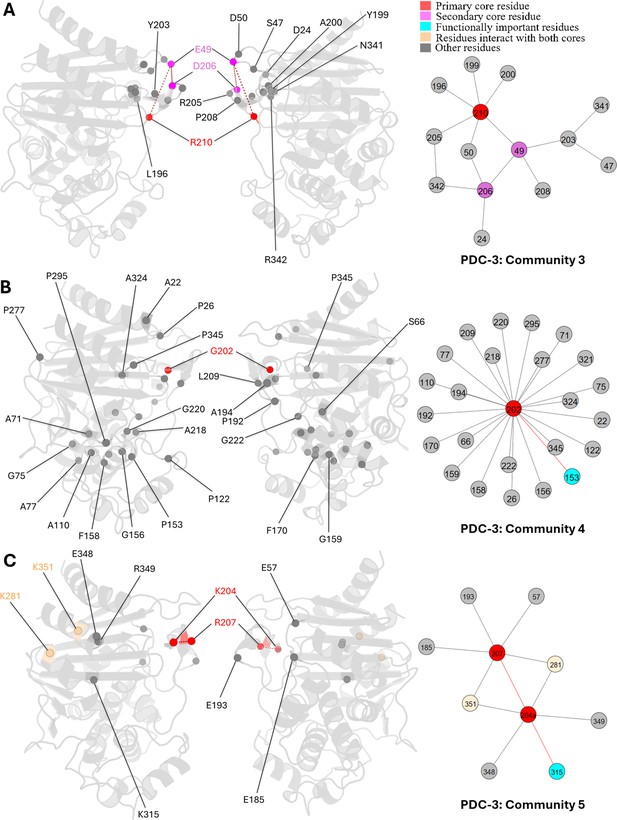
Communities 3, 4, and 5 of PDC-3 β-lactamase.
All the residues are depicted as spheres on the protein structure. The core residue for each community is highlighted in red, while purple is used to emphasize the secondary core residue. Residues that interact with both cores are coloured in light yellow. Functional important residues are marked in cyan. (A) Community 3 of PDC-3, comprising 14 residues with R210 being the primary core residue. (B) Community 4 of PDC-3, containing 25 residues and is centred by G202. (C) Community 5 of PDC-3, embracing 10 residues and having two core residues K204 and R207.
Additional files
-
Supplementary file 1
Features, descriptors and protocol summary.
(a) Hydrophobic nodes in SHV-1. (b) Dynamic descriptors and number of residue pairs detected. (c) Number of residues in each community detected using only the coevolution scores. (d) Summary of molecular dynamics simulation systems.
- https://cdn.elifesciences.org/articles/105005/elife-105005-supp1-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/105005/elife-105005-mdarchecklist1-v1.docx