The Drosophila EGF domain protein uninflatable sets the switch between wrapping glia growth and axon wrapping instructed by Notch
Figures
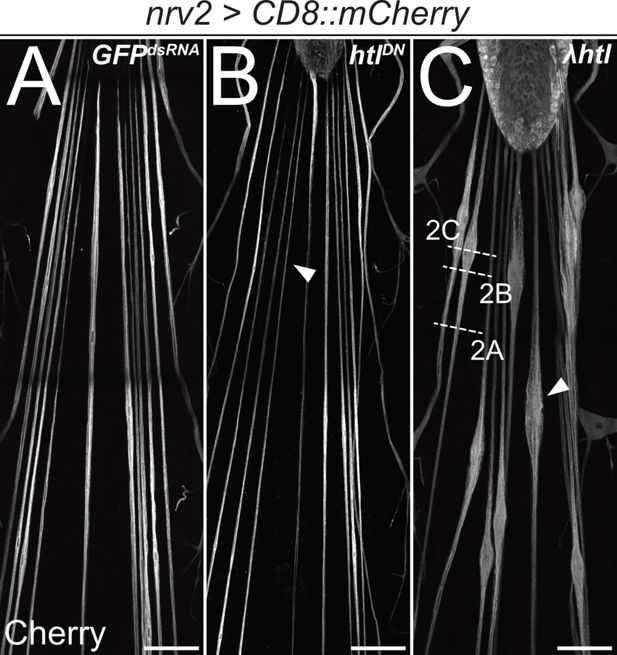
Heartless is required for wrapping glia development.
Confocal images of third instar larval filet preparations, stained for CD8::mCherry expression. The segmental nerves posterior to the ventral nerve cord are shown. Wrapping glial morphology is (A) not changed in control animals expressing (nrv2-Gal4) mock GFPdsRNA. (B) Differentiation of wrapping glial cells is affected following expression of a dominant negative form of Htl and thin wrapping glial cells are detected (arrowhead). (C) Expression of a constitutively active form of Htl [nrv2-Gal4; UAS- λhtl] leads to nerve bulges around the wrapping glial nuclei (C, arrowhead). Scale bars 100 µm. The white dashed lines indicate the level of sections shown in Figure 2.

Suppressor screen for genes downstream of activated heartless.
Exemplary images of living third instar larvae expressing repo-stinger::GFP. Analyzed genes were classified according to the changes in bulge size. 0=no change; 1, 2=slight changes in the bulge size. 3=strong suppression of the bulge size; 4=complete suppression of the bulging phenotype. For the list of genes screened, please see Supplementary file 1.
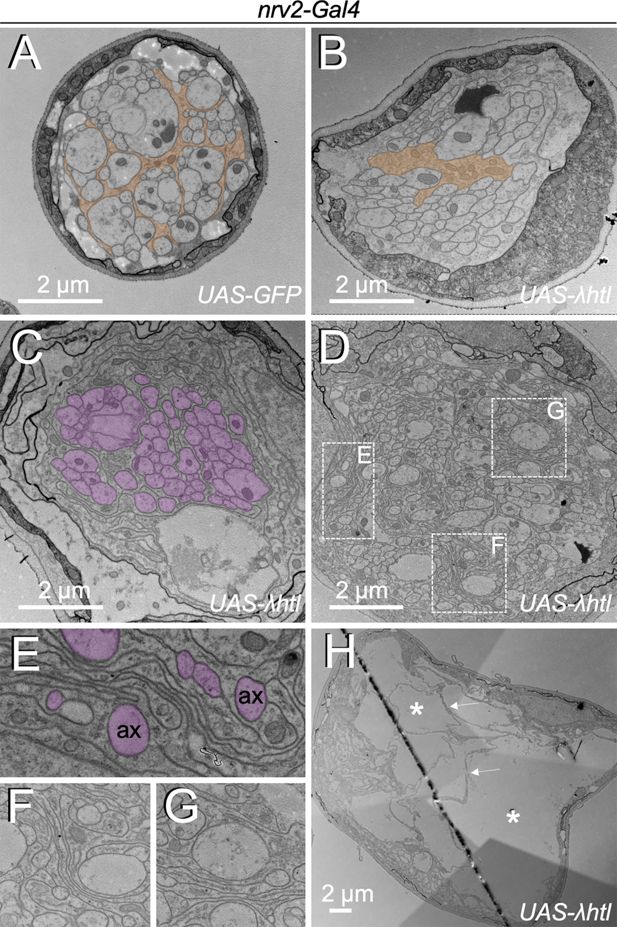
Heartless controls growth of wrapping glia cells.
Electron microscopic images of segmental nerves from wandering third instar larvae. (A) Nerve of a control larva expressing GFP in wrapping glia sectioned 150 µm posterior to the ventral nerve cord. Normally differentiated wrapping glia can be seen. (B–H) Nerves of larvae expressing activated Heartless in wrapping glia [nrv2-Gal4; UAS-λhtl] sectioned at the positions indicated in Figure 1C. (B) Note the poorly differentiated wrapping glial cells distant from the nerve bulge. (C,D) At the beginning of the nerve bulge, excessive differentiation of wrapping glial cell processes starts to be detected that do not always grow around axons (magenta). (E–G) Higher magnifications of the boxed areas in (D). Note the formation of wrapping glial cell processes that do not contact axons (ax, magenta) but rather contact glial cell processes. (H) In the central area of the nerve bulge, liquid-filled vacuolar structures (asterisks) can be detected. Thin wrapping glial cell processes (arrows) span the bulged area.
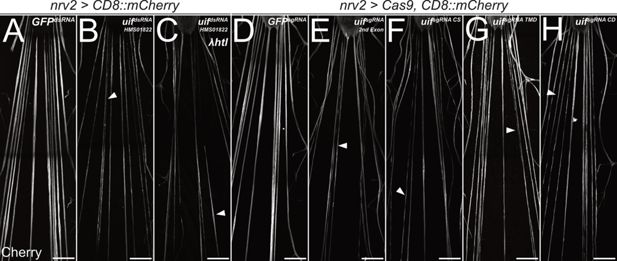
uninflatable affects differentiation of wrapping glia.
Confocal images of third instar larval filet preparations, stained for CD8::mCherry expression. The segmental nerves posterior to the ventral nerve cord are shown. (A) Filet preparation of a control third instar larva. (B) uif knockdown in wrapping glial cells [nrv2-Gal4; UAS-uifdsRNA-HMS01822] impairs their development, which (C) cannot be rescued by co-expression of activated Htl. (D–H) Ubiquitous expression of (D) mock control GFPsgRNA, (E) uifsgRNA 2nd Exon, (F) uifsgRNA CS, (G) uifsgRNA TMD, or (H) uifsgRNA CD in Cas9-expressing larvae. All uif sgRNAs disrupt wrapping glial cell development, and wrapping glia appear thin and fragmented (arrowheads). n=5 larvae for all genotypes. Scale bars 100 µm.
-
Figure 3—source data 1
Excel file giving the number of axons in the analyzed nerves to calculate the wrapping index.
- https://cdn.elifesciences.org/articles/105759/elife-105759-fig3-data1-v1.xlsx
-
Figure 3—source data 2
Excel file giving the number of axons in the analyzed nerves to calculate the wrapping index.
- https://cdn.elifesciences.org/articles/105759/elife-105759-fig3-data2-v1.xlsx

Schematic representation of sgRNA target sites used for CRISPR-mediated uif knockout.
Target sites (black arrowhead) and predicted protein residues (Uif isoform B; present domains shown in color, absent domains shown in gray). Sequence of domains from N- to C-terminus: signal sequence (pink); one C-type lectin-like (CLECT) domain (turquoise), proposed to be a calcium-dependent carbohydrate binding module Cambi and Figdor, 2003; one low-density lipoprotein receptor protein class A (LDLa) domain (dark yellow), predicted to have a protein binding function May et al., 2007; three CUB domains (light yellow), a structural domain of unknown function; eight CCP domains (beige), predicted to be adhesion protein interaction domains; 18 EGF-like repeats (10 are Ca2+ binding; light/dark petrol), which are protein interaction domains involved in cell signaling Appella et al., 1988; two coagulation factor 5/8 C-terminal (FA58C) domains (light red), which are carbohydrate binding motifs Zhang and Ward, 2009; three hyaline repeat (HYR) domains (dark red), with a putative function in cell adhesion Carroll et al., 2008; five ephrin-like domains (orange) and one laminin G domain (light blue), also likely involved in cell adhesion and signaling Banerjee et al., 2011; proteolytic cleavage site (scissors) at amino acid 2991 Zhang and Ward, 2009; antibody binding site (AB) at amino acid 2882–3157 Zhang and Ward, 2009; additional exon isoform C indicated by red line; transmembrane domain (dark blue); intracellular C-terminus does not contain any conserved domains (Zhang and Ward, 2009).
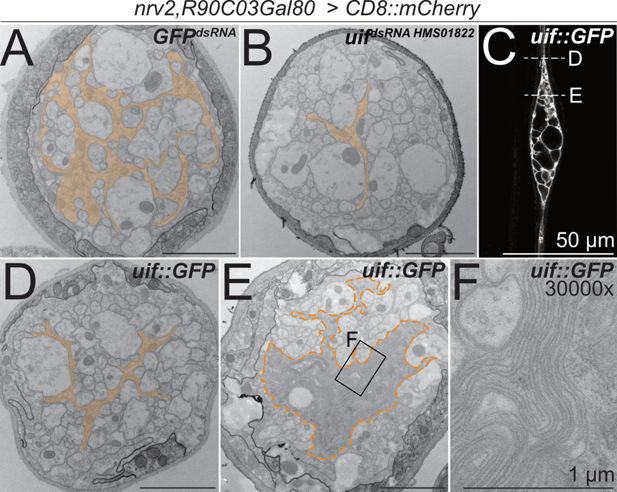
uif affects axonal ensheathment of wrapping glia.
(A+B, D–F) Electron microscopic cross sections of third instar larval abdominal peripheral nerves with wrapping glia-specific expression [nrv2-Gal4; R90C03-Gal80] of (A) GFPdsRNA mock control transgene and (B) uifdsRNA-HMS01822. Upon knockdown of uif, wrapping glial cell complexity is reduced. (C) Single plane of a confocal image of a third instar larval nerve stained for CD8::mCherry expression. Expression of uif::GFP in wrapping glial cells causes bulge formation, while outside the bulge, wrapping glia appear thin (arrowheads). The dashed lines indicate the plane of section in relation to the bulge of the images shown in (D, E). (D) Upon uif::GFP overexpression specifically in wrapping glia, glial morphology is reduced outside the bulge region. (E) Within the nerve bulge, wrapping glial membrane increases in size while most axons lack proper wrapping. (F) Close-up of the region indicated in (E). GFPdsRNA n=4 larvae, 4–7 nerves per specimen; uifdsRNA n=3 larvae, 5–9 nerves per specimen; uif::GFP n=3 larvae, 5–6 nerves per specimen. Scale bars 2 µm unless indicated otherwise.

Quantification of the wrapping index (WI).
WI quantified from electron microscopic cross sections of third instar larval abdominal peripheral nerves with (A) wrapping glia-specific (nrv2-Gal4;R90C03-Gal80) expression of mock control GFPdsRNA (WI = 0.17), uif dsRNAHMS01822, uif::GFP, Su(H)dsRNA, and mamdsRNA is plotted. Both uif knockdown (WI = 0.03) and overexpression (WI = 0.08) lead to a significant reduction of axonal wrapping (comparing the WI of control and uifdsRNA larvae: p=2.88 × 10–7; comparing the WI of control and uif::GFP larvae: p=2.56 × 10–9). Similarly, knockdown of the Notch downstream components Su(H) and mam reduces the WI significantly (mamdsRNA, WI = 0.07, p=4.29 × 10–9; Su(H)dsRNA, WI = 0.075, p=4.21 × 10–11). (B) Neuronal knockdown (nSyb-Gal4) of Contactin leads to a significant reduction in the WI from 0.15 to 0.11 (for both RNAi lines p=0,006). For statistical analysis, a t-test was performed for normally distributed data (Shapiro test), and a Mann-Whitney U test was performed for not normally distributed data. GFPdsRNA n=4 larvae with 4–7 nerves per specimen; uifdsRNA n=3 larvae, 5–9 nerves per specimen; uif::GFP n=3 larvae, 5–6 nerves per specimen; mamdsRNA n=3 larvae with 5–9 nerves per specimen; Su(H)dsRNA n=4 larvae with 6–8 nerves per specimen; ContdsRNA (both) n=5 larvae with 10 nerves per specimen. **p≤0.01, ***p≤0.001.
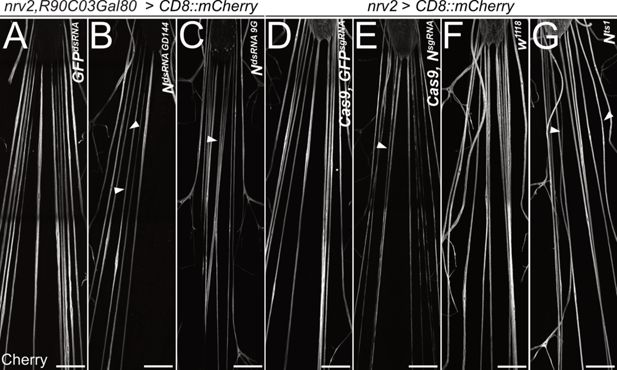
Notch is required for wrapping glial development.
Filet preparations of wandering third instar larvae stained for CD8::mCherry expression. The segmental nerves just posterior to the ventral nerve cord are shown. (A) Control larvae expressing the mock control GFPdsRNA in wrapping glial cells specifically [nrv2-Gal4; R90C03-Gal80]. Upon expression of dsRNA targeting Notch mRNAs (B) NGD144, (C) N9G. (D) Control larva expressing sgRNA directed against the GFP open reading frame in [nrv2-Gal4; UAS-Cas9]. (E) Conditional knockout of Notch leads to dramatically altered morphology of wrapping glial cells. (F) Control larva cultured at the same temperature regime as the Nts1 larva shown in (G). Wrapping glial cells appear smaller compared to the control. Scale bars 100 µm.

Notch activation impairs wrapping glial development at higher temperatures.
Filet preparations of wandering third instar larvae stained for CD8::mCherry expression. (A) Nuclear LacZ as mock control for (B) overexpression of Notch intracellular domain in wrapping glia specifically (nrv2-Gal4;R90C03-Gal80). NICD overexpression does not affect wrapping glial morphology. n=5 larvae for all genotypes. Scale bars 100 µm.
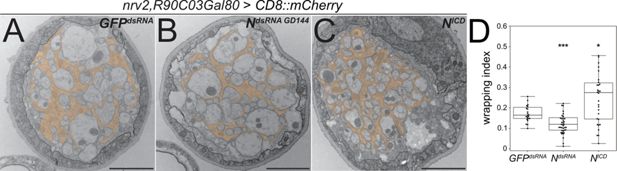
Notch function affects axonal wrapping.
Electron microscopic images of segmental nerves from wandering third instar larvae sectioned 100 µm posterior to the ventral nerve cord. (A) Control nerve of an animal expressing dsRNA directed against GFP [nrv2-Gal4; R90C03-Gal80]. Axons are wrapped by processes of the wrapping glia. The entire nerve is engulfed by the perineurial glia and the subperineurial glia. For quantification of glial wrapping, see (D). (B) Upon expression of dsRNA targeting Notch mRNA (NGD144), glial wrapping is reduced. (C) Upon expression of an activated form of Notch (Nintra), glial wrapping is increased. (D) Quantification of wrapping index (WI). While Notch knockdown significantly decreases the WI (from 0.17 to 0.12, p=0.00079), activation of Notch signaling significantly increases glial wrapping and the WI (from 0.17 to 0.27, p=0.014). For statistical analysis, a t-test was performed for normally distributed data (Shapiro test), and Mann-Whitney U test was performed for not normally distributed data. GFPdsRNA n=4 larvae with 4–7 nerves each; NdsRNA n=5, larvae with 7–9 nerves each; NICD n=5 larvae with 3–8 nerves each. α=0.05, *p≤0.05, ***p≤0.001. Scale bars 2 µm.
-
Figure 6—source data 1
Excel file giving the number of axons in the analyzed nerves to calculate the wrapping index.
- https://cdn.elifesciences.org/articles/105759/elife-105759-fig6-data1-v1.xlsx

The Notch activity reporter is active in differentiated glia.
Maximum intensity projection of a confocal image stack of third instar larval filet and adult brain preparations stained for anti-LacZ, representing Notch activity (green), glial nuclei (anti-Repo, magenta), and neuronal membranes (anti-HRP, blue). Bacterial LacZ gene under the control of three copies of palindromic Grainyhead (Grh) binding sites (Gbe), and Su(H) binding is active specifically in cells where Notch signaling is present. (A, B) Expression can be detected in neuroblasts of the brain lobes (asterisk and close-up) and the thoracic neuromeres (unfilled arrowhead). Additional Notch activity is detected in the imaginal discs (filled arrowheads); scale bar 200 µm. (C–C’’) No Notch activity is found in larval peripheral glial nuclei; scale bar 100 µm. (D, E) LacZ staining overlaps with Repo staining, indicating expression in adult glia (close-ups, filled arrowheads), which is also detectable in non-glial cells (close-ups, unfilled arrowheads). Not all Repo-positive cells show LacZ staining (close-ups, asterisk), indicating that Notch signaling is not active in all glia. Scale bar 100 µm. (F–F’’’) Single plane of Z-stack is shown. Some peripheral glial cells show LacZ staining (arrowheads); scale bar 100 µm. n=3 animals for both developmental stages.
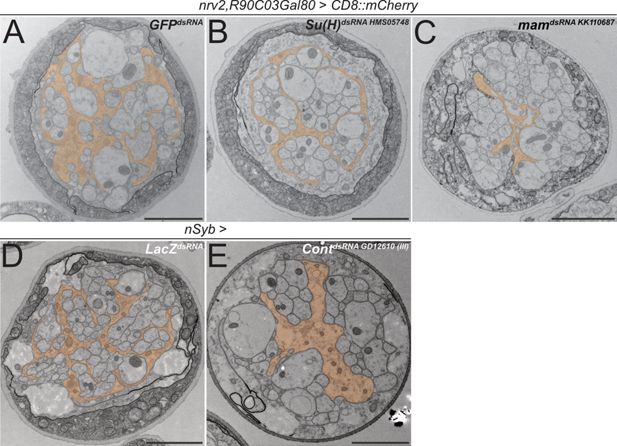
Knockdown of mam, Su(H), and contactin impairs axonal wrapping.
Electron microscopic cross sections of third instar larval abdominal peripheral nerves. Glial cell morphology is indicated in orange color. (A–C) Wrapping glia-specific [nrv2-Gal4; R90C03-Gal80] expression of GFPdsRNA as mock control, (B) Suppressor of Hairless (Su(H)) dsRNAHMS05748, or (C) mastermind (mam) dsRNAKK110687. Note reduced complexity of glial cell processes. (D) Neuron-specific [nSyb-Gal4] expression of LacZdsRNA as mock control and (E) Contactin (Cont) dsRNAGD12610 inserted on the third chromosome. Upon neuronal knockdown of Cont, glial wrapping of peripheral axons is impaired. Scale bars 2 µm.

Knockdown of mam and Su(H) impairs wrapping glial development.
Filet preparations of wandering third instar larvae stained for CD8::mCherry expression. (A) Wrapping glia-specific (nrv2-Gal4;R90C03-Gal80) expression of GFPdsRNA as mock control, (B) mastermind (mam) dsRNAKK110687, or (C) Suppressor of Hairless (Su(H)) dsRNAHMS05748. Note impaired wrapping glial morphology (B, C, arrowheads). n=5 larvae for all genotypes. Scale bars 100 µm.

Neuronal knockdown of Notch ligands Dl and Ser does not affect wrapping glial morphology.
Filet preparations of wandering third instar larvae stained for (A) CD8::mCherry expression or (B) Repo (stains glial nuclei, green) and H2B-mRFP expression. A Trojan-Gal4 insertion in the Delta (Dl) locus [DlMI04868-TG4.1] drives expression of (A) CD8::mCherry targeting membranes or (B) H2B::mRFP targeting nuclei. Two Dl-expressing neurons are found in each hemineuromere and show no repo expression (B and B’, close-up). n=5 larvae for all genotypes. Scale bars 100 µm. (C–E) Filet preparations stained for nrv2::GFP expression. (C) Neuron-specific expression of mock control Cherry dsRNA (D) Serrate (Ser) dsRNAGD14442 and (E) Delta (Dl) dsRNAGD2642. Neither Ser nor Dl knockdown in neurons affects wrapping glial morphology. n=5 larvae for all genotypes. Scale bars 100 µm.

Neuronal knockout of Notch ligands Dl and Ser does not affect wrapping glial morphology.
Filet preparations of wandering third instar larvae with the genotypes indicated were sectioned 150 µm distant from the tip of the ventral nerve cord. Electron microscopic images were analyzed for their wrapping index. Cas9-mediated neuronal knockout of neither Delta nor Serrate significantly affects the wrapping index.
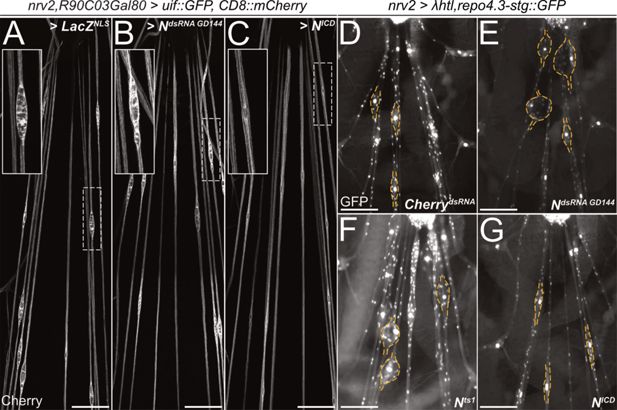
Notch counteracts heartless and uninflatable function.
Squeeze preparations of wandering third instar larvae. The segmental nerves posterior to the ventral nerve cord are shown. (A–C) Larvae expressing uninflatable and CD8::mCherry in the wrapping glia. The nerve bulge shown in the white dashed box is enlarged. (A) Control larvae co-expressing LacZ. Uif expression results in nerve bulges. (B) Upon co-expression of NotchdsRNA (GD144), the nerve bulging phenotype is not suppressed. (C) Upon activation of Notch signaling by expression of NICD, the nerve bulging phenotype is suppressed. (D–G) Larvae expressing an activated Heartless receptor (λhtl) in the wrapping glia [nrv2-Gal4] carrying a repo3.4-stinger::GFP element leading to a panglial nuclear GFP expression. (D) Control larvae co-expressing dsRNA directed against mCherry. Note the prominent nerve bulges. (E) Upon co-expression of NotchdsRNA, the nerve bulging phenotype is enhanced. (F) Likewise, the nerve bulging phenotype is enhanced in a Notchts1 mutant background when the larvae are kept at the restrictive temperature. (G) Upon activation of Notch signaling by expression of NICD, the nerve bulging phenotype is significantly rescued. n=7 larvae for all genotypes. (A-C) Scale bars 100 µm, (D-G) Scale bars 75 µm.
-
Figure 8—source data 1
Excel file giving the number of axons in the analyzed nerves to calculate the wrapping index.
- https://cdn.elifesciences.org/articles/105759/elife-105759-fig8-data1-v1.xlsx
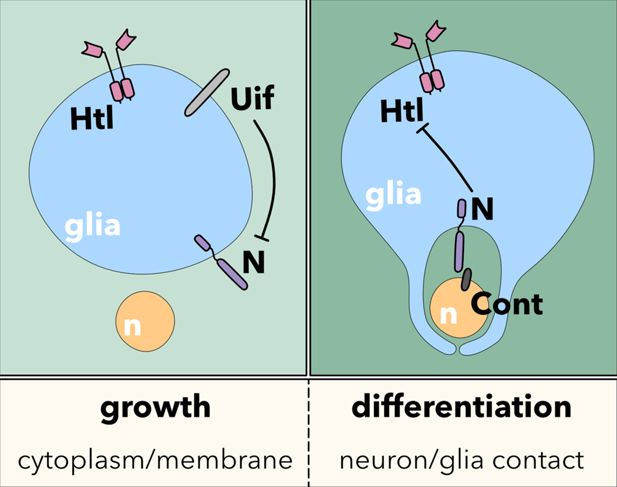
Model.
The interplay of FGF-receptor, Uninflatable, and Notch signaling controls the switch between glial cell growth and glial cell differentiation, leading to extensive neuron-glia contact. For details, please see the text.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | E. coli OneShot TOP10 | Invitrogen | C404003 | Chemically competent cells |
| Genetic reagent (D. melanogaster) | UAS-CD8::mCherry | Bloomington Drosophila Stock Center | RRID:BDSC_27391 | |
| Genetic reagent (D. melanogaster) | UAS-mCherrydsRNA | Bloomington Drosophila Stock Center | RRID:BDSC_35785 | |
| Genetic reagent (D. melanogaster) | w1118 | Bloomington Drosophila Stock Center | RRID:BDSC_3605 | |
| Genetic reagent (D. melanogaster) | UAS-uifdsRNA HMS01822 | Bloomington Drosophila Stock Center | RRID:BDSC_38354 | |
| Genetic reagent (D. melanogaster) | nSyb-Gal4 | Bloomington Drosophila Stock Center | RRID:BDSC_51635 | |
| Genetic reagent (D. melanogaster) | UAS-htlDN | Bloomington Drosophila Stock Center | RRID:BDSC_5366 | |
| Genetic reagent (D. melanogaster) | UAS-Cas9 | Bloomington Drosophila Stock Center | RRID:BDSC_58985 | |
| Genetic reagent (D. melanogaster) | UAS-Su(H)dsRNA HMS05748 | Bloomington Drosophila Stock Center | RRID:BDSC_67928 | |
| Genetic reagent (D. melanogaster) | nrv2-Gal4 | Bloomington Drosophila Stock Center | RRID:BDSC_6800 | |
| Genetic reagent (D. melanogaster) | nrv2::GFP | Bloomington Drosophila Stock Center | RRID:BDSC_6828 | |
| Genetic reagent (D. melanogaster) | UAS-NdsRNA 9G | Bloomington Drosophila Stock Center | RRID:BDSC_7077 | |
| Genetic reagent (D. melanogaster) | DlMI04868-TG4.1 | Bloomington Drosophila Stock Center | RRID:BDSC_77753 | |
| Genetic reagent (D. melanogaster) | U6-DlsgRNA | Bloomington Drosophila Stock Center | RRID:BDSC_83095 | |
| Genetic reagent (D. melanogaster) | NsgRNA | Bloomington Drosophila Stock Center | RRID:BDSC_84168 | |
| Genetic reagent (D. melanogaster) | U6-SersgRNS | Bloomington Drosophila Stock Center | RRID:BDSC_84169 | |
| Genetic reagent (D. melanogaster) | UAS-GFPdsRNA | Bloomington Drosophila Stock Center | RRID:BDSC_9331 | |
| Genetic reagent (D. melanogaster) | UAS-H2B-mRFP | Bloomington Drosophila Stock Center | RRID:BDSC_94270 | |
| Genetic reagent (D. melanogaster) | UAS-mamdsRNA KK110687 | Vienna Drosophila Resource Center | VDRC102091 | |
| Genetic reagent (D. melanogaster) | UAS-NdsRNA GD144 | Vienna Drosophila Resource Center | VDRC1112 | |
| Genetic reagent (D. melanogaster) | UAS-SerdsRNA GD14442 | Vienna Drosophila Resource Center | VDRC27172 | |
| Genetic reagent (D. melanogaster) | UAS-ContdsRNA GD12610 | Vienna Drosophila Resource Center | VDRC28294 | |
| Genetic reagent (D. melanogaster) | UAS-DldsRNA GD2642 | Vienna Drosophila Resource Center | VDRC37287 | |
| Genetic reagent (D. melanogaster) | UAS-ContdsRNA GD12610 | Vienna Drosophila Resource Center | VDRC40613 | |
| Genetic reagent (D. melanogaster) | N[ts]1 | Shellenbarger and Mohler, 1975 | ||
| Genetic reagent (D. melanogaster) | UAS-LacZNLS | Hummel et al., 2002 | ||
| Genetic reagent (D. melanogaster) | UAS-λhtl | Michelson et al., 1998 | ||
| Genetic reagent (D. melanogaster) | UAS-NICD | Klein, University of Düsseldorf | ||
| Genetic reagent (D. melanogaster) | UAS-uif::GFP | Gonzalez-Gaitan, Loubéry et al., 2014 | ||
| Genetic reagent (D. melanogaster) | UAS-LacZdsRNA | Schirmeier, University of Dresden | ||
| Genetic reagent (D. melanogaster) | GFPsgRNA | Schirmeier | ||
| Genetic reagent (D. melanogaster) | uifsgRNA 2nd Exon | This work | See Figure 3—figure supplement 1 | |
| Genetic reagent (D. melanogaster) | uifsgRNA CS | This work | See Figure 3—figure supplement 1 | |
| Genetic reagent (D. melanogaster) | uifsgRNA TMD | This work | See Figure 3—figure supplement 1 | |
| Genetic reagent (D. melanogaster) | uifsgRNA CD | This work | See Figure 3—figure supplement 1 | |
| Genetic reagent (D. melanogaster) | nrv2-Gal4;R90C03-Gal80,UAS-CD8::Cherry | Kottmeier et al., 2020 | ||
| Genetic reagent (D. melanogaster) | Gbe +Su(H)-lacZ | Furriols and Bray, 2001 | ||
| Genetic reagent (D. melanogaster) | repo4.3-stg-GFP | This work | Figure 1 | |
| Antibody | anti-dsRed 1:1000 | Clontech Labs 3P | 632496 | IF(1:1000) |
| Antibody | anti-β-galactosidase (Mouse monoclonal) | Developmental Studies Hybridoma Bank | 40-1a | IF(1:10) |
| Antibody | anti-Repo (Mouse monoclonal) | Developmental Studies Hybridoma Bank | 8D12 | IF(1:5) |
| Antibody | anti-HRP-DyLight 649 (Goat polyclonal) | Dianova | 123-165-021 | IF(1:500) |
| Antibody | anti-GFP (Rabbit polyclonal) | Invitrogen | A6455 | IF(1:1000) |
| Antibody | anti-mouse 488 (Goat polyclonal) | Invitrogen | A10680 | IF(1:1000) |
| Antibody | anti-mouse 568 (Goat polyclonal) | Invitrogen | A11031 | IF(1:1000) |
| Antibody | anti-rabbit 488 (Goat polyclonal) | Invitrogen | A11008 | IF(1:1000) |
| Antibody | anti-rabbit 568 (Goat polyclonal) | Invitrogen | A11011 | F(1:1000) |
| Recombinant DNA reagent | pUAST-dU63gRNA vector carrying a ubiquitous U6:3 promoter | Schirmeier, University of Dresden | ||
| Sequence-based reagent | Uif2ndExon_gRNA2_fw | This work | sgRNA for uif cleavage 3’ to signal sequence | GTCGTTTCAATATCAAGCACTCGT |
| Sequence-based reagent | Uif2ndExon_gRNA2_rev | This work | sgRNA for uif cleavage 3’ to signal sequence | AAACACGAGTGCTTGATATTGAAA |
| Sequence-based reagent | UifCS_gRNA2_fw | This work | sgRNA for uif cleavage 5’ to cleavage site | GTCGTGTTCTGCGTACCTCGGTAG |
| Sequence-based reagent | UifCS_gRNA2_rev | This work | sgRNA for uif cleavage 5’ to cleavage site | AAATCTACCGAGGTACGCAGAACA |
| Sequence-based reagent | UifTMD_gRNA4_fw | This work | sgRNA for uif cleavage 5’ to transmembrane domain | GTCGCGCTGTGTGGGCTCCTTTAC |
| Sequence-based reagent | UifTMD_gRNA4_rev | This work | sgRNA for uif cleavage 5’ to transmembrane domain | AAACGTAAAGGAGCCCACACAGCG |
| Sequence-based reagent | UifCD_gRNA1_fw | This work | sgRNA for cleavage of uif cytoplasmic domain | GTCGCTACAATGAAACGTACATGA |
| Sequence-based reagent | UifCD_gRNA1_rev | This work | sgRNA for cleavage of uif cytoplasmic domain | AAACTCATGTACGTTTCATTGTAG |
| Software, algorithm | Fiji | Schindelin et al., 2012 |
Additional files
-
Supplementary file 1
Summary of Heartless modifier screen, related to STAR methods>Method details>Heartless modifier screen.
- https://cdn.elifesciences.org/articles/105759/elife-105759-supp1-v1.xlsx
-
Supplementary file 2
Rescue of the Htl gain of function wrapping glial phenotype by two independent double-stranded RNA (dsRNA) transgenes, related to STAR methods >Method details>Heartless modifier screen (Figure 3).
- https://cdn.elifesciences.org/articles/105759/elife-105759-supp2-v1.xlsx
-
Supplementary file 3
Rescue of Htl gain of function wrapping glial phenotype by two overlapping double-stranded RNA (dsRNA) transgenes, related to STAR methods>Method details>Heartless modifier screen.
- https://cdn.elifesciences.org/articles/105759/elife-105759-supp3-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/105759/elife-105759-mdarchecklist1-v1.docx






