Catalysis-dependent selenium incorporation and migration in the nitrogenase active site iron-molybdenum cofactor
Figures
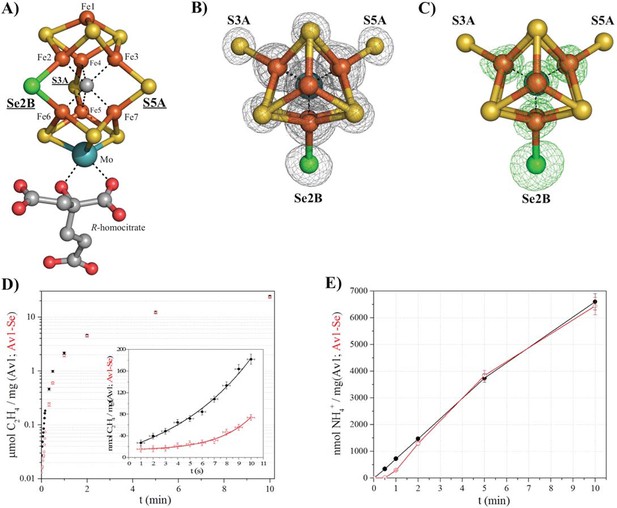
Selective Se-incorporation into the active site of the MoFe-protein.
(A) Side view of FeMoSe-cofactor ([7Fe:8S:1Se:Mo:C]-R-homocitrate) in Av1-Se2B at a resolution of 1.60 Å, highlighting the S2B replacement by Se. (B) View along the Fe1-C-Mo axis. The electron density (2Fo-Fc) map is contoured at 5.0 σ and represented as grey mesh. The 2Fo-Fc density at the Se2B site is significantly increased compared to the S5A and S3A sites. (C) Same orientation as B) superimposed with the anomalous difference Fourier map calculated at 12,662 eV (green) at a resolution of 1.60 Å contoured at 5.0 σ showing the presence of anomalous electron density arising from Se. Fe atoms are shown in orange, S in yellow, Se in green, Mo in turquoise, C in grey, and O in red. (D) Acetylene reduction activity of Av1 (black) compared to Av-Se (red). (E) Ammonia formation from reduction of the natural substrate, N2, was determined for Av1 (black) and Av1-Se (red). Error bars represent standard deviations from three measurements.
-
Figure 1—source data 1
Numerical data for the graphs depicted in Figure 1D and 1E.
- https://doi.org/10.7554/eLife.11620.004
-
Figure 1—source data 2
Numerical data for the graphs depicted in Figure 1—figure supplement 1.
- https://doi.org/10.7554/eLife.11620.005
-
Figure 1—source data 3
Numerical data for the graphs depicted in Figure 1—figure supplement 2.
- https://doi.org/10.7554/eLife.11620.006
-
Figure 1—source data 4
Numerical data for the graphs depicted in Figure 1—figure supplement 3.
- https://doi.org/10.7554/eLife.11620.007
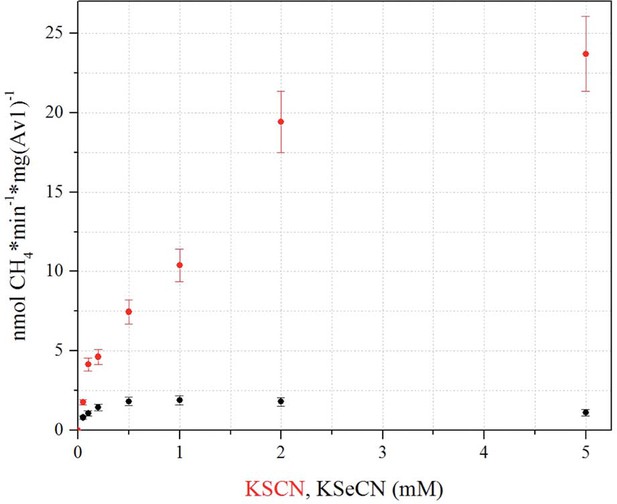
CH4 production based on KSeCN and KSCN as substrates.
Methane production was determined based on 0.05, 0.1, 0.2, 0.5, 1, 2, 5 mM KSCN (red) or KSeCN (black) as substrates. Maximum CH4 production from KSeCN was obtained at a concentration of 1 mM, whereas CH4 production from KSCN does not reach maximum within the tested range. Error bars represent standard deviations from three measurements.
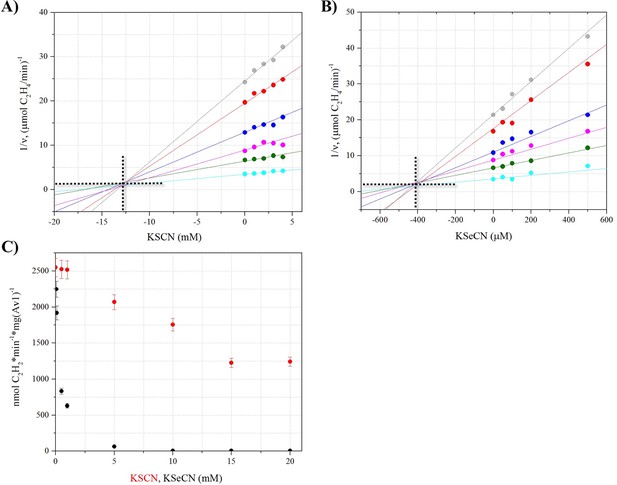
Inhibition of acetylene reduction by KSeCN and KSCN.
Inhibitory properties of KSeCN and KSCN were determined using a modified acetylene reduction assay. Concentrations for substrate (C2H2) were below saturation and concentrations for inhibitors (KSCN, KSeCN) were at low inhibition to allow for analysis. Dixon plots were prepared by plotting 1/v versus inhibitor concentration. Ki was determined from the intersection point derived from unrestrained linear fits of data points. (A) Dixon plot for KSCN, showing a Ki of 12.7 ± 1.2 mM KSCN. Concentrations of C2H2 were varied as follows: 20 (grey), 30 (red), 40 (blue), 60 (magenta), 100 (green), 500 (teal) μL per 9 mL total headspace volume. Concentrations of KSCN were: 0, 1, 2, 3, 4 mM. (B) Dixon plot for KSeCN, showing a Ki of 410 ± 30 uM KSeCN. Concentrations of C2H2 were varied as follows: 20 (grey), 30 (red), 40 (blue), 60 (magenta), 100 (green), 500 (teal) μL per 9 mL total headspace volume. Concentrations of KSeCN were: 0, 50, 100, 200, 500 μM. (C) Acetylene reduction activity in the presence of KSeCN (black) or KSCN (red) at varied concentrations: 50 μM, 75 μM, 100 μM, 500 μM, 1 mM, 5 mM, 10 mM, 15 mM and 20 mM. Error bars represent standard deviations from three measurements.
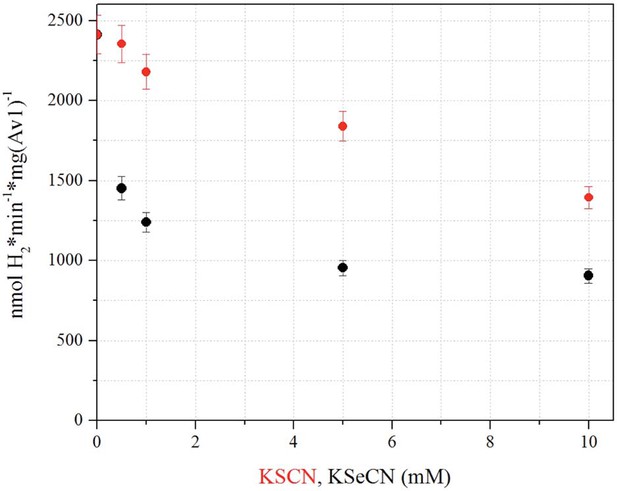
Influence of KSeCN and KSCN on proton reduction.
Proton reduction activity as a function of KSeCN (black) or KSCN (red) concentrations (0, 0.5, 1, 5, 10 mM). H2 production in the presence of 10 mM KSeCN is approximately 65% when compared to 10 mM KSCN, and approximately 38% in comparison to the KSCN/KSeCN free reduction activity. Error bars represent standard deviations from three measurements.
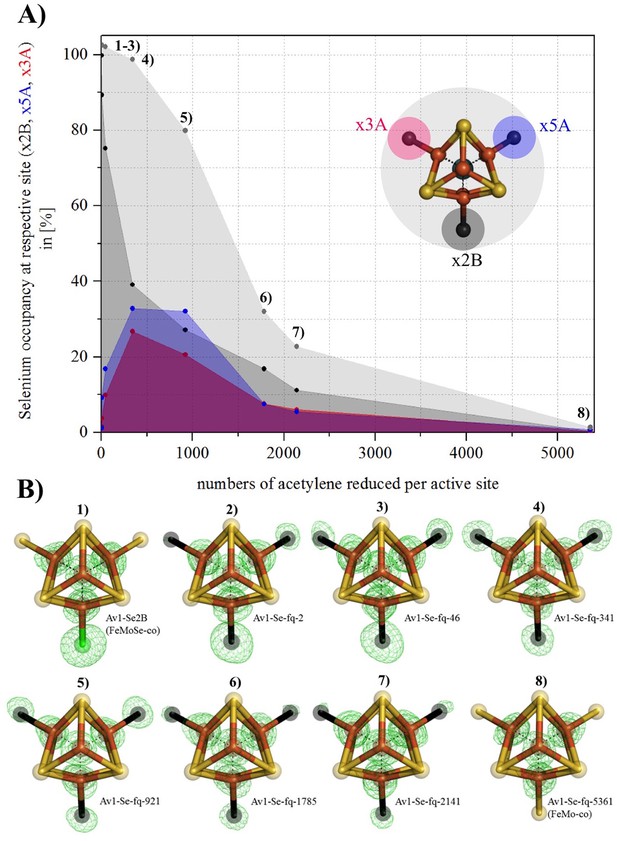
Se-migration in the active site during substrate reduction.
Se incorporation into all belt-S positions based on Av1-Se2B (FeMoSe-cofactor). (A) Se-occupancy in the active site as a function of numbers of acetylene reduced per cofactor. Se-occupancy of site x2B-position is shown in dark-grey, x5A in blue, and x3A in red. Sum of (x2B, x5A, x3A) is shown in light grey. (B) Structural models of Se-incorporated FeMo-cofactor during turnover. 1) FeMoSe-cofactor resting state in Av1-Se2B. 2–8) Cofactor structures obtained at seven time points according to numbers of acetylene reduced per active site: 2, 46, 341, 921, 1785, 2141 and 5361. Crystal structure resolutions in the order 1–8: 1.60, 1.50, 1.45, 1.32, 1.64, 1.66, 1.65 and 1.48 Å, respectively. Anomalous difference Fourier maps (calculated at 12,662 eV) allowing for the quantification of Se are shown as green mesh, and are contoured at 5.0 σ. Color scheme is according to Figure 1.
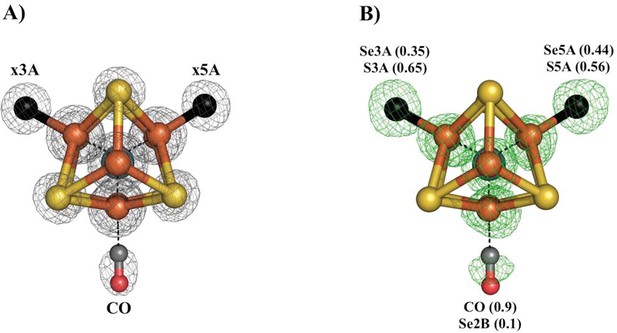
Se-migration upon CO-binding to Av1-Se2B.
Structure of Av1-Se-CO at a resolution of 1.53 Å, highlighting the Se2B replacement by CO and migration of Se to the remaining belt-S sites. (A) View along the Fe1-C-Mo axis of the metal center. The electron density (2Fo-Fc) map is contoured at 5.0 σ and represented as grey mesh. The electron density at the CO site is significantly decreased compared to the x5A and x3A sites and in excellent agreement with CO when residual Se-density is subtracted. (B) Same orientation as A) superimposed with the anomalous difference Fourier map calculated at 12,662 eV (green) at a resolution of 1.53 Å contoured at 5.0 σ showing the presence of anomalous electron density arising from Se. Numbers in parentheses indicate the fractional occupancies of the specified groups.
Additional files
-
Supplementary file 1
(A) Quantification of selenium occupancies at FeMoSe-cofactor belt x2B, x5A and x3A positions. (B) Data collection and refinement statistics.
- https://doi.org/10.7554/eLife.11620.013





